Forensic DNA Phenotyping: Genes and Genetic Variants for Eye Color Prediction
Abstract
1. Introduction
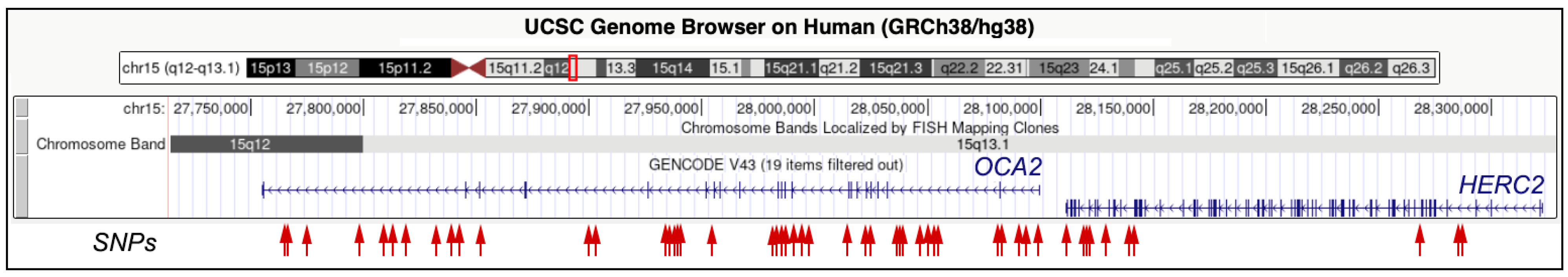
2. The Genetic Basis of Eye Color: OCA2 and HERC2 Polymorphisms
2.1. OCA2 Gene
| Gene | SNP-ID (a) | Allele | Variant | Chr. | Pos. (nt) | Variant Type |
|---|---|---|---|---|---|---|
| ASIP | rs1015362 | C | T | 20q11.22 | 34,150,806 | 5′UTR |
| ASIP | rs6119471 * | C | G | 20q11.22 | 34,197,406 | Intron |
| ASIP | rs6058017 | A | G | 20q11.22 | 34,269,192 | 3′UTR |
| HERC2 | rs1129038 * | C | T | 15q13.1 | 28,111,713 | Intron |
| HERC2 | rs191109490 | A | C | 15q13.1 | 28,120,403 | Intron |
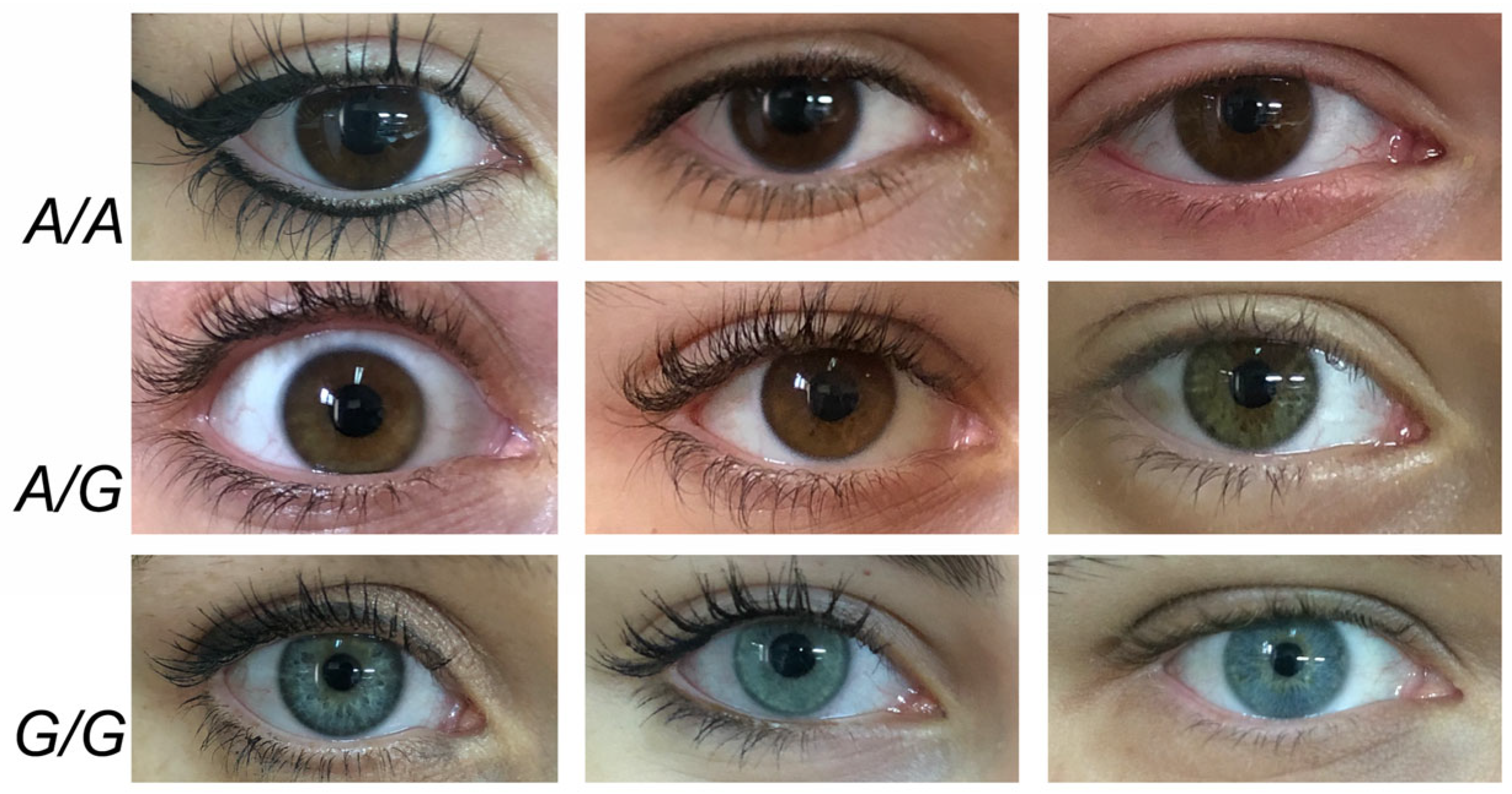
| HERC2 | rs12913832 * | A | G | 15q13.1 | 28,120,472 | Intron |
| HERC2 | rs7183877 * | C | A | 15q13.1 | 28,120,587 | Intron |
| HERC2 | rs3935591 | T | C | 15q13.1 | 28,128,866 | Intron |
| HERC2 | rs551217952 | A | C | 15q13.1 | 28,139,387 | Intron |
| HERC2 | rs11636232 * | C | T | 15q13.1 | 28,141,480 | Syn.: Gln3989Gln |
| HERC2 | rs916977 | T | C | 15q13.1 | 28,268,218 | Intron |
| HERC2 | rs1667394 * | C | T | 15q13.1 | 28,285,036 | Intron |
| HERC2 | rs12592730 | G | A | 15q13.1 | 28,285,213 | Intron |
| IRF4 | rs12203592 * | C | T | 6p25.3 | 396,321 | Intron |
| MC1R | rs2228479 | G | A | 16q24.3 | 89,919,532 | Missense: Val92Met |
| MC1R | rs1805007 | C | T | 16q24.3 | 89,919,709 | Missense:Arg151Cys |
| MC1R | rs1805008 | C | T | 16q24.3 | 89,919,736 | Missense: Arg160Trp |
| MC1R | rs885479 | G | A | 16q24.3 | 89,919,746 | Missense: Arg163Gln |
| MC1R | rs1805009 | A | C | 16q24.3 | 89,920,138 | Missense:Asp294His |
| OCA2 | rs1498519 | G | T | 15q13.1 | 27,766,505 | Intron |
| OCA2 | rs4778177 | C | T | 15q12 | 27,766,748 | Intron |
| OCA2 | rs8023340 | A | G | 15q12 | 27,775,242 | Intron |
| OCA2 | rs1874835 | G | T | 15q12 | 27,799,154 | Intron |
| OCA2 | rs924314 | G | A | 15q13.1 | 27,809,200 | Intron |
| OCA2 | rs924312 | C | G | 15q13.1 | 27,812,717 | Intron |
| OCA2 | rs74409036 | G | A | 15q13.1 | 27,818,606 | Intron |
| OCA2 | rs4778190 | T | C | 15q13.1 | 27,832,014 | Intron |
| OCA2 | rs2036213 | C | A | 15q13.1 | 27,839,034 | Intron |
| OCA2 | rs78544415 | C | T | 15q13.1 | 27,842,634 | Intron |
| OCA2 | rs1004611 | G | A | 15q13.1 | 27,852,132 | Intron |
| OCA2 | rs2871886 | T | C | 15q13.1 | 27,899,878 | Intron |
| OCA2 | rs3099645 | G | T | 15q13.1 | 27,902,213 | Intron |
| OCA2 | rs977588 | C | A | 15q13.1 | 27,934,160 | Intron |
| OCA2 | rs977589 | C | T | 15q13.1 | 27,934,457 | Intron |
| OCA2 | rs1375170 | G | T | 15q13.1 | 27,939,008 | Intron |
| OCA2 | rs1448490 | C | A | 15q13.1 | 27,939,381 | Intron |
| OCA2 | rs2594935 | G | A | 15q13.1 | 27,939,892 | Intron |
| OCA2 | rs1800414 | T | C | 15q13.1 | 27,951,891 | Missense: Hys615Arg |
| OCA2 | rs728405 | C | A | 15q13.1 | 27,954,707 | Intron |
| OCA2 | rs74653330 | C | T | 15q13.1 | 27,983,407 | Missense: Ala481Thr |
| OCA2 | rs1900758 | C | T | 15q13.1 | 27,984,951 | Intron |
| OCA2 | rs121918166 | C | T | 15q13.1 | 27,985,101 | Missense: Val443Ile |
| OCA2 | rs1800407 * | C | T | 15q13.1 | 27,985,172 | Missense: Arg419Gln |
| OCA2 | rs10852218 | T | C | 15q13.1 | 27,986,647 | Intron |
| OCA2 | rs1800404 | C | T | 15q13.1 | 27,990,627 | Syn: Ala355Ala |
| OCA2 | rs735066 | A | G | 15q13.1 | 27,993,749 | Intron |
| OCA2 | rs1800401 | G | A | 15q13.1 | 28,014,907 | Missense: Arg305Trp |
| OCA2 | rs2305252 | C | T | 15q13.1 | 28,022,192 | Intron |
| OCA2 | rs3794606 | A | C | 15q13.1 | 28,023,862 | Intron |
| OCA2 | rs4778232 * | C | T | 15q13.1 | 28,036,619 | Intron |
| OCA2 | rs1448485 | G | T | 15q13.1 | 28,037,595 | Intron |
| OCA2 | rs8024968 * | C | T | 15q13.1 | 28,038,543 | Intron |
| OCA2 | rs1375164 | C | T | 15q13.1 | 28,046,666 | Intron |
| OCA2 | rs1597196 | G | T | 15q13.1 | 28,049,776 | Intron |
| OCA2 | rs895828 | G | C | 15q13.1 | 28,052,887 | Intron |
| OCA2 | rs895829 | T | C | 15q13.1 | 28,052,911 | Intron |
| OCA2 | rs4778137 | C | G | 15q13.1 | 28,082,689 | Intron |
| OCA2 | rs72714116 | C | T | 15q13.1 | 28,083,061 | Intron |
| OCA2 | rs4778138 | A | G | 15q13.1 | 28,090,674 | Intron |
| OCA2 | rs4778241 * | A | C | 15q13.1 | 28,093,567 | Intron |
| OCA2 | rs7495174 | T | G | 15q13.1 | 28,099,092 | Intron |
| SLC24A4 | rs12896399 * | G | T | 14q32.12 | 92,307,319 | 5′UTR |
| SLC24A5 | rs1426654 | A | G | 15q21.1 | 48,134,287 | Missense: Thr111Ala |
| SLC45A2 | rs16891982 * | C | G | 5p13.2 | 33,951,588 | Missense: Leu374Phe |
| SLC45A2 | rs26722 | C | T | 5p13.2 | 33,963,765 | Missense: Glu272Lys |
| TYR | rs1393350 * | G | A | 11q14.3 | 89,277,878 | Intron |
| TYR | rs1126809 | G | A | 11q14.3 | 89,284,793 | Missense: Arg402Gln |
| TYRP1 | rs1325127 | C | T | 9p23 | 12,668,328 | 5′UTR |
| TYRP1 | rs1408799 | T | C | 9p23 | 12,672,097 | 5′UTR |
| TYRP1 | rs62538956 | T | C | 9p23 | 12,679,244 | 5′UTR |
| TYRP1 | rs35866166 | T | C | 9p23 | 12,698,471 | Syn: Ser243Ser |
| TYRP1 | rs2733832 | C | T | 9p23 | 12,704,725 | Intron |
| TYRP1 | rs683 | C | A | 9p23 | 12,709,305 | 3′UTR |
| Gene | SNP-ID | Walsh et al. [34] (a) | Allwood et al. [35] | Ruiz et al. [36] | Hart et al. [37] |
|---|---|---|---|---|---|
| ASIP | rs6119471 | X | |||
| HERC2 | rs1129038 | X | X | ||
| HERC2 | rs12913832 | X | X | X | |
| HERC2 | rs7183877 | X | |||
| HERC2 | rs11636232 | X | |||
| HERC2 | rs1667394 | X | |||
| IRF4 | rs12203592 | X | X | X | |
| OCA2 | rs1800407 | X | X | X | |
| OCA2 | rs4778232 | X | |||
| OCA2 | rs8024968 | X | |||
| OCA2 | rs4778241 | X | |||
| SLC24A4 | rs12896399 | X | X | X | X |
| SLC45A2 | rs16891982 | X | X | X | |
| TYR | rs1393350 | X | X | X |
2.2. HERC2 Gene
2.3. Model for OCA2 Gene Regulation
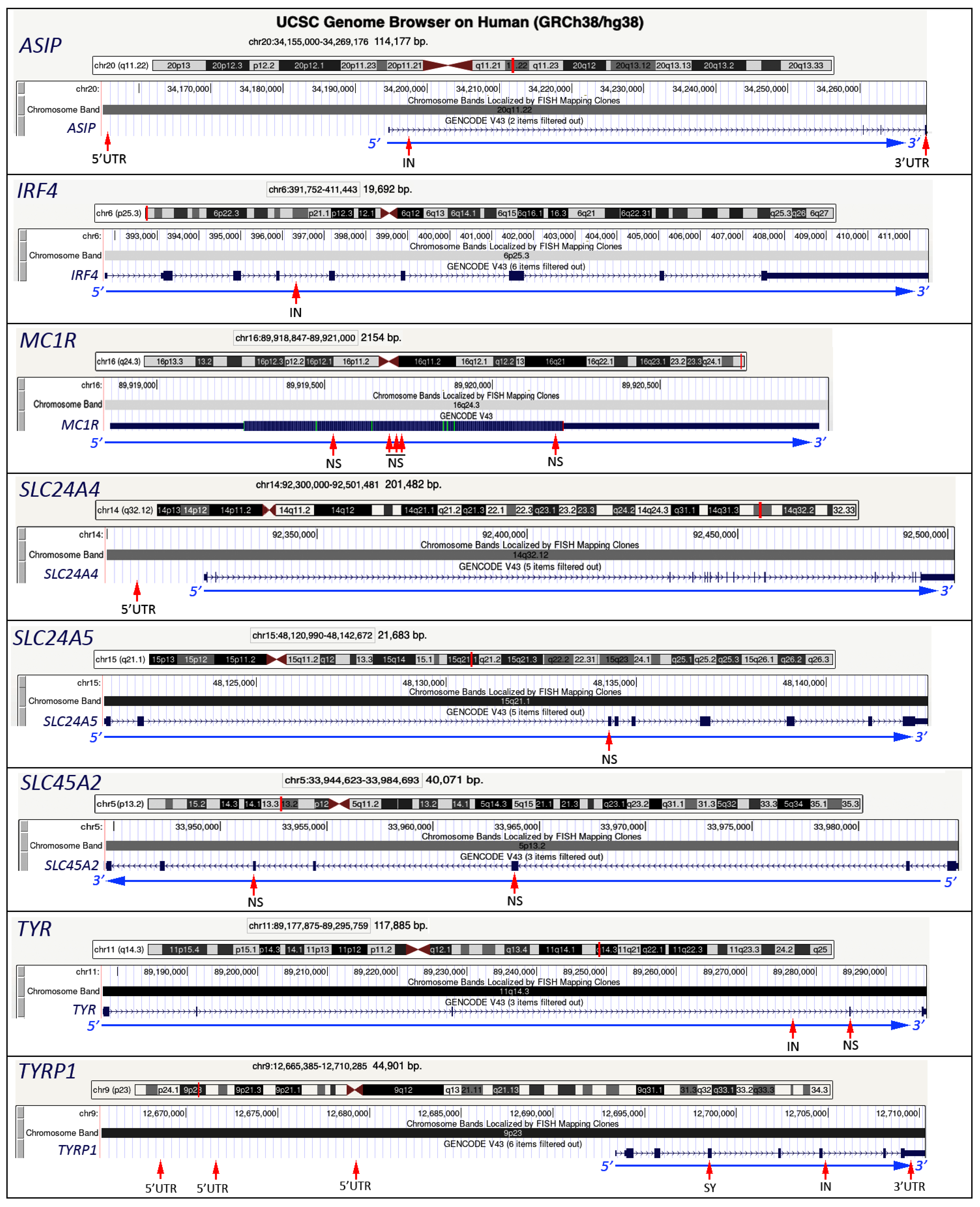
3. Other Genes Related to the Eye Color
3.1. TYR Gene
3.2. TYRP1 Gene
3.3. SLC45A2 Gene
3.4. SLC24A4 Gene
3.5. SLC24A5 Gene
3.6. ASIP Gene
3.7. MC1R Gene
3.8. IRF4 Gene
4. Gene Panels to Predict Eye Color
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Jobling, M.A.; Gill, P. Encoded evidence: DNA in forensic analysis. Nat. Rev. Genet. 2004, 5, 739–751. [Google Scholar] [CrossRef]
- Schneider, P.M.; Prainsack, B.; Kayser, M. The Use of Forensic DNA Phenotyping in Predicting Appearance and Biogeographic Ancestry. Dtsch. Arztebl. Int. 2019, 23, 873–880. [Google Scholar] [CrossRef]
- Wienroth, M. Governing anticipatory technology practices. Forensic DNA phenotyping and the forensic genetics community in Europe. New Genet. Soc. 2018, 37, 137–152. [Google Scholar] [CrossRef]
- Oosthuizen, T.; Howes, L.M. The development of forensic DNA analysis: New debates on the issue of fundamental human rights. Forensic Sci. Int. Genet. 2022, 6, 102606. [Google Scholar] [CrossRef]
- Hopman, R.; M’charek, A. Facing the unknown suspect: Forensic DNA phenotyping and the oscillation between the individual and the collective. Biosocieties 2020, 15, 438–462. [Google Scholar] [CrossRef]
- M’charek, A.; Toom, V.; Jong, L. The Trouble with Race in Forensic Identification. Sci. Technol. Hum. Values 2020, 45, 804–828. [Google Scholar] [CrossRef]
- Kayser, M.; Branicki, W.; Parson, W.; Phillips, C. Recent advances in Forensic DNA Phenotyping of appearance, ancestry and age. Forensic Sci. Int. Genet. 2023, 6, 102870. [Google Scholar] [CrossRef]
- Ballard, D.; Winkler-Galicki, J.; Wesoły, J. Massive parallel sequencing in forensics: Advantages, issues, technicalities, and prospects. Int. J. Leg. Med. 2020, 134, 1291–1303. [Google Scholar] [CrossRef]
- Butler, J.M. Recent advances in forensic biology and forensic DNA typing: INTERPOL review 2019–2022. Forensic Sci. Int. Synerg. 2022, 6, 100311. [Google Scholar] [CrossRef] [PubMed]
- Carratto, T.M.T.; Moraes, V.M.S.; Recalde, T.S.F.; de Oliveira, M.L.G.; Mendes-Junior, C.T. Applications of massively parallel sequencing in forensic genetics. Genet. Mol. Biol. 2022, 45, e20220077. [Google Scholar] [CrossRef] [PubMed]
- Fabbri, M.; Alfieri, L.; Mazdai, L.; Frisoni, P.; Gaudio, R.M.; Neri, M. Application of Forensic DNA Phenotyping for Prediction of Eye, Hair and Skin Colour in Highly Decomposed Bodies. Healthcare 2023, 5, 647. [Google Scholar] [CrossRef]
- Haddrill, P.R. Developments in forensic DNA analysis. Emerg. Top. Life Sci. 2021, 3, 381–393. [Google Scholar] [CrossRef]
- Pajnič, I.Z.; Zupanc, T.; Leskovar, T.; Črešnar, M.; Fattorini, P. Eye and Hair Color Prediction of Ancient and Second World War Skeletal Remains Using a Forensic PCR-MPS Approach. Genes 2022, 8, 1432. [Google Scholar] [CrossRef]
- Melchionda, F.; Silvestrini, B.; Robino, C.; Bini, C.; Fattorini, P.; Martinez-Labarga, C.; De Angelis, F.; Tagliabracci, A.; Turchi, C. Development and Validation of MPS-Based System for Human Appearance Prediction in Challenging Forensic Samples. Genes 2022, 10, 1688. [Google Scholar] [CrossRef]
- Diepenbroek, M.; Bayer, B.; Anslinger, K. Pushing the Boundaries: Forensic DNA Phenotyping Challenged by Single-Cell Sequencing. Genes 2021, 9, 1362. [Google Scholar] [CrossRef]
- Montesanto, A.; D’aquila, P.; Lagani, V.; Paparazzo, E.; Geracitano, S.; Formentini, L.; Giacconi, R.; Cardelli, M.; Provinciali, M.; Bellizzi, D.; et al. A New Robust Epigenetic Model for Forensic Age Prediction. J. Forensic Sci. 2020, 65, 1424–1431. [Google Scholar] [CrossRef] [PubMed]
- Onofri, M.; Delicati, A.; Marcante, B.; Carlini, L.; Alessandrini, F.; Tozzo, P.; Carnevali, E. Forensic Age Estimation through a DNA Methylation-Based Age Prediction Model in the Italian Population: A Pilot Study. Int. J. Mol. Sci. 2023, 6, 5381. [Google Scholar] [CrossRef] [PubMed]
- Alshehhi, A.; Almarzooqi, A.; Alhammadi, K.; Werghi, N.; Tay, G.K.; Alsafar, H. Advancement in Human Face Prediction Using DNA. Genes 2023, 1, 136. [Google Scholar] [CrossRef]
- Kaiser, M. Forensic DNA Phenotyping: Predicting human appearance from crime scene material for investigative purposes. Forensic Sci. Int. Genet. 2015, 18, 33–48. [Google Scholar] [CrossRef]
- Maroñas, O.; Söchtig, J.; Ruiz, Y.; Phillips, C.; Carracedo, A.; Lareu, M.V. The genetics of skin, hair, and eye color variation and its relevance to forensic pigmentation predictive tests. Forensic Sci. Rev. 2015, 27, 13–40. [Google Scholar]
- Dorgaleleh, S.; Naghipoor, K.; Barahouie, A.; Dastaviz, F.; Oladnabi, M. Molecular and biochemical mechanisms of human iris color: A comprehensive review. J. Cell. Physiol. 2020, 235, 8972–8982. [Google Scholar] [CrossRef]
- D’Mello, S.A.N.; Finlay, G.J.; Baguley, B.C.; Askarian-Amiri, M.E. Signaling Pathways in Melanogenesis. Int. J. Mol. Sci. 2016, 15, 1144. [Google Scholar] [CrossRef]
- Duffy, D.L.; Montgomery, G.W.; Chen, W.; Zhao, Z.Z.; Le, L.; James, M.R.; Hayward, N.K.; Martin, N.G.; Sturm, R.A. A three-single-nucleotide polymorphism haplotype in intron 1 of OCA2 explains most human eye-color variation. Am. J. Hum. Genet 2007, 80, 241–252. [Google Scholar] [CrossRef] [PubMed]
- Kayser, M.; Liu, F.; Janssens, A.C.J.; Rivadeneira, F.; Lao, O.; van Duijn, K.; Vermeulen, M.; Arp, P.; Jhamai, M.M.; van IJcken, W.F.; et al. Three genome-wide association studies and a linkage analysis identify HERC2 as a human iris color gene. Am. J. Hum. Genet. 2008, 82, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Sturm, R.A.; Duffy, D.L.; Zhao, Z.Z.; Leite, F.P.; Stark, M.S.; Hayward, N.K.; Martin, N.G.; Montgomery, G.W. A Single SNP in an Evolutionary Conserved Region within Intron 86 of the HERC2 Gene Determines Human Blue-Brown Eye Color. Am. J. Hum. Genet. 2008, 82, 424–431. [Google Scholar] [CrossRef]
- Sulem, P.; Gudbjartsson, D.F.; Stacey, S.N.; Helgason, A.; Rafnar, T.; Magnusson, K.P.; Manolescu, A.; Karason, A.; Palsson, A.; Thorleifsson, G.; et al. Genetic determinants of hair, eye and skin pigmentation in Europeans. Nat. Genet. 2007, 39, 1443–1452. [Google Scholar] [CrossRef]
- Visser, M.; Kayser, M.; Palstra, R.-J. HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Res. 2012, 22, 446–455. [Google Scholar] [CrossRef] [PubMed]
- Frudakis, T.; Terravainen, T.; Thomas, M. Multilocus OCA2 genotypes specify human iris colors. Hum. Genet. 2007, 122, 311–326. [Google Scholar] [CrossRef]
- Liu, F.; van Duijn, K.; Vingerling, J.R.; Hofman, A.; Uitterlinden, A.G.; Janssens, A.C.; Kayser, M. Eye color and the prediction of complex phenotypes from genotypes. Curr. Biol. 2009, 19, R192–R193. [Google Scholar] [CrossRef]
- Lee, S.-T.; Nicholls, R.D.; Jong, M.T.; Fukai, K.; Spritz, R.A. Organization and sequence of the human P gene and identification of a new family of transport proteins. Genomics 1995, 20, 354–363. [Google Scholar] [CrossRef]
- Toyofuku, K.; Valencia, J.C.; Kushimoto, T.; Costin, G.-E.; Virador, V.M.; Vieira, W.D.; Ferrans, V.J.; Hearing, V.J. The Etiology of Oculocutaneous Albinism (OCA) Type II: The Pink Protein Modulates the Processing and Transport of Tyrosinase. Pigment. Cell Res. 2002, 15, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Sturm, R.A.; Larsson, M. Genetics of human iris colour and patterns. Pigment. Cell Melanoma Res. 2009, 22, 544–562. [Google Scholar] [CrossRef]
- Sturm, R.A.; Frudakis, T.N. Eye colour: Portals into pigmentation genes and ancestry. Trends Genet. 2004, 20, 327–332. [Google Scholar] [CrossRef]
- Walsh, S.; Lindenbergh, A.; Zuniga, S.B.; Sijen, T.; de Knijff, P.; Kayser, M.; Ballantyne, K.N. Developmental validation of the IrisPlex system: Determination of blue and brown iris colour for forensic intelligence. Forensic Sci. Int. Genet. 2011, 5, 464–471. [Google Scholar] [CrossRef]
- Allwood, J.S.; Harbison, S. SNP model development for the prediction of eye colour in New Zealand. Forensic Sci. Int. Genet. 2013, 7, 444–452. [Google Scholar] [CrossRef]
- Ruiz, Y.; Phillips, C.; Gomez-Tato, A.; Alvarez-Dios, J.; de Cal, M.C.; Cruz, R.; Maroñas, O.; Söchtig, J.; Fondevila, M.; Rodriguez-Cid, M.; et al. Further development of forensic eye color predictive tests. Forensic Sci. Int. Genet. 2013, 7, 28–40. [Google Scholar] [CrossRef]
- Hart, K.L.; Kimura, S.L.; Mushailov, V.; Budimlija, Z.M.; Prinz, M.; Wurmbach, E. Improved eye- and skin-color prediction based on 8 SNPs. Croat. Med. J. 2013, 54, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Andersen, J.D.; Johansen, P.; Harder, S.; Christoffersen, S.R.; Delgado, M.C.; Henriksen, S.T.; Nielsen, M.M.; Sørensen, E.; Ullum, H.; Hansen, T.; et al. Genetic analyses of the human eye colours using a novel objective method for eye colour classification. Forensic Sci. Int. Genet. 2013, 7, 508–515. [Google Scholar] [CrossRef] [PubMed]
- Pośpiech, E.; Wojas-Pelc, A.; Walsh, S.; Liu, F.; Maeda, H.; Ishikawa, T.; Skowron, M.; Kayser, M.; Branicki, W. The common occurrence of epistasis in the determination of human pigmentation and its impact on DNA-based pigmentation phenotype prediction. Forensic Sci. Int. Genet. 2014, 11, 64–72. [Google Scholar] [CrossRef]
- Galligan, J.T.; Martinez-Noël, G.; Arndt, V.; Hayes, S.; Chittenden, T.W.; Harper, J.W.; Howley, P.M. Proteomic Analysis and Identification of Cellular Interactors of the Giant Ubiquitin Ligase HERC2. J. Proteome Res. 2015, 6, 953–966. [Google Scholar] [CrossRef]
- Eiberg, H.; Troelsen, J.; Nielsen, M.; Mikkelsen, A.; Mengel-From, J.; Kjaer, K.W.; Hansen, L. Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum. Genet. 2008, 123, 177–187. [Google Scholar] [CrossRef]
- Giebel, L.B.; Strunk, K.M.; Spritz, R.A. Organization and nucleotide sequences of the human tyrosinase gene and a truncated tyrosinase-related segment. Genomics 1991, 9, 435–445. [Google Scholar] [CrossRef]
- Wang, N.; Hebert, D.N. Tyrosinase maturation through the mammalian secretory pathway: Bringing color to life. Pigment. Cell Res. 2006, 19, 3–18. [Google Scholar] [CrossRef] [PubMed]
- Candille, S.I.; Absher, D.M.; Beleza, S.; Bauchet, M.; McEvoy, B.; Garrison, N.A.; Li, J.Z.; Myers, R.M.; Barsh, G.S.; Tang, H.; et al. Genome-Wide Association Studies of Quantitatively Measured Skin, Hair, and Eye Pigmentation in Four European Populations. PLoS ONE 2012, 7, e48294. [Google Scholar] [CrossRef] [PubMed]
- Alaluf, S.; Barrett, K.; Blount, M.; Carter, N. Ethnic Variation in Tyrosinase and TYRP1 Expression in Photoexposed and Photoprotected Human Skin. Pigment. Cell Res. 2003, 16, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Meyer, O.S.; Lunn, M.M.B.; Garcia, S.L.; Kjærbye, A.B.; Morling, N.; Børsting, C.; Andersen, J.D. Association between brown eye colour in rs12913832:GG individuals and SNPs in TYR, TYRP1, and SLC24A4. PLoS ONE 2020, 11, e0239131. [Google Scholar] [CrossRef]
- Harada, M.; Li, Y.F.; El-Gamil, M.; Rosenberg, S.A.; Robbins, P.F. Use of an in vitro immunoselected tumor line to identify shared melanoma antigens recognized by HLA-A*0201-restricted T cells. Cancer Res 2001, 61, 1089–1094. [Google Scholar]
- Graf, J.; Hodgson, R.; van Daal, A. Single nucleotide polymorphisms in theMATP gene are associated with normal human pigmentation variation. Hum. Mutat. 2005, 25, 278–284. [Google Scholar] [CrossRef]
- Newton, J.; Cohen-Barak, O.; Hagiwara, N.; Gardner, J.M.; Davisson, M.T.; King, R.A.; Brilliant, M.H. Mutations in the Human Orthologue of the Mouse underwhite Gene (uw) Underlie a New Form of Oculocutaneous Albinism, OCA4. Am. J. Hum. Genet. 2001, 69, 981–988. [Google Scholar] [CrossRef]
- Du, J.; Fisher, D.E. Identification of Aim-1 as the underwhiteMouse Mutant and Its Transcriptional Regulation by MITF. J. Biol. Chem. 2002, 277, 402–406. [Google Scholar] [CrossRef]
- Nakayama, K.; Fukamachi, S.; Kimura, H.; Koda, Y.; Soemantri, A.; Ishida, T. Distinctive distribution of AIM1 polymorphism among major human populations with different skin color. J. Hum. Genet. 2002, 47, 92–94. [Google Scholar] [CrossRef] [PubMed]
- Fernandez, L.; Milne, R.; Pita, G.; Avilés, J.; Lázaro, P.; Benítez, J.; Ribas, G. SLC45A2: A novel malignant melanoma-associated gene. Hum. Mutat. 2008, 29, 1161–1167. [Google Scholar] [CrossRef] [PubMed]
- Pośpiech, E.; Draus-Barini, J.; Kupiec, T.; Wojas-Pelc, A.; Branicki, W. Gene–gene interactions contribute to eye colour variation in humans. J. Hum. Genet. 2011, 56, 447–455. [Google Scholar] [CrossRef]
- Lamason, R.L.; Mohideen, M.A.P.; Mest, J.R.; Wong, A.C.; Norton, H.L.; Aros, M.C.; Jurynec, M.J.; Mao, X.; Humphreville, V.R.; Humbert, J.E.; et al. SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 2005, 16, 1782–1786. [Google Scholar] [CrossRef]
- Soejima, M.; Koda, Y. Population differences of two coding SNPs in pigmentation-related genes SLC24A5 and SLC45A2. Int. J. Leg. Med. 2007, 121, 36–39. [Google Scholar] [CrossRef] [PubMed]
- Cook, A.L.; Chen, W.; Thurber, A.E.; Smit, D.J.; Smith, A.G.; Bladen, T.G.; Brown, D.L.; Duffy, D.L.; Pastorino, L.; Bianchi-Scarra, G.; et al. Analysis of cultured human melanocytes based on polymorphisms within the SLC45A2/MATP, SLC24A5/NCKX5, and OCA2/P loci. J. Investig. Dermatol. 2009, 129, 392–405. [Google Scholar] [CrossRef] [PubMed]
- Canfield, V.A.; Berg, A.; Peckins, S.; Wentzel, S.M.; Ang, K.C.; Oppenheimer, S.; Cheng, K.C. Molecular Phylogeography of a Human Autosomal Skin Color Locus Under Natural Selection. G3 2013, 3, 2059–2067. [Google Scholar] [CrossRef][Green Version]
- Wilson, B.D.; Ollmann, M.M.; Kang, L.; Stoffel, M.; Bell, G.I.; Barsh, G.S. Structure and function of ASP, the human homolog of the mouse agouti gene. Hum. Mol. Genet. 1995, 4, 223–230. [Google Scholar] [CrossRef]
- Voisey, J.; Kelly, G.; Van Daal, A. Agouti Signal Protein Regulation in Human Melanoma Cells. Pigment. Cell Res. 2003, 16, 65–71. [Google Scholar] [CrossRef]
- Zeigler-Johnson, C.; Panossian, S.; Gueye, S.M.; Jalloh, M.; Ofori-Adjei, D.; Kanetsky, P.A. Population Differences in the Frequency of the Agouti Signaling Protein g.8818A>G Polymorphism. Pigment. Cell Res. 2004, 17, 185–187. [Google Scholar] [CrossRef]
- Kanetsky, P.A.; Swoyer, J.; Panossian, S.; Holmes, R.; Guerry, D.; Rebbeck, T.R. A Polymorphism in the Agouti Signaling Protein Gene Is Associated with Human Pigmentation. Am. J. Hum. Genet. 2002, 70, 770–775. [Google Scholar] [CrossRef]
- Harding, R.M.; Healy, E.; Ray, A.J.; Ellis, N.S.; Flanagan, N.; Todd, C.; Dixon, C.; Sajantila, A.; Jackson, I.J.; Birch-Machin, M.A.; et al. Evidence for variable selective pressures at MC1R. Am. J. Hum. Genet. 2000, 66, 1351–1361. [Google Scholar] [CrossRef]
- Palmer, J.S.; Duffy, D.L.; Box, N.F.; Aitken, J.F.; O’Gorman, L.E.; Green, A.C.; Hayward, N.K.; Martin, N.G.; Sturm, R.A. Melanocortin-1 receptor polymorphisms and risk of melanoma: Is the association explained solely by pigmentation phenotype? Am. J. Hum. Genet. 2000, 66, 176–186. [Google Scholar] [CrossRef] [PubMed]
- Valverde, P.; Healy, E.; Jackson, I.; Rees, J.L.; Thody, A.J. Variants of the melanocyte–stimulating hormone receptor gene are associated with red hair and fair skin in humans. Nat. Genet. 1995, 11, 328–330. [Google Scholar] [CrossRef] [PubMed]
- Frändberg, P.-A.; Doufexis, M.; Kapas, S.; Chhajlani, V. Human Pigmentation Phenotype: A Point Mutation Generates Nonfunctional MSH Receptor. Biochem. Biophys. Res. Commun. 1998, 245, 490–492. [Google Scholar] [CrossRef] [PubMed]
- Cone, R.D.; Lu, D.; Koppula, S.; Våge, D.I.; Klungland, H.; Boston, B.; Chen, W.; Orth, D.N.; Pouton, C.; Kesterson, R.A. The melanocortin receptors: Agonists, antagonists, and the hormonal control of pigmentation. Recent Prog. Horm. Res. 1996, 51, 287–317. [Google Scholar] [PubMed]
- Makova, K.; Norton, H. Worldwide polymorphism at the MC1R locus and normal pigmentation variation in humans. Peptides 2005, 26, 1901–1908. [Google Scholar] [CrossRef] [PubMed]
- Paun, A.; Pitha, P.M. The IRF family, revisited. Biochimie 2007, 89, 744–753. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Kraft, P.; Nan, H.; Guo, Q.; Chen, C.; Qureshi, A.; Hankinson, S.E.; Hu, F.B.; Duffy, D.L.; Zhao, Z.Z.; et al. A Genome-Wide Association Study Identifies Novel Alleles Associated with Hair Color and Skin Pigmentation. PLoS Genet. 2008, 4, e1000074. [Google Scholar] [CrossRef]
- Praetorius, C.; Grill, C.; Stacey, S.N.; Metcalf, A.M.; Gorkin, D.U.; Robinson, K.C.; Van Otterloo, E.; Kim, R.S.; Bergsteinsdottir, K.; Ogmundsdottir, M.H.; et al. A Polymorphism in IRF4 Affects Human Pigmentation through a Tyrosinase-Dependent MITF/TFAP2A Pathway. Cell 2013, 21, 1022–1033. [Google Scholar] [CrossRef]
- Visser, M.; Palstra, R.-J.; Kayser, M. Allele-specific transcriptional regulation of IRF4 in melanocytes is mediated by chromatin looping of the intronic rs12203592 enhancer to the IRF4 promoter. Hum. Mol. Genet. 2015, 1, 2649–2661. [Google Scholar] [CrossRef]
- Valenzuela, R.K.; Henderson, M.S.; Walsh, M.H.; Garrison, N.A.; Kelch, J.T.; Cohen-Barak, O.; Erickson, D.T.; Meaney, F.J.; Walsh, J.B.; Cheng, K.C.; et al. Predicting Phenotype from Genotype: Normal Pigmentation. J. Forensic Sci. 2010, 1, 315–322. [Google Scholar] [CrossRef]
- Mengel-From, J.; Børsting, C.; Sanchez, J.J.; Eiberg, H.; Morling, N. Human eye colour and HERC2, OCA2 and MATP. Forensic Sci. Int. Genet. 2010, 4, 323–328. [Google Scholar] [CrossRef]
- Walsh, S.; Liu, F.; Ballantyne, K.N.; van Oven, M.; Lao, O.; Kayser, M. IrisPlex: A sensitive DNA tool for accurate prediction of blue and brown eye colour in the absence of ancestry information. Forensic Sci. Int. Genet. 2011, 5, 170–180. [Google Scholar] [CrossRef]
- Walsh, S.; Wollstein, A.; Liu, F.; Chakravarthy, U.; Rahu, M.; Seland, J.H.; Soubrane, G.; Tomazzoli, L.; Topouzis, F.; Vingerling, J.R.; et al. DNA-based eye colour prediction across Europe with the IrisPlex system. Forensic Sci. Int. Genet. 2012, 6, 330–340. [Google Scholar] [CrossRef]
- Chaitanya, L.; Walsh, S.; Andersen, J.D.; Ansell, R.; Ballantyne, K.; Ballard, D.; Banemann, R.; Bauer, C.M.; Bento, A.M.; Brisighelli, F.; et al. Collaborative EDNAP exercise on the IrisPlex system for DNA-based prediction of human eye colour. Forensic Sci. Int. Genet. 2014, 11, 241–251. [Google Scholar] [CrossRef] [PubMed]
- Keating, B.; Bansal, A.T.; Walsh, S.; Millman, J.; Newman, J.; Kidd, K.; Budowle, B.; Eisenberg, A.; Donfack, J. First all-in-one diagnostic tool for DNA intelligence: Genome-wide inference of biogeographic ancestry, appearance, relatedness, and sex with the Identitas v1 Forensic Chip. Int. J. Leg. Med. 2013, 127, 559–572. [Google Scholar] [CrossRef] [PubMed]
- Walsh, S.; Chaitanya, L.; Clarisse, L.; Wirken, L.; Draus-Barini, J.; Kovatsi, L.; Maeda, H.; Ishikawa, T.; Sijen, T.; de Knijff, P.; et al. Developmental validation of the HIrisPlex system: DNA-based eye and hair colour prediction for forensic and anthropological usage. Forensic Sci. Int. Genet. 2014, 9, 150–161. [Google Scholar] [CrossRef]
- Sari, O.I.; Simsek, S.Z.; Filoglu, G.; Bulbul, O. Predicting Eye and Hair Color in a Turkish Population Using the HIrisPlex System. Genes 2022, 11, 2094. [Google Scholar] [CrossRef] [PubMed]
- Freire-Aradas, A.; Ruiz, Y.; Phillips, C.; Maroñas, O.; Söchtig, J.; Tato, A.G.; Dios, J.; de Cal, M.C.; Silbiger, V.; Luchessi, A.; et al. Exploring iris colour prediction and ancestry inference in admixed populations of South America. Forensic Sci. Int. Genet. 2014, 13, 3–9. [Google Scholar] [CrossRef]
- Xavier, C.; de la Puente, M.; Mosquera-Miguel, A.; Freire-Aradas, A.; Kalamara, V.; Ralf, A.; Revoir, A.; Gross, T.; Schneider, P.; Ames, C.; et al. Development and inter-laboratory evaluation of the VISAGE Enhanced Tool for Appearance and Ancestry inference from DNA. Forensic Sci. Int. Genet. 2022, 61, 102779. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ramírez, J.; de la Puente, M.; Xavier, C.; Ambroa-Conde, A.; Álvarez-Dios, J.; Freire-Aradas, A.; Mosquera-Miguel, A.; Ralf, A.; Amory, C.; Katsara, M.; et al. Development and evaluations of the ancestry informative markers of the VISAGE Enhanced Tool for Appearance and Ancestry. Forensic Sci. Int. Genet. 2023, 64, 102853. [Google Scholar] [CrossRef] [PubMed]
- Palencia-Madrid, L.; Xavier, C.; De La Puente, M.; Hohoff, C.; Phillips, C.; Kayser, M.; Parson, W. Evaluation of the VISAGE Basic Tool for Appearance and Ancestry Prediction Using PowerSeq Chemistry on the MiSeq FGx System. Genes 2020, 6, 708. [Google Scholar] [CrossRef] [PubMed]
- Xavier, C.; de la Puente, M.; Mosquera-Miguel, A.; Freire-Aradas, A.; Kalamara, V.; Vidaki, A.; Gross, T.E.; Revoir, A.; Pośpiech, E.; Kartasińska, E.; et al. Development and validation of the VISAGE AmpliSeq basic tool to predict appearance and ancestry from DNA. Forensic Sci. Int. Genet. 2020, 48, 102336. [Google Scholar] [CrossRef] [PubMed]
- Zieger, M. Forensic DNA phenotyping in Europe: How far may it go? J. Law. Biosci. 2022, 2, lsac024. [Google Scholar] [CrossRef]
- Paparazzo, E.; Gozalishvili, A.; Lagani, V.; Geracitano, S.; Bauleo, A.; Falcone, E.; Passarino, G.; Montesanto, A. A new approach to broaden the range of eye colour identifiable by IrisPlex in DNA phenotyping. Sci. Rep. 2022, 12, 12803. [Google Scholar] [CrossRef]
- Gross, T.E.; Fleckhaus, J.; Schneider, P.M. Progress in the implementation of massively parallel sequencing for forensic genetics: Results of a European-wide survey among professional users. Int. J. Leg. Med. 2021, 135, 1425–1432. [Google Scholar] [CrossRef]
- Kukla-Bartoszek, M.; Teisseyre, P.; Pośpiech, E.; Karłowska-Pik, J.; Zieliński, P.; Woźniak, A.; Boroń, M.; Dąbrowski, M.; Zubańska, M.; Jarosz, A.; et al. Searching for improvements in predicting human eye colour from DNA. Int. J. Leg. Med. 2021, 135, 2175–2187. [Google Scholar] [CrossRef]
- Ragazzo, M.; Puleri, G.; Errichiello, V.; Manzo, L.; Luzzi, L.; Potenza, S.; Strafella, C.; Peconi, C.; Nicastro, F.; Caputo, V.; et al. Evaluation of OpenArray™ as A Genotyping Method for Forensic DNA Phenotyping and Human Identification. Genes 2021, 12, 221. [Google Scholar] [CrossRef]
- Pajnič, I.Z.; Leskovar, T.; Črešnar, M. Eye and hair color prediction of an early medieval adult and subadult skeleton using massive parallel sequencing technology. Int. J. Leg. Med. 2023, in press. [Google Scholar] [CrossRef]
- Pośpiech, E.; Teisseyre, P.; Mielniczuk, J.; Branicki, W. Predicting Physical Appearance from DNA Data—Towards Genomic Solutions. Genes 2022, 1, 121. [Google Scholar] [CrossRef] [PubMed]
- Salvo, N.M.; Andersen, J.D.; Janssen, K.; Meyer, O.L.; Berg, T.; Børsting, C.; Olsen, G.-H. Association between Variants in the OCA2-HERC2 Region and Blue Eye Colour in HERC2 rs12913832 AA and AG Individuals. Genes 2023, 3, 698. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Visser, M.; Duffy, D.L.; Hysi, P.G.; Jacobs, L.C.; Lao, O.; Zhong, K.; Walsh, S.; Chaitanya, L.; Wollstein, A.; et al. Genetics of skin color variation in Europeans: Genome-wide association studies with functional follow-up. Hum. Genet. 2015, 134, 823–835. [Google Scholar] [CrossRef]
- Davenport, G.C.; Davenport, C.B. Heredity of eye-color in man. Science 1907, 1, 589–592. [Google Scholar] [CrossRef]
- d’Ischia, M.; Wakamatsu, K.; Cicoira, F.; Di Mauro, E.; Garcia-Borron, J.C.; Commo, S.; Galván, I.; Ghanem, G.; Kenzo, K.; Meredith, P.; et al. Melanins and melanogenesis: From pigment cells to human health and technological applications. Pigment. Cell Melanoma Res. 2015, 28, 520–544. [Google Scholar] [CrossRef]
- Wakamatsu, K.; Ito, S. Recent Advances in Characterization of Melanin Pigments in Biological Samples. Int. J. Mol. Sci. 2023, 24, 8305. [Google Scholar] [CrossRef] [PubMed]
| Gene | SNPs (No.) | SNP Location | ||||
|---|---|---|---|---|---|---|
| 5′-UTR | 3′-UTR | Intron | N.Syn. | Syn. | ||
| ASIP | 3 | 1 | 1 | 1 | ||
| HERC2 | 10 | 9 | 1 | |||
| IRF4 | 1 | 1 | ||||
| MC1R | 5 | 5 | ||||
| OCA2 | 42 | 36 | 5 | 1 | ||
| SLC24A4 | 1 | 1 | ||||
| SLC24A5 | 1 | 1 | ||||
| SLC45A2 | 2 | 2 | ||||
| TYR | 2 | 1 | 1 | |||
| TYRP1 | 6 | 3 | 1 | 1 | 1 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Brancato, D.; Coniglio, E.; Bruno, F.; Agostini, V.; Saccone, S.; Federico, C. Forensic DNA Phenotyping: Genes and Genetic Variants for Eye Color Prediction. Genes 2023, 14, 1604. https://doi.org/10.3390/genes14081604
Brancato D, Coniglio E, Bruno F, Agostini V, Saccone S, Federico C. Forensic DNA Phenotyping: Genes and Genetic Variants for Eye Color Prediction. Genes. 2023; 14(8):1604. https://doi.org/10.3390/genes14081604
Chicago/Turabian StyleBrancato, Desiree, Elvira Coniglio, Francesca Bruno, Vincenzo Agostini, Salvatore Saccone, and Concetta Federico. 2023. "Forensic DNA Phenotyping: Genes and Genetic Variants for Eye Color Prediction" Genes 14, no. 8: 1604. https://doi.org/10.3390/genes14081604
APA StyleBrancato, D., Coniglio, E., Bruno, F., Agostini, V., Saccone, S., & Federico, C. (2023). Forensic DNA Phenotyping: Genes and Genetic Variants for Eye Color Prediction. Genes, 14(8), 1604. https://doi.org/10.3390/genes14081604









