Transcriptome Signatures of Atlantic Salmon—Resistant Phenotypes against Sea Lice Infestation Are Associated with Tissue Repair
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Trial
2.2. High-Throughput Transcriptome Sequencing
2.3. RNA-Seq Data Analysis
2.4. Whole-Genome Transcript Expression Analysis
2.5. LncRNA Identification and Genome Localization
2.6. Functional Annotation and SNP Identification
3. Results
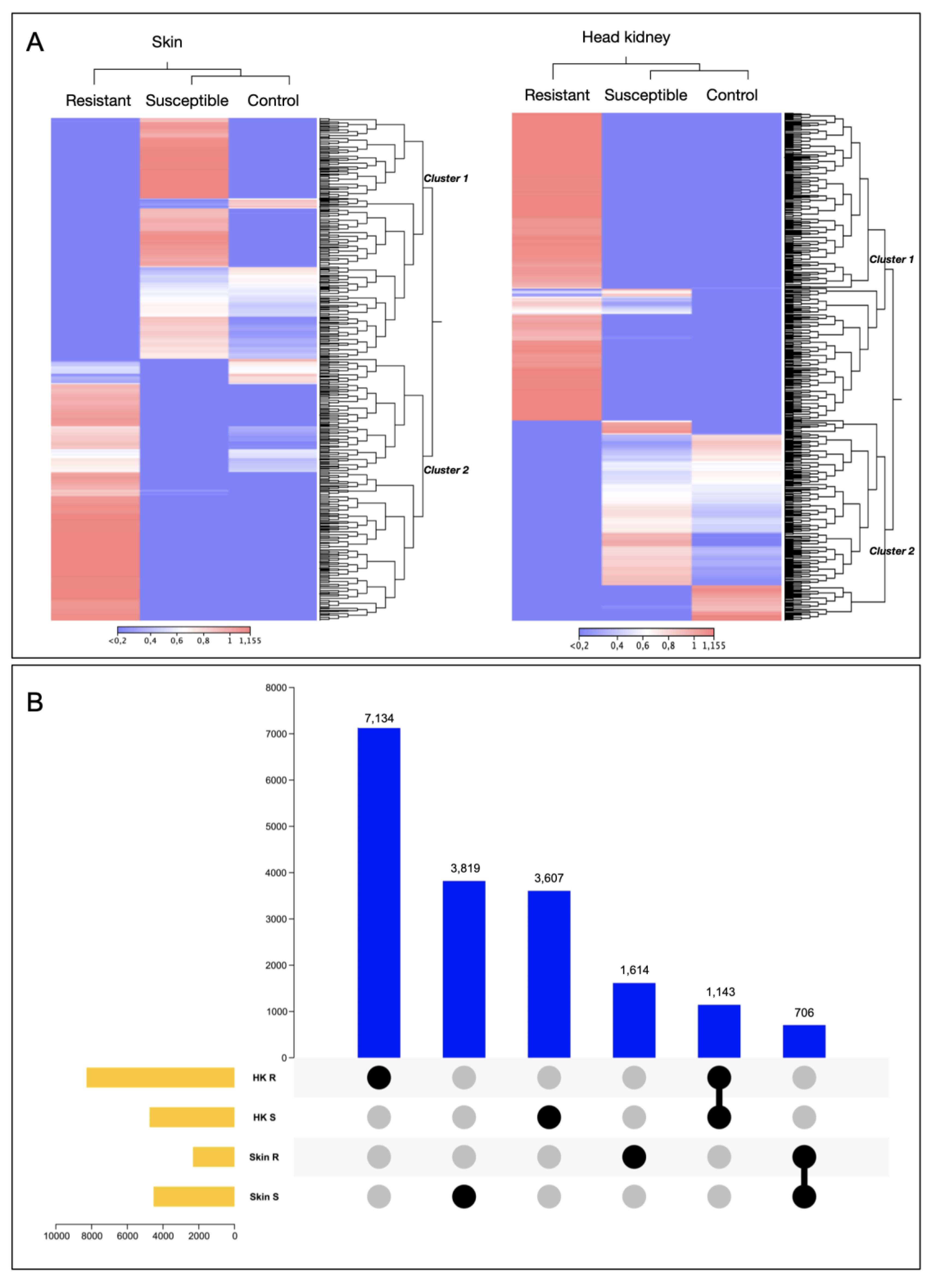
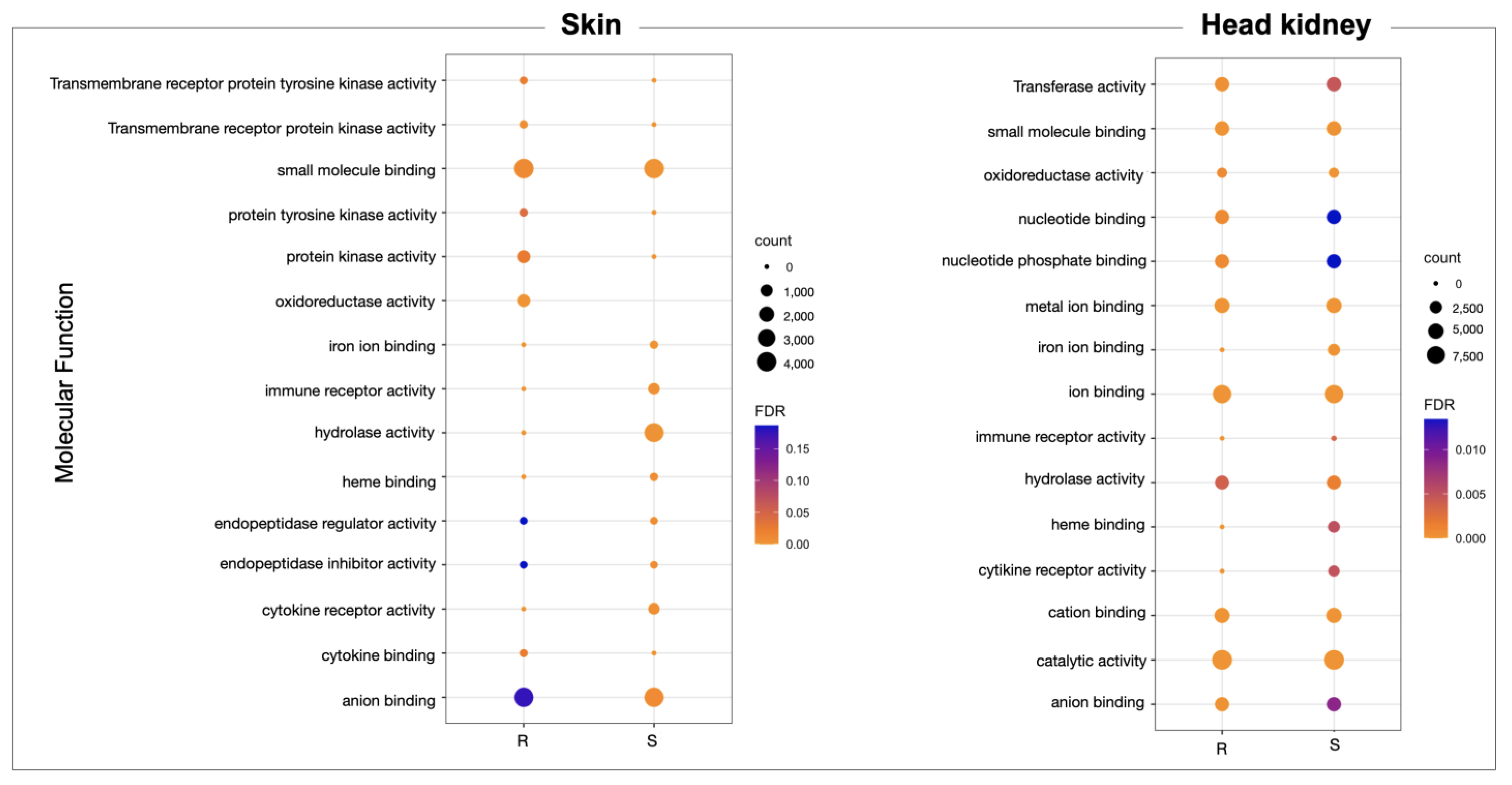
3.1. Transcriptomic Profile of Skin and Head Kidney Tissue of Atlantic Salmon R/S Families
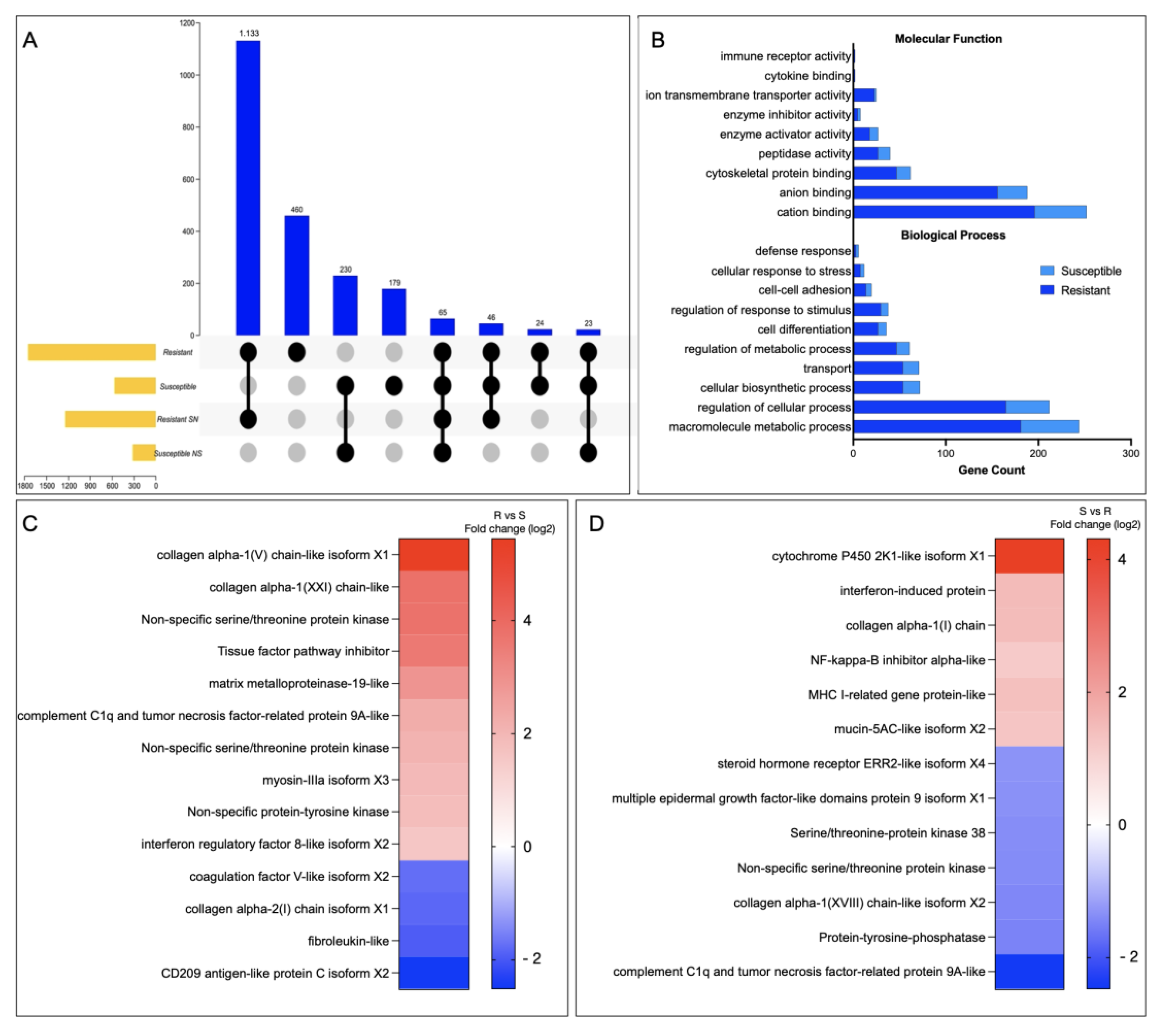
3.2. Whole-Genome Transcriptome Analysis of R/S Atlantic Salmon Families
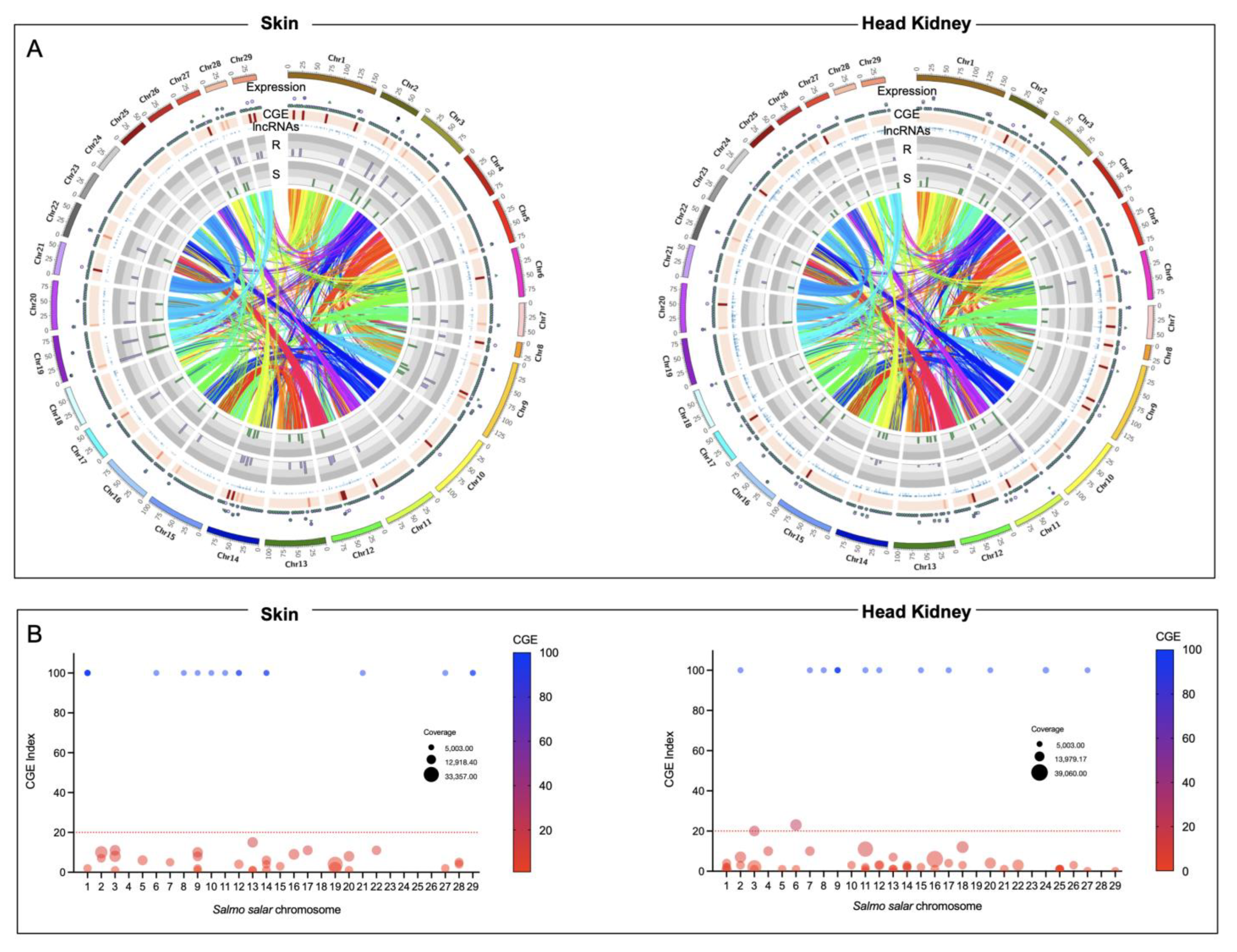
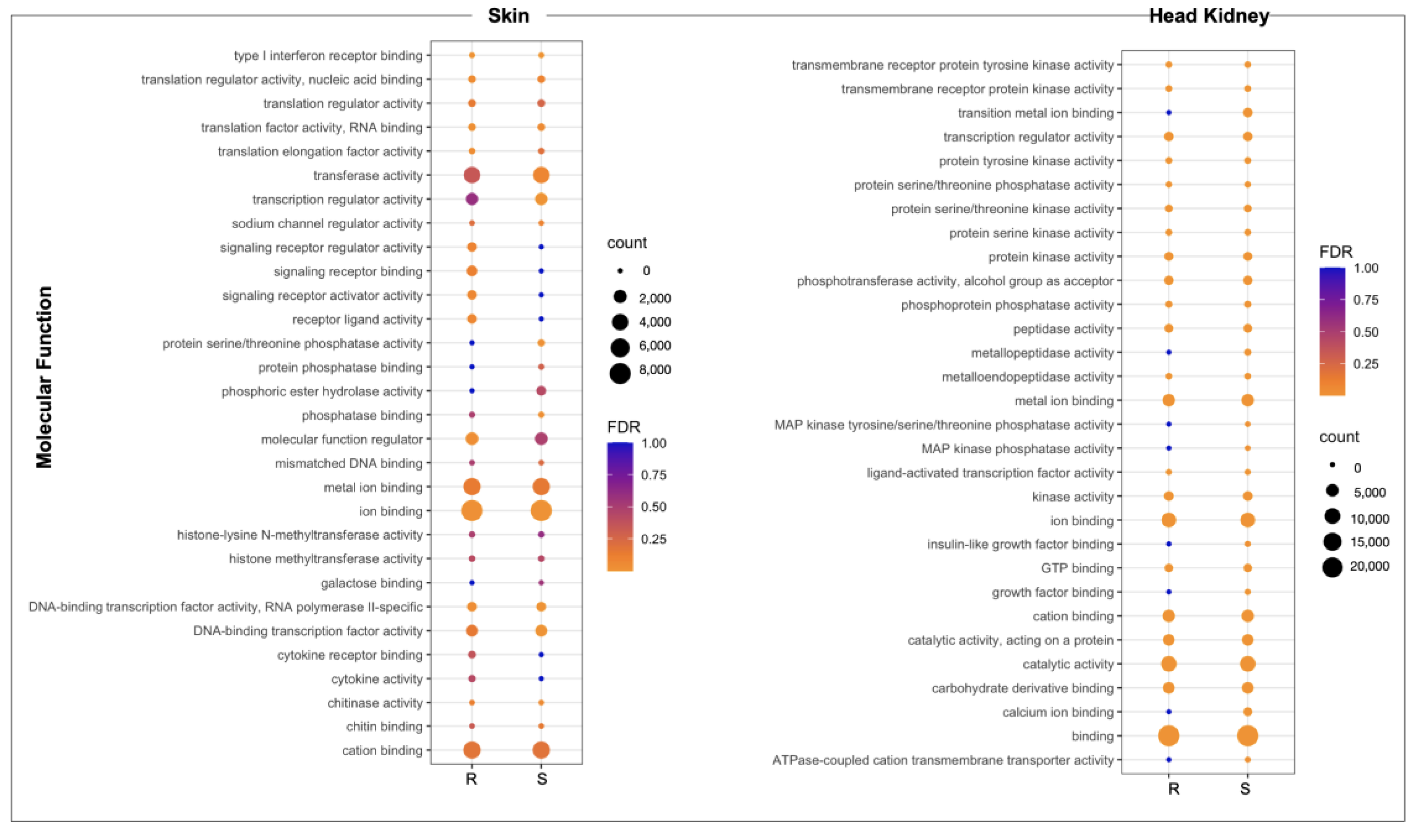
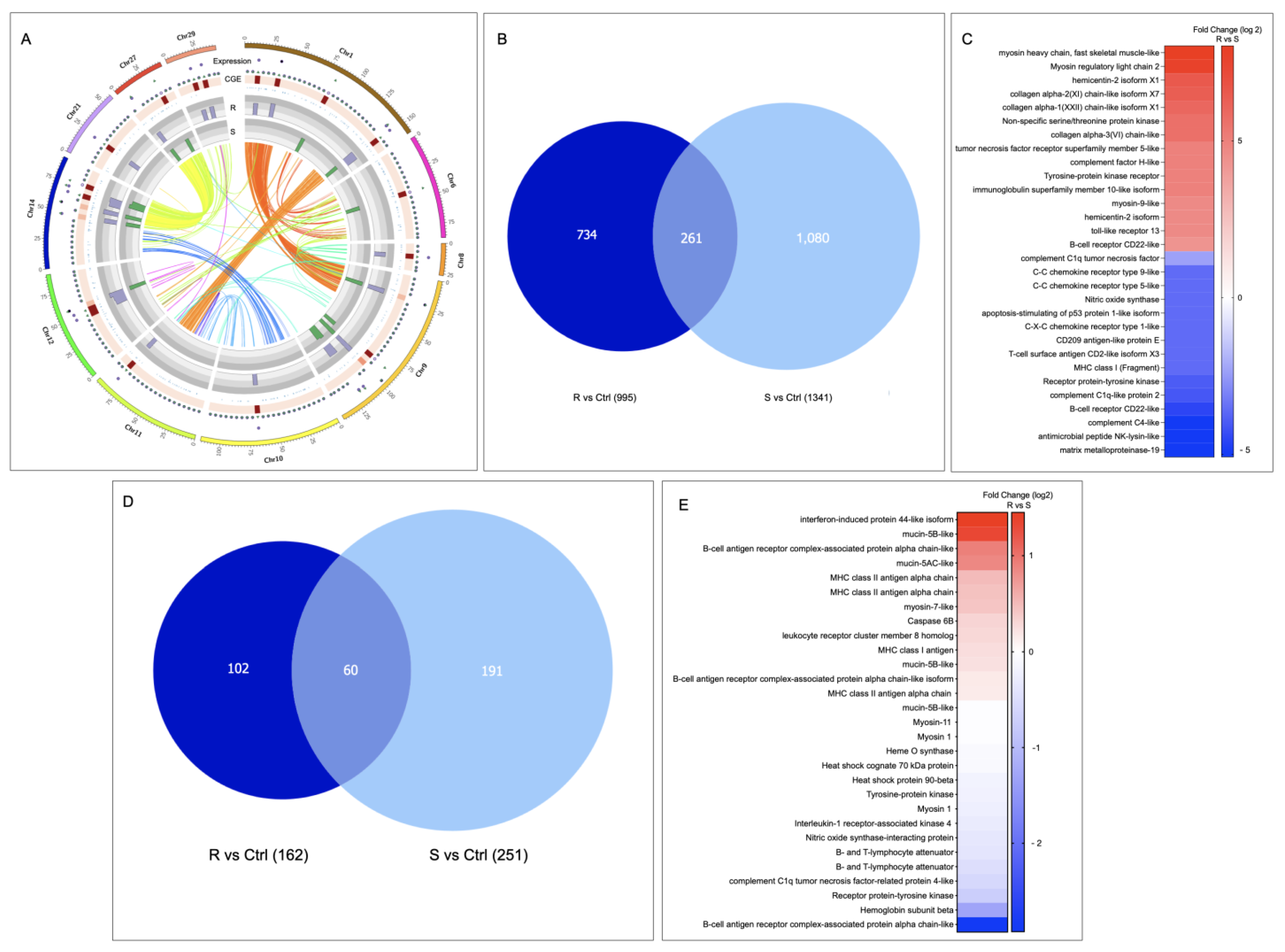
3.3. Looking for R/S Transcriptome Differences in CGE Areas of Atlantic Salmon Skin
3.4. LncRNAs Identification in R/S Atlantic Salmon Skin
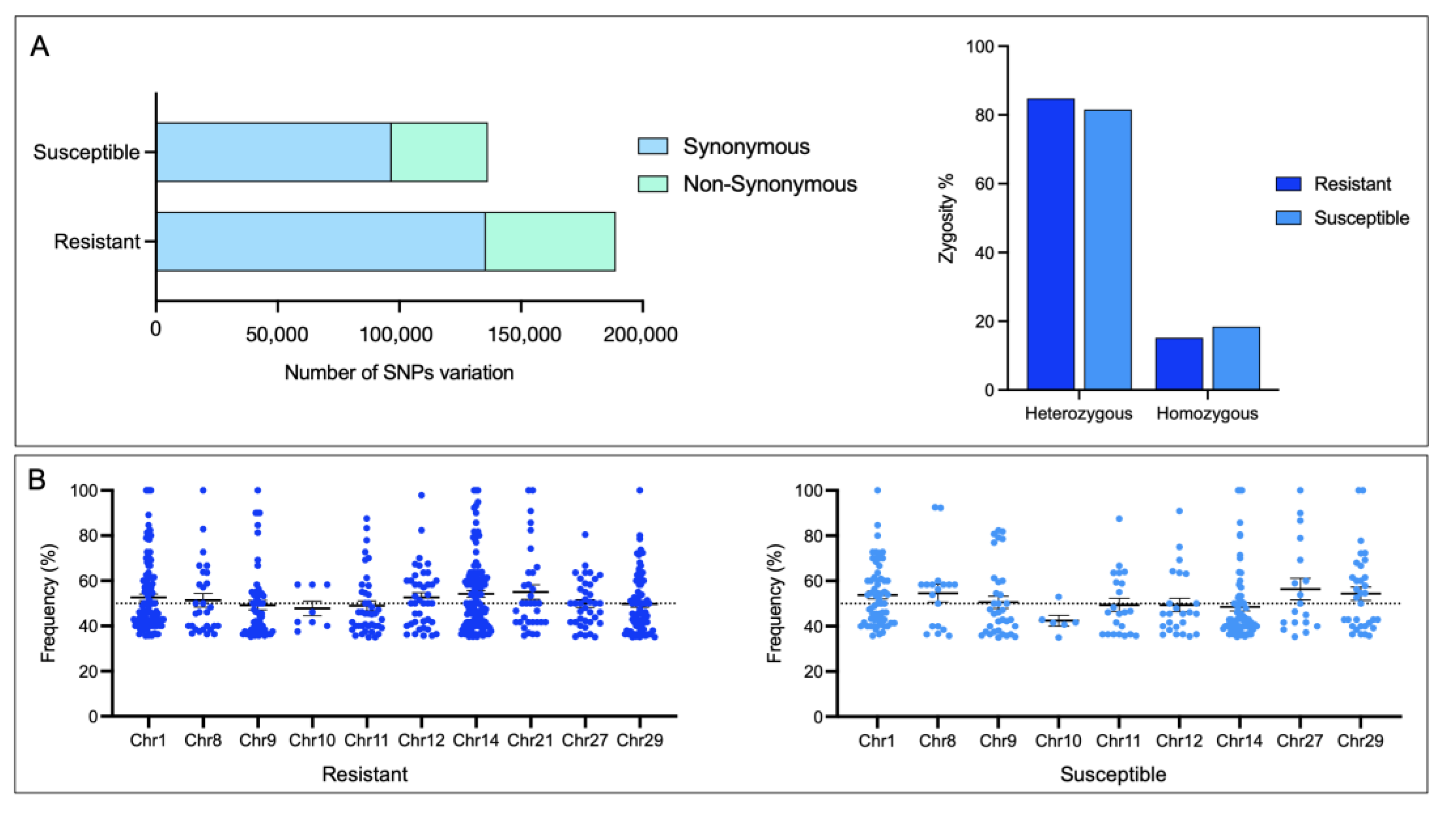
3.5. SNPs Variation in Skin CGE Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gallardo-Escárate, C.; Arriagada, G.; Carrera, C.; Gonçalves, A.T.; Nuñez-Acuña, G.; Valenzuela-Miranda, D.; Valenzuela-Muñoz, V. The race between host and sea lice in the Chilean salmon farming: A genomic approach. Rev. Aquac. 2019, 11, 325–339. [Google Scholar] [CrossRef]
- Abolofia, J.; Asche, F.; Wilen, J.E. The cost of lice: Quantifying the impacts of parasitic sea lice on farmed salmon. Mar. Resour. Econ. 2017, 32, 329–349. [Google Scholar] [CrossRef]
- Aaen, S.M.; Helgesen, K.O.; Bakke, M.J.; Kaur, K.; Horsberg, T.E. Drug resistance in sea lice: A threat to salmonid aquaculture. Trends Parasitol. 2015, 31, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Meuwissen, T.; Hayes, B.; Goddard, M. Genomic selection: A paradigm shift in animal breeding. Anim. Front. 2016, 6, 6–14. [Google Scholar] [CrossRef]
- Houston, R.D. Future directions in breeding for disease resistance in aquaculture species. Rev. Bras. Zootec. 2017, 46, 545–551. [Google Scholar] [CrossRef]
- Kolstad, K.; Heuch, P.A.; Gjerde, B.; Gjedrem, T.; Salte, R. Genetic variation in resistance of Atlantic salmon (Salmo salar) to the salmon louse Lepeophtheirus salmonis. Aquaculture 2005, 247, 145–151. [Google Scholar] [CrossRef]
- Tsai, H.-Y.; Hamilton, A.; Tinch, A.E.; Guy, D.R.; Bron, J.E.; Taggart, J.B.; Gharbi, K.; Stear, M.; Matika, O.; Pong-Wong, R. Genomic prediction of host resistance to sea lice in farmed Atlantic salmon populations. Genet. Sel. Evol. 2016, 48, 47. [Google Scholar] [CrossRef]
- Lhorente, J.P.; Gallardo, J.A.; Villanueva, B.; Araya, A.M.; Torrealba, D.A.; Toledo, X.E.; Neira, R. Quantitative genetic basis for resistance to Caligus rogercresseyi sea lice in a breeding population of Atlantic salmon (Salmo salar). Aquaculture 2012, 324, 55–59. [Google Scholar] [CrossRef]
- Correa, K.; Lhorente, J.P.; Bassini, L.; López, M.E.; Di Genova, A.; Maass, A.; Davidson, W.S.; Yáñez, J.M. Genome wide association study for resistance to Caligus rogercresseyi in Atlantic salmon (Salmo salar L.) using a 50K SNP genotyping array. Aquaculture 2017, 472, 61–65. [Google Scholar] [CrossRef]
- Yáñez, J.M.; Lhorente, J.P.; Bassini, L.N.; Oyarzún, M.; Neira, R.; Newman, S. Genetic co-variation between resistance against both Caligus rogercresseyi and Piscirickettsia salmonis, and body weight in Atlantic salmon (Salmo salar). Aquaculture 2014, 433, 295–298. [Google Scholar] [CrossRef]
- Fraslin, C.; Quillet, E.; Rochat, T.; Dechamp, N.; Bernardet, J.-F.; Collet, B.; Lallias, D.; Boudinot, P. Combining Multiple Approaches and Models to Dissect the Genetic Architecture of Resistance to Infections in Fish. Front. Genet. 2020, 11, 677. [Google Scholar] [CrossRef] [PubMed]
- Glover, K.A.; Grimholt, U.; Bakke, H.; Nilsen, F.; Storset, A.; Skaala, Ø. Major histocompatibility complex (MHC) variation and susceptibility to the sea louse Lepeophtheirus salmonis in Atlantic salmon Salmo salar. Dis. Aquat. Org. 2007, 76, 57–65. [Google Scholar] [CrossRef]
- Robledo, D.; Gutiérrez, A.P.; Barría, A.; Lhorente, J.P.; Houston, R.D.; Yáñez, J.M. Discovery and Functional Annotation of Quantitative Trait Loci Affecting Resistance to Sea Lice in Atlantic Salmon. Front. Genet. 2019, 10, 56. [Google Scholar] [CrossRef]
- Cáceres, P.; Barría, A.; Christensen, K.A.; Bassini, L.N.; Correa, K.; Garcia, B.; Lhorente, J.P.; Yáñez, J.M. Genome-scale comparative analysis for host resistance against sea lice between Atlantic salmon and rainbow trout. Sci. Rep. 2021, 11, 13231. [Google Scholar] [CrossRef] [PubMed]
- Holm, H.; Santi, N.; Kjøglum, S.; Perisic, N.; Skugor, S.; Evensen, Ø. Difference in skin immune responses to infection with salmon louse (Lepeophtheirus salmonis) in Atlantic salmon (Salmo salar L.) of families selected for resistance and susceptibility. Fish Shellfish Immunol. 2015, 42, 384–394. [Google Scholar] [CrossRef] [PubMed]
- Robledo, D.; Gutiérrez, A.P.; Barría, A.; Yáñez, J.M.; Houston, R.D. Gene Expression Response to Sea Lice in Atlantic Salmon Skin: RNA Sequencing Comparison Between Resistant and Susceptible Animals. Front. Genet. 2018, 9, 287. [Google Scholar] [CrossRef] [PubMed]
- Maurano, M.T.; Humbert, R.; Rynes, E.; Thurman, R.E.; Haugen, E.; Wang, H.; Reynolds, A.P.; Sandstrom, R.; Qu, H.; Brody, J. Systematic localization of common disease-associated variation in regulatory DNA. Science 2012, 337, 1190–1195. [Google Scholar] [CrossRef]
- Valenzuela-Muñoz, V.; Valenzuela-Miranda, D.; Gallardo-Escárate, C. Comparative analysis of long non-coding RNAs in Atlantic and Coho salmon reveals divergent transcriptome responses associated with immunity and tissue repair during sea lice infestation. Dev. Comp. Immunol. 2018, 87, 36–50. [Google Scholar] [CrossRef]
- Valenzuela-Muñoz, V.; Gallardo-Escárate, C.; Benavente, B.P.; Valenzuela-Miranda, D.; Núñez-Acuña, G.; Escobar-Sepulveda, H.; Váldes, J.A. Whole-genome transcript expression profiling reveals novel insights into transposon genes and non-coding RNAs during Atlantic salmon seawater adaptation. Biology 2021, 11, 1. [Google Scholar] [CrossRef]
- Gallardo-Escárate, C.; Valenzuela-Muñoz, V.; Nuñez-Acuña, G.; Valenzuela-Miranda, D.; Tapia, F.J.; Yévenes, M.; Gajardo, G.; Toro, J.E.; Oyarzún, P.A.; Arriagada, G.; et al. Chromosome-Level Genome Assembly of the Blue Mussel Mytilus chilensis Reveals Molecular Signatures Facing the Marine Environment. Genes 2023, 14, 876. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Valenzuela-Muñoz, V.; Váldes, J.A.; Gallardo-Escárate, C. Transcriptome profiling of long non-coding RNAs during the Atlantic salmon smoltification Process. Mar. Biotechnol. 2021, 23, 308–320. [Google Scholar] [CrossRef] [PubMed]
- Raudvere, U.; Kolberg, L.; Kuzmin, I.; Arak, T.; Adler, P.; Peterson, H.; Vilo, J. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 2019, 47, W191–W198. [Google Scholar] [CrossRef]
- Jones, C.S.; Lockyer, A.E.; Verspoor, E.; Secombes, C.J.; Noble, L.R. Towards selective breeding of Atlantic salmon for sea louse resistance: Approaches to identify trait markers. Pest Manag. Sci. 2002, 58, 559–568. [Google Scholar] [CrossRef]
- Lien, S.; Koop, B.F.; Sandve, S.R.; Miller, J.R.; Kent, M.P.; Nome, T.; Hvidsten, T.R.; Leong, J.S.; Minkley, D.R.; Zimin, A. The Atlantic salmon genome provides insights into rediploidization. Nature 2016, 533, 200. [Google Scholar] [CrossRef]
- Valenzuela-Muñoz, V.; Gallardo-Escárate, C. Iron metabolism modulation in Atlantic salmon infested with the sea lice Lepeophtheirus salmonis and Caligus rogercresseyi: A matter of nutritional immunity? Fish Shellfish Immunol. 2017, 60, 97–102. [Google Scholar] [CrossRef]
- Valenzuela-Muñoz, V.; Boltaña, S.; Gallardo-Escárate, C. Uncovering iron regulation with species-specific transcriptome patterns in Atlantic and coho salmon during a Caligus rogercresseyi infestation. J. Fish Dis. 2017, 40, 1169–1184. [Google Scholar] [CrossRef]
- Braden, L.M.; Koop, B.F.; Jones, S.R. Signatures of resistance to Lepeophtheirus salmonis include a T H 2-type response at the louse-salmon interface. Dev. Comp. Immunol. 2015, 48, 178–191. [Google Scholar] [CrossRef] [PubMed]
- Firth, K.J.; Johnson, S.C.; Ross, N.W. Characterization of proteases in the skin mucus of Atlantic salmon (Salmo salar) infected with the salmon louse (Lepeophtheirus salmonis) and in whole-body louse homogenate. J. Parasitol. 2000, 86, 1199–1205. [Google Scholar] [CrossRef]
- Valenzuela-Muñoz, V.; Boltaña, S.; Gallardo-Escárate, C. Comparative immunity of Salmo salar and Oncorhynchus kisutch during infestation with the sea louse Caligus rogercresseyi: An enrichment transcriptome analysis. Fish Shellfish Immunol. 2016, 59, 276–287. [Google Scholar] [CrossRef]
- Umasuthan, N.; Xue, X.; Caballero-Solares, A.; Kumar, S.; Westcott, J.D.; Chen, Z.; Fast, M.D.; Skugor, S.; Nowak, B.F.; Taylor, R.G. Transcriptomic profiling in fins of Atlantic salmon parasitized with sea lice: Evidence for an early imbalance between chalimus-induced immunomodulation and the host’s defense response. Int. J. Mol. Sci. 2020, 21, 2417. [Google Scholar] [CrossRef]
- Lang, T.; Hansson, G.C.; Samuelsson, T. Gel-Forming mucins appeared early in metazoan evolution. Proc. Natl. Acad. Sci. USA 2007, 104, 16209–16214. [Google Scholar] [CrossRef] [PubMed]
- Perez-Sanchez, J.; Estensoro, I.; Redondo, M.J.; Calduch-Giner, J.A.; Kaushik, S.; Sitja-Bobadilla, A. Mucins as diagnostic and prognostic biomarkers in a fish-parasite model: Transcriptional and functional analysis. PLoS ONE 2013, 8, e65457. [Google Scholar] [CrossRef] [PubMed]
- Marcos-López, M.; Calduch-Giner, J.A.; Mirimin, L.; MacCarthy, E.; Rodger, H.D.; O’Connor, I.; Sitjà-Bobadilla, A.; Pérez-Sánchez, J.; Piazzon, M.C. Gene expression analysis of Atlantic salmon gills reveals mucin 5 and interleukin 4/13 as key molecules during amoebic gill disease. Sci. Rep. 2018, 8, 13689. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, G.; O’Neill, S.; Fanning, J.; McCarthy, E.; Sekiya, M. Tissue migration by parasitic helminths—An immunoevasive strategy? Trends Parasitol. 2005, 21, 273–277. [Google Scholar] [CrossRef]
- Hunt, R.; Sauna, Z.E.; Ambudkar, S.V.; Gottesman, M.M.; Kimchi-Sarfaty, C. Silent (synonymous) SNPs: Should we care about them? Single Nucleotide Polymorph. 2009, 578, 23–39. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Valenzuela-Muñoz, V.; Gallardo-Escárate, C.; Valenzuela-Miranda, D.; Nuñez-Acuña, G.; Benavente, B.P.; Alert, A.; Arevalo, M. Transcriptome Signatures of Atlantic Salmon—Resistant Phenotypes against Sea Lice Infestation Are Associated with Tissue Repair. Genes 2023, 14, 986. https://doi.org/10.3390/genes14050986
Valenzuela-Muñoz V, Gallardo-Escárate C, Valenzuela-Miranda D, Nuñez-Acuña G, Benavente BP, Alert A, Arevalo M. Transcriptome Signatures of Atlantic Salmon—Resistant Phenotypes against Sea Lice Infestation Are Associated with Tissue Repair. Genes. 2023; 14(5):986. https://doi.org/10.3390/genes14050986
Chicago/Turabian StyleValenzuela-Muñoz, Valentina, Cristian Gallardo-Escárate, Diego Valenzuela-Miranda, Gustavo Nuñez-Acuña, Bárbara P. Benavente, Alejandro Alert, and Marta Arevalo. 2023. "Transcriptome Signatures of Atlantic Salmon—Resistant Phenotypes against Sea Lice Infestation Are Associated with Tissue Repair" Genes 14, no. 5: 986. https://doi.org/10.3390/genes14050986
APA StyleValenzuela-Muñoz, V., Gallardo-Escárate, C., Valenzuela-Miranda, D., Nuñez-Acuña, G., Benavente, B. P., Alert, A., & Arevalo, M. (2023). Transcriptome Signatures of Atlantic Salmon—Resistant Phenotypes against Sea Lice Infestation Are Associated with Tissue Repair. Genes, 14(5), 986. https://doi.org/10.3390/genes14050986







