Satellite DNAs—From Localized to Highly Dispersed Genome Components
Abstract
1. Introduction
2. Functional Importance of Satellite DNAs and Partnership with TEs
3. Diversity, Homogeneity and Evolution of satDNAs
4. Genomic Content or How Many satDNAs Are in the Genome
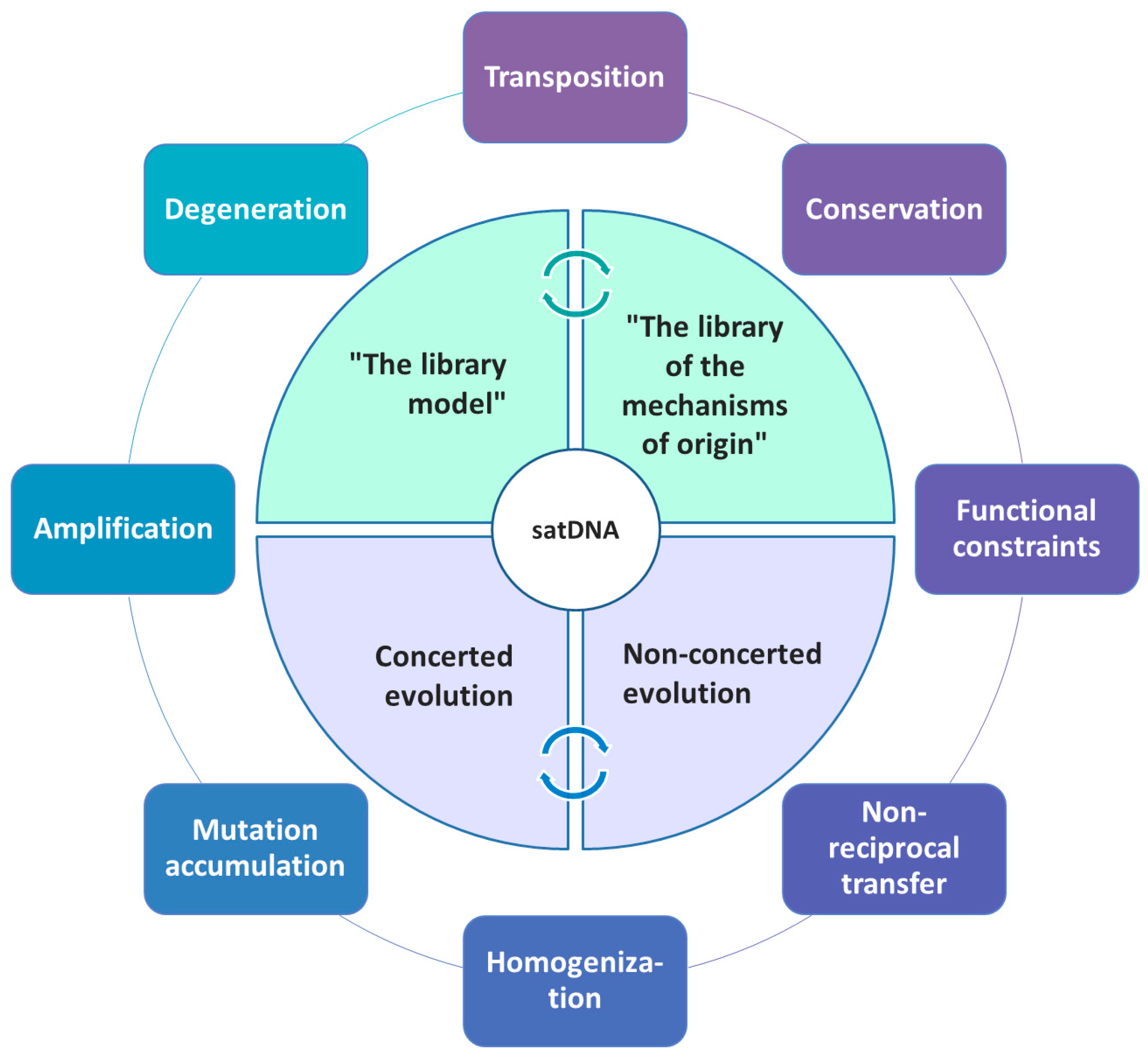
5. Comparative Satellitome Analysis: Satellitome and the Library Hypothesis
6. SatDNA Defying Predictions of the Library Model and the Concerted Evolution
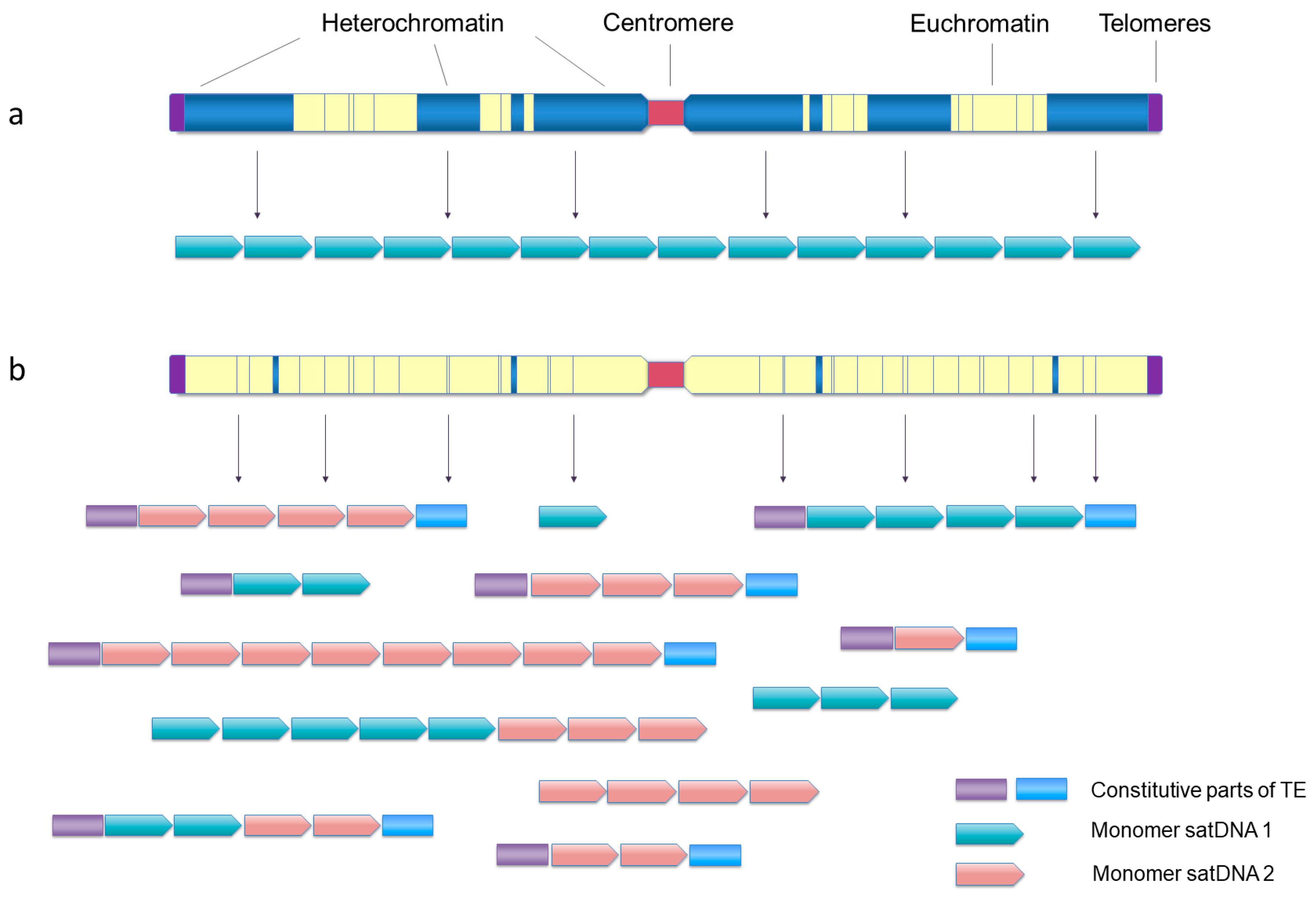
7. (Slow but Steady) Heterochromatin Mining
8. In and Out of Heterochromatin
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- López-Flores, I.; Garrido-Ramos, M.A. The repetitive DNA content of eukaryotic genomes. Genome Dyn. 2012, 7, 1–28. [Google Scholar]
- Biscotti, M.A.; Olmo, E.; Heslop-Harrison, J.S. Repetitive DNA in eukaryotic genomes. Chromosom. Res. 2015, 23, 415–420. [Google Scholar] [CrossRef] [PubMed]
- Peona, V.; Weissensteiner, M.H.; Suh, A. How complete are “complete” genome assemblies?—An avian perspective. Mol. Ecol. Resoure 2018, 18, 1188–1195. [Google Scholar] [CrossRef]
- Sedlazeck, F.J.; Lee, H.; Darby, C.A.; Schatz, M.C. Piercing the dark matter: Bioinformatics of long-range sequencing and mapping. Nat. Rev. Genet. 2018, 19, 329–346. [Google Scholar] [CrossRef]
- Ahmad, S.F.; Singchat, W.; Jehangir, M.; Suntronpong, A.; Panthum, T.; Malaivijitnond, S.; Srikulnath, K. Dark Matter of Primate Genomes: Satellite DNA Repeats and Their Evolutionary Dynamics. Cells 2020, 9, 2714. [Google Scholar] [CrossRef] [PubMed]
- Šatović, E.; Tunjić Cvitanić, M.; Plohl, M. Tools and databases for solving problems in detection and identification of repetitive DNA sequences. Period. Biol. 2020, 121–122, 7–14. [Google Scholar] [CrossRef]
- Charlesworth, B.; Sniegowski, P.; Stephan, W. The evolutionary dynamics of repetitive DNA in eukaryotes. Nature 1994, 371, 215–220. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. Satellite DNA: An evolving topic. Genes 2017, 8, 230. [Google Scholar] [CrossRef]
- Plohl, M.; Meštrović, N.; Mravinac, B. Satellite DNA evolution. Genome Dyn. 2012, 7, 126–152. [Google Scholar] [PubMed]
- Šatović Vukšić, E.; Plohl, M. Exploring satellite DNAs: Specificities of bivalve mollusks genomes. In Satellite DNAs in Physiology and Evolution; Ugarković, Đ., Ed.; Springer International Publishing: Cham, Switzerland, 2021; pp. 57–83. ISBN 978-3-030-74889-0. [Google Scholar]
- Kit, S. Equilibrium sedimentation in density gradients of DNA preparations from animal tissues. J. Mol. Biol. 1961, 3, 711–716. [Google Scholar] [CrossRef]
- Sueoka, N. Variation and heterogeneity of base composition of deoxyribonucleic acids: A compilation of old and new data. J. Mol. Biol. 1961, 3, 31–40. [Google Scholar] [CrossRef]
- Bennetzen, J.L.; Wang, H. The contributions of transposable elements to the structure, function, and evolution of plant genomes. Annu. Rev. Plant Biol. 2014, 65, 505–530. [Google Scholar] [CrossRef] [PubMed]
- Piacentini, L.; Fanti, L.; Specchia, V.; Bozzetti, M.P.; Berloco, M.; Palumbo, G.; Pimpinelli, S. Transposons, environmental changes, and heritable induced phenotypic variability. Chromosoma 2014, 123, 345–354. [Google Scholar] [CrossRef] [PubMed]
- Wells, J.N.; Feschotte, C. A Field Guide to Eukaryotic Transposable Elements. Annu. Rev. Genet. 2020, 54, 539–561. [Google Scholar] [CrossRef]
- Marsano, R.M.; Dimitri, P. Constitutive Heterochromatin in Eukaryotic Genomes: A Mine of Transposable Elements. Cells 2022, 11, 761. [Google Scholar] [CrossRef]
- Meštrović, N.; Mravinac, B.; Pavlek, M.; Vojvoda-Zeljko, T.; Šatović, E.; Plohl, M. Structural and functional liaisons between transposable elements and satellite DNAs. Chromosom. Res. 2015, 23, 583–596. [Google Scholar] [CrossRef]
- Louzada, S.; Lopes, M.; Ferreira, D.; Adega, F.; Escudeiro, A.; Gama-Carvalho, M.; Chaves, R. Decoding the Role of Satellite DNA in Genome Architecture and Plasticity—An Evolutionary and Clinical Affair. Genes 2020, 11, 72. [Google Scholar] [CrossRef]
- Ahmad, S.F.; Singchat, W.; Panthum, T.; Srikulnath, K. Impact of Repetitive DNA Elements on Snake Genome Biology and Evolution. Cells 2021, 10, 1707. [Google Scholar] [CrossRef]
- Zattera, M.L.; Bruschi, D.P. Transposable Elements as a Source of Novel Repetitive DNA in the Eukaryote Genome. Cells 2022, 11, 3373. [Google Scholar] [CrossRef]
- Sproul, J.S.; Hotaling, S.; Heckenhauer, J.; Powell, A.; Larracuente, A.M.; Kelley, J.L.; Pauls, S.U.; Frandsen, P.B. Repetitive elements in the era of biodiversity genomics: Insights from 600+ insect genomes. bioRxiv 2022. bioRxiv: 2022.06.02.494618. [Google Scholar] [CrossRef]
- Lower, S.S.; McGurk, M.P.; Clark, A.G.; Barbash, D.A. Satellite DNA evolution: Old ideas, new approaches. Curr. Opin. Genet. Dev. 2018, 49, 70–78. [Google Scholar] [CrossRef]
- Thakur, J.; Packiaraj, J.; Henikoff, S. Sequence, Chromatin and Evolution of Satellite DNA. Int. J. Mol. Sci. 2021, 22, 4309. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ruano, F.J.; López-León, M.D.; Cabrero, J.; Camacho, J.P.M. High-throughput analysis of the satellitome illuminates satellite DNA evolution. Sci. Rep. 2016, 6, 28333. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.B.; Oh, J.H.; McIver, L.J.; Rashkovetsky, E.; Michalak, K.; Garner, H.R.; Kang, L.; Nevo, E.; Korol, A.B.; Michalak, P. Divergence of Drosophila melanogaster repeatomes in response to a sharp microclimate contrast in Evolution Canyon, Israel. Proc. Natl. Acad. Sci. USA 2014, 111, 10630–10635. [Google Scholar] [CrossRef]
- van Dijk, E.L.; Jaszczyszyn, Y.; Naquin, D.; Thermes, C. The Third Revolution in Sequencing Technology. Trends Genet. 2018, 34, 666–681. [Google Scholar] [CrossRef]
- Wang, X.; Xu, W.; Wei, L.; Zhu, C.; He, C.; Song, H.; Cai, Z.; Yu, W.; Jiang, Q.; Li, L.; et al. Nanopore Sequencing and De Novo Assembly of a Black-Shelled Pacific Oyster (Crassostrea gigas) Genome. Front. Genet. 2019, 10, 1211. [Google Scholar] [CrossRef] [PubMed]
- Bai, C.M.; Xin, L.S.; Rosani, U.; Wu, B.; Wang, Q.C.; Duan, X.K.; Liu, Z.H.; Wang, C.M. Chromosomal-level assembly of the blood clam, Scapharca (Anadara) broughtonii, using long sequence reads and Hi-C. Gigascience 2019, 8, giz067. [Google Scholar] [CrossRef]
- Skanderson, D.; Meauley, M.; Garrett, J.; Nurk, S.; Koren, S.; Rhie, A.; Rautiainen, M.; Bzikadze, A.V.; Mikheenko, A.; Vollger, M.R.; et al. The complete sequence of a human genome. Science 2022, 376, 44–53. [Google Scholar] [CrossRef]
- Vondrak, T.; Robledillo, L.A.; Novák, P.; Koblížková, A.; Neumann, P.; Macas, J. Characterization of repeat arrays in ultra-long nanopore reads reveals frequent origin of satellite DNA from retrotransposon-derived tandem repeats. Plant J. 2020, 101, 484–500. [Google Scholar] [CrossRef] [PubMed]
- Vondrak, T.; Oliveira, L.; Novák, P.; Koblíz, A.; Neumann, P.; Macas, J. Complex sequence organization of heterochromatin in the holocentric plant Cuscuta europaea elucidated by the computational analysis of nanopore reads. Comput. Struct. Biotechnol. J. 2021, 19, 2179–2189. [Google Scholar] [CrossRef]
- Peñaloza, C.; Gutierrez, A.P.; Eory, L.; Wang, S.; Guo, X.; Archibald, A.L.; Bean, T.P.; Houston, R.D. A chromosome-level genome assembly for the Pacific oyster (Crassostrea gigas). Gigascience 2021, 10, giab020. [Google Scholar] [CrossRef] [PubMed]
- Talbert, P.B.; Henikoff, S. The genetics and epigenetics of satellite centromeres. Genome Res. 2022, 32, 608–615. [Google Scholar] [CrossRef] [PubMed]
- Kirov, I.; Kolganova, E.; Dudnikov, M.; Yurkevich, O.Y.; Amosova, A.V.; Muravenko, O.V. A Pipeline NanoTRF as a New Tool for De Novo Satellite DNA Identification in the Raw Nanopore Sequencing Reads of Plant Genomes. Plants 2022, 11, 2103. [Google Scholar] [CrossRef]
- Slamovits, C.H.; Cook, J.A.; Lessa, E.P.; Rossi, M.S. Recurrent amplifications and deletions of satellite DNA accompanied chromosomal diversification in South American tuco-tucos (genus Ctenomys, Rodentia: Octodontidae): A phylogenetic approach. Mol. Biol. Evol. 2001, 18, 1708–1719. [Google Scholar] [CrossRef] [PubMed]
- Adega, F.; Guedes-Pinto, H.; Chaves, R. Satellite DNA in the karyotype evolution of domestic animals—Clinical considerations. Cytogenet. Genome Res. 2009, 126, 12–20. [Google Scholar] [CrossRef]
- Shatskikh, A.S.; Kotov, A.A.; Adashev, V.E.; Bazylev, S.S.; Olenina, L.V. Functional Significance of Satellite DNAs: Insights from Drosophila. Front. Cell Dev. Biol. 2020, 8, 312. [Google Scholar] [CrossRef]
- Plohl, M.; Meštrović, N.; Mravinac, B. Centromere identity from the DNA point of view. Chromosoma 2014, 123, 313–325. [Google Scholar] [CrossRef]
- Hartley, G.; O’Neill, R. Centromere Repeats: Hidden Gems of the Genome. Genes 2019, 10, 223. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Ahmad, K.; Malik, H.S. The Centromere Paradox: Stable Inheritance with Rapidly Evolving DNA. Science 2001, 293, 1098–1102. [Google Scholar] [CrossRef]
- Larracuente, A.M. The organization and evolution of the Responder satellite in species of the Drosophila melanogaster group: Dynamic evolution of a target of meiotic drive. BMC Evol. Biol. 2014, 14, 233. [Google Scholar] [CrossRef]
- Ferree, P.M.; Barbash, D.A. Species-specific heterochromatin prevents mitotic chromosome segregation to cause hybrid lethality in Drosophila. PLoS Biol. 2009, 7, e1000234. [Google Scholar] [CrossRef]
- Podgornaya, O.I. Nuclear organization by satellite DNA, SAF-A/hnRNPU and matrix attachment regions. Semin. Cell Dev. Biol. 2022, 128, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Biscotti, M.A.; Canapa, A.; Forconi, M.; Olmo, E.; Barucca, M. Transcription of tandemly repetitive DNA: Functional roles. Chromosom. Res. 2015, 23, 463–477. [Google Scholar] [CrossRef] [PubMed]
- Ting, D.T.; Lipson, D.; Paul, S.; Brannigan, B.W.; Akhavanfard, S.; Coffman, E.J.; Contino, G.; Deshpande, V.; Iafrate, J.A.; Letovsky, S.; et al. Aberrant overexpression of satellite repeats in pancreatic and other epithelial cancers. Science 2011, 331, 593–596. [Google Scholar] [CrossRef]
- Feliciello, I.; Akrap, I.; Ugarković, Đ. Satellite DNA Modulates Gene Expression in the Beetle Tribolium castaneum after Heat Stress. PLoS Genet. 2015, 11, e1005466. [Google Scholar] [CrossRef]
- Ferreira, D.; Meles, S.; Escudeiro, A.; Mendes-da-Silva, A.; Adega, F.; Chaves, R. Satellite non-coding RNAs: The emerging players in cells, cellular pathways and cancer. Chromosom. Res. 2015, 23, 479–493. [Google Scholar] [CrossRef]
- Miga, K.H. Centromeric satellite DNAs: Hidden sequence variation in the human population. Genes 2019, 10, 352. [Google Scholar] [CrossRef] [PubMed]
- Porokhovnik, L.N.; Veiko, N.N.; Ershova, E.S.; Kostyuk, S.V. The role of human satellite iii (1q12) copy number variation in the adaptive response during aging, stress, and pathology: A pendulum model. Genes 2021, 12, 1524. [Google Scholar] [CrossRef]
- Ugarković, Đ.; Sermek, A.; Ljubić, S.; Feliciello, I. Satellite DNAs in Health and Disease. Genes 2022, 13, 1154. [Google Scholar] [CrossRef]
- Lopes, M.; Louzada, S.; Ferreira, D.; Veríssimo, G.; Eleutério, D.; Carvalho, M.G.; Chaves, R. Human Satellite 1A analysis provides evidence of pericentromeric transcription. BMC Biol. 2023, 21, 28. [Google Scholar] [CrossRef]
- Paço, A.; Freitas, R.; Vieira-Da-Silva, A. Conversion of DNA sequences: From a transposable element to a tandem repeat or to a gene. Genes 2019, 10, 1014. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, G.C.S.; Heringer, P.; Dias, G.B. Structure, Organization, and Evolution of Satellite DNAs: Insights from the Drosophila repleta and D. virilis Species Groups. In Satellite DNAs in Physiology and Evolution. Progress in Molecular and Subcellular Biology; Ugarković, Đ., Ed.; Springer: Cham, Switzerland, 2021; pp. 27–56. [Google Scholar]
- McGurk, M.P.; Barbash, D.A. Double insertion of transposable elements provides a substrate for the evolution of satellite DNA. Genome Res. 2018, 28, 714–725. [Google Scholar] [CrossRef] [PubMed]
- Kapitonov, V.V.; Jurka, J. Helitrons on a roll: Eukaryotic rolling-circle transposons. Trends Genet. 2007, 23, 521–529. [Google Scholar] [CrossRef] [PubMed]
- Thomas, J.; Pritham, E.J. Helitrons, the Eukaryotic Rolling-circle Transposable Elements. Microbiol. Spectr. 2015, 3, 1–32. [Google Scholar] [CrossRef]
- Gaffney, P.M.; Pierce, J.C.; Mackinley, A.G.; Titchen, D.A.; Glenn, W.K. Pearl, a novel family of putative transposable elements in bivalve mollusks. J. Mol. Evol. 2003, 56, 308–316. [Google Scholar] [CrossRef]
- Yang, H.-P.; Barbash, D.A. Abundant and species-specific DINE-1 transposable elements in 12 Drosophila genomes. Genome Biol. 2008, 9, R39. [Google Scholar] [CrossRef]
- Šatović, E.; Plohl, M. Tandem repeat-containing MITE elements in the clam Donax trunculus. Genome Biol. Evol. 2013, 5, 2549–2559. [Google Scholar] [CrossRef]
- Dias, G.B.; Heringer, P.; Svartman, M.; Kuhn, G.C.S.S. Helitrons shaping the genomic architecture of Drosophila: Enrichment of DINE-TR1 in α- and β-heterochromatin, satellite DNA emergence, and piRNA expression. Chromosom. Res. 2015, 23, 597–613. [Google Scholar] [CrossRef]
- Šatović, E.; Vojvoda Zeljko, T.; Luchetti, A.; Mantovani, B.; Plohl, M. Adjacent sequences disclose potential for intra-genomic dispersal of satellite DNA repeats and suggest a complex network with transposable elements. BMC Genom. 2016, 17, 997. [Google Scholar] [CrossRef]
- Tunjić-Cvitanić, M.; Pasantes, J.J.; García-Souto, D.; Cvitanić, T.; Plohl, M.; Šatović-Vukšić, E. Satellitome Analysis of the Pacific Oyster Crassostrea gigas Reveals New Pattern of Satellite DNA Organization, Highly Scattered across the Genome. Int. J. Mol. Sci. 2021, 22, 6798. [Google Scholar] [CrossRef]
- Cohen, S.; Agmon, N.; Sobol, O.; Segal, D. Extrachromosomal circles of satellite repeats and 5S ribosomal DNA in human cells. Mob. DNA 2010, 1, 11. [Google Scholar] [CrossRef] [PubMed]
- Grabundzija, I.; Messing, S.A.; Thomas, J.; Cosby, R.L.; Bilic, I.; Miskey, C.; Gogol-Döring, A.; Kapitonov, V.; Diem, T.; Dalda, A.; et al. A Helitron transposon reconstructed from bats reveals a novel mechanism of genome shuffling in eukaryotes. Nat. Commun. 2016, 7, 10716. [Google Scholar] [CrossRef] [PubMed]
- Scalvenzi, T.; Pollet, N. Insights on genome size evolution from a miniature inverted repeat transposon driving a satellite DNA. Mol. Phylogenet. Evol. 2014, 81, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Plohl, M.; Luchetti, A.; Mestrović, N.; Mantovani, B. Satellite DNAs between selfishness and functionality: Structure, genomics and evolution of tandem repeats in centromeric (hetero)chromatin. Gene 2008, 409, 72–82. [Google Scholar] [CrossRef]
- Šatović-Vukšić, E.; Plohl, M. Classification Problems of Repetitive DNA Sequences. DNA 2021, 1, 84–90. [Google Scholar] [CrossRef]
- Talbert, P.B.; Henikoff, S. What makes a centromere? Exp. Cell Res. 2020, 389, 111895. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. The Genomics of Plant Satellite DNA. In Satellite DNAs in Physiology and Evolution. Progress in Molecular and Subcellular Biology; Ugarković, Đ., Ed.; Springer: Cham, Switzerland, 2021; pp. 103–143. [Google Scholar]
- Smith, G. Evolution of repeated DNA sequences by unequal crossover. Science 1976, 191, 528–535. [Google Scholar] [CrossRef]
- Stephan, W. Recombination and the evolution of satellite DNA. Genet. Res. 1986, 47, 167–174. [Google Scholar] [CrossRef]
- Elder, J.F.; Turner, B.J. Concerted evolution of repetitive DNA sequences in eukaryotes. Q. Rev. Biol. 1995, 70, 297–320. [Google Scholar] [CrossRef]
- Dover, G. Molecular drive. Trends Genet. 2002, 18, 587–589. [Google Scholar] [CrossRef]
- Dover, G. Molecular drive: A cohesive mode of species evolution. Nature 1982, 299, 111–117. [Google Scholar] [CrossRef]
- Dover, G.A. Molecular drive in multigene families: How biological novelties arise, spread and are assimilated. Trends Genet. 1986, 2, 159–165. [Google Scholar] [CrossRef]
- Strachan, T.; Webb, D.; Dover, G.A. Transition stages of molecular drive in multiple-copy DNA families in Drosophila. EMBO J. 1985, 4, 1701–1708. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, L.; Sperlich, D. Gradual evolution of a specific satellite DNA family in Drosophila ambigua, D. tristis, and D. obscura. Mol. Biol. Evol. 1993, 10, 647–659. [Google Scholar] [CrossRef] [PubMed]
- Dover, G.A. DNA turnover and the molecular clock. J. Mol. Evol. 1987, 26, 47–58. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A.; De La Herrán, R.; Jamilena, M.; Lozano, R.; Ruiz Rejón, C.; Ruiz Rejón, M. Evolution of Centromeric Satellite DNA and Its Use in Phylogenetic Studies of the Sparidae Family (Pisces, Perciformes). Mol. Phylogenet. Evol. 1999, 12, 200–204. [Google Scholar] [CrossRef]
- Nijman, I.J.; Lenstra, J.A. Mutation and recombination in cattle satellite DNA: A feedback model for the evolution of satellite DNA repeats. J. Mol. Evol. 2001, 52, 361–371. [Google Scholar] [CrossRef]
- McAllister, B.F.; Werren, J.H. Evolution of tandemly repeated sequences: What happens at the end of an array? J. Mol. Evol. 1999, 48, 469–481. [Google Scholar] [CrossRef]
- Mravinac, B.; Plohl, M. Satellite DNA junctions identify the potential origin of new repetitive elements in the beetle Tribolium madens. Gene 2007, 394, 45–52. [Google Scholar] [CrossRef]
- Camacho, J.P.M.; Cabrero, J.; López-León, M.D.; Martín-Peciña, M.; Perfectti, F.; Garrido-Ramos, M.A.; Ruiz-Ruano, F.J. Satellitome comparison of two oedipodine grasshoppers highlights the contingent nature of satellite DNA evolution. BMC Biol. 2022, 20, 36. [Google Scholar] [CrossRef]
- Ugarković, Đ.; Plohl, M. Variation in satellite DNA profiles-causes and effects. EMBO J. 2002, 21, 5955–5959. [Google Scholar] [CrossRef] [PubMed]
- Fry, K.; Salser, W. Nucleotide sequences of HS-α satellite DNA from kangaroo rat Dipodomys ordii and characterization of similar sequences in other rodents. Cell 1977, 12, 1069–1084. [Google Scholar] [CrossRef] [PubMed]
- Macas, J.; Neumann, P.; Navrátilová, A. Repetitive DNA in the pea (Pisum sativum L.) genome: Comprehensive characterization using 454 sequencing and comparison to soybean and Medicago truncatula. BMC Genom. 2007, 8, 427. [Google Scholar] [CrossRef] [PubMed]
- Macas, J.; Novak, P.; Pellicer, J.; Cizkova, J.; Koblizkova, A.; Neumann, P.; Fukova, I.; Dolezel, J.; Kelly, L.J.; Leitch, I.J. In depth characterization of repetitive DNA in 23 plant genomes reveals sources of genome size variation in the legume tribe FABEAE. PLoS ONE 2015, 10, e0143424. [Google Scholar] [CrossRef]
- Novák, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; MacAs, J. RepeatExplorer: A Galaxy-based web server for genome-wide characterization of eukaryotic repetitive elements from next-generation sequence reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef] [PubMed]
- Novák, P.; Robledillo, L.Á.; Koblížková, A.; Vrbová, I.; Neumann, P.; Macas, J. TAREAN: A computational tool for identification and characterization of satellite DNA from unassembled short reads. Nucleic Acids Res. 2017, 45, e111. [Google Scholar] [CrossRef]
- Novák, P.; Neumann, P. Global analysis of repetitive DNA from unassembled sequence reads using RepeatExplorer2. Nat. Protoc. 2020, 15, 3745–3776. [Google Scholar] [CrossRef]
- Bardella, V.B.; Milani, D.; Cabral-de-Mello, D.C. Analysis of Holhymenia histrio genome provides insight into the satDNA evolution in an insect with holocentric chromosomes. Chromosom. Res. 2020, 28, 369–380. [Google Scholar] [CrossRef]
- Cabral-de-Mello, D.C.; Zrzavá, M.; Kubíčková, S.; Rendón, P.; Marec, F. The Role of Satellite DNAs in Genome Architecture and Sex Chromosome Evolution in Crambidae Moths. Front. Genet. 2021, 12, 661417. [Google Scholar] [CrossRef]
- Milani, D.; Ruiz-Ruano, F.J.; Camacho, J.P.M.; Cabral-de-Mello, D.C. Out of patterns, the euchromatic B chromosome of the grasshopper Abracris flavolineata is not enriched in high-copy repeats. Heredity 2021, 127, 475–483. [Google Scholar] [CrossRef]
- Gomes Goes, C.A.; dos Santos, R.Z.; Clemente Aguiar, W.R.; Vieira Alves, D.C.; Zerbinato de Andrade Silva, D.M.; Foresti, F.; Oliveira, C.; Utsunomia, R.; Porto-Foresti, F. Revealing the Satellite DNA History in Psalidodon and Astyanax Characid Fish by Comparative Satellitomics. Front. Genet. 2022, 13, 884072. [Google Scholar] [CrossRef] [PubMed]
- de Silva, D.M.Z.A.; Utsunomia, R.; Ruiz-Ruano, F.J.; Daniel, S.N.; Porto-Foresti, F.; Hashimoto, D.T.; Oliveira, C.; Camacho, J.P.M.; Foresti, F.; Silva, D.M.Z.d.A.; et al. High-throughput analysis unveils a highly shared satellite DNA library among three species of fish genus Astyanax. Sci. Rep. 2017, 7, 12726. [Google Scholar] [CrossRef] [PubMed]
- Serrano-Freitas, É.A.; Silva, D.M.Z.A.; Ruiz-Ruano, F.J.; Utsunomia, R.; Araya-Jaime, C.; Oliveira, C.; Camacho, J.P.M.; Foresti, F. Satellite DNA content of B chromosomes in the characid fish Characidium gomesi supports their origin from sex chromosomes. Mol. Genet. Genom. 2020, 295, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Sena, R.S.; Heringer, P.; Valeri, M.P.; Pereira, V.S.; Kuhn, G.C.S.; Svartman, M. Identification and characterization of satellite DNAs in two-toed sloths of the genus Choloepus (Megalonychidae, Xenarthra). Sci. Rep. 2020, 10, 19202. [Google Scholar] [CrossRef]
- Navarro-Dominguez, B.; Cabrero, J.; López-León, M.D.; Ruiz-Ruano, F.J.; Pita, M.; Bella, J.L.; Camacho, J.P.M. Tandem Repeat DNA Provides Many Cytological Markers for Hybrid Zone Analysis in Two Subspecies of the Grasshopper Chorthippus parallelus. Genes 2023, 14, 397. [Google Scholar] [CrossRef]
- Gomes Goes, C.A.; dos Santos, N.; de Mira Rodrigues, P.H.; Forte Stornioli, J.H.; Bueno da Silva, A.; dos Santos, R.Z.; Dziechciarz Vidal, J.A.; Zerbinato de Andrade Silva, D.M.; Artoni, R.F.; Foresti, F.; et al. The Satellite DNA Catalogues of Two Serrasalmidae (Teleostei, Characiformes): Conservation of General satDNA Features over 30 Million Years. Genes 2023, 14, 91. [Google Scholar] [CrossRef]
- Gasparotto, A.E.; Milani, D.; Martí, E.; Ferretti, A.B.; Bardella, V.B.; Hickmann, F.; Zrzavá, M.; Marec, F.; Cabral-de-Mello, D.C. A step forward in the genome characterization of the sugarcane borer, Diatraea saccharalis: Karyotype analysis, sex chromosome system and repetitive DNAs through a cytogenomic approach. Chromosoma 2022, 131, 253–267. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Dias, G.B.; De Lima, L.G.; Kuhn, G.C.E.S.; Ramos, É.; Martins, C.; Cabral-De-Mello, D.C. High-throughput analysis of the satellitome revealed enormous diversity of satellite DNAs in the neo-Y chromosome of the cricket Eneoptera surinamensis. Sci. Rep. 2017, 7, 6422. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; Cabrero, J.; López-León, M.D.; Camacho, J.P.M. Satellite DNA content illuminates the ancestry of a supernumerary (B) chromosome. Chromosoma 2017, 126, 487–500. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Bardella, V.B.; Lemos, B.; Cabral-De-Mello, D.C. Satellite DNAs are conserved and differentially transcribed among Gryllus cricket species. DNA Res. 2018, 25, 137–147. [Google Scholar] [CrossRef]
- Mora, P.; Vela, J.; Ruiz-Ruano, F.J.; Ruiz-Mena, A.; Montiel, E.E.; Palomeque, T.; Lorite, P. Satellitome analysis in the ladybird beetle Hippodamia variegata (Coleoptera, coccinellidae). Genes 2020, 11, 783. [Google Scholar] [CrossRef] [PubMed]
- Crepaldi, C.; Martí, E.; Gonçalves, É.M.; Martí, D.A.; Parise-Maltempi, P.P. Genomic Differences between the Sexes in a Fish Species Seen through Satellite DNAs. Front. Genet. 2021, 12, 728670. [Google Scholar] [CrossRef] [PubMed]
- Utsunomia, R.; Silva, D.M.Z.d.A.; Ruiz-Ruano, F.J.; Goes, C.A.G.; Melo, S.; Ramos, L.P.; Oliveira, C.; Porto-Foresti, F.; Foresti, F.; Hashimoto, D.T. Satellitome landscape analysis of Megaleporinus macrocephalus (Teleostei, Anostomidae) reveals intense accumulation of satellite sequences on the heteromorphic sex chromosome. Sci. Rep. 2019, 9, 5856. [Google Scholar] [CrossRef]
- Pereira, J.A.; Milani, D.; Ferretti, A.B.S.M.; Bardella, V.B.; Cabral-de-Mello, D.C.; Lopes, D.M. The extensive amplification of heterochromatin in Melipona bees revealed by high throughput genomic and chromosomal analysis. Chromosoma 2021, 130, 251–262. [Google Scholar] [CrossRef]
- Despot-Slade, E.; Širca, S.; Mravinac, B.; Castagnone-Sereno, P.; Plohl, M.; Meštrović, N. Satellitome analyses in nematodes illuminate complex species history and show conserved features in satellite DNAs. BMC Biol. 2022, 20, 259. [Google Scholar] [CrossRef]
- Boštjančić, L.L.; Bonassin, L.; Anušić, L.; Lovrenčić, L.; Besendorfer, V.; Maguire, I.; Grandjean, F.; Austin, C.M.; Greve, C.; Hamadou, A.B.; et al. The Pontastacus leptodactylus (Astacidae) Repeatome Provides Insight Into Genome Evolution and Reveals Remarkable Diversity of Satellite DNA. Front. Genet. 2021, 11, 611745. [Google Scholar] [CrossRef]
- da Silva, M.J.; Fogarin Destro, R.; Gazoni, T.; Narimatsu, H.; Pereira Dos Santos, P.S.; Haddad, C.F.B.; Parise-Maltempi, P.P. Great Abundance of Satellite DNA in Proceratophrys (Anura, Odontophrynidae) Revealed by Genome Sequencing. Cytogenet. Genome Res. 2020, 160, 141–147. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; Castillo-Martínez, J.; Cabrero, J.; Gómez, R.; Pedro, J.; Camacho, M.; López-León, M.D.; Camacho, J.P.M.; López-León, M.D. High-throughput analysis of satellite DNA in the grasshopper Pyrgomorpha conica reveals abundance of homologous and heterologous higher-order repeats. Chromosoma 2018, 127, 323–340. [Google Scholar] [CrossRef]
- Milani, D.; Bardella, V.B.; Ferretti, A.B.S.M.; Palacios-Gimenez, O.M.; Melo, A.d.S.; Moura, R.C.; Loreto, V.; Song, H.; Cabral-De-Mello, D.C. Satellite DNAs unveil clues about the ancestry and composition of B chromosomes in three grasshopper species. Genes 2018, 9, 523. [Google Scholar] [CrossRef]
- Montiel, E.E.; Panzera, F.; Palomeque, T.; Lorite, P.; Pita, S. Satellitome Analysis of Rhodnius prolixus, One of the Main Chagas Disease Vector Species. Int. J. Mol. Sci. 2021, 22, 6052. [Google Scholar] [CrossRef]
- Montiel, E.E.; Mora, P.; Rico-porras, J.M.; Palomeque, T.; Lorite, P. Satellitome of the Red Palm Weevil, Rhynchophorus ferrugineus (Coleoptera: Curculionidae), the Most Diverse Among Insects. Front. Ecol. Evol. 2022, 10, 826808. [Google Scholar] [CrossRef]
- Ferretti, A.B.S.M.; Milani, D.; Palacios-Gimenez, O.M.; Ruiz-Ruano, F.J.; Cabral-de-Mello, D.C. High dynamism for neo-sex chromosomes: Satellite DNAs reveal complex evolution in a grasshopper. Heredity 2020, 125, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Haq, I.U.; Muhammad, M.; Yuan, H.; Ali, S.; Abbasi, A.; Asad, M.; Ashraf, H.J.; Khurshid, A.; Zhang, K.; Zhang, Q.; et al. Satellitome Analysis and Transposable Elements Comparison in Geographically Distant Populations of Spodoptera frugiperda. Life 2022, 12, 521. [Google Scholar] [CrossRef]
- Gutiérrez, J.; Aleix-Mata, G.; Montiel, E.E.; Cabral-de-Mello, D.C.; Marchal, J.A.; Sanchez, A. Satellitome Analysis on Talpa aquitania Genome and Inferences about the satDNAs Evolution on Some Talpidae. Genes 2022, 14, 117. [Google Scholar] [CrossRef]
- Mora, P.; Montiel, E.E.; Rico-porras, J.M.; Palomeque, T.; Panzera, F.; Lorite, P. Making the Genome Huge: The Case of Triatoma delpontei, a Triatominae Species with More than 50 % of Its Genome Full of Satellite DNA. Genes 2023, 14, 371. [Google Scholar] [CrossRef]
- Pita, S.; Panzera, F.; Mora, P.; Vela, J.; Cuadrado, Á.; Sánchez, A.; Palomeque, T.; Lorite, P. Comparative repeatome analysis on Triatoma infestans Andean and Non-Andean lineages, main vector of Chagas disease. PLoS ONE 2017, 12, e0181635. [Google Scholar] [CrossRef]
- Pereira, J.A.; Cabral-de-Mello, D.C.; Lopes, D.M. The Satellite DNAs Populating the Genome of Trigona hyalinata and the Sharing of a Highly Abundant satDNA in Trigona Genus. Genes 2023, 14, 418. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Koelman, J.; Flores, M.P.; Bradford, T.M.; Jones, K.K.; Cooper, S.J.B.; Kawakami, T.; Suh, A. Comparative analysis of morabine grasshopper genomes reveals highly abundant transposable elements and rapidly proliferating satellite DNA repeats. BMC Biol. 2020, 18, 199. [Google Scholar] [CrossRef]
- Kroupin, P.Y.; Badaeva, E.D.; Sokolova, V.M.; Chikida, N.N.; Belousova, M.K.; Surzhikov, S.A.; Kocheshkova, A.A. Aegilops crassa Boiss. repeatome characterized using low-coverage NGS as a source of new FISH markers: Application in phylogenetic studies of the Triticeae. Front. Plant Sci. 2022, 13, 980764. [Google Scholar] [CrossRef]
- Amosova, A.V.; Yurkevich, O.Y.; Bolsheva, N.L.; Samatadze, T.E.; Zoshchuk, S.A.; Muravenko, O.V. Repeatome Analyses and Satellite DNA Chromosome Patterns in Deschampsia sukatschewii, D. cespitosa, and D. antarctica (Poaceae). Genes 2022, 13, 762. [Google Scholar] [CrossRef]
- Heitkam, T.; Schulte, L.; Weber, B.; Liedtke, S.; Breitenbach, S.; Kögler, A.; Morgenstern, K.; Brückner, M.; Tröber, U.; Wolf, H.; et al. Comparative Repeat Profiling of Two Closely Related Conifers (Larix decidua and Larix kaempferi) Reveals High Genome Similarity With Only Few Fast-Evolving Satellite DNAs. Front. Genet. 2021, 12, 683668. [Google Scholar] [CrossRef] [PubMed]
- Mascagni, F.; Barghini, E.; Ceccarelli, M.; Baldoni, L.; Trapero, C.; Díez, C.M.; Natali, L.; Cavallini, A.; Giordani, T. The Singular Evolution of Olea Genome Structure. Front. Plant Sci. 2022, 13, 869048. [Google Scholar] [CrossRef] [PubMed]
- Sader, M.; Vaio, M.; Cauz-Santos, L.A.; Dornelas, M.C.; Vieira, M.L.C.; Melo, N.; Pedrosa-Harand, A. Large vs small genomes in Passiflora: The influence of the mobilome and the satellitome. Planta 2021, 253, 86. [Google Scholar] [CrossRef]
- Pamponét, V.C.C.; Souza, M.M.; Silva, G.S.; Micheli, F.; De Melo, C.A.F.; De Oliveira, S.G.; Costa, E.A.; Corrêa, R.X. Low coverage sequencing for repetitive DNA analysis in Passiflora edulis Sims: Citogenomic characterization of transposable elements and satellite DNA. BMC Genom. 2019, 20, 262. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; Navarro-Domínguez, B.; Camacho, J.P.M.; Garrido-Ramos, M.A. Characterization of the satellitome in lower vascular plants: The case of the endangered fern Vandenboschia speciosa. Ann. Bot. 2019, 123, 587–599. [Google Scholar] [CrossRef]
- Robledillo, L.A.; Koblížková, A.; Novák, P.; Böttinger, K.; Vrbová, I.; Neumann, P.; Schubert, I.; Macas, J. Satellite DNA in Vicia faba is characterized by remarkable diversity in its sequence composition, association with centromeres, and replication timing. Sci. Rep. 2018, 8, 5838. [Google Scholar] [CrossRef]
- de Lima, L.G.; Ruiz-Ruano, F.J. In-Depth Satellitome Analyses of 37 Drosophila Species Illuminate Repetitive DNA Evolution in the Drosophila Genus. Genome Biol. Evol. 2022, 14, evac064. [Google Scholar] [CrossRef]
- Silva, B.S.M.L.; Picorelli, A.C.R.; Kuhn, G.C.S. In Silico Identification and Characterization of Satellite DNAs in 23 Drosophila Species from the Montium Group. Genes 2023, 14, 300. [Google Scholar] [CrossRef]
- Liu, Q.; Li, X.; Zhou, X.; Li, M.; Zhang, F.; Schwarzacher, T.; Heslop-Harrison, J.S. The repetitive DNA landscape in Avena (Poaceae): Chromosome and genome evolution defined by major repeat classes in whole-genome sequence reads. BMC Plant Biol. 2019, 19, 226. [Google Scholar] [CrossRef]
- Utsunomia, R.; Ruiz-Ruano, F.J.; Silva, D.M.Z.A.; Serrano, É.A.; Rosa, I.F.; Scudeler, P.E.S.; Hashimoto, D.T.; Oliveira, C.; Camacho, J.P.M.; Foresti, F. A glimpse into the satellite DNA library in characidae fish (Teleostei, Characiformes). Front. Genet. 2017, 8, 103. [Google Scholar] [CrossRef]
- Pita, S.; Mora, P.; Vela, J.; Palomeque, T.; Sánchez, A.; Panzera, F.; Lorite, P. Comparative analysis of repetitive DNA between the main vectors of chagas disease: Triatoma infestans and Rhodnius prolixus. Int. J. Mol. Sci. 2018, 19, 1277. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Liu, S.; Zhou, T.; Tian, C.; Bao, L.; Dunham, R.; Liu, Z. Comparative genome analysis of 52 fish species suggests differential associations of repetitive elements with their living aquatic environments. BMC Genom. 2018, 19, 141. [Google Scholar] [CrossRef] [PubMed]
- Shah, A.; Hoffman, J.I.; Schielzeth, H. Comparative analysis of genomic repeat content in gomphocerine grasshoppers reveals expansion of satellite DNA and helitrons in species with unusually large genomes. Genome Biol. Evol. 2020, 12, 1180–1193. [Google Scholar] [CrossRef]
- Valeri, M.P.; Dias, G.B.; Moreira, C.N.; Yonenaga-Yassuda, Y.; Stanyon, R.; e Silva Kuhn, G.C.; Svartman, M. Characterization of Satellite DNAs in Squirrel Monkeys genus Saimiri (Cebidae, Platyrrhini). Sci. Rep. 2020, 10, 7783. [Google Scholar] [CrossRef] [PubMed]
- Dias, C.A.R.; Kuhn, G.C.S.; Svartman, M.; Dos Santos Júnior, J.E.; Santos, F.R.; Pinto, C.M.; Perini, F.A. Identification and characterization of repetitive DNA in the genus Didelphis Linnaeus, 1758 (Didelphimorphia, Didelphidae) and the use of satellite DNAs as phylogenetic markers. Genet. Mol. Biol. 2021, 44, e20200384. [Google Scholar] [CrossRef] [PubMed]
- Peona, V.; Kutschera, V.E.; Blom, M.P.K.; Irestedt, M.; Suh, A. Satellite DNA evolution in Corvoidea inferred from short and long reads. Mol. Ecol. 2022, 32, 1288–1305. [Google Scholar] [CrossRef]
- Kretschmer, R.; Goes, C.A.G.; Bertollo, L.A.C.; Ezaz, T.; Porto-Foresti, F.; Toma, G.A.; Utsunomia, R.; de Bello Cioffi, M. Satellitome analysis illuminates the evolution of ZW sex chromosomes of Triportheidae fishes (Teleostei: Characiformes). Chromosoma 2022, 131, 29–45. [Google Scholar] [CrossRef]
- Meštrović, N.; Plohl, M.; Mravinac, B.; Ugarković, D. Evolution of satellite DNAs from the genus Palorus-experimental evidence for the “library” hypothesis. Mol. Biol. Evol. 1998, 15, 1062–1068. [Google Scholar] [CrossRef]
- Cesari, M.; Luchetti, A.; Passamonti, M.; Scali, V.; Mantovani, B. Polymerase chain reaction amplification of the Bag320 satellite family reveals the ancestral library and past gene conversion events in Bacillus rossius (Insecta Phasmatodea). Gene 2003, 312, 289–295. [Google Scholar] [CrossRef]
- Meštrović, N.; Mravinac, B.; Plohl, M.; Ugarković, Đ.; Bruvo, B. Preliminary phylogeny of Tribolium beetles (Coleoptera: Tenebrionidae) resolved by combined analysis of mitochondrial genes. Eur. J. Entomol. 2006, 103, 709–715. [Google Scholar] [CrossRef]
- Koukalova, B.; Moraes, A.P.; Renny-Byfield, S.; Matyasek, R.; Leitch, A.R.; Kovarik, A. Fall and rise of satellite repeats in allopolyploids of Nicotiana over c. 5 million years. New Phytol. 2010, 186, 148–160. [Google Scholar] [CrossRef] [PubMed]
- del Bosque, M.E.Q.; López-Flores, I.; Suárez-Santiago, V.N.; Garrido-Ramos, M.A. Satellite-DNA diversification and the evolution of major lineages in Cardueae (Carduoideae, Asteraceae). J. Plant Res. 2014, 127, 575–583. [Google Scholar] [CrossRef] [PubMed]
- Samoluk, S.S.; Robledo, G.; Bertioli, D.; Seijo, J.G. Evolutionary dynamics of an at-rich satellite DNA and its contribution to karyotype differentiation in wild diploid Arachis species. Mol. Genet. Genom. 2017, 292, 283–296. [Google Scholar] [CrossRef] [PubMed]
- Palacios-Gimenez, O.M.; Milani, D.; Song, H.; Marti, D.A.; López-León, M.D.; Ruiz-Ruano, F.J.; Camacho, J.P.M.; Cabral-De-Mello, D.C.; O’Neill, R. Eight Million Years of Satellite DNA Evolution in Grasshoppers of the Genus Schistocerca Illuminate the Ins and Outs of the Library Hypothesis. Genome Biol. Evol. 2020, 12, 88–102. [Google Scholar] [CrossRef] [PubMed]
- Anjos, A.; Milani, D.; Bardella, V.B.; Paladini, A.; Cabral-de-Mello, D.C. Evolution of satDNAs on holocentric chromosomes: Insights from hemipteran insects of the genus Mahanarva. Chromosom. Res. 2023, 31, 5. [Google Scholar] [CrossRef]
- Costa, L.; Marques, A.; Buddenhagen, C.E.; Pedrosa-Harand, A.; Souza, G. Investigating the diversification of holocentromeric satDNA Tyba in Rhynchospora (Cyperaceae). Ann. Bot. 2023, mcad036. [Google Scholar] [CrossRef]
- Belyayev, A.; Jandová, M.; Josefiová, J.; Kalendar, R.; Mahelka, V.; Mandák, B.; Krak, K. The major satellite DNA families of the diploid Chenopodium album aggregate species: Arguments for and against the “library hypothesis”. PLoS ONE 2020, 15, e0241206. [Google Scholar] [CrossRef]
- Plohl, M.; Petrović, V.; Luchetti, A.; Ricci, A.; Šatović, E.; Passamonti, M.; Mantovani, B. Long-term conservation vs high sequence divergence: The case of an extraordinarily old satellite DNA in bivalve mollusks. Heredity 2010, 104, 543–551. [Google Scholar] [CrossRef]
- Šatović, E.; Plohl, M. Distribution of DTHS3 satellite DNA across 12 bivalve species. J. Genet. 2018, 97, 575–580. [Google Scholar] [CrossRef]
- Petraccioli, A.; Odierna, G.; Capriglione, T.; Barucca, M.; Forconi, M.; Olmo, E.; Biscotti, M.A. A novel satellite DNA isolated in Pecten jacobaeus shows high sequence similarity among molluscs. Mol. Genet. Genom. 2015, 290, 1717–1725. [Google Scholar] [CrossRef]
- Abad, J.P.; Carmena, M.; Baars, S.; Saunders, R.D.C.; Glover, D.M.; Ludeña, P.; Sentis, C.; Tyler-Smith, C.; Villasante, A. Dodeca satellite: A conserved G+C-rich satellite from the centromeric heterochromatin of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1992, 89, 4663–4667. [Google Scholar] [CrossRef]
- de la Herrán, R.; Fontana, F.; Lanfredi, M.; Congiu, L.; Leis, M.; Rossi, R.; Ruiz Rejón, C.; Ruiz Rejón, M.; Garrido-Ramos, M.A. Slow Rates of Evolution and Sequence Homogenization in an Ancient Satellite DNA Family of Sturgeons. Mol. Biol. Evol. 2001, 18, 432–436. [Google Scholar] [CrossRef] [PubMed]
- Mravinac, B.; Plohl, M.; Mestrović, N.; Ugarković, D. Sequence of PRAT satellite DNA “frozen” in some Coleopteran species. J. Mol. Evol. 2002, 54, 774–783. [Google Scholar] [CrossRef] [PubMed]
- Chaves, R.; Ferreira, D.; Mendes-Da-Silva, A.; Meles, S.; Adega, F. FA-SAT is an old satellite DNA frozen in several bilateria genomes. Genome Biol. Evol. 2017, 9, 3073–3087. [Google Scholar] [CrossRef] [PubMed]
- Lorite, P.; Carrillo, J.A.; Tinaut, A.; Palomeque, T. Evolutionary dynamics of satellite DNA in species of the Genus Formica (Hymenoptera, Formicidae). Gene 2004, 332, 159–168. [Google Scholar] [CrossRef]
- Luchetti, A.; Marini, M.; Mantovani, B. Non-concerted evolution of the RET76 satellite DNA family in Reticulitermes taxa (Insecta, Isoptera). Genetica 2006, 128, 123–132. [Google Scholar] [CrossRef]
- Belyayev, A.; Josefiová, J.; Jandová, M.; Kalendar, R.; Krak, K.; Mandák, B. Natural History of a Satellite DNA Family: From the Ancestral Genome Component to Species-Specific Sequences, Concerted and Non-Concerted Evolution. Int. J. Mol. Sci. 2019, 20, 1201. [Google Scholar] [CrossRef]
- Smalec, B.M.; Heider, T.N.; Flynn, B.L.; O’Neill, R.J. A centromere satellite concomitant with extensive karyotypic diversity across the Peromyscus genus defies predictions of molecular drive. Chromosom. Res. 2019, 27, 237–252. [Google Scholar] [CrossRef]
- Heitz, E. Das Heterochromatin der Moose. Jahrb. Wiss. Bot. 1928, 69, 762–818. [Google Scholar]
- Allshire, R.C.; Madhani, H.D. Ten principles of heterochromatin formation and function. Nat. Rev. Mol. Cell Biol. 2018, 19, 229–244. [Google Scholar] [CrossRef]
- Wallrath, L.L.; Rodriguez-Tirado, F.; Geyer, P.K. Shining Light on the Dark Side of the Genome. Cells 2022, 11, 330. [Google Scholar] [CrossRef] [PubMed]
- Bouilly, K.; Chaves, R.; Leitao, A.; Benabdelmouna, A.; Guedes-Pinto, H. Chromosomal organization of simple sequence repeats in chromosome patterns. J. Genet. 2008, 87, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Cross, I.; Díaz, E.; Sánchez, I.; Rebordinos, L. Molecular and cytogenetic characterization of Crassostrea angulata chromosomes. Aquaculture 2005, 247, 135–144. [Google Scholar] [CrossRef]
- Bressa, M.J.; Franco, M.J.; Toscani, M.A.; Papeschi, A.G. Heterochromatin heteromorphism in Holhymenia rubiginosa (Heteroptera: Coreidae). Eur. J. Entomol. 2008, 105, 65–72. [Google Scholar] [CrossRef]
- Singer, M.F. Highly Repeated Sequences in Mammalian Genomes. Int. Rev. Cytol. 1982, 76, 67–112. [Google Scholar]
- Schmidt, T.; Heslop-Harrison, J.S. Genomes, genes and junk: The large-scale organization of plant chromosomes. Trends Plant Sci. 1998, 3, 195–199. [Google Scholar] [CrossRef]
- DiBartolomeis, S.M.; Tartof, K.D.; Rob Jackson, F.; Jackson, F.R. A superfamily of Drosophila satellite related (SR) DNA repeats restricted to the X chromosome euchromatin. Nucleic Acids Res. 1992, 20, 1113–1116. [Google Scholar] [CrossRef]
- Kuhn, G.C.S.; Küttler, H.; Moreira-Filho, O.; Heslop-Harrison, J.S. The 1.688 repetitive DNA of Drosophila: Concerted evolution at different genomic scales and association with genes. Mol. Biol. Evol. 2012, 29, 7–11. [Google Scholar] [CrossRef]
- Palomeque, T.; Antonio Carrillo, J.; Muñoz-López, M.; Lorite, P. Detection of a mariner-like element and a miniature inverted-repeat transposable element (MITE) associated with the heterochromatin from ants of the genus Messor and their possible involvement for satellite DNA evolution. Gene 2006, 371, 194–205. [Google Scholar] [CrossRef]
- Langdon, T.; Seago, C.; Mende, M.; Leggett, M.; Thomas, H.; Forster, J.W.; Jones, R.N.; Jenkins, G. Retrotransposon evolution in diverse plant genomes. Genetics 2000, 156, 313–325. [Google Scholar] [CrossRef]
- Tek, A.L.; Song, J.; Macas, J.; Jiang, J. Sobo, a recently amplified satellite repeat of potato, and its implications for the origin of tandemly repeated sequences. Genetics 2005, 170, 1231–1238. [Google Scholar] [CrossRef]
- Biscotti, M.A.; Barucca, M.; Capriglione, T.; Odierna, G.; Olmo, E.; Canapa, A. Molecular and cytogenetic characterization of repetitive DNA in the Antarctic polyplacophoran Nuttallochiton mirandus. Chromosom. Res. 2008, 16, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Macas, J.; Koblízková, A.; Navrátilová, A.; Neumann, P. Hypervariable 3′ UTR region of plant LTR-retrotransposons as a source of novel satellite repeats. Gene 2009, 448, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Wolfgruber, T.K.; Presting, G.G. Tandem repeats derived from centromeric retrotransposons. BMC Genom. 2013, 14, 142. [Google Scholar] [CrossRef] [PubMed]
- Dias, G.B.; Svartman, M.; Delprat, A.; Ruiz, A.; Kuhn, G.C.S.S. Tetris is a foldback transposon that provided the building blocks for an emerging satellite DNA of Drosophila virilis. Genome Biol. Evol. 2014, 6, 1302–1313. [Google Scholar] [CrossRef]
- Luchetti, A. terMITEs: Miniature inverted-repeat transposable elements (MITEs) in the termite genome (Blattodea: Termitoidae). Mol. Genet. Genom. 2015, 290, 1499–1509. [Google Scholar] [CrossRef] [PubMed]
- Durajlija-Žinić, S.; Ugarković, D.; Cornudella, L.; Plohl, M. A novel interspersed type of organization of satellite DNAs in Tribolium madens heterochromatin. Chromosom. Res. 2000, 8, 201–212. [Google Scholar] [CrossRef]
- Liu, J.; Seetharam, A.S.; Chougule, K.; Ou, S.; Swentowsky, K.W.; Gent, J.I.; Llaca, V.; Woodhouse, M.R.; Manchanda, N.; Presting, G.G.; et al. Gapless assembly of maize chromosomes using long-read technologies. Genome Biol. 2020, 21, 121. [Google Scholar] [CrossRef]
- Tunjić Cvitanić, M.; Vojvoda Zeljko, T.; Pasantes, J.J.; García-Souto, D.; Gržan, T.; Despot-Slade, E.; Plohl, M.; Šatović, E. Sequence Composition Underlying Centromeric and Heterochromatic Genome Compartments of the Pacific Oyster Crassostrea gigas. Genes 2020, 11, 695. [Google Scholar] [CrossRef]
- Kowar, T.; Zakrzewski, F.; Macas, J.; Kobližková, A.; Viehoever, P.; Weisshaar, B.; Schmidt, T. Repeat Composition of CenH3-chromatin and H3K9me2-marked heterochromatin in Sugar Beet (β vulgaris). BMC Plant Biol. 2016, 16, 120. [Google Scholar] [CrossRef]
- Feliciello, I.; Pezer, Ž.; Kordiš, D.; Mađarić, B.B.; Ugarković, Đ. Evolutionary history of α satellite DNA repeats dispersed within human genome euchromatin. Genome Biol. Evol. 2020, 12, 2125–2138. [Google Scholar] [CrossRef] [PubMed]
- Feliciello, I.; Chinali, G.; Ugarković, D. Structure and population dynamics of the major satellite DNA in the red flour beetle Tribolium castaneum. Genetica 2011, 139, 999–1008. [Google Scholar] [CrossRef] [PubMed]
- Brajković, J.; Feliciello, I.; Bruvo-Mađarić, B.; Ugarković, D. Satellite DNA-Like Elements Associated With Genes Within Euchromatin of the Beetle Tribolium castaneum. G3 Genes|Genomes|Genet. 2012, 2, 931–941. [Google Scholar] [CrossRef] [PubMed]
- Pavlek, M.; Gelfand, Y.; Plohl, M.; Meštrović, N. Genome-wide analysis of tandem repeats in Tribolium castaneum genome reveals abundant and highly dynamic tandem repeat families with satellite DNA features in euchromatic chromosomal arms. DNA Res. 2015, 22, 387–401. [Google Scholar] [CrossRef]
- Kuhn, G.C.S.; Heslop-Harrison, J.S. Characterization and genomic organization of PERI, a repetitive DNA in the Drosophila buzzatii cluster related to DINE-1 transposable elements and highly abundant in the sex chromosomes. Cytogenet. Genome Res. 2011, 132, 79–88. [Google Scholar] [CrossRef]
- Vojvoda Zeljko, T.; Pavlek, M.; Meštrović, N.; Plohl, M. Satellite DNA—Like repeats are dispersed throughout the genome of the Pacific oyster Crassostrea gigas carried by Helentron non-autonomous mobile elements. Sci. Rep. 2020, 10, 15107. [Google Scholar] [CrossRef]
- Hofstatter, P.G.; Thangavel, G.; Lux, T.; Neumann, P.; Vondrak, T.; Novak, P.; Zhang, M.; Costa, L.; Castellani, M.; Scott, A.; et al. Repeat-based holocentromeres influence genome architecture and karyotype evolution. Cell 2022, 185, 3153–3168.e18. [Google Scholar] [CrossRef]
- Marques, A.; Ribeiro, T.; Neumann, P.; Macas, J.; Novák, P.; Schubert, V.; Pellino, M.; Fuchs, J.; Ma, W.; Kuhlmann, M.; et al. Holocentromeres in Rhynchospora are associated with genome-wide centromere-specific repeat arrays interspersed among euchromatin. Proc. Natl. Acad. Sci. USA 2015, 112, 13633–13638. [Google Scholar] [CrossRef]
- Despot-Slade, E.; Mravinac, B.; Širca, S.; Castagnone-Sereno, P.; Plohl, M.; Meštrović, N. The Centromere Histone Is Conserved and Associated with Tandem Repeats Sharing a Conserved 19-bp Box in the Holocentromere of Meloidogyne Nematodes. Mol. Biol. Evol. 2021, 38, 1943–1965. [Google Scholar] [CrossRef]
| Species | Number of satDNAs | % of the Genome | Most Abundant satDNA (%) | Reference |
|---|---|---|---|---|
| animals | ||||
| Abracris flavolineata | 53 | 4.52 | 1.73 | [93] |
| Astyanax lacustris | 33 | - | 0.001 | [94] |
| Astyanax paranae | 45 | 8.39 | 6.30 | [95] |
| Characidium gomesi | 59 | 3.98 | 0.58 | [96] |
| Choloepus didactylus | 2 | 13.62 | 13 | [97] |
| Choloepus hoffmanni | 2 | 2.83 | 2.6 | [97] |
| Chorthippus parallelus | 110 | - | 1 | [98] |
| Colossoma macropomum | 46 | <1 | 0.013 | [99] |
| Crassostrea gigas | 52 | 6.33 | 1.29 | [62] |
| Cydalima perspectalis | 1 | 0.14 | 0.14 | [92] |
| Diatraea postlineella | 2 | 0.06 | 0.04 | [92] |
| Diatraea saccharalis | 14 | 0.215 | 0.057 | [100] |
| Eneoptera surinamensis | 45 | 14 | 1.41 | [101] |
| Eumigus monticola | 27 | 1.91 | 0.55 | [102] |
| Gryllus assimilis | 13 | 4 | 1.35 | [103] |
| Hippodamia variegata | 30 | 14.93 | 9.37 | [104] |
| Locusta migratoria | 62 | 2.39 | 0.98 | [24] |
| Megaleporinus elongatus | 140 | ~5 | 0.48 | [105] |
| Megaleporinus macrocephalus | 164 | 13.47 | 2.78 | [106] |
| Melipona quadrifasciata | 13 | 2.83 | 0.94 | [107] |
| Melipona scutellaris | 13 | 38.4 | 38.2 | [107] |
| Meloidogyne arenaria | 81 | 5.07 | 0.31 | [108] |
| Meloidogyne floridensis | 38 | 1.55 | 0.14 | [108] |
| Meloidogyne incognita | 56 | 3.52 | 0.24 | [108] |
| Meloidogyne javanica | 73 | 4.27 | 0.21 | [108] |
| Oedaleus decorus | 58 | 2.53 | 0.62 | [83] |
| Ostrinia nubilalis | 4 | 0.27 | 0.15 | [92] |
| Piaractus mesopotamicus | 30 | <1 | 0.009 | [99] |
| Pontastacus leptodactylus | 258 | 27.57 | 10.91 | [109] |
| Proceratophrys boiei | 28 | 15.87 | 8.0 | [110] |
| Psalidodon bockmanni | 50 | - | 0.016 | [94] |
| Psalidodon fasciatus | 57 | - | 0.091 | [94] |
| Pyrgomorpha conica | 76 | 9.4 | 5.66 | [111] |
| Rhammatocerus brasiliensis | 12 | 1.49 | 0.76 | [112] |
| Rhodnius prolixus | 39 | 8 | 2.13 | [113] |
| Rhynchophorus ferrugineus | 112 | 25 | 20.4 | [114] |
| Ronderosia bergii | 53 | 2.44 | 0.43 | [115] |
| Schistocerca rubiginosa | 9 | 2.17 | 0.73 | [112] |
| Spodoptera frugiperda | 7 | 0.65 | 0.23 | [116] |
| Talpa aquitania | 15 | 1.24 | 0.55 | [117] |
| Triatoma delpontei | 160 | 18.15 | 53.92 | [118] |
| Triatoma infestans | 42 | 25 | 10.04 | [119] |
| Trigona hyalinata | 8 | 16.56 | 13.77 | [120] |
| Vandiemenella viatica | 129 | - | 1.48 | [121] |
| Xyleus discoideus angulatus | 18 | 2.32 | 0.62 | [112] |
| plants | ||||
| Aegilops crassa | 19 | - | 0.95 | [122] |
| Deschampsia antarctica | 20 | 2.07 | 0.21 | [123] |
| Deschampsia cespitosa | 27 | 2.85 | 0.69 | [123] |
| Deschampsia sukatschewii | 21 | 1.61 | 0.22 | [123] |
| Larix decidua | 5 | 3.2 | 1.28 | [124] |
| Larix kaempferi | 4 | 2.0 | 0.81 | [124] |
| Olea europaea cuspidata | 11 | 50.43 | 22.95 | [125] |
| Olea europaea europaea | 11 | 23.89 | 7.89 | [125] |
| Olea europaea guanchica | 11 | 23.35 | 9.23 | [125] |
| Olea exasperata | 11 | 26.43 | 15.74 | [125] |
| Olea paniculata | 11 | 1.93 | 0.79 | [125] |
| Passiflora cincinnata | 2 | - | 0.10 | [126] |
| Passiflora edulis | 2 | 0.22 | 0.16 | [127] |
| Passiflora organensis | 37 | - | 3.50 | [126] |
| Passiflora quadrangularis | 6 | - | 0.13 | [126] |
| Thinopyrum bessarabicum | 12 | - | 1.39 | [122] |
| Vandenboschia speciosa | 11 | 0.43 | 0.08 | [128] |
| Vicia faba | 23 | - | 2.72 | [129] |
| Species | Number of satDNAs | % of the Genome | Most Abundant satDNA (%) | Reference |
|---|---|---|---|---|
| Drosophila affinis | 4 | 2.07 | 0.911 | [130] |
| Drosophila albomicans | 6 | 38.8 | 36.946 | |
| Drosophila americana | 8 | 19.75 | 9.501 | |
| Drosophila ananassae | 6 | 3.68 | 1.41 | |
| Drosophila arizonae | 2 | 0.54 | 0.348 | |
| Drosophila biarmipes | 7 | 1.27 | 0.31 | |
| Drosophila bipectinata | 7 | 4.72 | 1.31 | |
| Drosophila burlai | 5 | 3.12 | 1.86 | |
| Drosophila busckii | 5 | 1.1 | 0.503 | |
| Drosophila buzzatii | 2 | 1.9 | 1.71 | |
| Drosophila elegans | 6 | 4.01 | 1.39 | |
| Drosophila erecta | 3 | 1.62 | 1.138 | |
| Drosophila eugracilis | 3 | 10.89 | 5.691 | |
| Drosophila ficusphila | 2 | 1.76 | 1.682 | |
| Drosophila hydei | 5 | 2.16 | 0.733 | |
| Drosophila kikkawai | 3 | 4.85 | 2.493 | |
| Drosophila leontia | 6 | 1.81 | 1.34 | |
| Drosophila malerkotliana | 6 | 6.04 | 2.40 | |
| Drosophila mauritiana | 7 | 4.86 | 3.58 | |
| Drosophila melanogaster | 5 | 6.6 | 1.75 | |
| Drosophila mojavensis baja | 2 | 1.76 | 1.06 | |
| Drosophila mojavensis wrigley | 2 | 2.49 | 1.63 | |
| Drosophila montana | 6 | 27.41 | 19.70 | |
| Drosophila nasuta | 7 | 33.93 | 32.68 | |
| Drosophila novamexicana | 7 | 6.82 | 3.03 | |
| Drosophila orena | 3 | 12.31 | 10.40 | |
| Drosophila persimilis | 4 | 5.87 | 5.20 | |
| Drosophila pseudoobscura | 4 | 5.48 | 1.93 | |
| Drosophila rhopaloa | 3 | 4.67 | 4.34 | |
| Drosophila santomea | 7 | 2.7 | 1.82 | |
| Drosophila sechellia | 7 | 7.72 | 6.04 | |
| Drosophila seriema | 4 | 2.9 | 1.93 | |
| Drosophila simulans | 8 | 4.53 | 3.18 | |
| Drosophila subobscura | 6 | 1.4 | 0.72 | |
| Drosophila takahashii | 5 | 3.95 | 0.98 | |
| Drosophila teissieri | 4 | 2.09 | 0.57 | |
| Drosophila virilis | 7 | 21.63 | 15.9 | |
| Drosophila yakuba | 5 | 2.83 | 1.18 | |
| Drosophila asahinai | 3 | 6.87 | 6.30 | [131] |
| Drosophila auraria | 2 | 4.72 | 4.69 | |
| Drosophila bakoue | 9 | 1.88 | 0.36 | |
| Drosophila birchii | 8 | 5.01 | 3.23 | |
| Drosophila bocki | 7 | 1.89 | 0.41 | |
| Drosophila bunnanda | 14 | 15.69 | 5.98 | |
| Drosophila burlai | 7 | 7.67 | 3.12 | |
| Drosophila jambulina | 6 | 12.24 | 5.87 | |
| Drosophila kanapiae | 4 | 1.98 | 0.78 | |
| Drosophila lacteicornis | 3 | 7.20 | 6.62 | |
| Drosophila leontia | 4 | 1.99 | 0.90 | |
| Drosophila mayri | 13 | 17.77 | 9.54 | |
| Drosophila nikananu | 3 | 8.33 | 2.80 | |
| Drosophila pectinifera | 10 | 21.65 | 15.56 | |
| Drosophila punjabiensis | 6 | 2.12 | 0.94 | |
| Drosophila rufa | 3 | 5.78 | 5.46 | |
| Drosophila seguyi | 12 | 10.73 | 4.58 | |
| Drosophila serrata | 6 | 14.50 | 10.46 | |
| Drosophila tani | 4 | 5.99 | 4.57 | |
| Drosophila triauraria | 3 | 5.70 | 5.43 | |
| Drosophila truncata | 5 | 8.92 | 4.17 | |
| Drosophila vulcana | 3 | 5.97 | 5.59 | |
| Drosophila watanabei | 3 | 1.40 | 0.74 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Šatović-Vukšić, E.; Plohl, M. Satellite DNAs—From Localized to Highly Dispersed Genome Components. Genes 2023, 14, 742. https://doi.org/10.3390/genes14030742
Šatović-Vukšić E, Plohl M. Satellite DNAs—From Localized to Highly Dispersed Genome Components. Genes. 2023; 14(3):742. https://doi.org/10.3390/genes14030742
Chicago/Turabian StyleŠatović-Vukšić, Eva, and Miroslav Plohl. 2023. "Satellite DNAs—From Localized to Highly Dispersed Genome Components" Genes 14, no. 3: 742. https://doi.org/10.3390/genes14030742
APA StyleŠatović-Vukšić, E., & Plohl, M. (2023). Satellite DNAs—From Localized to Highly Dispersed Genome Components. Genes, 14(3), 742. https://doi.org/10.3390/genes14030742







