No Association between ABCB1 G2677T/A or C3435T Polymorphisms and Survival of Breast Cancer Patients—A 10-Year Follow-Up Study in the Polish Population
Abstract
:1. Introduction
1.1. Cancer as a Therapy Target
1.2. Drug Resistance
2. Materials and Methods
2.1. Patients’ Characteristics
2.2. Histological Assessment
2.2.1. Preparation of Histological Material
2.2.2. Immunoperoxidase Methods
- Mouse monoclonal antibodies raised against human α estrogen receptors (Dako Denmark A/S, Clone 1D5, No: M 7047);
- Mouse monoclonal antibodies raised against human progesterone receptors (Dako Denmark A/S, Clone PgR 636, No: M 3569).
- Negative [−], no staining was observed;
- Positive [+], staining of up to 10% of cells was seen;
- Strongly positive [++], staining was seen in more than 10% of cells;
- Very strongly positive [+++], staining was observed in more than 75% of tested structures.
- Negative results [−];
- Positive results [+,++,+++].
2.3. The Cancer Genome Atlas (TCGA) Database
2.4. Statistical Analysis
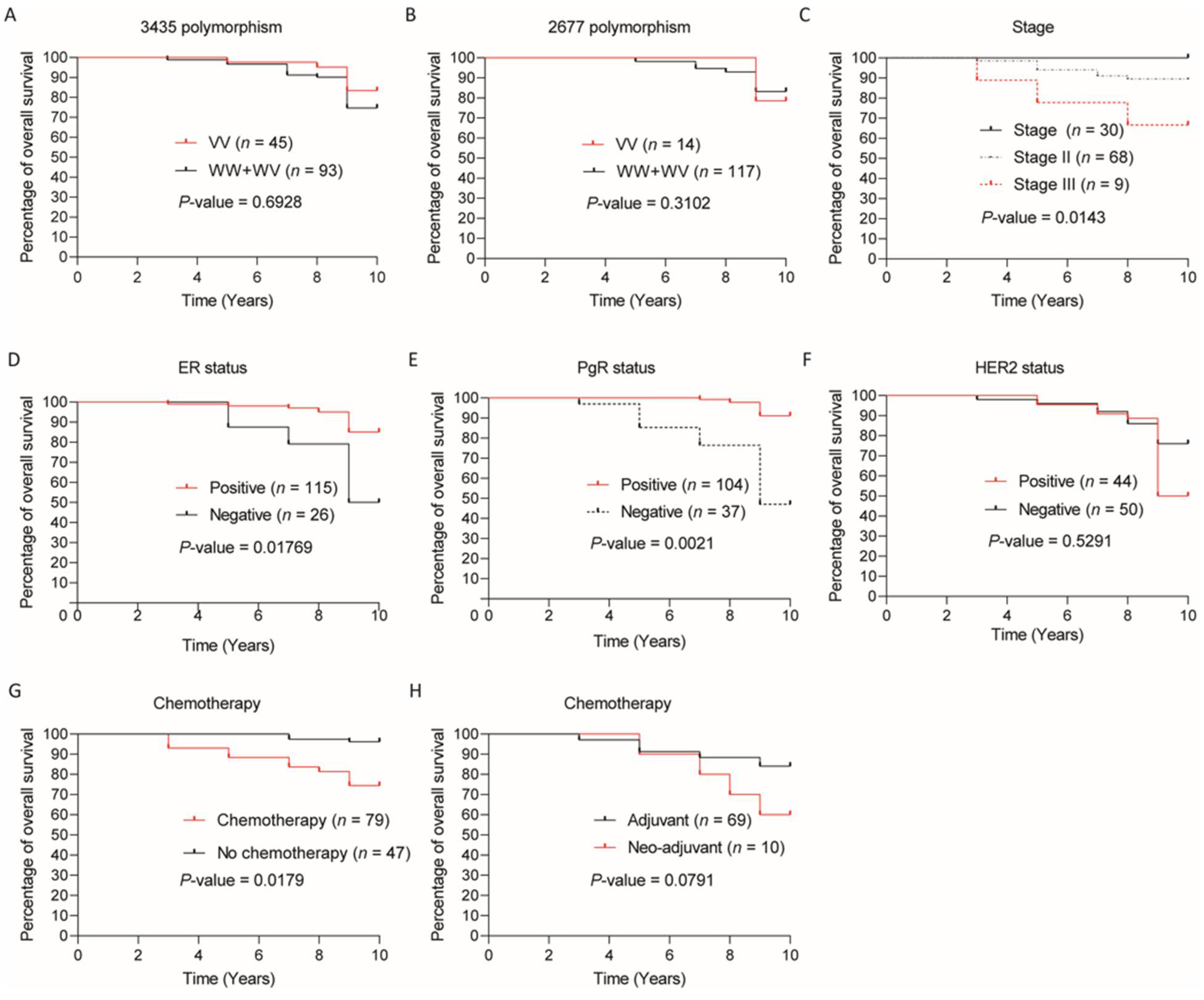
3. Results
4. Discussion
4.1. Polymorphisms and Survival Assessment
4.2. Response Prediction
4.3. Receptor Status
5. Summary and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Al-Alem, U.; Mahmoud, A.M.; Batai, K.; Shah-Williams, E.; Gann, P.H.; Kittles, R.; Rauscher, G.H. Genetic Variation and Immunohistochemical Localization of the Glucocorticoid Receptor in Breast Cancer Cases from the Breast Cancer Care in Chicago Cohort. Cancers 2021, 13, 2261. [Google Scholar] [CrossRef] [PubMed]
- Stierer, M.; Rosen, H.; Weber, R.; Hanak, H.; Spona, J.; Tuchler, H. Immunohistochemical and Biochemical Measurement of Estrogen and Progesterone Receptors in Primary Breast-Cancer-Correlation of Histopathology and Prognostic Factors. Ann. Surg. 1993, 218, 13–21. [Google Scholar] [CrossRef] [PubMed]
- Yarden, Y.; Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2001, 2, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Abe, O.; Abe, R.; Enomoto, K.; Kikuchi, K.; Koyama, H.; Masuda, H.; Nomura, Y.; Sakai, K.; Sugimachi, K.; Tominaga, T.; et al. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: An overview of the randomised trials. Lancet 2005, 365, 1687–1717. [Google Scholar]
- Jordan, V.C. Tamoxifen: A most unlikely pioneering medicine. Nat. Rev. Drug Discov. 2003, 2, 205–213. [Google Scholar] [CrossRef]
- Martínez-Sáez, O.; Prat, A. Current and Future Management of HER2-Positive Metastatic Breast Cancer. JCO Oncol. Pract. 2021, 17, 594–604. [Google Scholar] [CrossRef]
- Waks, A.G.; Winer, E.P. Breast Cancer Treatment A Review. JAMA-J. Am. Med. Assoc. 2019, 321, 288–300. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Smith, I.E.; Reis, J.S. Triple-Negative Breast Cancer. N. Engl. J. Med. 2010, 363, 1938–1948. [Google Scholar] [CrossRef] [Green Version]
- Bergin, A.R.T.; Loi, S. Triple-negative breast cancer: Recent treatment advances. F1000Research 2019, 8, 1342. [Google Scholar] [CrossRef]
- Catalano, A.; Iacopetta, D.; Ceramella, J.; Scumaci, D.; Giuzio, F.; Saturnino, C.; Aquaro, S.; Rosano, C.; Sinicropi, M.S. Multidrug Resistance (MDR): A Widespread Phenomenon in Pharmacological Therapies. Molecules 2022, 27, 616. [Google Scholar] [CrossRef]
- Mansoori, B.; Mohammadi, A.; Davudian, S.; Shirjang, S.; Baradaran, B. The Different Mechanisms of Cancer Drug Resistance: A Brief Review. Adv. Pharm. Bull. 2017, 7, 339–348. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z. Drug Resistance and Novel Therapies in Cancers in 2019. Cancers 2021, 13, 924. [Google Scholar] [CrossRef] [PubMed]
- Settleman, J. Bet on drug resistance. Nature 2016, 529, 289–290. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.H.; Yu, A.M. ABC Transporters in Multidrug Resistance and Pharmacokinetics, and Strategies for Drug Development. Curr. Pharm. Des. 2014, 20, 793–807. [Google Scholar] [CrossRef]
- Schinkel, A.H. The physiological function of drug-transporting P-glycoproteins. Semin. Cancer Biol. 1997, 8, 161–170. [Google Scholar] [CrossRef]
- Borst, P.; Elferink, R.O. Mammalian ABC transporters in health and disease. Annu. Rev. Biochem. 2002, 71, 537–592. [Google Scholar] [CrossRef] [Green Version]
- Longo, R.; D’Andrea, M.; Sarmiento, R.; Gasparini, G. Pharmacogenetics in breast cancer: Focus on hormone therapy, taxanes, trastuzumab and bevacizumab. Expert Opin. Investig. Drug 2010, 19, S41–S50. [Google Scholar] [CrossRef]
- Grimm, C.; Polterauer, S.; Zeillinger, R.; Tong, D.; Heinze, G.; Wolf, A.; Natter, C.; Reinthaller, A.; Hefler, L.A. Two Multidrug-resistance (ABCB1) Gene Polymorphisms as Prognostic Parameters in Women with Ovarian Cancer. Anticancer Res. 2010, 30, 3487–3491. [Google Scholar]
- Green, H.; Soderkvist, P.; Rosenberg, P.; Horvath, G.; Peterson, C. mdr-1 single nucleotide polymorphisms in ovarian cancer tissue: G2677T/A correlates with response to paclitaxel chemotherapy. Clin. Cancer Res. 2006, 12, 854–859. [Google Scholar] [CrossRef] [Green Version]
- Kafka, A.; Sauer, G.; Jaeger, C.; Grundmann, R.; Kreienberg, R.; Zeillinger, R.; Deissler, H. Polymorphism C3435T of the MDR-1 gene predicts response to preoperative chemotherapy in locally advanced breast cancer. Int. J. Oncol. 2003, 22, 1117–1121. [Google Scholar] [CrossRef]
- Wu, X.F.; Gu, J.; Wu, T.T.; Swisher, S.G.; Liao, Z.X.; Correa, A.M.; Liu, J.; Etzel, C.J.; Amos, C.I.; Huang, S.S.; et al. Genetic variations in radiation and chemotherapy drug action pathways predict clinical outcomes in esophageal cancer. J. Clin. Oncol. 2006, 24, 3789–3798. [Google Scholar] [CrossRef] [PubMed]
- Monzo, M.; Brunet, S.; Urbano-Ispizua, A.; Navarro, A.; Perea, G.; Esteve, J.; Artells, R.; Granell, M.; Berlanga, J.; Ribera, J.M.; et al. Genomic polymorphisms provide prognostic information in intermediate-risk acute myeloblastic leukemia. Blood 2006, 107, 4871–4879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buda, G.; Maggini, V.; Galimberti, S.; Martino, A.; Giuliani, N.; Morabito, F.; Genestreti, G.; Iacopino, P.; Rizzoli, V.; Barale, R.; et al. MDR1 polymorphism influences the outcome of multiple myeloma patients. Br. J. Haematol. 2007, 137, 454–456. [Google Scholar] [CrossRef] [PubMed]
- Rubis, B.; Holysz, H.; Barczak, W.; Gryczka, R.; Lacinski, M.; Jagielski, P.; Czernikiewicz, A.; Półrolniczak, A.; Wojewoda, A.; Perz, K.; et al. Study of ABCB1 polymorphism frequency in breast cancer patients from Poland. Pharmacol. Rep. 2012, 64, 1560–1566. [Google Scholar] [CrossRef]
- Green, H.; Peterson, C.; Soderkvist, P.; Rosenberg, P.; Horvath, G. ABCB1 2677G > T/A genotype and paclitaxel pharmacogenetics in ovarian cancer-Response. Clin. Cancer Res. 2006, 12, 4127–4129. [Google Scholar]
- Marsh, S.; Paul, J.; King, C.R.; Gifford, G.; McLeod, H.L.; Brown, R. Pharmacogenetic assessment of toxicity and outcome after platinum plus taxane chemotherapy in ovarian cancer: The Scottish Randomised Trial in Ovarian Cancer. J. Clin. Oncol. 2007, 25, 4528–4535. [Google Scholar] [CrossRef]
- Li, W.; Zhang, D.; Du, F.; Xing, X.M.; Wu, Y.; Xiao, D.; Liang, M.; Fan, Z.; Zhao, P.; Liu, T.; et al. ABCB1 3435TT and ABCG2 421CC genotypes were significantly associated with longer progression-free survival in Chinese breast cancer patients. Oncotarget 2017, 8, 111041–111052. [Google Scholar] [CrossRef]
- Kim, H.J.; Im, S.A.; Keam, B.; Ham, H.S.; Lee, K.H.; Kim, T.Y.; Oh, D.Y.; Kim, J.H.; Han, W.; Jang, I.J.; et al. ABCB1 polymorphism as prognostic factor in breast cancer patients treated with docetaxel and doxorubicin neoadjuvant chemotherapy. Cancer Sci. 2015, 106, 86–93. [Google Scholar] [CrossRef] [Green Version]
- Madrid-Paredes, A.; Canadas-Garre, M.; Sanchez-Pozo, A.; Segura-Perez, A.M.; Chamorro-Santos, C.; Vergara-Alcaide, E.; Castillo-Portellano, L.; Calleja-Hernandez, M.A. ABCB1 C3435T gene polymorphism as a potential biomarker of clinical outcomes in HER2-positive breast cancer patients. Pharmacol. Res. 2016, 108, 111–118. [Google Scholar] [CrossRef]
- Johnatty, S.E.; Beesley, J.; Paul, J.; Fereday, S.; Spurdle, A.B.; Webb, P.M.; Byth, K.; Marsh, S.; McLeod, H.; AOCS Study Group; et al. ABCB1 (MDR 1) polymorphisms and progression-free survival among women with ovarian cancer following paclitaxel/carboplatin chemotherapy. Clin. Cancer Res. 2008, 14, 5594–5601. [Google Scholar] [CrossRef] [Green Version]
- Chen, G.D.; Quan, S.; Hu, Q.G.; Wang, L.C.; Xia, X.P.; Wu, J. Lack of association between MDR1 C3435T polymorphism and chemotherapy response in advanced breast cancer patients: Evidence from current studies. Mol. Biol. Rep. 2012, 39, 5161–5168. [Google Scholar] [CrossRef] [PubMed]
- Lévy, P.; Gligorov, J.; Antoine, M.; Rezai, K.; Lévy, E.; Selle, F.; Saintigny, P.; Lokiec, F.; Avenin, D.; Beerblock, K.; et al. Influence of ABCB1 polymorphisms and docetaxel pharmacokinetics on pathological response to neoadjuvant chemotherapy in breast cancer patients. Breast Cancer Res. Treat. 2013, 139, 421–428. [Google Scholar] [CrossRef] [PubMed]
- Madrid-Paredes, A.; Canadas-Garre, M.; Sanchez-Pozo, A.; Exposito-Ruiz, M.; Calleja-Hernandez, M.A. ABCB1 gene polymorphisms and response to chemotherapy in breast cancer patients: A meta-analysis. Surg. Oncol. 2017, 26, 473–482. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Wei, Z.; Shi, K.; Zhang, Y. Association between ABCB1 G2677T/A Polymorphism and Breast Cancer Risk: A Meta-Analysis. Crit. Rev. Eukaryot Gene Expr. 2019, 29, 243–249. [Google Scholar] [CrossRef] [PubMed]
- Priyadarshini, R.; Raj, G.M.; Kayal, S.; Ramesh, A.; Shewade, D.G. Influence of ABCB1 C3435T and C1236T gene polymorphisms on tumour response to docetaxel-based neo-adjuvant chemotherapy in locally advanced breast cancer patients of South India. J. Clin. Pharm. Ther. 2019, 44, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Priyadarshini, R.; Raj, G.M.; Sundaram, R.; Kayal, S.; Ramesh, A.; Shewade, D.G. Association of plasma docetaxel levels with ABCB1 gene polymorphisms and tumour response in locally advanced breast cancer patients of South India on neo-adjuvant chemotherapy. Breast Cancer 2021, 28, 414–423. [Google Scholar] [CrossRef] [PubMed]
- Poorolajal, J.; Nafissi, N.; Akbari, M.E.; Mahjub, H.; Esmailnasab, N.; Babaee, E. Breast Cancer Survival Analysis Based on Immunohistochemistry Subtypes (ER/PR/HER2): A Retrospective Cohort Study. Arch. Iran. Med. 2016, 19, 680–686. [Google Scholar]
- Ding, Y.; Ding, K.; Qian, H.; Yu, X.; Zou, D.; Yang, H.; Mo, W.; He, X.; Zhang, F.; Qin, C.; et al. Impact on survival of estrogen receptor, progesterone receptor and Ki-67 expression discordance pre- and post-neoadjuvant chemotherapy in breast cancer. PLoS ONE 2020, 15, e0231895. [Google Scholar] [CrossRef] [Green Version]
- Kawaguchi, H.; Masuda, N.; Nakayama, T.; Aogi, K.; Anan, K.; Ito, Y.; Ohtani, S.; Sato, N.; Saji, S.; Takano, T.; et al. Factors associated with prolonged overall survival in patients with postmenopausal estrogen receptor-positive advanced breast cancer using real-world data: A follow-up analysis of the JBCRG-C06 Safari study. Breast Cancer 2020, 27, 389–398. [Google Scholar] [CrossRef] [Green Version]
- Slamon, D.J.; Clark, G.M.; Wong, S.G.; Levin, W.J.; Ullrich, A.; Mcguire, W.L. Human-Breast Cancer-Correlation of Relapse and Survival with Amplification of the Her-2 Neu Oncogene. Science 1987, 235, 177–182. [Google Scholar] [CrossRef] [Green Version]

| Characteristic | No. of Patients | % of Patients |
|---|---|---|
| 3435 polymorphism | ||
| VV | 45 | 28.7 |
| WV | 60 | 38.2 |
| WW | 33 | 21 |
| Missing | 19 | 12.1 |
| 2677 polymorphism | ||
| VV | 14 | 8.9 |
| WV | 57 | 36.3 |
| WW | 60 | 38.2 |
| Missing | 26 | 16.6 |
| Grading status | ||
| G1 | 46 | 29.3 |
| G2 | 59 | 37.6 |
| G3 | 32 | 20.4 |
| Gx | 5 | 3.2 |
| Missing | 15 | 9.5 |
| T status | ||
| T1 | 65 | 41.4 |
| T2 | 58 | 36.9 |
| T3 | 4 | 2.5 |
| T4 | 3 | 1.9 |
| Tx | 3 | 1.9 |
| Tis | 1 | 0.6 |
| Missing | 23 | 14.6 |
| N status | ||
| N0 | 50 | 31.8 |
| N1 | 74 | 47.1 |
| N2 | 7 | 4.5 |
| N3 | 1 | 0.6 |
| Nx | 2 | 1.3 |
| Missing | 23 | 14.6 |
| M status | ||
| M0 | 134 | 85.4 |
| Mx | 2 | 1.3 |
| Missing | 21 | 13.4 |
| Stage | ||
| I | 30 | 19.1 |
| II | 68 | 43.3 |
| III | 9 | 5.7 |
| Missing | 50 | 31.9 |
| PgR status | ||
| “+” | 104 | 66.2 |
| “−” | 37 | 23.6 |
| Missing | 16 | 10.2 |
| ER status | ||
| “+” | 115 | 73.2 |
| “−” | 26 | 16.6 |
| Missing | 16 | 10.2 |
| HER2 status | ||
| “+” | 44 | 28 |
| “−” | 50 | 31.8 |
| Missing | 63 | 40.2 |
| Chemotherapy | ||
| Adjuvant (AC or CMF) | 69 | 87.3 |
| Neoadjuvant (AC or CMF) | 10 | 12.7 |
| Yes | 79 | 50.4 |
| No | 47 | 29.9 |
| Missing | 31 | 19.7 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Totoń, E.; Jacczak, B.; Barczak, W.; Jagielski, P.; Gryczka, R.; Hołysz, H.; Grodecka-Gazdecka, S.; Rubiś, B. No Association between ABCB1 G2677T/A or C3435T Polymorphisms and Survival of Breast Cancer Patients—A 10-Year Follow-Up Study in the Polish Population. Genes 2022, 13, 729. https://doi.org/10.3390/genes13050729
Totoń E, Jacczak B, Barczak W, Jagielski P, Gryczka R, Hołysz H, Grodecka-Gazdecka S, Rubiś B. No Association between ABCB1 G2677T/A or C3435T Polymorphisms and Survival of Breast Cancer Patients—A 10-Year Follow-Up Study in the Polish Population. Genes. 2022; 13(5):729. https://doi.org/10.3390/genes13050729
Chicago/Turabian StyleTotoń, Ewa, Barbara Jacczak, Wojciech Barczak, Paweł Jagielski, Robert Gryczka, Hanna Hołysz, Sylwia Grodecka-Gazdecka, and Błażej Rubiś. 2022. "No Association between ABCB1 G2677T/A or C3435T Polymorphisms and Survival of Breast Cancer Patients—A 10-Year Follow-Up Study in the Polish Population" Genes 13, no. 5: 729. https://doi.org/10.3390/genes13050729
APA StyleTotoń, E., Jacczak, B., Barczak, W., Jagielski, P., Gryczka, R., Hołysz, H., Grodecka-Gazdecka, S., & Rubiś, B. (2022). No Association between ABCB1 G2677T/A or C3435T Polymorphisms and Survival of Breast Cancer Patients—A 10-Year Follow-Up Study in the Polish Population. Genes, 13(5), 729. https://doi.org/10.3390/genes13050729








