Highlights
- Canonical TGF-ß signalling regulates both CTGF and MMP-1 expression in CRC progression.
- Upregulation of Smad2 and Smad4 proteins was found to be significantly correlated with lymph node metasta-sis, distant metastasis, advanced stage, and poor survival in CRC.
- TGF-ß signalling and its downstream genes could be used as novel biomarkers and new approaches for target-ed therapy in CRC.
Abstract
TGF-β signaling pathways promote tumour development and control several downstream genes such as CTGF and MMPs. This study aimed to investigate the association between CTGF and MMP-1 mRNA expressions with clinicopathological status and survival rate in colorectal cancer patients. We investigated expression levels of CTGF and MMP-1 genes in paraffin-embedded tumours and adjacent normal tissue blocks (ADJ) by Real Time-PCR. Then, the expression of Smad2 and Smad4 proteins in the TGF-β canonical pathway was evaluated by immunohistochemistry. Finally, the correlation between CTGF, MMP-1, and the canonical TGF-β-signalling pathway with the clinicopathological features was investigated. Expression levels of MMP-1and CTGF were higher in tumours compared with adjacent normal tissues. Overexpression levels of MMP-1 and CTGF were associated with lymph node metastasis, distant metastasis, tumour histopathological grading, advanced stage, and poor survival (p < 0.05). Additionally, a significant association between the upregulation of MMP-1 and tumour location was noted. Upregulation of Smad2 and Smad4 proteins were also significantly correlated with lymph node metastasis, distant metastasis, advanced stage, and poor survival (p < 0.0001). This study showed that canonical TGF-β signalling regulates both CTGF and MMP-1 expression and CRC progression. Moreover, TGF-β signalling and its downstream genes could be used as novel biomarkers and novel approaches for targeted therapy in CRC.
1. Introduction
Colorectal cancer (CRC) is the third leading cause of cancer mortality and the fourth most frequently occurring cancer in the world [1,2]. Many signalling pathways are involved in CRC initiation and progression, one of which is transforming growth factor-β (TGF-β). It has been shown previously that TGF-β signalling plays a crucial role in cancer deployment via both Smad-dependent and Smad-independent pathways [3,4,5,6]. At the early stages of CRC, TGF-β inhibits tumorigenesis by stimulating apoptosis in cancerous cells. However, in the advanced stages of CRC the upregulation of TGF-β induces metastasis and malignancy features. Previous studies have demonstrated the TGF-β pathway affects CRC pathogenesis through upregulation of CTGF via the canonical Smad-dependent pathway [3,7,8,9]. CTGF is a transcriptional target of TGF-β signalling and induces angiogenesis and invasion by the regulation of matrix metalloproteinases (MMPs) [10,11], which are extracellular matrix (ECM) degradation enzymes and play key roles in tumour progression through digestion of ECM and providing a suitable condition for tumour angiogenesis, invasion, and metastasis [12,13]. Overexpression levels of MMP-1 have been reported in various cancer tissues, where MMP-1 overexpression was associated with angiogenesis, lymph node metastasis, and poor prognosis in CRC [14,15,16,17,18].
The discovery of potential new biomarkers for CRC is essential for improvements in early diagnosis and improvements in patient outcomes. This study aimed to investigate the association between the TGF-β/Smad signalling pathways and CTGF and MMP-1 expression, with clinical status and survival rates in CRC patients.
2. Materials and Methods
2.1. Patients and Sample Collection
In this prospective study, we utilized 122 Formalin-Fixed Paraffin-Embedded (FFPE) tissue blocks of surgical samples from CRC patients who were referred to Taleghani and Shohada Hospitals, Tehran, Iran, from 2015 to 2018. Additionally, 20 FFPE samples with normal pathologic reports assigned were added as the control group. Tumour and ADJ normal resected sections are all confirmed by two expert pathologists. The inclusion criteria were considered as: (I) All entered CRC patients underwent colorectal surgery at the Taleghani or Shohada Hospitals; (II) All participants were followed up for 3 years and their clinicopathological reports were available; (III) None of those patients included received chemotherapy or radiotherapy before surgery. The exclusion criteria were determined as: (I) A history of preoperative radio-chemotherapy before surgery; (II) A history of familial polyposis and hereditary nonpolyposis CRC syndrome; (III) No valid clinical and pathological information was available, or the specimen quality or quantity was not acceptable.
The study was approved by the local research ethics committee of the Research Centre for Gastroenterology and Liver Diseases of Shahid Beheshti, University of Medical Sciences, Tehran (No. 136-1395). The Declaration of Helsinki as a statement of ethical principles for medical research was considered. Written informed consent was obtained from all patients and donors included in the study.
2.2. Clinicopathological Data
The present study consisted of 77 (63.1%) men and 45 (36.9%) women, with an age cut-off at 60 years, where 45% were designated under 60 and near 55% over. All patient’s demographic and clinicopathological parameters are presented in detail in Table 1. All clinical and histopathological reports were assigned by two expert pathologists.

Table 1.
Clinicopathological data of the CRC included patients.
2.3. RNA Isolation and cDNA Synthesis
Then, 12 µm thick sections were prepared from both tumour and normal paraffin-embedded tissue blocks. Sections were deparaffinized in 180 µL xylene and then rehydrated by twice incubating in 850 µL absolute ethanol for 3 min followed by PBS washing. Total RNA was extracted from each air-dried deparaffinized section of the tumour and adjacent normal tissues, 10 cm away from the canonical tumour region via RNeasy FFPE kit (Qiagen, Hilden, Germany) as per the manufacturer’s protocol and was stored at −80 °C until use. The yield of extracted RNA and the A260/A280 purity ratios were measured using Nanodrop (ND-1000 spectrophotometer-Thermo Fisher, Waltham, MA, USA). RNA integrity was determined by electrophoresis on a 1% agarose gel. QIAxcel capillary electrophoresis with a high-resolution cartridge, 25–500 nucleotide molecular marker, and 15–156 nucleotide align marker for detachment of segments generated by PCR was used for RNA quality measurement with new RNA Integrity Score. cDNA was synthesized using the PrimeScript 1st strand cDNA Synthesis Kit (Takara, Dalian, China) as per the manufacturer’s instructions.
2.4. Real Time PCR Analysis
For gene expression quantification, real-time-PCR was performed in an Applied Biosystems 7500 Real-Time PCR System using an SYBR Green Real-Time PCR Master Mix (Takara, Dalian, China). Specific primers are listed in Table 2. Thermal cycling conditions were as follows: 30 s at 95 °C, 95 °C for 5 s, 58 °C for 34 s, and a primer extension 60 °C for 34 s and 15 s at 95 °C, for 40 cycles. Relative expression of the selected genes was measured by normalizing to β-globin via the 2−ΔΔCt method.

Table 2.
Real-time primer sequences.
2.5. Immunohistochemistry and Evaluation of Staining
Immunohistochemistry (IHC) was used to evaluate the canonical TGF-β signalling pathway proteins including TGF-β, Smad2, and Smad4. In brief, 4-micron thick sections of paraffin-embedded tumoral tissues were laid on glass slides coated with Poly-L-lysine and incubated at 60 °C for 15 min; then the sections were deparaffinized and dehydrated by xylene and ethanol, and retrieval antigen was performed in a microwave (900 W, 27 min) by adding HCL to change pH to the extreme alkaline (pH = 9) and coolness. Hydrogen peroxidase 10% was used to block endogenous peroxidase activity and, thereafter, was rinsed in deionized water. The samples were blocked using blocking serum. Next, the prepared slides were incubated with the Smad2 (phospho S467, Abcam, Cambridge, UK), Smad4 (Anti Smad4-Phospho-T277Abgent), and anti-TGF-β (antibody-ab92486) antibodies for 30 min at 37 °C. The Mouse/Rabbit PolyVue Plus HRP/DAB Detection System (Diagnostic BioSystems, Pleasanton, CA, USA) was used for primary antibodies visualization according to the manufacturer’s instructions.
The slides were examined under a light microscope (Nikon, Tokyo, Japan) by two pathologists who were unaware of the clinical and genetic data. Mean values were approximated through the scanning of the entire tissue parts of all samples via two graded scales: negative, <10%, and positive, >10%. The positive controls were the following: (a) a healthy colonic tissue was taken as an internal control for β-globin IHC, and (b) a histologically diagnosed part of colon cancer tissues for nuclear positivity by β-globin IHC. Negative controls were obtained by omitting the primary antibody.
The immunoreactivity of each slide was scored based on the Allred scoring system. Expression of TGF-β in tumour stroma and simultaneous expression of pSmad2 and Smad4 in the nucleus were assessed. Tumours were thus divided into two groups, including TGF-β positive, in which all the three genes were upregulated, and TGF-β negative, in which all the genes or at least two were downregulated.
2.6. Statistical Analysis
All gene expression experiments were performed in triplicate and repeated three times. The Relative Expression Software Tool (REST) was applied for the calculation of fold changes in gene expression. Statistical Package for Social Sciences SPSS (version 21) (SPSS Inc., Chicago, IL, USA) and Graph Pad Prism 8.0 software were used for analysing released data. The efficiency of gene expressions was assessed by the linear regression method. The Pearson, Chi-squared test (χ2), and Fisher’s exact test were used to determine the correlations between expressions and clinicopathological parameters in colorectal cancer patients. Non-parametric analysis (Mann–Whitney u-test) was used to compare gene expression in two different categories and one-way ANOVA (Kruskal–Wallis tests) was compared among more than two series. Survival analysis graphs were drawn by the Kaplan–Meier method. The p-value below 0.05 (p < 0.05) was considered statistically significant.
3. Results
3.1. High Expression of CTGF Contributes to Poor Prognosis in Colorectal Cancer Patients
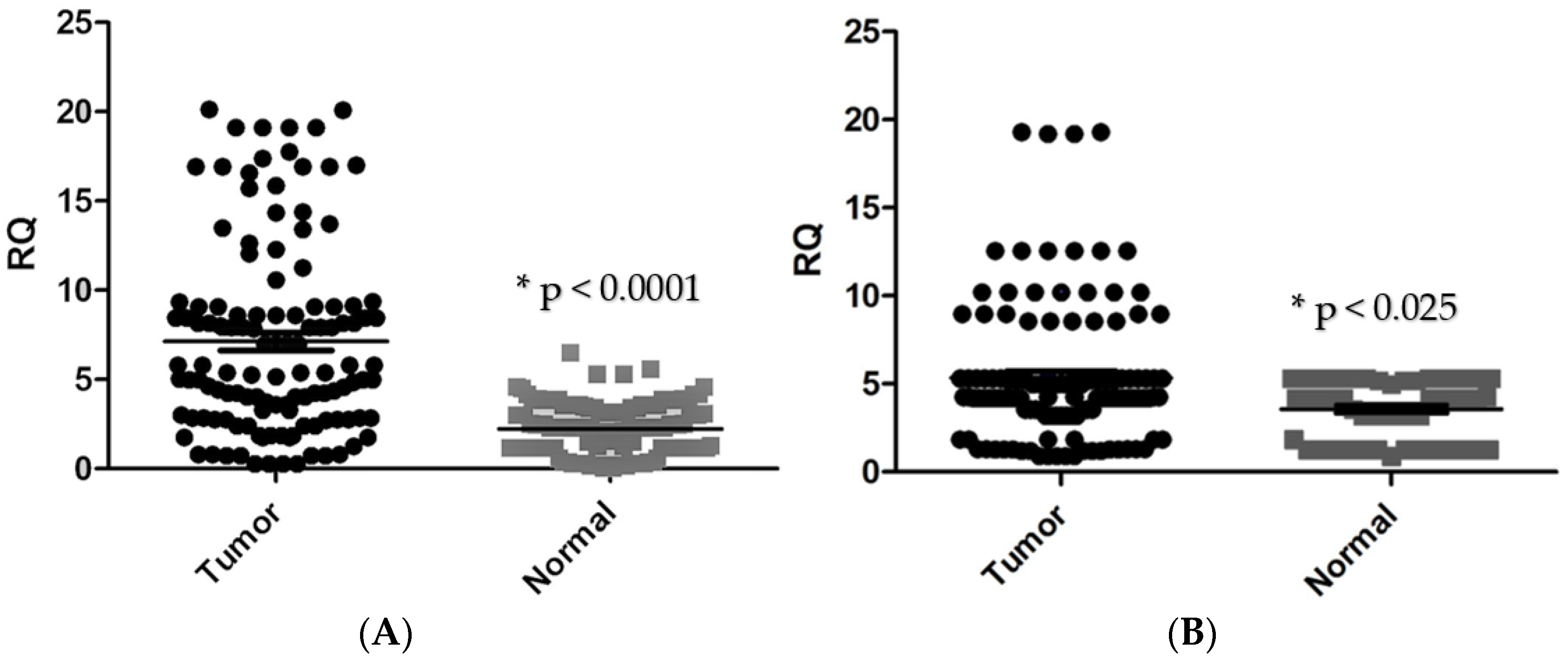
CTGF mRNA expression in tumour tissues was found to be significantly higher in CRC tissues in comparison with adjacent normal tissues (p < 0.0001, Figure 1A). CTGF expression was significantly correlated with tumour location (p < 0.003), tumour differentiation (p < 0.0002), lymph node metastasis (p < 0.005), distant metastasis (p < 0.044), and tumour stage (p < 0.0001). However, no significant association was found between the expression level of CTGF and tumour size. Additionally, significant upregulation of CTGF was found in stage IV of disease and CRC samples with positive lymph node metastasis (Figure 2, Table 3).
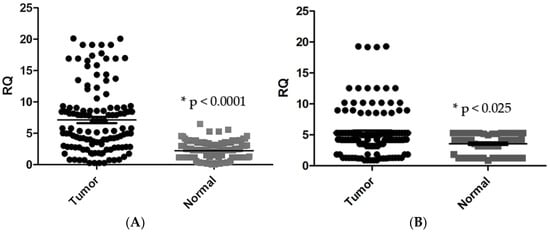
Figure 1.
(A) Comparison of CTGF mRNA expression in tumour and normal tissues (* p < 0.0001). (B) MMP-1 mRNA expression is compared in tumour and normal tissues. (* p < 0.025). * RQ related to relative quantification.
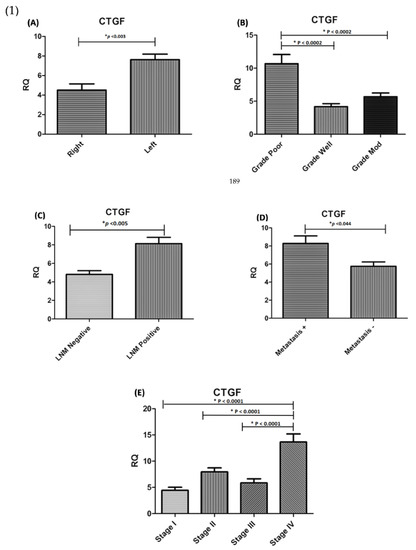
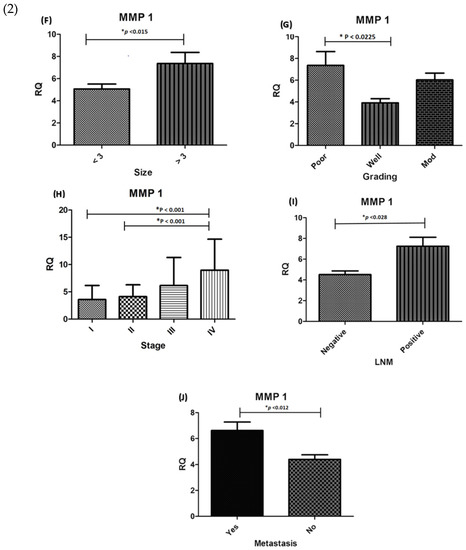
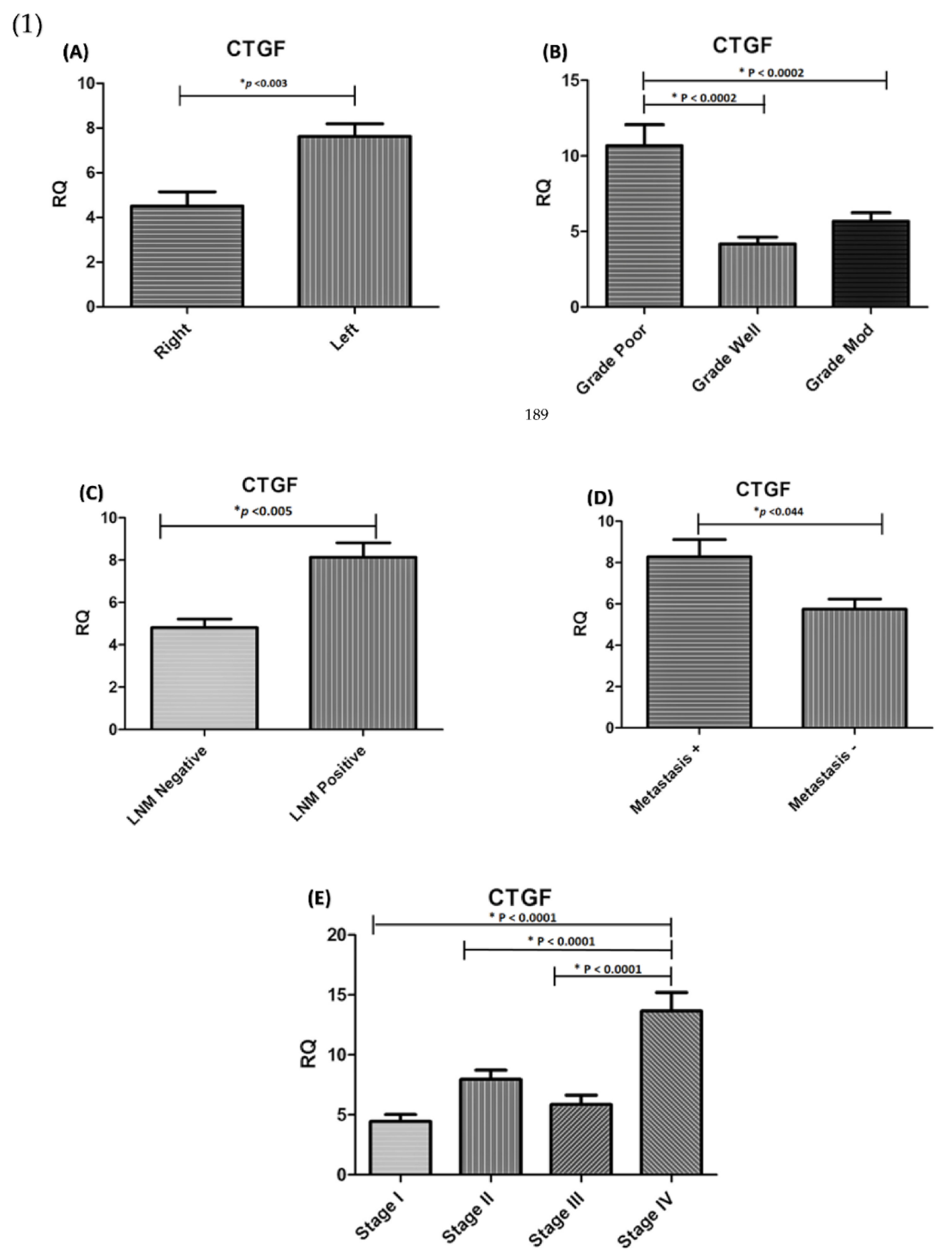
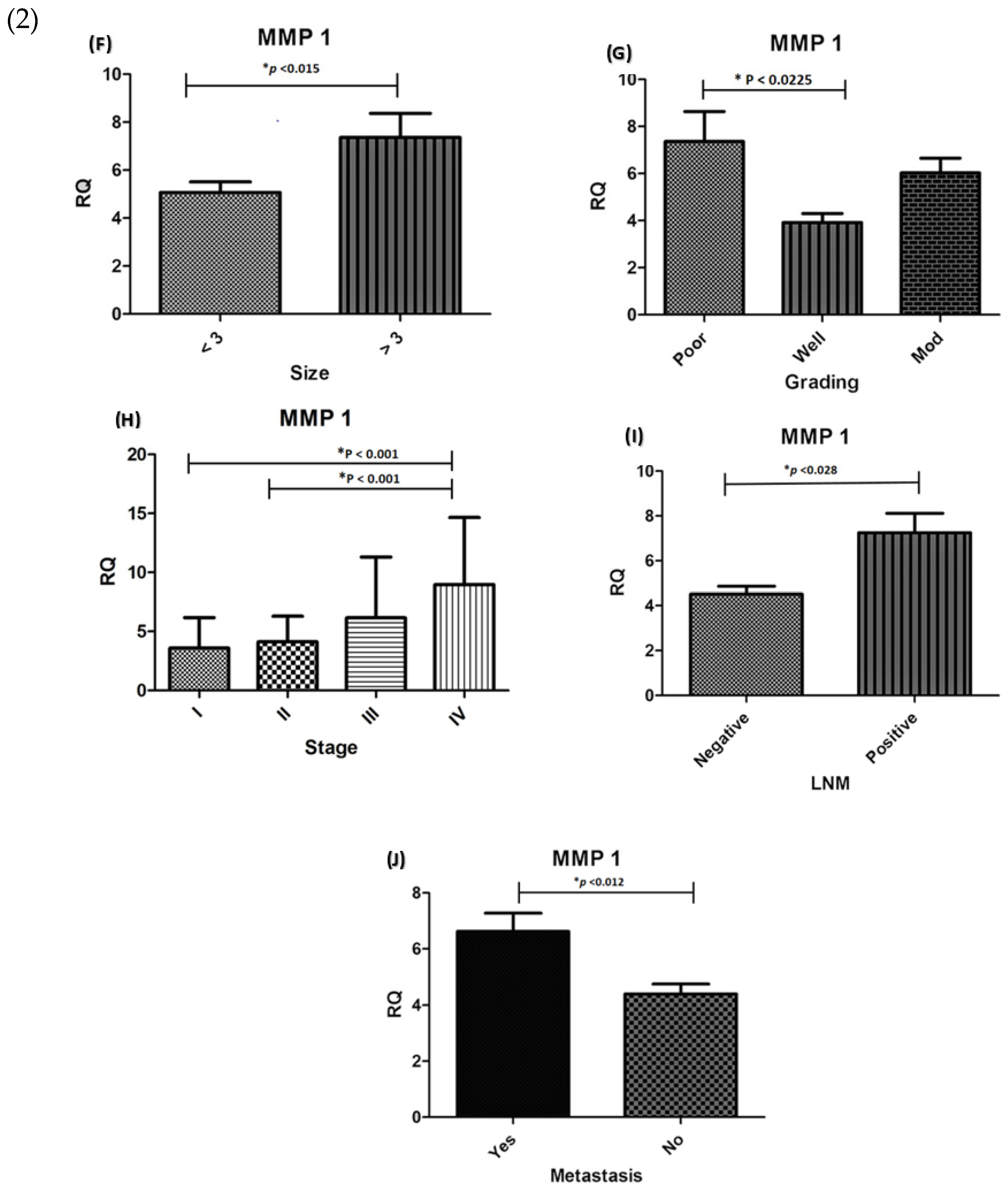
Figure 2.
(1) Expression association between CTGF and clinicopathological parameters in CRC. Significant differences were seen in (A) Tumour location (p < 0.003); (B) Tumour grading (p < 0.0002); (C) lymph node metastasis (p < 0.005) and (D) distant metastasis (p < 0.044); (E) Tumour stage (p < 0.001).). (2) Association between MMP-1 expression level and clinicopathological parameters in CRC patients. Significant correlation was observed between MMP-1 expression and (F) Tumour size (p < 0.015) (G) Tumour differentiation (p < 0.022), (H) Tumour stage (p < 0.001), (I) lymph node metastasis (p < 0.028), and (J) Metastasis (p < 0.012). * RQ related to Relative quantification.

Table 3.
Relationship between the expression of TGF-β signalling pathway and clinicopathological features.
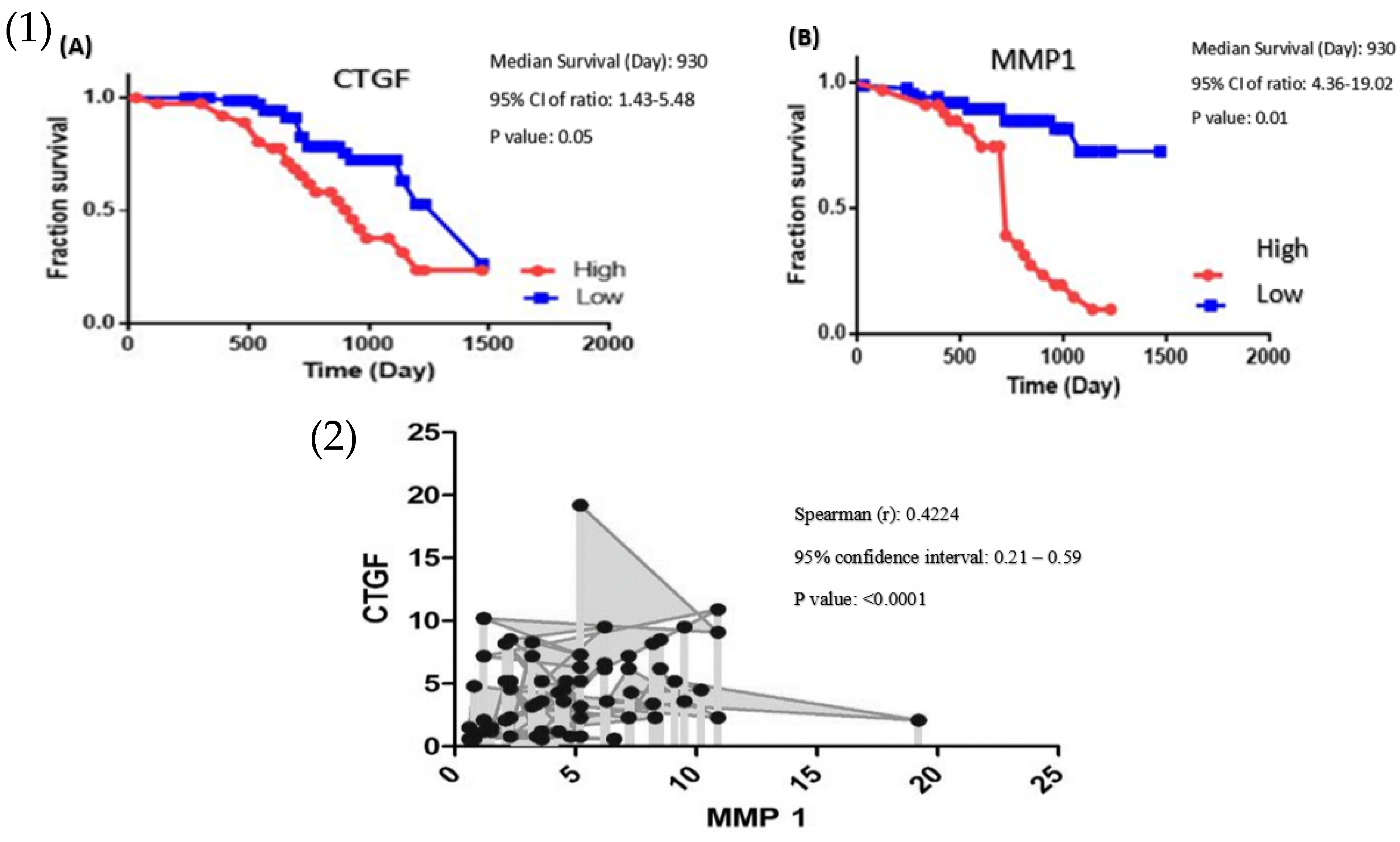
Kaplan–Meier analysis for plotting survival curve data was used for CRC patients’ survival rate, these data documented that high CTGF expression was significantly related to poor survival (p < 0.01) (Figure 3).
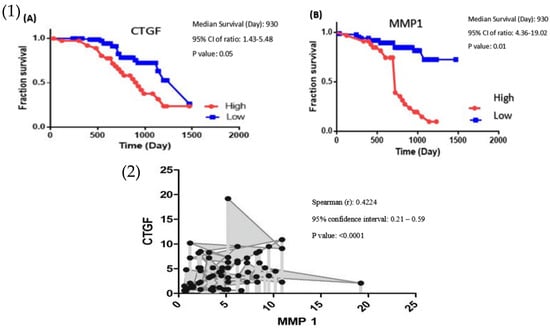
Figure 3.
(1) Log Rank Test demonstrate the association between survival rate and the expression level of (A) CTGF and (B) MMP-1 in patients with colon cancer. (2) Association between the CTGF and MMP-1 expression level (Spearman’s rank test, r = 0.4224, p < 0.0001, n = 81). r = Spearman’s rank, n = Number of XY Pairs.
3.2. MMP-1 Is Overexpressed in Colorectal Carcinoma Samples and Is Related to Poor Prognosis in CRC Patients
The mRNA expression level of MMP-1 was significantly higher in tumour tissues compared with adjacent normal tissues (p < 0.0253, Figure 1B). Our data demonstrated a significant association between MMP-1 mRNA level and clinicopathological parameters, including tumour size (p < 0.0153), cellular differentiation (p < 0.0225), tumour stage (p < 0.0016), lymph node metastasis (p < 0.0285), and distant metastasis (p < 0.0128, Figure 2). However, no significant relationship was observed between MMP-1 expression and tumour location (p < 0.593). The MMP-1 expression was significantly higher in CRC cases at stage IV and the up-regulation of MMP-1 was found in cases with lymph node metastasis as well as distant metastasis. The Kaplan–Meier analysis graph was plotted to correlate patient survival through the expression level of MMP-1. The overexpression of MMP-1 was significantly linked to a lower survival rate compared with cases with downregulation of MMP-1 (p < 0.05) (Figure 3).
3.3. CTGF Increases MMP-1 Expression in Colorectal Carcinomas
Spearman test for determining the correlation between designated genes was applied. A significant correlation was observed between the expression of levels MMP-1 and CTGF in CRC patients (Spearman’s rank test, r = 0.4224, p < 0.0001, Figure 3).
3.4. TGF-β Signalling Pathway Has a Multifaceted Role in Colorectal Cancer
We analysed and scored expression levels of TGF-β, Smad2, and Smad4 proteins in 122 FFPE specimens by IHC staining and investigated the association between protein expression and the clinicopathologic characteristics of CRC patients.
Based on our data, 66.40% (81 cases) of tumours had high expression of Smad2 followed by positive TGF-β signalling (described above and 33.60% (41 cases) of tumours represented a low expression of Smad2 with negative TGF-β signalling through cytoplasmic staining. All positive samples had a homogeneous pattern in staining. Figure 4A,B illustrates the low and high expression of Smad2 in the cytoplasm.
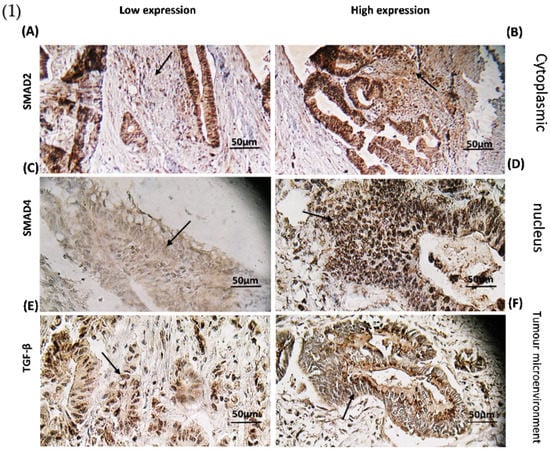
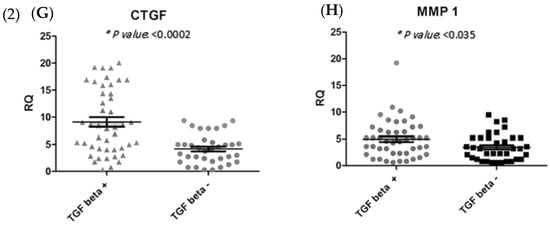
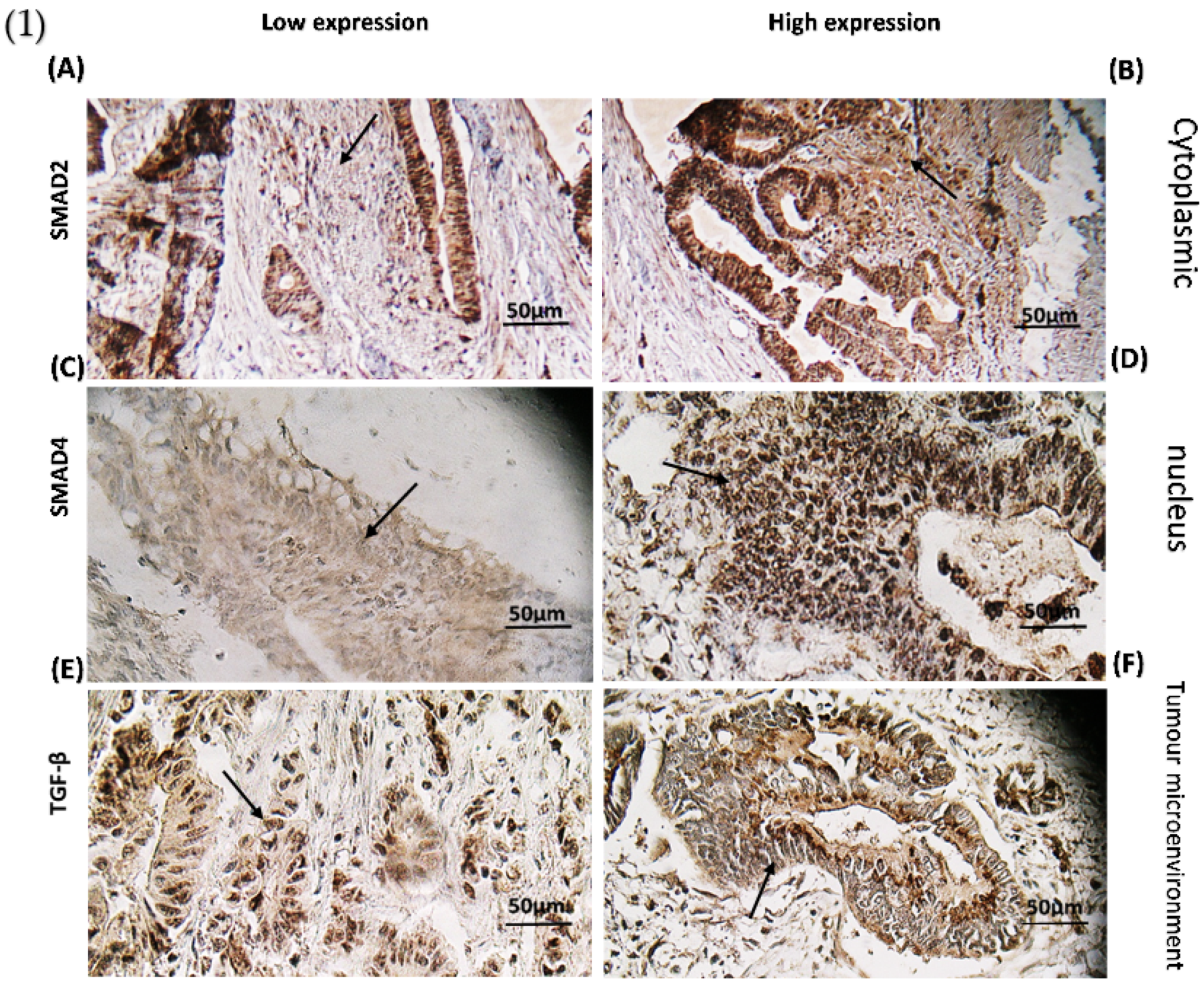
Figure 4.
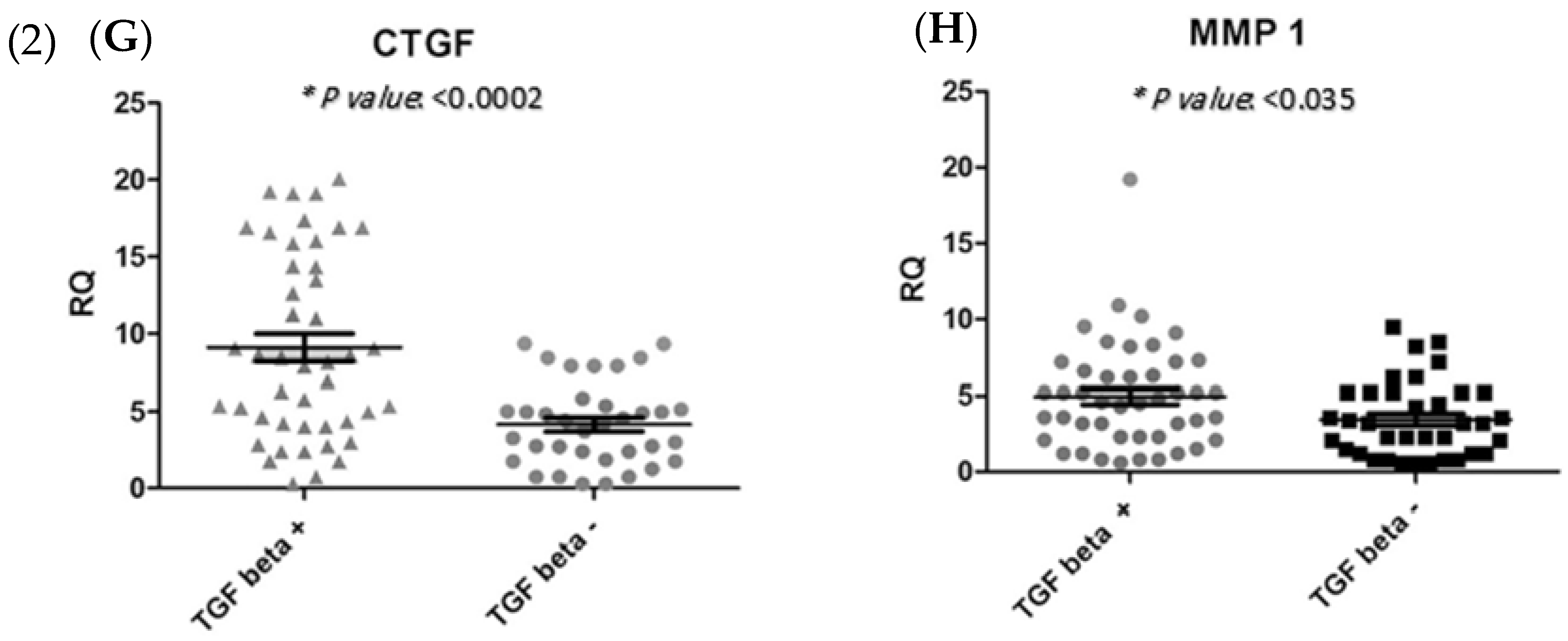
(1) Immunohistochemical staining (A) Smad2 cytoplasmic staining low expression; (B) Smad2 high expression specimen; (C) Smad4 nucleus staining low expression; (D) Smad4 high expression specimen; (E) IHC staining of TGF-β in Low expressed sample; and (F) High TGF-β expression. (2) Association between (G) CTGF expression level (* p < 0.0002) and (H) MMP-1 (* p < 0.035) through TGF-β pathway. * RQ related to Relative quantification.
Moreover, 82.52% (104 cases) of tumours with positive TGF-β signalling showed a high expression of Smad4 in the nucleus and 17.48 % (18 cases) of tumours indicated low expression of Smad4 with negative TGF-β signalling. All positive samples had a homogeneous pattern of staining. Figure 4C,D shows differences in Smad4 expression levels between low and high groups.
TGF-β as a nucleocytoplasmic shuttle was expressed in the cytoplasm and nucleus. Figure 4E,F depicts images of TGF-β protein expression 72.13% (88 cases) of participating patients illustrated high expression of TGF-β protein in the tumour microenvironment, while the rest 27.87% (34 cases) are categorized in low TGF-β expression category.
The obtained data from the current study revealed a significant correlation between positive TGF-β signalling and lymph node metastasis, distant metastasis, tumour stage, and overall survival (p < 0.001). However, no significant association was seen between age, gender, tumour location (p = 0.826), tumour size (p = 0.653), and tumour differentiation (p = 0.847).
3.5. The Effect of CTGF and MMP-1 mRNA Expression on TGF-β Protein Expression
As shown in Figure 4, mRNA expression levels of CTGF and MMP-1 were determined in groups with increased TGF-β protein and decreased TGF-β protein levels. The gene expressions levels of CTGF and MMP-1 were significantly increased (p = 0.0002, p = 0.0356, respectively) in tumours with a high level of TGF-β protein (Figure 4).
4. Discussion
The detection of CRC progression using biomarkers and novel approaches for early diagnosis/improved treatment is essential to reducing mortality rates in CRC [19]. In this study, we investigated the correlation of the canonical TGF-β signalling pathway with its downstream genes, including CTGF and MMP-1, and their effect on clinicopathological features in CRC patients. TGF- β signalling is known as a critical signalling pathway that has a tumour-suppressive role in the early stages of cancer and pro-metastatic function in advanced stages [20,21,22,23]. The Smad proteins, such as Smad2 and Smad4, are master regulators for the canonical TGF-β signalling [24,25,26]. Furthermore, the TGF-β signalling pathway regulates many downstream target genes, which are involved in cell growth, differentiation, apoptosis, and invasion [27]. TGF-β signalling activation in tumour microenvironments induces CMS4 mesenchymal phenotype (Malignant CRC phenotype), which is linked to worsening outcomes, via differentiation of mesenchymal stem cells to cancer-associated fibroblast, induction of JAK/STAT pathway, and suppression of tumour immunity [28,29]. Moreover, there are many challenges in cancer treatment using TGF-β signalling inhibition because of some side effects on patients. Hence, downstream pathways of TGF-β signalling could be utilized for targeted therapy with fewer side effects [30,31]. In the current investigation, obtained results confirmed previous findings of the association of activated TGF- β signalling with lymph nodes metastasis, distant metastasis, advanced stages, and poor survival [32,33,34]. CTGF gene is a downstream target of TGF-β signalling in the canonical pathway and a regulator of cell proliferation in pathological conditions [35,36]. The upregulation of CTGF leads cancerous cells to angiogenesis and metastasis, also the high expression level of CTGF has been detected in CMS4 colorectal cancer [10,37,38]. In this study, we found significant correlations between the upregulation of CTGF, tumour location, cellular differentiation, lymph nodes metastasis, distant metastasis, advanced stages, and poor survival, also upregulation of CTGF at stage IV was associated with lymph nodes metastasis; these results agree with previous studies [10,39]. Indicating that TGF-β signalling is an important regulator for CTGF expression and CRC development.
The MMP-1 protein is one of the most important matrix metalloproteinases, and a member of the collagenases, with a key role in CRC initiation and development [13,40]. In addition, CTGF can stimulate angiogenesis in cancerous cells via the regulation of MMPs [41,42,43,44]. In the current investigation, our data illustrated the overexpression of MMP-1 in CRC specimens, which confirmed the oncogenic role of MMP-1 in colorectal cancers [45]. Additionally, we observed that upregulation of MMP-1 was associated with tumour size, cellular differentiation, lymph nodes metastasis, distant metastasis, and poor survival. These results are again in agreement with previous findings [14,40,46,47,48]. It is worth mentioning that we found a significant correlation in MMP-1 overexpression among stage IV, lymph nodes metastasis, and distant metastasis. Therefore, it seems that MMP-1 plays a critical role in metastasis in advanced stages of CRC. It is interesting to note that, CTGF and MMP-1 presented a significant positive association with each other also. This result was reported for the first time in the Iranian population. Additionally, statistical analysis showed that the expression levels of CTGF and MMP-1 have been affected by TGF-β signalling. Based on the data presented here, CTGF and MMP-1 are regulated by TGF-β signalling, and TGF-β plays a crucial function through angiogenesis and malignancy in CRC.
5. Conclusions
In conclusion, this study showed that canonical TGF-β signalling is a master regulator for MMP-1 and CTGF and can control malignancy features in CRC. Moreover, downstream genes of TGF-β signalling, such as MMP-1 and CTGF, could be used as biomarkers for CRC detection to improve early diagnosis and patient outcomes.
Author Contributions
Conceptualization, S.H., Z.P., M.A.B. and E.N.M.; data curation, N.P. and E.N.M.; formal analysis, N.P. and L.R.; funding acquisition, M.A.B. and H.A.A.; investigation, E.N.M.; methodology, M.A.B., N.P., L.R., Z.P. and E.N.M.; project administration, E.N.M.; resources, H.A.A.; software, N.P.; supervision, E.N.M.; validation, N.F. and Z.P.; Lab working, S.H., Z.P. and M.M.; writing—original draft, N.P., Z.M., B.K., G.S., M.A.B. and E.N.M.; writing—review and editing, N.P., Z.P., M.H., L.R., M.F., M.A.B. and E.N.M. All authors have read and agreed to the published version of the manuscript.
Funding
This project was completely supported and funded by the Gastroenterology and Liver Diseases Research Center, Research Institute for Gastroenterology and Liver Diseases, Shahid Beheshti University of Medical Sciences, with Grant Nos. 858 and 988, and the Medical Ethical Committee of the RCGLD, with Ethics No. IR.SBMU.RIGLD.REC.1396.947.
Institutional Review Board Statement
This study was approved by the ethical committee (Ethics No. IR.SBMU.RIGLD.REC.1396.947) of the Research Institute for Gastroenterology and Liver Disease, Shahid Beheshti University of Medical Sciences, Tehran, Iran.
Informed Consent Statement
Informed consent was obtained from all subjects involved in the study.
Acknowledgments
The authors would like to thank all the staff of the Cancer Department in the Research Institute for Gastroenterology and Liver Diseases, Shahid Beheshti University of Medical Sciences, Tehran, Iran.
Conflicts of Interest
The authors declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
References
- Rawla, P.; Sunkara, T.; Barsouk, A. Epidemiology of colorectal cancer: Incidence, mortality, survival, and risk factors. Prz. Gastroenterol. 2019, 14, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Bhandari, A.; Woodhouse, M.; Gupta, S. Colorectal cancer is a leading cause of cancer incidence and mortality among adults younger than 50 years in the USA: A SEER-based analysis with comparison to other young-onset cancers. J. Investig. Med. 2017, 65, 311–315. [Google Scholar] [CrossRef] [PubMed]
- Syed, V. TGF-β Signaling in Cancer. J. Cell. Biochem. 2016, 117, 1279–1287. [Google Scholar] [CrossRef]
- Ji, Q.; Liu, X.; Han, Z.; Zhou, L.; Sui, H.; Yan, L.; Jiang, H.; Ren, J.; Cai, J.; Li, Q. Resveratrol suppresses epithelial-to-mesenchymal transition in colorectal cancer through TGF-β1/Smads signaling pathway mediated Snail/E-cadherin expression. BMC Cancer 2015, 15, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Soleimani, A.; Pashirzad, M.; Avan, A.; Ferns, G.A.; Khazaei, M.; Hassanian, S.M. Role of the transforming growth factor-β signaling pathway in the pathogenesis of colorectal cancer. J. Cell. Biochem. 2019, 120, 8899–8907. [Google Scholar] [CrossRef] [PubMed]
- Katz, L.H.; Likhter, M.; Jogunoori, W.; Belkin, M.; Ohshiro, K.; Mishra, L. TGF-β signaling in liver and gastrointestinal cancers. Cancer Lett. 2016, 379, 166–172. [Google Scholar] [CrossRef] [PubMed]
- Cheruku, H.; Mohamedali, A.; Cantor, D.; Tan, S.H.; Nice, E.; Baker, M. Transforming Growth Factor-β, MAPK and Wnt Signaling Interactions in Colorectal Cancer. EuPA Open Proteom. 2015, 8, 104–115. [Google Scholar] [CrossRef]
- Wells, J.E.; Howlett, M.; Cole, C.H.; Kees, U.R. Deregulated expression of connective tissue growth factor (CTGF/CCN2) is linked to poor outcome in human cancer. Int. J. Cancer 2015, 137, 504–511. [Google Scholar] [CrossRef]
- Inman, G.J. Switching TGFβ from a tumor suppressor to a tumor promoter. Curr. Opin. Genet. Dev. 2011, 21, 93–99. [Google Scholar] [CrossRef]
- Ubink, I.; Verhaar, E.R.; Kranenburg, O.; Goldschmeding, R. A potential role for CCN2/CTGF in aggressive colorectal cancer. J. Cell Commun. Signal. 2016, 10, 223–227. [Google Scholar] [CrossRef]
- Shirin, M.; Madadi, S.; Peyravian, N.; Pezeshkian, Z.; Rejali, L.; Hosseini, M.; Moradi, A.; Khanabadi, B.; Sherkat, G.; Aghdaei, H.A.; et al. A linkage between effectual genes in progression of CRC through canonical and non-canonical TGF-β signaling pathways. Med. Oncol. 2022, 39, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Conlon, G.A.; Murray, G.I. Recent advances in understanding the roles of matrix metalloproteinases in tumour invasion and metastasis. J. Pathol. 2019, 247, 629–640. [Google Scholar] [CrossRef] [PubMed]
- Said, A.H.; Raufman, J.-P.; Xie, G. The role of matrix metalloproteinases in colorectal cancer. Cancers 2014, 6, 366–375. [Google Scholar] [CrossRef]
- Bendardaf, R.; Buhmeida, A.; Ristamäki, R.; Syrjänen, K.; Pyrhönen, S. MMP-1 (collagenase-1) expression in primary colorectal cancer and its metastases. Scand. J. Gastroenterol. 2007, 42, 1473–1478. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Zheng, J.; Yu, J.; Wu, Y.; Guo, J.; Xu, Z.; Sun, X. Knockdown of MMP-1 inhibits the progression of colorectal cancer by suppressing the PI3K/Akt/c-myc signaling pathway and EMT. Oncol. Rep. 2020, 43, 1103–1112. [Google Scholar] [CrossRef]
- Lyall, M.S.; Dundas, S.R.; Curran, S.; Murray, G.I. Profiling markers of prognosis in colorectal cancer. Clin. Cancer Res. 2006, 12, 1184–1191. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.W.; Ko, Y.T.; Kim, N.K.; Chung, H.C.; Min, B.S.; Lee, K.Y.; Park, J.-P.; Kim, H. A comparative study of protein expression in primary colorectal cancer and synchronous hepatic metastases: The significance of matrix metalloproteinase-1 expression as a predictor of liver metastasis. Scand. J. Gastroenterol. 2010, 45, 217–225. [Google Scholar] [CrossRef]
- Wagenaar-Miller, R.A.; Gorden, L.; Matrisian, L.M. Matrix metalloproteinases in colorectal cancer: Is it worth talking about? Cancer Metastasis Rev. 2004, 23, 119–135. [Google Scholar] [CrossRef]
- Peyravian, N.; Larki, P.; Gharib, E.; Nazemalhosseini-Mojarad, E.; Anaraki, F.; Young, C.; McClellan, J.; Bonab, M.A.; Asadzadeh-Aghdaei, H.; Zali, M.R. The Application of Gene Expression Profiling in Predictions of Occult Lymph Node Metastasis in Colorectal Cancer Patients. Biomedicines 2018, 6, 27. [Google Scholar] [CrossRef]
- Muñoz, N.M.; Upton, M.; Rojas, A.; Washington, M.K.; Lin, L.; Chytil, A.; Sozmen, E.G.; Madison, B.B.; Pozzi, A.; Moon, R.T.; et al. Transforming growth factor beta receptor type II inactivation induces the malignant transformation of intestinal neoplasms initiated by Apc mutation. Cancer Res. 2006, 66, 9837–9844. [Google Scholar] [CrossRef]
- Itatani, Y.; Kawada, K.; Fujishita, T.; Kakizaki, F.; Hirai, H.; Matsumoto, T.; Iwamoto, M.; Inamoto, S.; Hatano, E.; Hasegawa, S.; et al. Loss of SMAD4 from colorectal cancer cells promotes CCL15 expression to recruit CCR1+ myeloid cells and facilitate liver metastasis. Gastroenterology 2013, 145, 1064–1075.e11. [Google Scholar] [CrossRef] [PubMed]
- Ramamoorthi, G.; Sivalingam, N. Molecular mechanism of TGF-β signaling pathway in colon carcinogenesis and status of curcumin as chemopreventive strategy. Tumor Biol. 2014, 35, 7295–7305. [Google Scholar] [CrossRef] [PubMed]
- Kumar Pandurangan, A.; Divya, T.; Kumar, K.; Dineshbabu, V.; Velavan, B.; Sudhandiran, G. Colorectal carcinogenesis: Insights into the cell death and signal transduction pathways: A review. World J. Gastrointest. Oncol. 2018, 10, 244. [Google Scholar]
- Zhao, M.; Mishra, L.; Deng, C.X. The role of TGF-β/SMAD4 signaling in cancer. Int. J. Biol. Sci. 2018, 14, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Samanta, D.; Datta, P.K. Alterations in the Smad pathway in human cancers. Front. Biosci. 2012, 17, 1281. [Google Scholar] [CrossRef] [PubMed]
- Sameer, A.S.; Abdullah, S.; Syeed, N.; Siddiqi, M.A. Colorectal cancer, TGF-signaling and SMADs. Int. J. Genet. Mol. Biol. 2010, 2, 101–111. [Google Scholar]
- Jung, B.; Staudacher, J.J.; Beauchamp, D. Transforming Growth Factor β Superfamily Signaling in Development of Colorectal Cancer. Gastroenterology 2017, 152, 36–52. [Google Scholar] [CrossRef]
- Tan, H.X.; Cao, Z.B.; He, T.T.; Huang, T.; Xiang, C.L.; Liu, Y. TGFβ1 is essential for MSCs-CAFs differentiation and promotes HCT116 cells migration and invasion via JAK/STAT3 signaling. Onco Targets Ther. 2019, 12, 5323–5334. [Google Scholar] [CrossRef]
- Otegbeye, F.; Ojo, E.; Moreton, S.; Mackowski, N.; Lee, D.A.; de Lima, M.; Wald, D.N. Inhibiting TGF-beta signaling preserves the function of highly activated, in vitro expanded natural killer cells in AML and colon cancer models. PLoS ONE. 2018, 13, e0191358. [Google Scholar]
- Huynh, L.K.; Hipolito, C.J.; Ten Dijke, P. A Perspective on the Development of TGF-β Inhibitors for Cancer Treatment. Biomolecules 2019, 9, 743. [Google Scholar] [CrossRef]
- Gachpazan, M.; Kashani, H.; Hassanian, S.M.; Khazaei, M.; Khorrami, S.; Ferns, G.A.; Avan, A. Therapeutic potential of targeting transforming growth factor-beta in colorectal cancer: Rational and progress. Curr. Pharm. Des. 2019, 25, 4085–4089. [Google Scholar] [CrossRef] [PubMed]
- Lampropoulos, P.; Zizi-Sermpetzoglou, A.; Rizos, S.; Kostakis, A.; Nikiteas, N.; Papavassiliou, A.G. Prognostic significance of transforming growth factor beta (TGF-β) signaling axis molecules and E-cadherin in colorectal cancer. Tumour Biol. 2012, 33, 1005–1014. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Liu, Z.; Tan, J.; Dong, H.; Zhang, X. Multispectral imaging reveals hyperactive TGF-β signaling in colorectal cancer. Cancer Biol. Ther. 2018, 19, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Fessler, E.; Drost, J.; Van Hooff, S.R.; Linnekamp, J.F.; Wang, X.; Jansen, M.; De Sousa E Melo, F.; Prasetyanti, P.R.; Ijspeert, J.E.; Franitza, M.; et al. TGFβ signaling directs serrated adenomas to the mesenchymal colorectal cancer subtype. EMBO Mol. Med. 2016, 8, 745–760. [Google Scholar] [CrossRef]
- Parada, C.; Li, J.; Iwata, J.; Suzuki, A.; Chai, Y. CTGF mediates Smad-dependent transforming growth factor β signaling to regulate mesenchymal cell proliferation during palate development. Mol. Cell. Biol. 2013, 33, 3482–3493. [Google Scholar] [CrossRef]
- Ihn, H. Pathogenesis of fibrosis: Role of TGF-beta and CTGF. Curr. Opin. Rheumatol. 2002, 14, 681–685. [Google Scholar] [CrossRef]
- Zhang, S.D.; McCrudden, C.M.; Yuen, H.F.; Leung, K.L.; Hong, W.J.; Kwok, H.F. Association between the expression levels of TAZ, AXL and CTGF and clinicopathological parameters in patients with colon cancer. Oncol. Lett. 2016, 11, 1223–1229. [Google Scholar] [CrossRef][Green Version]
- Lau, L.F.; Lam, S.C. The CCN family of angiogenic regulators: The integrin connection. Exp. Cell Res. 1999, 248, 44–57. [Google Scholar] [CrossRef]
- Ladwa, R.; Pringle, H.; Kumar, R.; West, K. Expression of CTGF and Cyr61 in colorectal cancer. J. Clin. Pathol. 2011, 64, 58–64. [Google Scholar] [CrossRef]
- Sunami, E.; Tsuno, N.; Osada, T.; Saito, S.; Kitayama, J.; Tomozawa, S.; Tsuruo, T.; Shibata, Y.; Muto, T.; Nagawa, H. MMP-1 is a prognostic marker for hematogenous metastasis of colorectal cancer. Oncology 2000, 5, 108–114. [Google Scholar] [CrossRef]
- Chu, C.Y.; Chang, C.C.; Prakash, E.; Kuo, M.L. Connective tissue growth factor (CTGF) and cancer progression. J. Biomed. Sci. 2008, 15, 675–685. [Google Scholar] [CrossRef] [PubMed]
- Chien, W.; O’Kelly, J.; Lu, D.; Leiter, A.; Sohn, J.; Yin, D.; Karlan, B.; Vadgama, J.; Lyons, K.M.; Koeffler, H.P. Expression of connective tissue growth factor (CTGF/CCN2) in breast cancer cells is associated with increased migration and angiogenesis. Int. J. Oncol. 2011, 38, 1741–1747. [Google Scholar] [CrossRef] [PubMed]
- Cutler, S.J.; Doecke, J.D.; Ghazawi, I.; Yang, J.; Griffiths, L.R.; Spring, K.; Ralph, S.J.; Mellick, A.S. Novel STAT binding elements mediate IL-6 regulation of MMP-1 and MMP-3. Sci. Rep. 2017, 7, 8526. [Google Scholar] [CrossRef]
- Quintero-Fabián, S.; Arreola, R.; Becerril-Villanueva, E.; Torres-Romero, J.C.; Arana-Argáez, V.; Lara-Riegos, J.; Ramírez-Camacho, M.A.; Alvarez-Sánchez, M.E. Role of Matrix Metalloproteinases in Angiogenesis and Cancer. Front. Oncol. 2019, 9, 1370. [Google Scholar] [CrossRef] [PubMed]
- Yeh, Y.-C.; Sheu, B.-S. Matrix metalloproteinases and their inhibitors in the gastrointestinal cancers: Current knowledge and clinical potential. Met. Med. 2014, 1, 3–13. [Google Scholar]
- Langenskiöld, M.; Ivarsson, M.L.; Holmdahl, L.; Falk, P.; Kåbjörn-Gustafsson, C.; Angenete, E. Intestinal mucosal MMP-1—A prognostic factor in colon cancer. Scand. J. Gastroenterol. 2013, 48, 563–569. [Google Scholar] [CrossRef]
- Shiozawa, J.; Ito, M.; Nakayama, T.; Nakashima, M.; Kohno, S.; Sekine, I. Expression of matrix metalloproteinase-1 in human colorectal carcinoma. Mod. Pathol. 2000, 13, 925–933. [Google Scholar] [CrossRef] [PubMed]
- Jonsson, A.; Falk, P.; Angenete, E.; Hjalmarsson, C.; Ivarsson, M.-L. Plasma MMP-1 Expression as a Prognostic Factor in Colon Cancer. J. Surg. Res. 2021, 266, 254–260. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).