Population Genetics of Sillago japonica Among Five Populations Based on Mitochondrial Genome Sequences
Abstract
1. Introduction
2. Materials and Methods
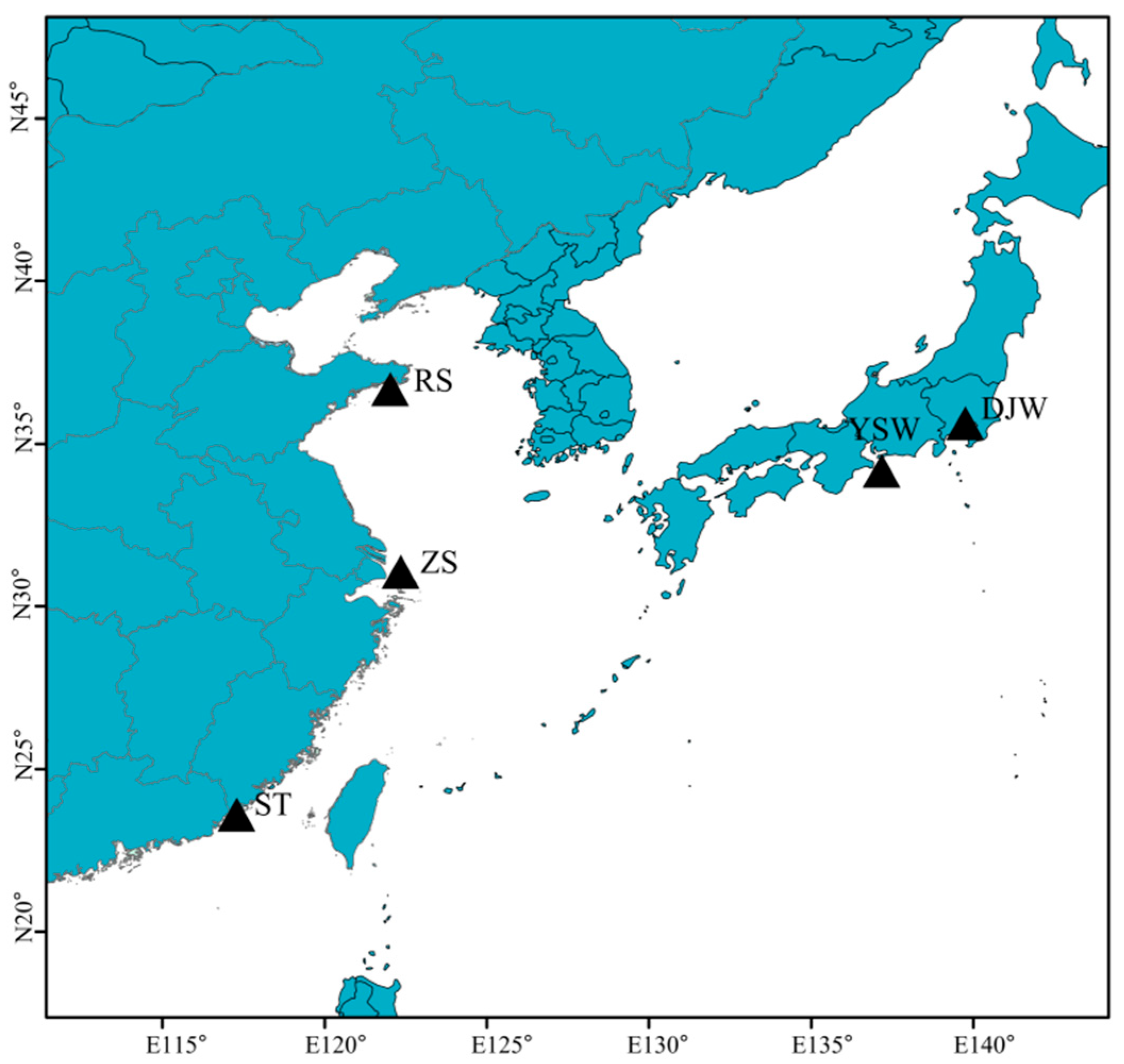
2.1. Sample Collection and DNA Extraction
2.2. Library Construction, Illumina Sequencing, and Mitochondrial Genome Assembly
2.3. Variant Calling, Population Structure, Diversity, and Divergence
2.4. Selection Pressure Analysis
3. Results
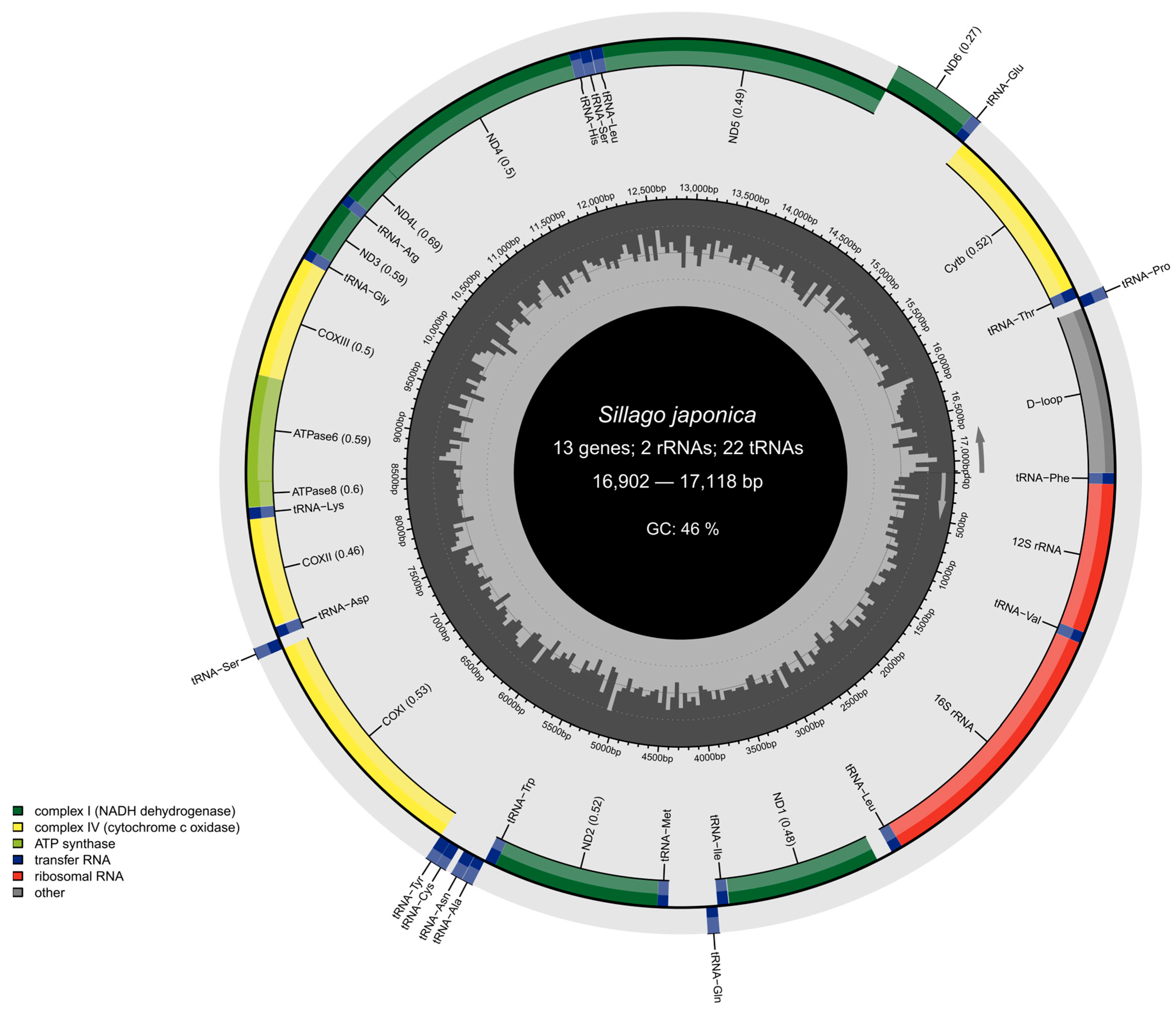
3.1. Mitochondrial Genome Assembly and Genetic Variant Mining
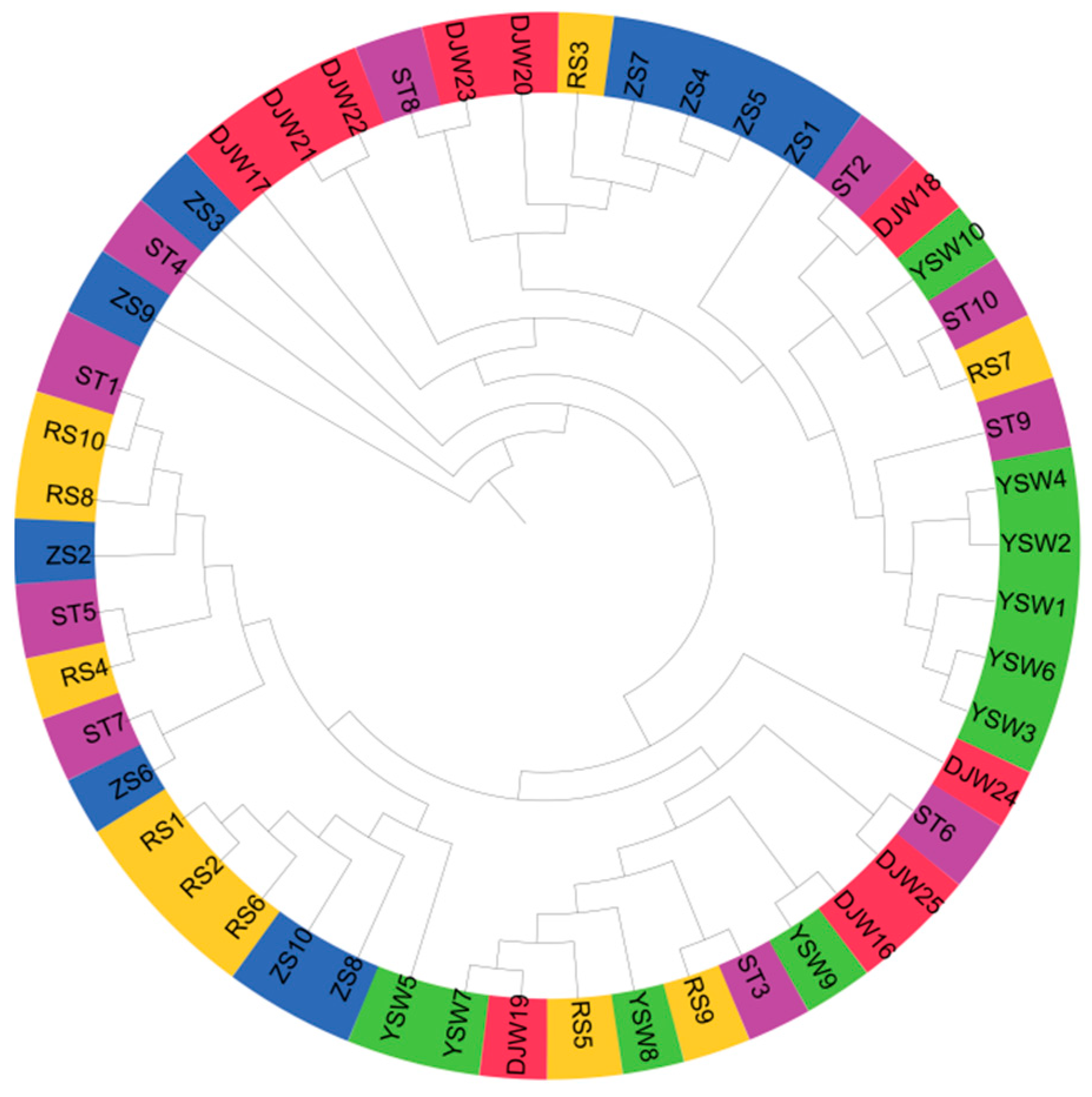
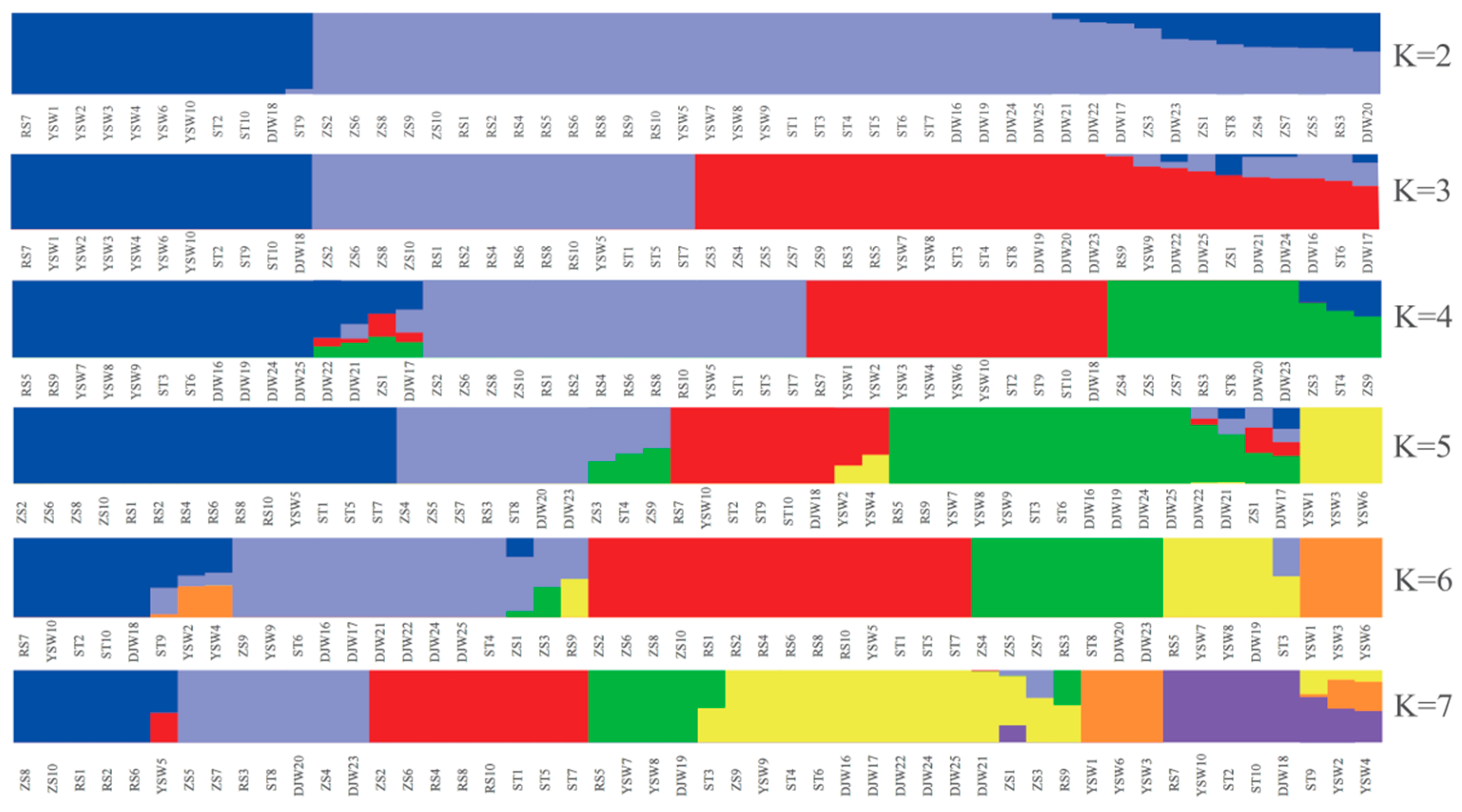
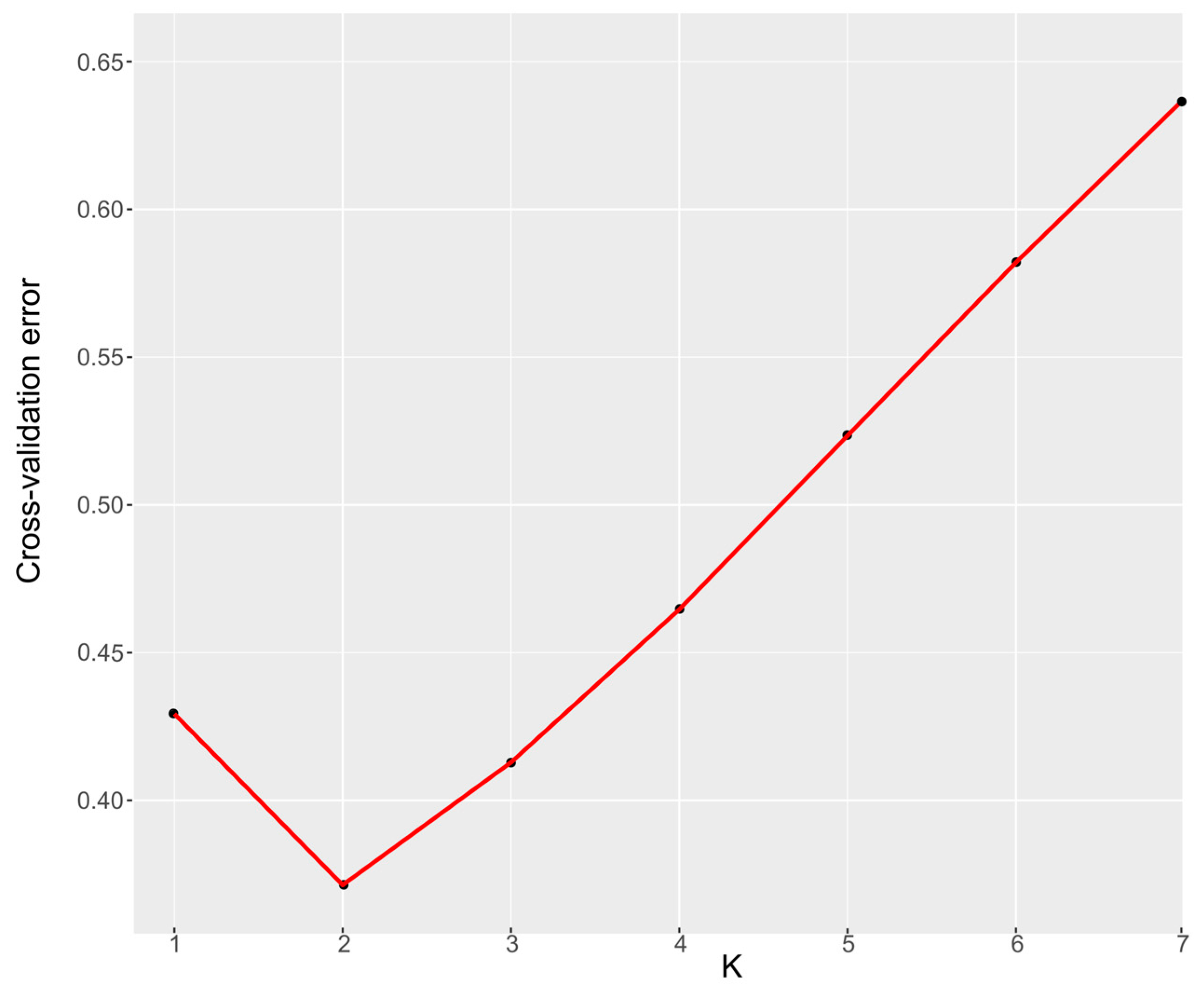
3.2. Population Structure of S. japonica
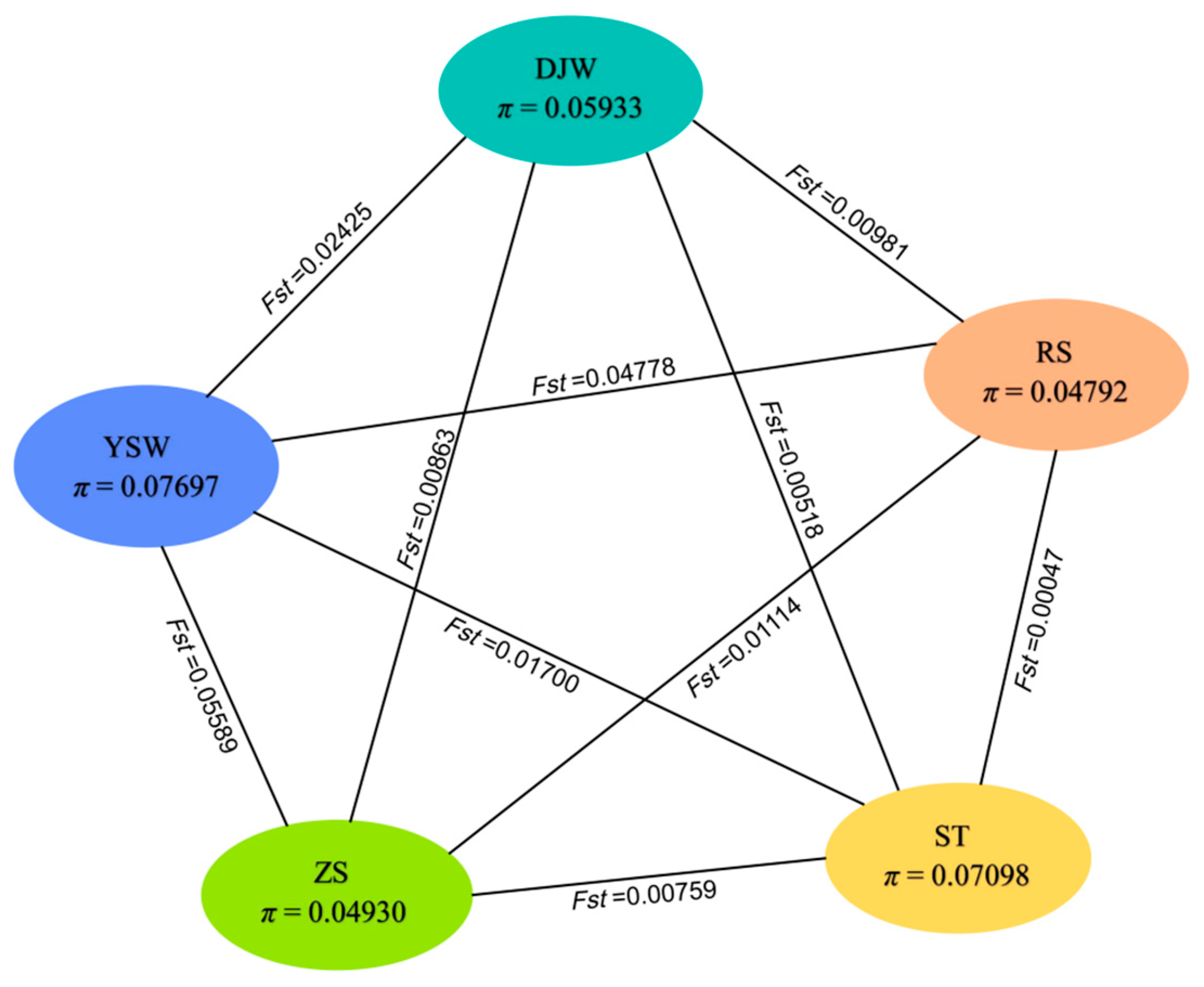
3.3. Genetic Diversity and Divergence of S. japonica Population
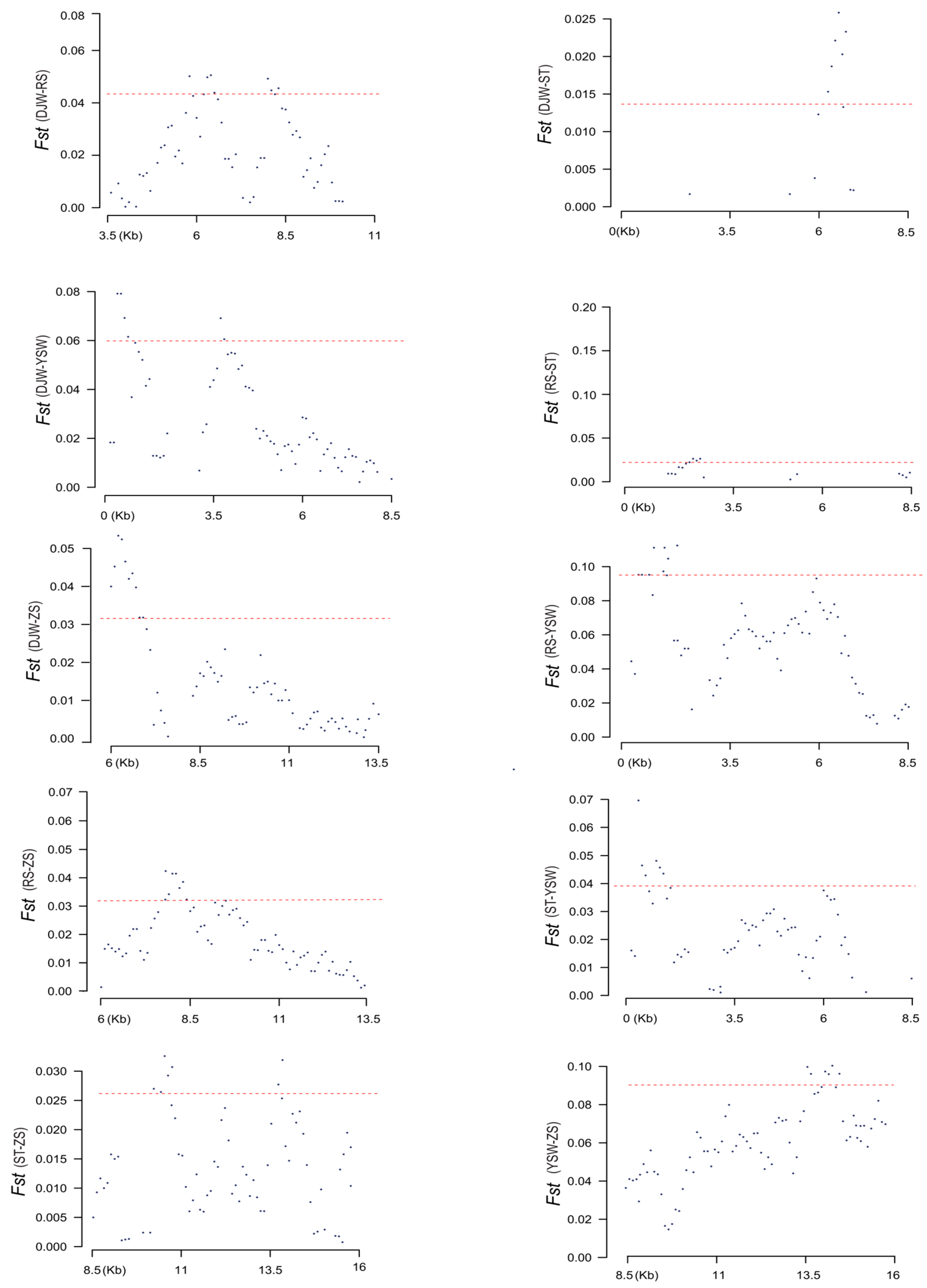
3.4. Genome-Wide Selection Pressure Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Oozeki, Y.; Hwang, P.P.; Hirano, R. Larval development of the Japanese whiting, Sillago japonica. Jpn. J. Ichthyol. 1992, 39, 59–66. [Google Scholar] [CrossRef]
- Wang, L.; Song, N.; Gao, T. Isolation and characterization of 24 polymorphic microsatellite loci in Japanese sillago (Sillago japonica). Conserv. Genet. Resour. 2014, 6, 581–584. [Google Scholar] [CrossRef]
- Khan, M.S.; Zakaria, M.S.; Ambak, M.A.; Alam, M.J.; Kashiwagi, M.; Iwai, T. Effect of temperature on the hatching success of the eggs of Japanese whiting, Sillago japonica Temminck & Schlegel, during the spawning season. Aquac. Res. 1991, 22, 317–322. [Google Scholar] [CrossRef]
- Sulistiono; Watanabe, S.; Yokota, M. Reproduction of the Japanese whiting, Sillago japonica, in Tateyama Bay. Aquacult. Sci. 1999, 47, 209–214. [Google Scholar]
- Shimasaki, Y.; Oshima, Y.; Inoue, S.; Inoue, Y.; Kang, I.J.; Nakayama, K.; Imoto, H.; Honjo, T. Effect of tributyltin on reproduction in Japanese whiting, Sillago japonica. Mar. Environ. Res. 2006, 62, S245–S248. [Google Scholar] [CrossRef]
- Gao, T.X.; Yang, T.Y.; Yanagimoto, T.; Xiao, Y.S. Levels and patterns of genetic variation in Japanese whiting (Sillago japonica) based on mitochondrial DNA control region. Mitochondrial DNA A 2019, 30, 172–183. [Google Scholar] [CrossRef]
- Yang, T.Y.; Gao, T.X.; Meng, W.; Jiang, Y.L. Genome-wide population structure and genetic diversity of Japanese whiting (Sillago japonica) inferred from genotyping-by-sequencing (GBS): Implications for fisheries management. Fish. Res. 2020, 225, 105501. [Google Scholar] [CrossRef]
- Castellana, S.; Vicario, S.; Saccone, C. Evolutionary patterns of the mitochondrial genome in Metazoa: Exploring the role of mutation and selection in mitochondrial protein–coding genes. Genome Biol. Evol. 2011, 3, 1067–1079. [Google Scholar] [CrossRef]
- Nagpure, N.S.; Rashid, I.; Pathak, A.K.; Singh, M.; Pati, R.; Singh, S.P.; Sarkar, U.K. FMiR: A curated resource of mitochondrial DNA information for fish. PLoS One 2015, 10, e0136711. [Google Scholar] [CrossRef] [PubMed]
- Ruan, H.; Li, M.; Li, Z.; Huang, J.; Chen, W.; Sun, J.; Liu, L.; Zou, K. Comparative analysis of complete mitochondrial genomes of three Gerres fishes (Perciformes: Gerreidae) and primary exploration of their evolution history. Int. J. Mol. Sci. 2020, 21, 1874. [Google Scholar] [CrossRef] [PubMed]
- Hao, C.; Liu, Y.; Wei, N.; Arken, K.; Shi, C.; Yue, C. The complete mitochondrial genomes of the Leuciscus baicalensis and Rutilus rutilus: A detailed genomic comparison among closely related species of the Leuciscinae subfamily. Gene 2023, 877, 147535. [Google Scholar] [CrossRef] [PubMed]
- Corral-Lou, A.; Perea, S.; Doadrio, I. High genetic differentiation in the endemic and endangered freshwater fish Achondrostoma salmantinum Doadrio and Elvira, 2007 from Spain, as revealed by mitochondrial and SNP markers. Conserv. Genet. 2021, 22, 585–600. [Google Scholar] [CrossRef]
- Parubrub, A.; Reyes, R.; Smallridge, C.J.; Woods, B.; Haymer, D. mitochondrial single nucleotide polymorphisms in Ceratitis capitata (Diptera: Tephritidae) can distinguish sterile, released flies from wild flies in various regions of the world. J. Econ. Entom. 2015, 108, 301–306. [Google Scholar] [CrossRef]
- Gong, L.; Jiang, H.; Zhu, K.; Lu, X.; Liu, L.; Liu, B.; Jiang, L.; Ye, Y.; Lü, Z. Large-scale mitochondrial gene rearrangements in the hermit crab Pagurus nigrofascia and phylogenetic analysis of the Anomura. Gene 2019, 695, 75–83. [Google Scholar] [CrossRef]
- Elmerot, C.; Arnason, U.; Gojobori, T.; Janke, A. The mitochondrial genome of the pufferfish, Fugu rubripes, and ordinal teleostean relationships. Gene 2002, 295, 163–172. [Google Scholar] [CrossRef]
- Laurimäe, T.; Kinkar, L.; Romig, T.; Omer, R.A.; Casulli, A.; Umhang, G.; Saarma, U. The benefits of analysing complete mitochondrial genomes: Deep insights into the phylogeny and population structure of Echinococcus granulosus sensu lato genotypes G6 and G7. Infect. Genet. Evol. 2018, 64, 85–94. [Google Scholar] [CrossRef]
- Jeon, H.B.; Choi, S.H.; Suk, H.Y. Exploring the utility of partial cytochrome c oxidase subunit 1 for DNA barcoding of gobies. Anim. Syst. Evol. Divers. 2012, 28, 269–278. [Google Scholar] [CrossRef]
- Wang, L.; Shi, X.; Su, Y.; Meng, Z.; Lin, H. Loss of genetic diversity in the cultured stocks of the large yellow croaker, Larimichthys crocea, revealed by microsatellites. Int. J. Mol. Sci. 2012, 13, 5584–5597. [Google Scholar] [CrossRef]
- Mazaris, A.D.; Kallimanis, A.; Gissi, E.; Pipitone, C.; Danovaro, R.; Claudet, J.; Fraschetti, S. Threats to marine biodiversity in European protected areas. Sci. Total Environ. 2019, 677, 418–426. [Google Scholar] [CrossRef]
- Worm, B.; Barbier, E.B.; Beaumont, N.; Duffy, J.E.; Folke, C.; Halpern, B.S.; Watson, R. Impacts of biodiversity loss on ocean ecosystem services. Science 2006, 314, 787–790. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Farrell, A.P.; Matala, A.; Hoffman, N.; Narum, S.R. Physiological and genomic signatures of evolutionary thermal adaptation in redband trout from extreme climates. Evol. Appl. 2018, 11, 1686–1699. [Google Scholar] [CrossRef]
- Zhang, Z.; Xu, S.; Capinha, C.; Weterings, R.; Gao, T. Using species distribution model to predict the impact of climate change on the potential distribution of Japanese whiting Sillago japonica. Ecol. Indic. 2019, 104, 333–340. [Google Scholar] [CrossRef]
- Yona, G.; Madalla, N.; Mwandya, A.; Ishimatsu, A.; Bwathondi, P. Effects of elevated carbon dioxide and temperature on survival and morphology of Japanese whiting Sillago japonica. Int. J. Fish. Aquat. Stud. 2016, 4, 48–55. [Google Scholar]
- Kramvis, A.; Bukofzer, S.; Kew, M.C. Comparison of hepatitis B virus DNA extractions from serum by the QIAamp blood kit, GeneReleaser, and the phenol-chloroform method. J. Clin. Microbiol. 1996, 34, 2731–2733. [Google Scholar] [CrossRef] [PubMed]
- de Sena Brandine, G.; Smith, A.D. Falco: High-speed FastQC emulation for quality control of sequencing data. F1000Research 2021, 8, 1874. [Google Scholar] [CrossRef] [PubMed]
- Giannoulatou, E.; Park, S.H.; Humphreys, D.T.; Ho, J.W. Verification and validation of bioinformatics software without a gold standard: A case study of BWA and Bowtie. BMC Bioinform. 2014, 15, S15. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Dierckxsens, N.; Mardulyn, P.; Smits, G. NOVOPlasty: De novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 2017, 45, e18. [Google Scholar]
- McKenna, A.; Hanna, M.; Banks, E.; Sivachenko, A.; Cibulskis, K.; Kernytsky, A.; Garimella, K.; Altshuler, D.; Gabriel, S.; Daly, M.; et al. The Genome Analysis Toolkit: A MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010, 20, 1297–1303. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Yang, J.; Lee, S.H.; Goddard, M.E.; Visscher, P.M. GCTA: A tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 2011, 88, 76–82. [Google Scholar] [CrossRef]
- Catchen, J.; Hohenlohe, P.A.; Bassham, S.; Amores, A.; Cresko, W.A. Stacks: An analysis tool set for population genomics. Mol. Ecol. 2013, 22, 3124–3140. [Google Scholar] [CrossRef]
- Danecek, P.; Auton, A.; Abecasi, G.; Albers, C.A.; Banks, E.; DePristo, M.A.; Handsaker, R.E.; Lunter, G.; Marth, G.T.; Sherry, S.T.; et al. The variant call format and VCFtools. Bioinformatics 2011, 27, 2156–2158. [Google Scholar] [CrossRef] [PubMed]
- Lujan, N.K.; Colm, J.E.; Weir, J.T.; Montgomery, F.A.; Noonan, B.P.; Lovejoy, N.R.; Mandrak, N.E. Genomic population structure of Grass Pickerel (Esox americanus vermiculatus) in Canada: Management guidance for an at-risk fish at its northern range limit. Conserv. Genet. 2022, 23, 713–725. [Google Scholar] [CrossRef]
- Lopez-Marquez, V.; Cushman, S.A.; Templado, J.; Wan, H.Y.; Bothwell, H.M.; Machordom, A. Genetic connectivity of two marine gastropods in the Mediterranean Sea: Seascape genetics reveals species-specific oceanographic drivers of gene flow. Mol. Ecol. 2021, 30, 4608–4629. [Google Scholar] [CrossRef]
- Zhang, B.D.; Li, Y.L.; Xue, D.X.; Liu, J.X. Population genomics reveals shallow genetic structure in a connected and ecologically important fish from the Northwestern Pacific Ocean. Front. Mar. Sci. 2020, 7, 374. [Google Scholar] [CrossRef]
- Sun, C.H.; Yang, F.; Huang, Q.; Zeng, X.S.; Zhang, Y.N.; Li, S.; Yu, J.F.; Zhang, Q. Genetic population structure and demographic history of the endemic fish Paralichthys olivaceus of the Northwest Pacific Ocean. Ecol. Evol. 2022, 12, e9506. [Google Scholar] [CrossRef]
- Han, Z.Q.; Guo, X.Y.; Liu, Q.; Liu, S.S.; Zhang, Z.X.; Xiao, S.J.; Gao, T.X. Whole-genome resequencing of Japanese whiting (Sillago japonica) provide insights into local adaptations. Zool. Res. 2021, 42, 548–561. [Google Scholar] [CrossRef]
- Fridjonsson, O.; Olafsson, K.; Tompsett, S.; Bjornsdottir, S.; Consuegra, S.; Knox, D.; de Leaniz, C.G.; Magnusdottir, S.; Olafsdottir, G.; Verspoor, E.; et al. Detection and map of mtDNA SNPs in Atlantic salmon using high throughput DNA sequencing. BMC Genom. 2011, 12, 179. [Google Scholar] [CrossRef]
- Hu, X.; Luan, P.; Cao, C.; Li, C.; Jia, Z.; Ge, Y.; Shi, L.; Shang, M.; Wang, S.; Meng, Z.; et al. Characterization of the mitochondrial genome of Megalobrama terminalis in the Heilong River and a clearer phylogeny of the genus Megalobrama. Sci. Rep. 2019, 9, 8509. [Google Scholar] [CrossRef]
- Kivisild, T. Maternal ancestry and population history from whole mitochondrial genomes. Investig. Genet. 2015, 6, 3. [Google Scholar] [CrossRef]
- Toews, D.P.; Brelsford, A. The biogeography of mitochondrial and nuclear discordance in animals. Mol. Ecol. 2012, 21, 3907–3930. [Google Scholar] [CrossRef]
- Vandewoestijne, S.; Schtickzelle, N.; Baguette, M. Positive correlation between genetic diversity and fitness in a large, well-connected metapopulation. BMC Biol. 2008, 6, 46. [Google Scholar] [CrossRef]
- Johnson, D.W.; Freiwald, J.; Bernardi, G. Genetic diversity affects the strength of population regulation in a marine fish. Ecology 2016, 97, 627–639. [Google Scholar] [CrossRef]
- Yan, H.F.; Kyne, P.M.; Jabado, R.W.; Leeney, R.H.; Davidson, L.N.; Derrick, D.H.; Finucci, B.; Freckleton, R.P.; Fordham, S.V.; Dulvy, N.K. Overfishing and habitat loss drive range contraction of iconic marine fishes to near extinction. Sci. Adv. 2021, 7, eabb6026. [Google Scholar] [CrossRef] [PubMed]
- McClenachan, L.; Cooper, A.B.; Dulvy, N.K. Rethinking trade-driven extinction risk in marine and terrestrial megafauna. Curr. Biol. 2016, 26, 1640–1646. [Google Scholar] [CrossRef] [PubMed]
- Sulistiono; Yokota, M.; Kitada, S.; Watanabe, S. Age and growth of Japanese whiting Sillago japonica in Tateyama Bay. Fish. Sci. 1999, 65, 117–122. [Google Scholar] [CrossRef]
- Wright, S. Evolution and the Genetics of Populations, Volume 4: Variability Within and Among Natural Populations; University of Chicago Press: Chicago, IL, USA, 1984; Volume 4. [Google Scholar]

| Category | S. japonica Populations |
|---|---|
| SNPs | 2966 |
| Indels | 414 |
| Synonymous variants | 88 |
| Upstream | 1458 |
| Downstream | 1650 |
| Exon | 148 |
| Intergenic variants | 10 |
| Intragenic variants | 106 |
| Frameshift variants | 5 |
| Noncoding transcript exon variants | 33 |
| Population | Observed Heterozygosity (Ho) | Expected Heterozygosity (He) | Percentages of Polymorphic Loci (PPB) % |
|---|---|---|---|
| DJW | 0.00019 | 0.05636 | 26.54360 |
| RS | 0.00006 | 0.04552 | 19.85996 |
| ST | 0.00001 | 0.06743 | 30.36283 |
| YSW | 0.00001 | 0.07311 | 28.38956 |
| ZS | 0.00528 | 0.04683 | 19.41439 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, B.; Gao, T.; Qu, Y.; Zhang, X. Population Genetics of Sillago japonica Among Five Populations Based on Mitochondrial Genome Sequences. Genes 2025, 16, 978. https://doi.org/10.3390/genes16080978
Zhu B, Gao T, Qu Y, Zhang X. Population Genetics of Sillago japonica Among Five Populations Based on Mitochondrial Genome Sequences. Genes. 2025; 16(8):978. https://doi.org/10.3390/genes16080978
Chicago/Turabian StyleZhu, Beiyan, Tianxiang Gao, Yinquan Qu, and Xiumei Zhang. 2025. "Population Genetics of Sillago japonica Among Five Populations Based on Mitochondrial Genome Sequences" Genes 16, no. 8: 978. https://doi.org/10.3390/genes16080978
APA StyleZhu, B., Gao, T., Qu, Y., & Zhang, X. (2025). Population Genetics of Sillago japonica Among Five Populations Based on Mitochondrial Genome Sequences. Genes, 16(8), 978. https://doi.org/10.3390/genes16080978






