Clinical Experience with Genome-Wide Noninvasive Prenatal Screening in a Large Cohort of Twin Pregnancies
Abstract
1. Introduction
2. Materials and Methods
3. Results
3.1. Details of Study Cohort
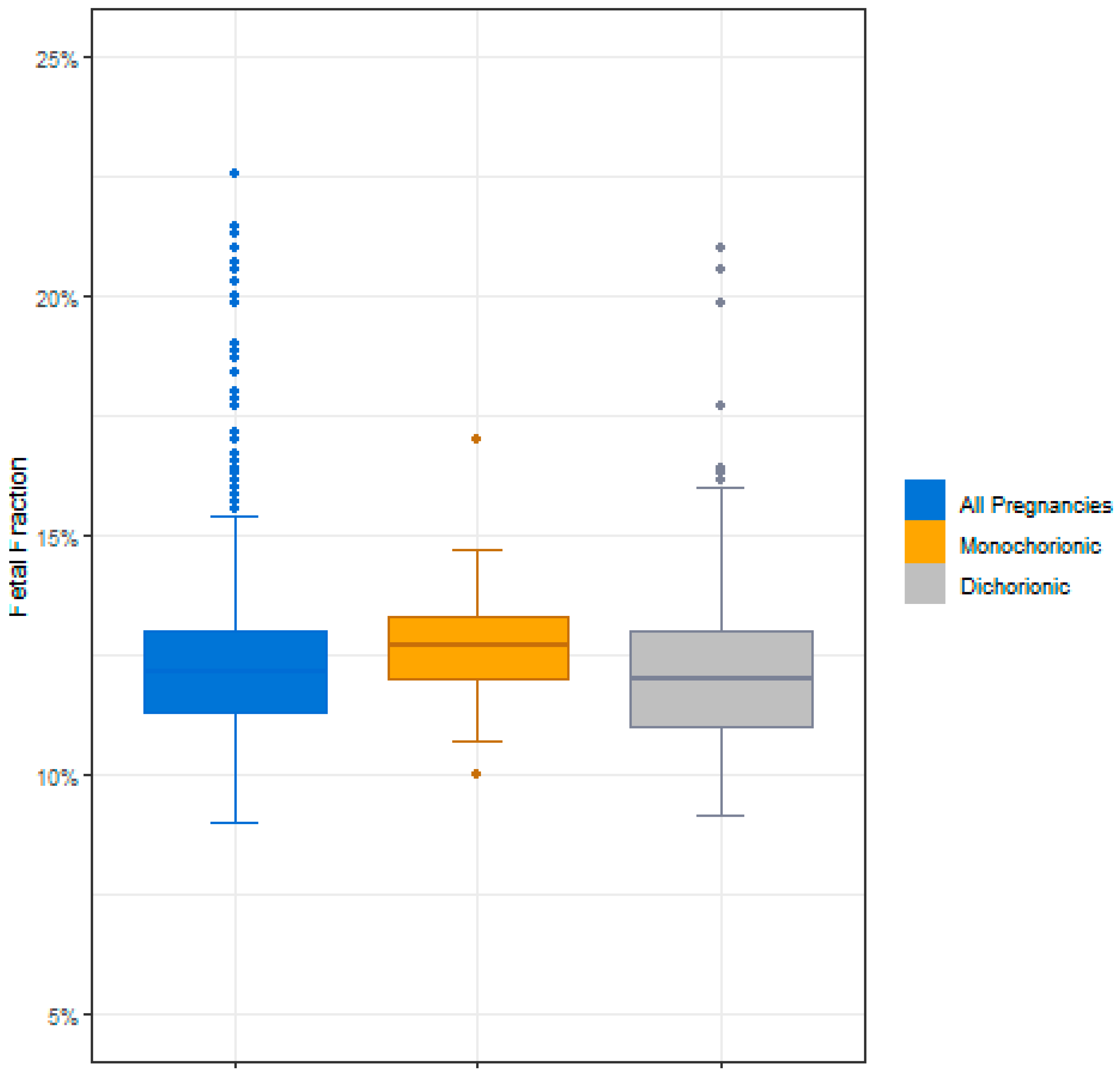
3.2. cfDNA Screening Results for Twin Samples
3.3. Clinical Outcomes for High-Risk NIPT Cases
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Palomaki, G.E.; Chiu, R.W.K.; Pertile, M.D.; Sistermans, E.A.; Yaron, Y.; Vermeesch, J.R.; Vora, N.L.; Best, R.G.; Wilkins-Haug, L. International Society for Prenatal Diagnosis Position Statement: Cell free (cf) DNA screening for Down syndrome in multiple pregnancies. Prenat. Diagn. 2020, 41, 1222–1232. [Google Scholar] [CrossRef] [PubMed]
- Pison, G.; Monden, C.; Smits, J. Twinning Rates in Developed Countries: Trends and Explanations. Popul. Dev. Rev. 2015, 41, 629–649. [Google Scholar] [CrossRef]
- Audibert, F.; Gagnon, A. No. 262-Prenatal Screening for and Diagnosis of Aneuploidy in Twin Pregnancies. J. Obstet. Gynaecol. Can. 2017, 39, e347–e361. [Google Scholar] [CrossRef]
- Yeaton-Massey, A.; Sparks, T.N.; Norton, M.E.; Jelliffe-Pawlowski, L.; Currier, R.J. Twin chorionicity and zygosity both vary with maternal age. Prenat. Diagn. 2021, 41, 1074–1079. [Google Scholar] [CrossRef]
- Committee on Practice Bulletins—Obstetrics; Society for Maternal–Fetal Medicine. Practice Bulletin No. 169: Multifetal Gestations: Twin, Triplet, and Higher-Order Multifetal Pregnancies. Obstet. Gynecol. 2016, 128, e131–e146. [Google Scholar] [CrossRef]
- Benn, P.; Rebarber, A. Non-invasive prenatal testing in the management of twin pregnancies. Prenat. Diagn. 2021, 41, 1233–1240. [Google Scholar] [CrossRef]
- Hopkins, M.K.; Dugoff, L. Screening for aneuploidy in twins. Am. J. Obstet. Gynecol. MFM 2021, 4, 100499. [Google Scholar] [CrossRef] [PubMed]
- Prats, P.; Rodríguez, I.; Comas, C.; Puerto, B. Systematic review of screening for trisomy 21 in twin pregnancies in first trimester combining nuchal translucency and biochemical markers: A meta-analysis. Prenat. Diagn. 2014, 34, 1077–1083. [Google Scholar] [CrossRef]
- Lo, Y.M.; Corbetta, N.; Chamberlain, P.F.; Rai, V.; Sargent, I.L.; Redman, C.W.; Wainscoat, J.S. Presence of fetal DNA in maternal plasma and serum. Lancet 1997, 350, 485–487. [Google Scholar] [CrossRef]
- Canick, J.A.; Kloza, E.M.; Lambert-Messerlian, G.M.; Haddow, J.E.; Ehrich, M.; van den Boom, D.; Bombard, A.T.; Deciu, C.; Palomaki, G.E. DNA sequencing of maternal plasma to identify Down syndrome and other trisomies in multiple gestations. Prenat. Diagn. 2012, 32, 730–734. [Google Scholar] [CrossRef]
- Rose, N.C.; Barrie, E.S.; Malinowski, J.; Jenkins, G.P.; McClain, M.R.; LaGrave, D.; Leung, M.L. Systematic evidence-based review: The application of noninvasive prenatal screening us-ing cell-free DNA in general-risk pregnancies. Genet. Med. 2022, 24, 1379–1391. [Google Scholar] [CrossRef] [PubMed]
- Dungan, J.S.; Klugman, S.; Darilek, S.; Malinowski, J.; Akkari, Y.M.N.; Monaghan, K.G.; Erwin, A.; Best, R.G. ACMG Board of Directors, Noninvasive prenatal screening (NIPS) for fetal chromosome abnormalities in a general-risk population: An evidence-based clinical guideline of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2023, 25, 100336. [Google Scholar] [CrossRef] [PubMed]
- van Prooyen Schuurman, L.; Sistermans, E.A.; Van Opstal, D.; Henneman, L.; Bekker, M.N.; Bax, C.J.; Pieters, M.J.; Bouman, K.; de Munnik, S.; den Hollander, N.S.; et al. Clinical impact of additional findings detected by genome-wide non-invasive prenatal testing: Follow-up results of the TRIDENT-2 study. Am. J. Hum. Genet. 2022, 109, 1140–1152. [Google Scholar] [CrossRef]
- American College of Obstetricians and Gynecologists (ACOG); Society for Maternal-Fetal Medicine (SMFM). Screening for Fetal Chromosomal Abnormalities: ACOG Practice Bulletin Summary, Number 226. Obstet. Gynecol. 2020, 136, 859–867. [Google Scholar] [CrossRef]
- Audibert, F.; De Bie, I.; Johnson, J.A.; Okun, N.; Wilson, R.D.; Armour, C.; Chitayat, D.; Kim, R. No. 348-Joint SOGC-CCMG Guideline: Update on Prenatal Screening for Fetal Aneuploidy, Fetal Anomalies, and Adverse Pregnancy Outcomes. J. Obstet. Gynaecol. Can. 2017, 39, 805–817. [Google Scholar] [CrossRef]
- Gregg, A.R.; Skotko, B.G.; Benkendorf, J.L.; Monaghan, K.G.; Bajaj, K.; Best, R.G.; Klugman, S.; Watson, M.S. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: A position statement of the American College of Medical Genetics and Genomics. Genet. Med. 2016, 18, 1056–1065. [Google Scholar] [CrossRef] [PubMed]
- Marteau, T.; Drake, H.; Bobrow, M. Counselling following diagnosis of a fetal abnormality: The differing approaches of obstetricians, clinical geneticists, and genetic nurses. J. Med. Genet. 1994, 31, 864–867. [Google Scholar] [CrossRef]
- La Verde, M.; De Falco, L.; Torella, A.; Savarese, G.; Savarese, P.; Ruggiero, R.; Conte, A.; Fico, V.; Torella, M.; Fico, A. Performance of cell-free DNA sequencing-based non-invasive prenatal testing: Experience on 36,456 singleton and multiple pregnancies. BMC Med. Genom. 2021, 14, 93. [Google Scholar] [CrossRef] [PubMed]
- Illumina, Inc. VeriSeq NIPT Solution v2 Package Insert. 2021. Available online: https://support.illumina.com/downloads/veriseq-nipt-solution-v2-package-insert-1000000078751.html (accessed on 1 December 2022).
- Pertile, M.D.; Flowers, N.; Vavrek, D.; Andrews, D.; Kalista, T.; Craig, A.; Deciu, C.; Duenwald, S.; Meier, K.; Bhatt, S. Performance of a Paired-End Sequencing-Based Noninvasive Prenatal Screening Test in the Detection of Genome-Wide Fetal Chromosomal Anomalies. Clin. Chem. 2021, 67, 1210–1219. [Google Scholar] [CrossRef]
- Borth, H.; Teubert, A.; Glaubitz, R.; Knippenberg, S.; Kutur, N.; Winkler, T.; Eiben, B. Analysis of cell-free DNA in a consecutive series of 13,607 routine cases for the detection of fetal chromosomal aneuploidies in a single center in Germany. Arch. Gynecol. Obs. 2021, 303, 1407–1414. [Google Scholar] [CrossRef]
- Mann, K.; Fox, S.P.; Abbs, S.J.; Yau, S.C.; Scriven, P.N.; Docherty, Z.; Ogilvie, C.M. Development and implementation of a new rapid aneuploidy diagnostic service within the UK National Health Service and implications for the future of prenatal diagnosis. Lancet 2001, 358, 1057–1061. [Google Scholar] [CrossRef] [PubMed]
- Newcombe, R.G. Two-sided confidence intervals for the single proportion: Comparison of seven methods. Stat. Med. 1998, 17, 857–872. [Google Scholar] [CrossRef]
- Mossfield, T.; Soster, E.; Menezes, M.; Agenbag, G.; Dubois, M.-L.; Gekas, J.; Hardy, T.; Jurkowska, M.; Kleinfinger, P.; Loggenberg, K.; et al. Multisite assessment of the impact of cell-free DNA-based screening for rare autosomal aneuploidies on pregnancy management and outcomes. Front. Genet. 2022, 13, 975987. [Google Scholar] [CrossRef]
- Khalil, A.; Archer, R.; Hutchinson, V.; Mousa, H.A.; Johnstone, E.D.; Cameron, M.J.; Cohen, K.E.; Ioannou, C.; Kelly, B.; Reed, K.; et al. Non-invasive Prenatal Screening in Twin Pregnancies with cell-free DNA using the IONA Test: A prospective multicentre study. Am. J. Obstet. Gynecol. 2021, 225, 79.e1–79.e13. [Google Scholar] [CrossRef]
- Judah, H.; Gil, M.M.; Syngelaki, A.; Galeva, S.; Jani, J.; Akolekar, R.; Nicolaides, K.H. Cell-free DNA testing of maternal blood in screening for trisomies in twin pregnancy: Updated cohort study at 10–14 weeks and meta-analysis. Ultrasound Obstet. Gynecol. 2021, 58, 178–189. [Google Scholar] [CrossRef] [PubMed]
- Gil, M.M.; Accurti, V.; Santacruz, B.; Plana, M.N.; Nicolaides, K.H. Analysis of cell-free DNA in maternal blood in screening for aneuploidies: Updated meta-analysis. Ultrasound Obstet. Gynecol. 2017, 50, 302–314. [Google Scholar] [CrossRef]
- Chibuk, J.; Rafalko, J.; Boomer, T.; McCullough, R.; McLennan, G.; Wyatt, P.; Almasri, E. Cell-free DNA screening in twin pregnancies: A more accurate and reliable screening tool. Prenat. Diagn. 2020, 40, 1321–1329. [Google Scholar] [CrossRef]
- van Riel, M.; Brison, N.; Baetens, M.; Blaumeiser, B.; Boemer, F.; Bourlard, L.; Bulk, S.; De Leener, A.; Désir, J.; Devriendt, K.; et al. Performance and Diagnostic Value of Genome-Wide Noninvasive Prenatal Testing in Multiple Gestations. Obstet. Gynecol. 2021, 137, 1102–1108. [Google Scholar] [CrossRef]
- Eggenhuizen, G.M.; Go, A.; Koster, M.P.H.; Baart, E.B.; Galjaard, R.J. Confined placental mosaicism and the association with pregnancy outcome and fetal growth: A review of the literature. Hum. Reprod. Update 2021, 27, 885–903. [Google Scholar] [CrossRef]

| Characteristic | Value |
|---|---|
| Indications for NIPT, n (%) Advanced maternal age Based on traditional screening 1 Based on pregnancy history 2 Parental carrier of balanced translocation 3 Patient preference Unknown | 628 (50.5) 25 (2.0) 12 (1.0) 2 (0.2) 288 (23.1) 289 (23.2) |
| Maternal age/Egg donor age, y Mean Median IQR | 35.22 35.05 6.48 |
| Gestational age at NIPT, days Mean Median IQR | 12.4 12.14 1.71 |
| Trimester of screening, n (%) First (up to 14 wk) Second (14–27 wk) | 1111(89.3) 133 (10.7) |
| ART pregnancies, n (%) Total IVF ICSI IUI Egg donation 4 PGT-A transfer euploid embryo Not specified | 370 (30.5) 12 (1.0) 15 (1.3) 5 (0.5) 35 (2.6) 7 (0.6) 282 (24.5) |
| Chorionicity, n (%) Dichorionic pregnancy 5 Monochorionic pregnancy 6 Chorionicity unknown | 1080 (86.8) 66 5.3) 98 (7.8) |
| NIPT Result | n | Lost to Follow-Up | No Confirmatory Testing | True Positives | Discordant Result |
| Trisomy 21 | 18 | 0 | 1 * | 17 | 1 |
| Trisomy 18 | 1 | 0 | 0 | 1 | 0 |
| RAA | 6 | 1 | 1 * | 0 | 4 |
| CNV | 4 | 1 | 0 | 0 | 2 |
| Case | Indication for NIPT | GA at Blood Draw, wk | NIPT Result | FFE, % | Confirmatory Testing | Pregnancy Outcomes |
|---|---|---|---|---|---|---|
| RAA Cases | ||||||
| 1 | AMA | 10 | Trisomy 3 | 9 | Amnio; RAA not confirmed | Two healthy babies |
| 2 | Patient preference | 13 + 6 | Trisomy 3 | 12 | Amnio; RAA not confirmed | Two healthy babies |
| 3 | AMA | 15 + 2 | Trisomy 5 | 11 | ------ | ------ |
| 4 | AMA | 13 + 1 | Trisomy 5 | 7 | None | Two healthy babies |
| 5 | AMA | 11 + 4 | Trisomy 8 | 13 | Postnatal karyotype; RAA not confirmed | Two healthy babies |
| 6 | Patient preference | 11 + 5 | Trisomy 9 | 10 | None * | One healthy baby |
| CNV Cases | ||||||
| 7 | Patient preference | 12 + 6 | dup(9)(p24.3p13.1) | 15 | ----- | ----- |
| 8 | AMA | 12 + 6 | dup(18)(p11.31p11.1) | 20 | Amnio; CNV not confirmed | Two healthy babies |
| 9 | AMA | 11 + 4 | del(1)(p36.32p32.3)(15) | 9 | Amnio; CNV not confirmed | Two healthy babies |
| 10 | AMA | 13 | dup(7)(p15.2p14.2) | 8 | None | Two healthy babies |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
De Falco, L.; Savarese, G.; Savarese, P.; Petrillo, N.; Ianniello, M.; Ruggiero, R.; Suero, T.; Barbato, C.; Mori, A.; Ramiro, C.; et al. Clinical Experience with Genome-Wide Noninvasive Prenatal Screening in a Large Cohort of Twin Pregnancies. Genes 2023, 14, 982. https://doi.org/10.3390/genes14050982
De Falco L, Savarese G, Savarese P, Petrillo N, Ianniello M, Ruggiero R, Suero T, Barbato C, Mori A, Ramiro C, et al. Clinical Experience with Genome-Wide Noninvasive Prenatal Screening in a Large Cohort of Twin Pregnancies. Genes. 2023; 14(5):982. https://doi.org/10.3390/genes14050982
Chicago/Turabian StyleDe Falco, Luigia, Giovanni Savarese, Pasquale Savarese, Nadia Petrillo, Monica Ianniello, Raffaella Ruggiero, Teresa Suero, Cosimo Barbato, Alessio Mori, Cristina Ramiro, and et al. 2023. "Clinical Experience with Genome-Wide Noninvasive Prenatal Screening in a Large Cohort of Twin Pregnancies" Genes 14, no. 5: 982. https://doi.org/10.3390/genes14050982
APA StyleDe Falco, L., Savarese, G., Savarese, P., Petrillo, N., Ianniello, M., Ruggiero, R., Suero, T., Barbato, C., Mori, A., Ramiro, C., Della Corte, L., Saccone, G., Di Spiezio Sardo, A., & Fico, A. (2023). Clinical Experience with Genome-Wide Noninvasive Prenatal Screening in a Large Cohort of Twin Pregnancies. Genes, 14(5), 982. https://doi.org/10.3390/genes14050982







