Identification and Somatic Characterization of the Germline PTEN Promoter Variant rs34149102 in a Family with Gastrointestinal and Breast Tumors
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Recruitment
2.2. DNA Extraction and Sanger Sequencing
2.3. Next-Generation Sequencing
2.4. Sample Collection and Immunohistochemical (IHC) Analysis
2.5. Second-Hit Somatic Alteration Analysis
2.6. In Silico Analysis
3. Results
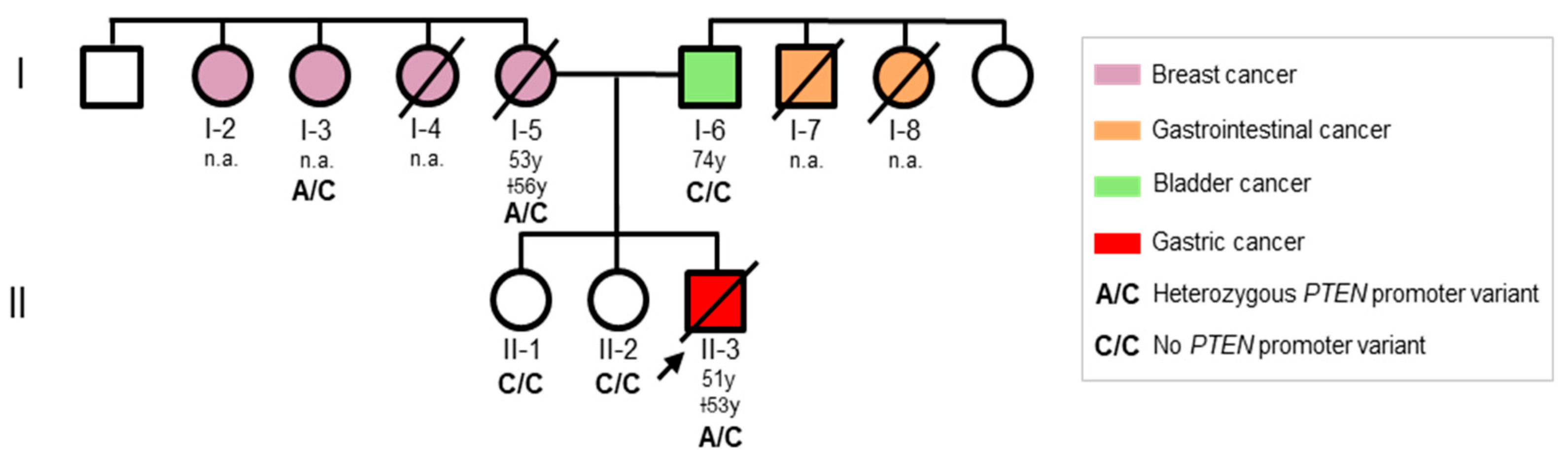
3.1. Medical History and Genetic Findings
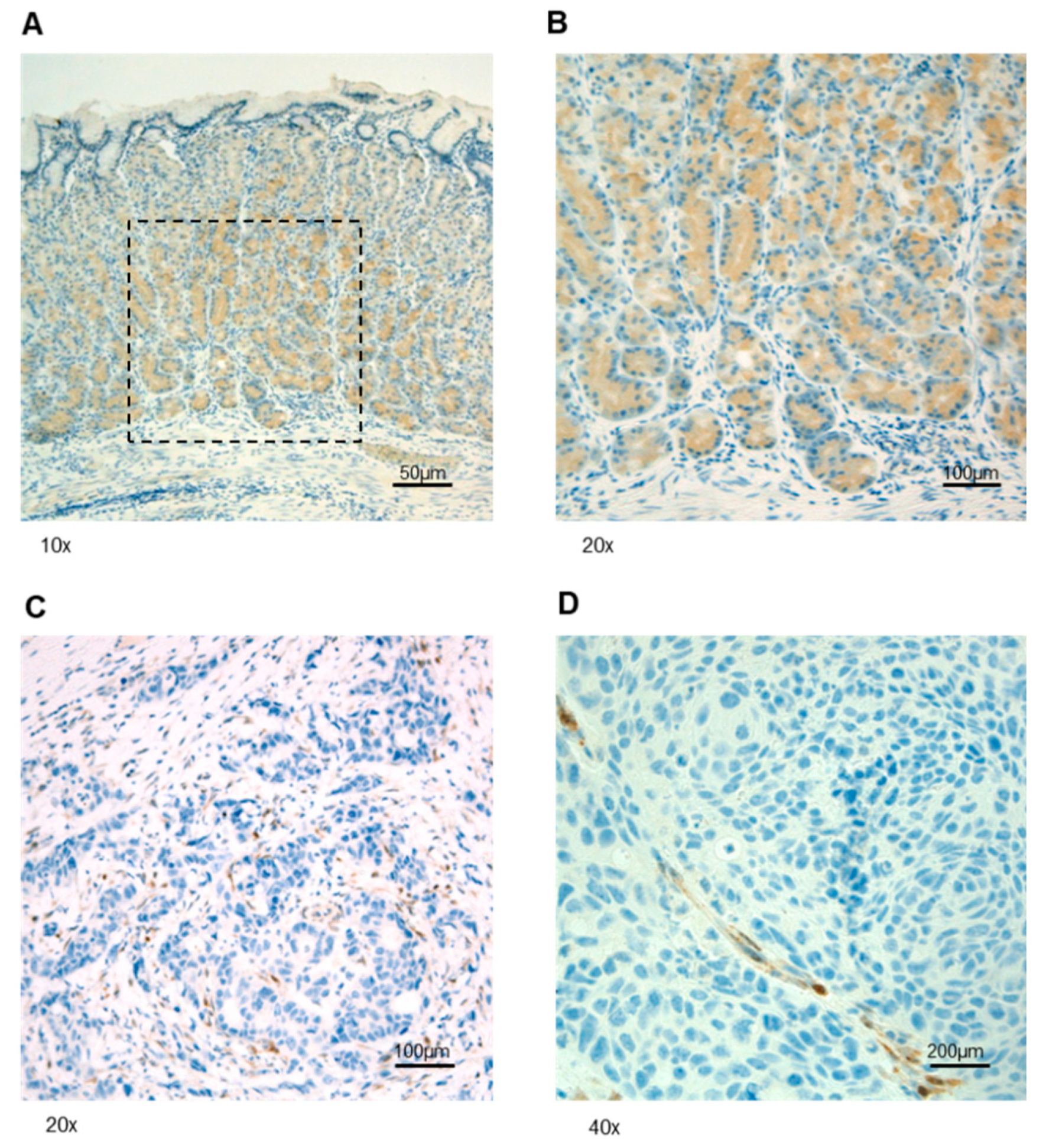
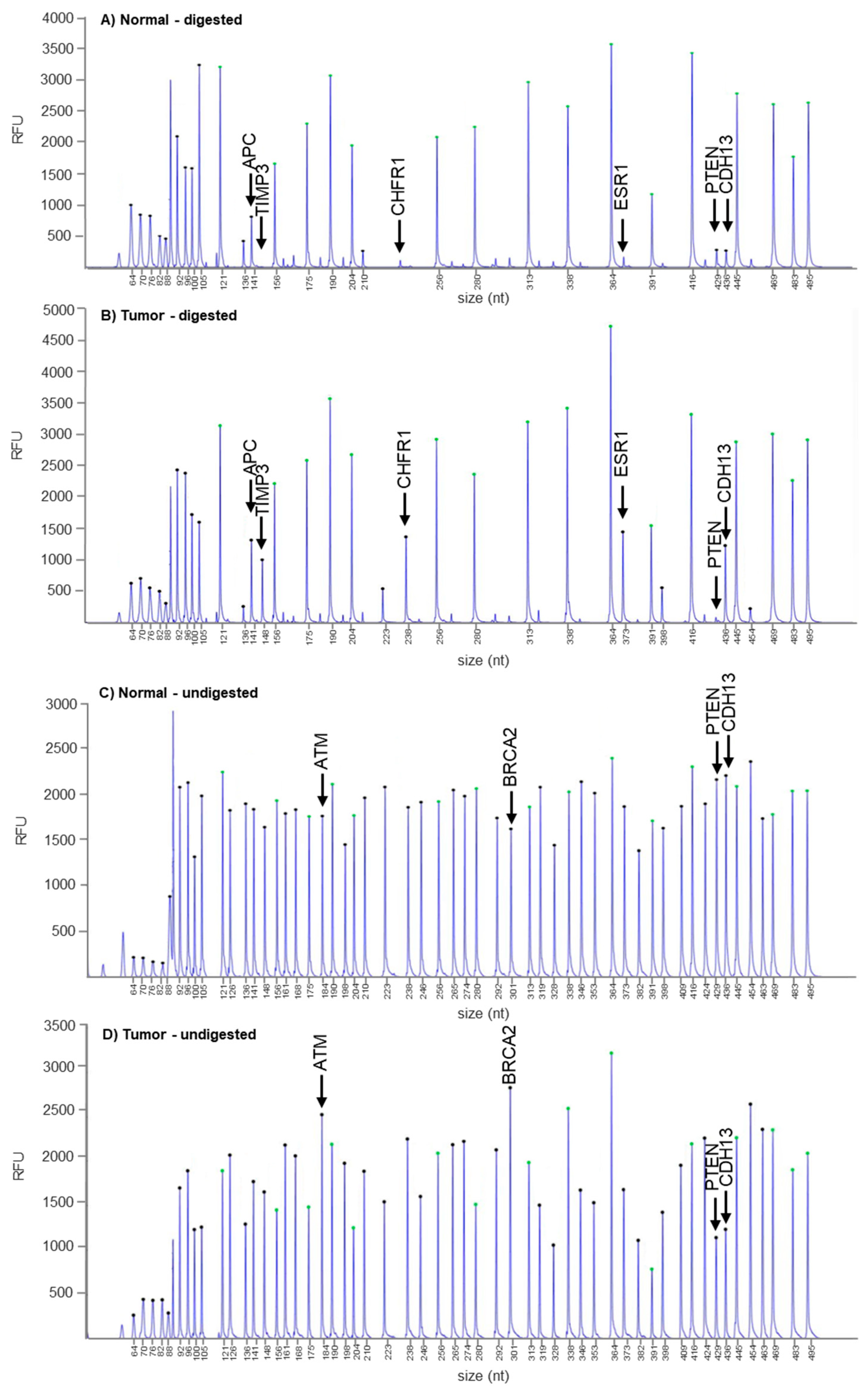
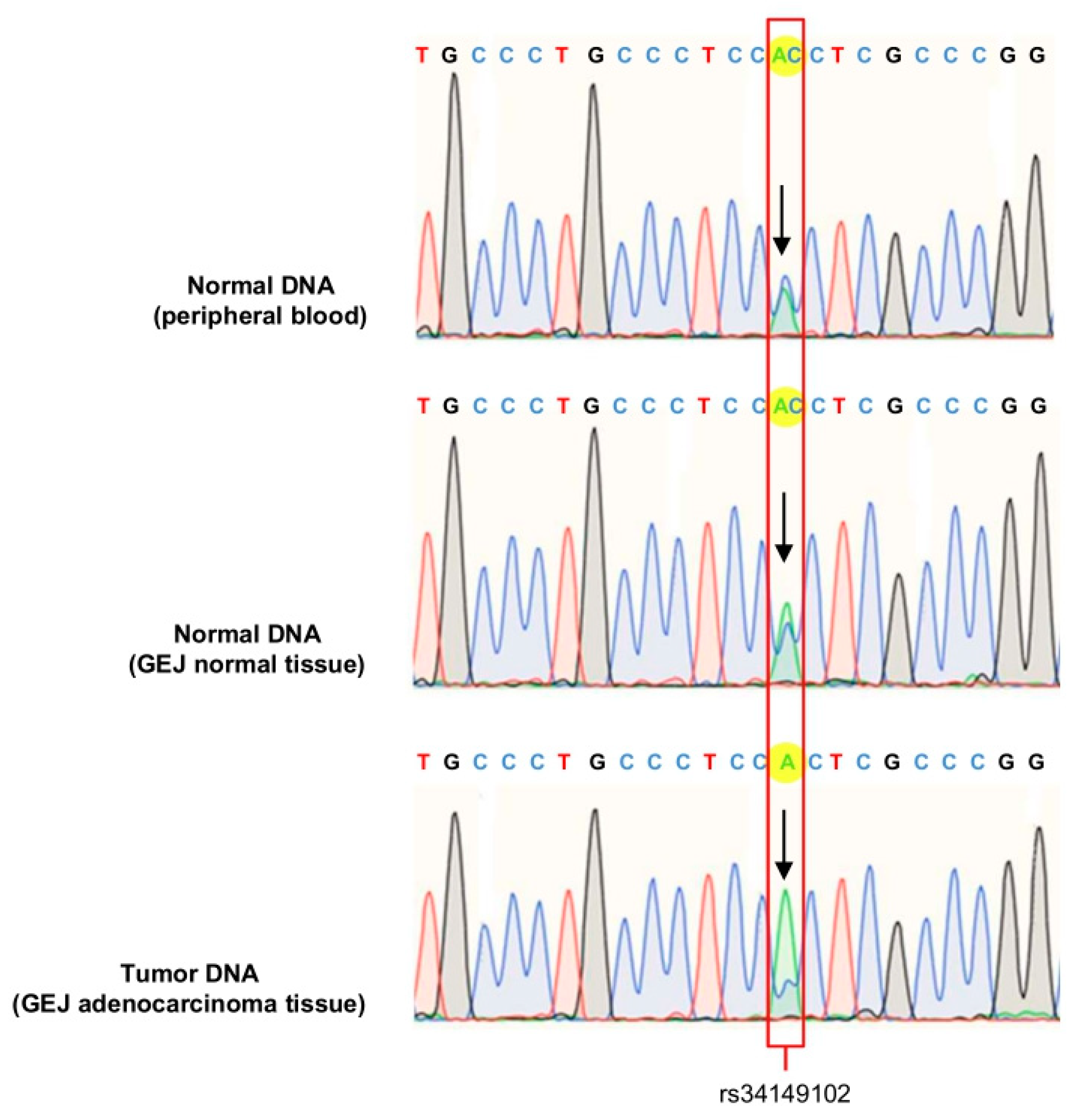
3.2. PTEN Somatic Alterations in GEJ Adenocarcinoma Samples
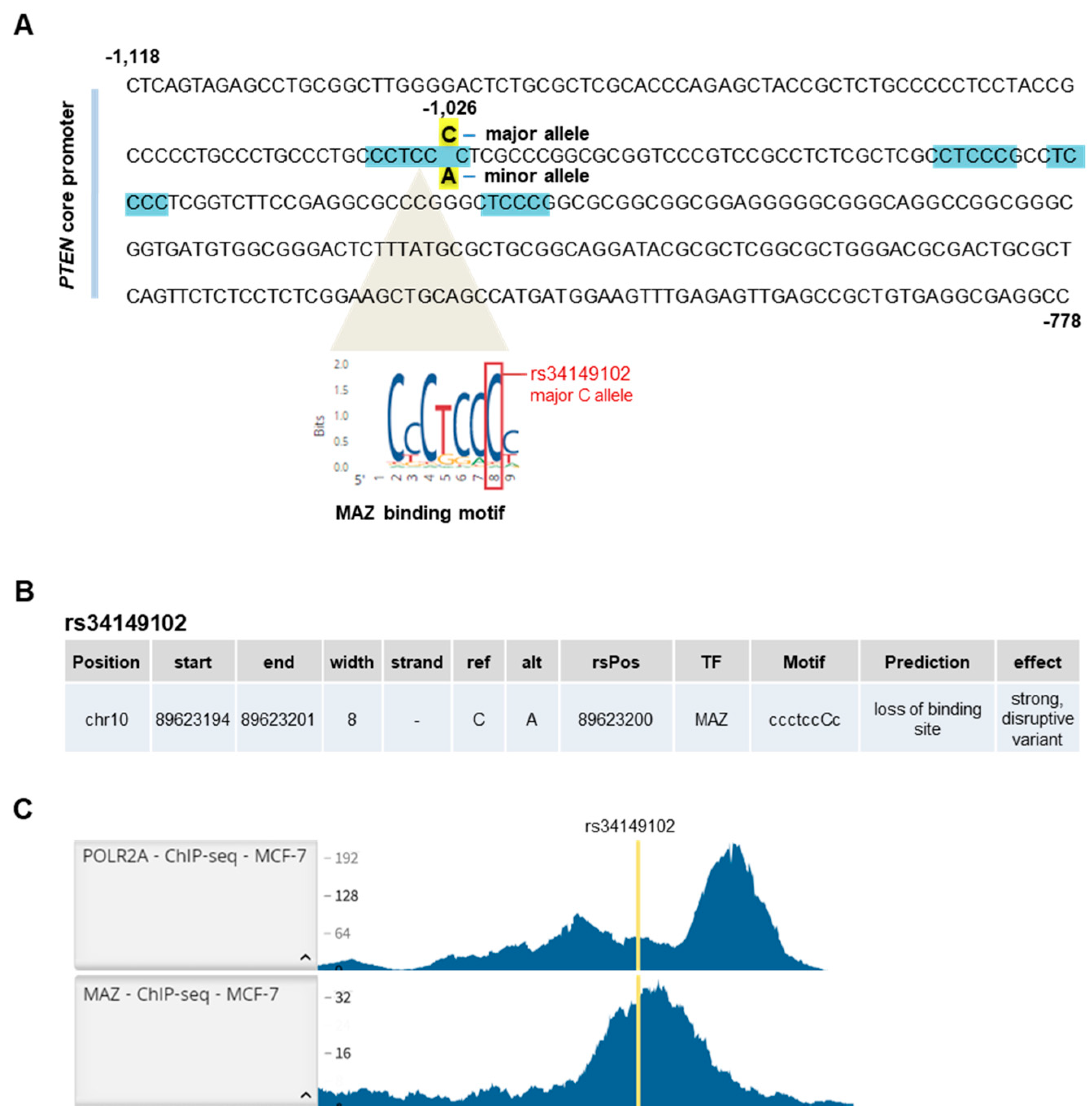
3.3. In Silico Analysis
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Claussnitzer, M.; Cho, J.H.; Collins, R.; Cox, N.J.; Dermitzakis, E.T.; Hurles, M.E.; Kathiresan, S.; Kenny, E.E.; Lindgren, C.M.; MacArthur, D.G.; et al. A Brief History of Human Disease Genetics. Nature 2020, 577, 179–189. [Google Scholar] [CrossRef] [PubMed]
- Loomans-Kropp, H.A.; Umar, A. Cancer Prevention and Screening: The next Step in the Era of Precision Medicine. NPJ Precis. Oncol. 2019, 3, 3. [Google Scholar] [CrossRef] [PubMed]
- Garber, J.E.; Offit, K. Hereditary Cancer Predisposition Syndromes. J. Clin. Oncol. 2005, 23, 276–292. [Google Scholar] [CrossRef] [PubMed]
- Disciglio, V.; Fasano, C.; Cariola, F.; Forte, G.; Grossi, V.; Sanese, P.; Lepore Signorile, M.; Resta, N.; Lotesoriere, C.; Stella, A.; et al. Gastric Polyposis and Desmoid Tumours as a New Familial Adenomatous Polyposis Clinical Variant Associated with APC Mutation at the Extreme 3’-End. J. Med. Genet. 2020, 57, 356–360. [Google Scholar] [CrossRef] [PubMed]
- Lepore Signorile, M.; Disciglio, V.; Di Carlo, G.; Pisani, A.; Simone, C.; Ingravallo, G. From Genetics to Histomolecular Characterization: An Insight into Colorectal Carcinogenesis in Lynch Syndrome. Int. J. Mol. Sci. 2021, 22, 6767. [Google Scholar] [CrossRef] [PubMed]
- Hemminki, K.; Sundquist, K.; Sundquist, J.; Försti, A.; Hemminki, A.; Li, X. Familial Risks and Proportions Describing Population Landscape of Familial Cancer. Cancers 2021, 13, 4385. [Google Scholar] [CrossRef] [PubMed]
- Khurana, E.; Fu, Y.; Chakravarty, D.; Demichelis, F.; Rubin, M.A.; Gerstein, M. Role of Non-Coding Sequence Variants in Cancer. Nat. Rev. Genet. 2016, 17, 93–108. [Google Scholar] [CrossRef]
- Diederichs, S.; Bartsch, L.; Berkmann, J.C.; Fröse, K.; Heitmann, J.; Hoppe, C.; Iggena, D.; Jazmati, D.; Karschnia, P.; Linsenmeier, M.; et al. The Dark Matter of the Cancer Genome: Aberrations in Regulatory Elements, Untranslated Regions, Splice Sites, Non-Coding RNA and Synonymous Mutations. EMBO Mol. Med. 2016, 8, 442–457. [Google Scholar] [CrossRef]
- Tan, M.-H.; Mester, J.; Peterson, C.; Yang, Y.; Chen, J.-L.; Rybicki, L.A.; Milas, K.; Pederson, H.; Remzi, B.; Orloff, M.S.; et al. A Clinical Scoring System for Selection of Patients for PTEN Mutation Testing Is Proposed on the Basis of a Prospective Study of 3042 Probands. Am. J. Hum. Genet. 2011, 88, 42–56. [Google Scholar] [CrossRef]
- Genome Browsers 1000 Genomes. Available online: https://www.internationalgenome.org/1000-genomes-browsers/index.html (accessed on 16 November 2021).
- GnomAD Browser. Available online: https://gnomad.broadinstitute.org/ (accessed on 16 November 2021).
- Jeuken, J.W.M.; Cornelissen, S.J.B.; Vriezen, M.; Dekkers, M.M.G.; Errami, A.; Sijben, A.; Boots-Sprenger, S.H.E.; Wesseling, P. MS-MLPA: An Attractive Alternative Laboratory Assay for Robust, Reliable, and Semiquantitative Detection of MGMT Promoter Hypermethylation in Gliomas. Lab. Investig. 2007, 87, 1055–1065. [Google Scholar] [CrossRef]
- MatInspector. Available online: https://www.genomatix.de/online_help/help_matinspector/matinspector_help.html (accessed on 16 November 2021).
- PROMO. Available online: http://alggen.lsi.upc.es/cgi-bin/promo_v3/promo/promoinit.cgi?dirdb=TF_8.3 (accessed on 16 November 2021).
- OncoBase. Available online: http://www.oncobase.biols.ac.cn/ (accessed on 16 November 2021).
- RegulomeDB. Available online: https://regulomedb.org/regulome-search/ (accessed on 16 November 2021).
- Daly, M.B.; Pilarski, R.; Berry, M.; Buys, S.S.; Farmer, M.; Friedman, S.; Garber, J.E.; Kauff, N.D.; Khan, S.; Klein, C.; et al. NCCN Guidelines Insights: Genetic/Familial High-Risk Assessment: Breast and Ovarian, Version 2.2017. J. Natl. Compr. Canc. Netw. 2017, 15, 9–20. [Google Scholar] [CrossRef] [PubMed]
- Pilarski, R.; Burt, R.; Kohlman, W.; Pho, L.; Shannon, K.M.; Swisher, E. Cowden Syndrome and the PTEN Hamartoma Tumor Syndrome: Systematic Review and Revised Diagnostic Criteria. J. Natl. Cancer Inst. 2013, 105, 1607–1616. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Li, X.; Yuan, Y.; Dong, C.; Yang, M. APC Promoter Methylation in Gastrointestinal Cancer. Front. Oncol. 2021, 11, 1291. [Google Scholar] [CrossRef] [PubMed]
- Kirn, V.; Strake, L.; Thangarajah, F.; Richters, L.; Eischeid, H.; Koitzsch, U.; Odenthal, M.; Fries, J. ESR1-Promoter-Methylation Status in Primary Breast Cancer and Its Corresponding Metastases. Clin. Exp. Metastasis 2018, 35, 707–712. [Google Scholar] [CrossRef]
- Kornegoor, R.; Moelans, C.B.; Verschuur-Maes, A.H.; Hogenes, M.C.; de Bruin, P.C.; Oudejans, J.J.; van Diest, P.J. Promoter Hypermethylation in Male Breast Cancer: Analysis by Multiplex Ligation-Dependent Probe Amplification. Breast Cancer Res. 2012, 14, R101. [Google Scholar] [CrossRef] [PubMed]
- Lui, E.L.H.; Loo, W.T.Y.; Zhu, L.; Cheung, M.N.B.; Chow, L.W.C. DNA Hypermethylation of TIMP3 Gene in Invasive Breast Ductal Carcinoma. Biomed. Pharm. 2005, 59 (Suppl. 2), S363–S365. [Google Scholar] [CrossRef]
- Maleva Kostovska, I.; Jakimovska, M.; Popovska-Jankovic, K.; Kubelka-Sabit, K.; Karagjozov, M.; Plaseska-Karanfilska, D. TIMP3 Promoter Methylation Represents an Epigenetic Marker of BRCA1ness Breast Cancer Tumours. Pathol. Oncol. Res. 2018, 24, 937–940. [Google Scholar] [CrossRef]
- Zhou, X.-P.; Waite, K.A.; Pilarski, R.; Hampel, H.; Fernandez, M.J.; Bos, C.; Dasouki, M.; Feldman, G.L.; Greenberg, L.A.; Ivanovich, J.; et al. Germline PTEN Promoter Mutations and Deletions in Cowden/Bannayan-Riley-Ruvalcaba Syndrome Result in Aberrant PTEN Protein and Dysregulation of the Phosphoinositol-3-Kinase/Akt Pathway. Am. J. Hum. Genet. 2003, 73, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, G.; Chen, R.; Yang, X.; Zhao, X.; Zhao, H. A Novel PTEN Gene Promoter Mutation and Untypical Cowden Syndrome. Chin. J. Cancer Res. 2013, 25, 306–311. [Google Scholar] [CrossRef] [PubMed]
- Teresi, R.E.; Zbuk, K.M.; Pezzolesi, M.G.; Waite, K.A.; Eng, C. Cowden Syndrome-Affected Patients with PTEN Promoter Mutations Demonstrate Abnormal Protein Translation. Am. J. Hum. Genet. 2007, 81, 756–767. [Google Scholar] [CrossRef] [PubMed]
- Sarquis, M.S.; Agrawal, S.; Shen, L.; Pilarski, R.; Zhou, X.-P.; Eng, C. Distinct Expression Profiles for PTEN Transcript and Its Splice Variants in Cowden Syndrome and Bannayan-Riley-Ruvalcaba Syndrome. Am. J. Hum. Genet. 2006, 79, 23–30. [Google Scholar] [CrossRef]
- Ma, D.-Z.; Xu, Z.; Liang, Y.-L.; Su, J.-M.; Li, Z.-X.; Zhang, W.; Wang, L.-Y.; Zha, X.-L. Down-Regulation of PTEN Expression Due to Loss of Promoter Activity in Human Hepatocellular Carcinoma Cell Lines. World J. Gastroenterol. 2005, 11, 4472–4477. [Google Scholar] [CrossRef]
- Ramalingam, V.; Natarajan, M.; Johnston, J.; Zeitlinger, J. TATA and Paused Promoters Active in Differentiated Tissues Have Distinct Expression Characteristics. Mol. Syst. Biol. 2021, 17, e9866. [Google Scholar] [CrossRef] [PubMed]
- Spitz, F.; Furlong, E.E.M. Transcription Factors: From Enhancer Binding to Developmental Control|Nature Reviews Genetics. Available online: https://www.nature.com/articles/nrg3207 (accessed on 23 November 2021).
- Boyle, A.P.; Hong, E.L.; Hariharan, M.; Cheng, Y.; Schaub, M.A.; Kasowski, M.; Karczewski, K.J.; Park, J.; Hitz, B.C.; Weng, S.; et al. Annotation of Functional Variation in Personal Genomes Using RegulomeDB. Genome. Res. 2012, 22, 1790–1797. [Google Scholar] [CrossRef]
- Horn, S.; Figl, A.; Rachakonda, P.S.; Fischer, C.; Sucker, A.; Gast, A.; Kadel, S.; Moll, I.; Nagore, E.; Hemminki, K.; et al. TERT Promoter Mutations in Familial and Sporadic Melanoma. Science 2013, 339, 959–961. [Google Scholar] [CrossRef] [PubMed]
- French, J.D.; Ghoussaini, M.; Edwards, S.L.; Meyer, K.B.; Michailidou, K.; Ahmed, S.; Khan, S.; Maranian, M.J.; O’Reilly, M.; Hillman, K.M.; et al. Functional Variants at the 11q13 Risk Locus for Breast Cancer Regulate Cyclin D1 Expression through Long-Range Enhancers. Am. J. Hum. Genet. 2013, 92, 489–503. [Google Scholar] [CrossRef] [PubMed]
- Sandoval-Bórquez, A.; Saavedra, K.; Carrasco-Avino, G.; Garcia-Bloj, B.; Fry, J.; Wichmann, I.; Corvalán, A.H. Noncoding Genomics in Gastric Cancer and the Gastric Precancerous Cascade: Pathogenesis and Biomarkers. Dis. Markers 2015, 2015, e503762. [Google Scholar] [CrossRef] [PubMed]
- Huyghe, J.R.; Bien, S.A.; Harrison, T.A.; Kang, H.M.; Chen, S.; Schmit, S.L.; Conti, D.V.; Qu, C.; Jeon, J.; Edlund, C.K.; et al. Discovery of Common and Rare Genetic Risk Variants for Colorectal Cancer. Nat. Genet. 2019, 51, 76–87. [Google Scholar] [CrossRef]
- Chen, C.-Y.; Chen, J.; He, L.; Stiles, B.L. PTEN: Tumor Suppressor and Metabolic Regulator. Front. Endocrinol. 2018, 9, 338. [Google Scholar] [CrossRef]
- Salmena, L.; Carracedo, A.; Pandolfi, P.P. Tenets of PTEN Tumor Suppression. Cell 2008, 133, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Carracedo, A.; Alimonti, A.; Pandolfi, P.P. PTEN Level in Tumor Suppression: How Much Is Too Little? Cancer Res. 2011, 71, 629–633. [Google Scholar] [CrossRef]
- Alimonti, A.; Carracedo, A.; Clohessy, J.G.; Trotman, L.C.; Nardella, C.; Egia, A.; Salmena, L.; Sampieri, K.; Haveman, W.J.; Brogi, E.; et al. Subtle Variations in Pten Dose Determine Cancer Susceptibility. Nat. Genet. 2010, 42, 454–458. [Google Scholar] [CrossRef]
- Shi, Y.; Paluch, B.E.; Wang, X.; Jiang, X. PTEN at a Glance. J. Cell Sci. 2012, 125, 4687–4692. [Google Scholar] [CrossRef]
- Kang, Y.-H.; Lee, H.S.; Kim, W.H. Promoter Methylation and Silencing of PTEN in Gastric Carcinoma. Lab. Investig. 2002, 82, 285–291. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.-M.; Cheng, F.; Teng, L.-S. The Association between Phosphatase and Tensin Homolog Hypermethylation and Patients with Breast Cancer, a Meta-Analysis and Literature Review. Sci. Rep. 2016, 6, 32723. [Google Scholar] [CrossRef]
- Goel, A.; Arnold, C.N.; Niedzwiecki, D.; Carethers, J.M.; Dowell, J.M.; Wasserman, L.; Compton, C.; Mayer, R.J.; Bertagnolli, M.M.; Boland, C.R. Frequent Inactivation of PTEN by Promoter Hypermethylation in Microsatellite Instability-High Sporadic Colorectal Cancers. Cancer Res. 2004, 64, 3014–3021. [Google Scholar] [CrossRef]
- Pilarski, R. PTEN Hamartoma Tumor Syndrome: A Clinical Overview. Cancers 2019, 11, 844. [Google Scholar] [CrossRef]
- Yehia, L.; Keel, E.; Eng, C. The Clinical Spectrum of PTEN Mutations. Annu. Rev. Med. 2020, 71, 103–116. [Google Scholar] [CrossRef] [PubMed]
- Al-Thihli, K.; Palma, L.; Marcus, V.; Cesari, M.; Kushner, Y.B.; Barkun, A.; Foulkes, W.D. A Case of Cowden’s Syndrome Presenting with Gastric Carcinomas and Gastrointestinal Polyposis. Nat. Rev. Gastroenterol. Hepatol. 2009, 6, 184–189. [Google Scholar] [CrossRef] [PubMed]
- Marques, M.; Ramalho, R.; Baldaque-Silva, F.; Macedo, G. Novel Mutation Identified in Cowden Syndrome Presenting as a Gastric Adenocarcinoma. Clin. Res. Hepatol. Gastroenterol. 2013, 37, e131–e132. [Google Scholar] [CrossRef]
- Makis, W.; Ciarallo, A.; Hickeson, M. The Use of 18F-FDG PET/CT in Cowden Syndrome to Differentiate Multifocal Gastric Carcinoma From Extensive Benign Gastric Polyposis. Clin. Nucl. Med. 2012, 37, 311–314. [Google Scholar] [CrossRef]
- Sherman, S.K.; Maxwell, J.E.; Qian, Q.; Bellizzi, A.M.; Braun, T.A.; Iannettoni, M.D.; Darbro, B.W.; Howe, J.R. Esophageal Cancer in a Family with Hamartomatous Tumors and Germline PTEN Frameshift and SMAD7 Missense Mutations. Cancer Genet. 2015, 208, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Pezzolesi, M.G.; Zbuk, K.M.; Waite, K.A.; Eng, C. Comparative Genomic and Functional Analyses Reveal a Novel Cis-Acting PTEN Regulatory Element as a Highly Conserved Functional E-Box Motif Deleted in Cowden Syndrome. Hum. Mol. Genet. 2007, 16, 1058–1071. [Google Scholar] [CrossRef] [PubMed]
- Heikkinen, T.; Greco, D.; Pelttari, L.M.; Tommiska, J.; Vahteristo, P.; Heikkilä, P.; Blomqvist, C.; Aittomäki, K.; Nevanlinna, H. Variants on the Promoter Region of PTEN Affect Breast Cancer Progression and Patient Survival. Breast Cancer Res. 2011, 13, R130. [Google Scholar] [CrossRef]
- Ray, A.; Ray, B.K. Induction of Ras by SAF-1/MAZ through a Feed-Forward Loop Promotes Angiogenesis in Breast Cancer. Cancer Med. 2015, 4, 224–234. [Google Scholar] [CrossRef] [PubMed]
- Cogoi, S.; Zorzet, S.; Rapozzi, V.; Géci, I.; Pedersen, E.B.; Xodo, L.E. MAZ-Binding G4-Decoy with Locked Nucleic Acid and Twisted Intercalating Nucleic Acid Modifications Suppresses KRAS in Pancreatic Cancer Cells and Delays Tumor Growth in Mice. Nucleic Acids Res. 2013, 41, 4049–4064. [Google Scholar] [CrossRef] [PubMed]
- Smits, M.; Wurdinger, T.; van het Hof, B.; Drexhage, J.A.R.; Geerts, D.; Wesseling, P.; Noske, D.P.; Vandertop, W.P.; de Vries, H.E.; Reijerkerk, A. Myc-Associated Zinc Finger Protein (MAZ) Is Regulated by MiR-125b and Mediates VEGF-Induced Angiogenesis in Glioblastoma. FASEB J. 2012, 26, 2639–2647. [Google Scholar] [CrossRef] [PubMed]
- Jiao, L.; Li, Y.; Shen, D.; Xu, C.; Wang, L.; Huang, G.; Chen, L.; Yang, Y.; Yang, C.; Yu, Y.; et al. The Prostate Cancer-up-Regulated Myc-Associated Zinc-Finger Protein (MAZ) Modulates Proliferation and Metastasis through Reciprocal Regulation of Androgen Receptor. Med. Oncol. 2013, 30, 570. [Google Scholar] [CrossRef]
- Wang, X.; Southard, R.C.; Allred, C.D.; Talbert, D.R.; Wilson, M.E.; Kilgore, M.W. MAZ drives tumor specific expression of PPAR gamma 1 in breast cancer cells. Breast Cancer Res. Treat 2008, 111, 103–111. [Google Scholar] [CrossRef][Green Version]
- Maity, G.; Haque, I.; Ghosh, A.; Dhar, G.; Gupta, V.; Sarkar, S.; Azeem, I.; McGregor, D.; Choudhary, A.; Campbell, D.R.; et al. The MAZ Transcription Factor Is a Downstream Target of the Oncoprotein Cyr61/CCN1 and Promotes Pancreatic Cancer Cell Invasion via CRAF-ERK Signaling. J. Biol. Chem. 2018, 293, 4334–4349. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Disciglio, V.; Sanese, P.; Fasano, C.; Lotesoriere, C.; Valentini, A.M.; Forte, G.; Lepore Signorile, M.; De Marco, K.; Grossi, V.; Lolli, I.; et al. Identification and Somatic Characterization of the Germline PTEN Promoter Variant rs34149102 in a Family with Gastrointestinal and Breast Tumors. Genes 2022, 13, 644. https://doi.org/10.3390/genes13040644
Disciglio V, Sanese P, Fasano C, Lotesoriere C, Valentini AM, Forte G, Lepore Signorile M, De Marco K, Grossi V, Lolli I, et al. Identification and Somatic Characterization of the Germline PTEN Promoter Variant rs34149102 in a Family with Gastrointestinal and Breast Tumors. Genes. 2022; 13(4):644. https://doi.org/10.3390/genes13040644
Chicago/Turabian StyleDisciglio, Vittoria, Paola Sanese, Candida Fasano, Claudio Lotesoriere, Anna Maria Valentini, Giovanna Forte, Martina Lepore Signorile, Katia De Marco, Valentina Grossi, Ivan Lolli, and et al. 2022. "Identification and Somatic Characterization of the Germline PTEN Promoter Variant rs34149102 in a Family with Gastrointestinal and Breast Tumors" Genes 13, no. 4: 644. https://doi.org/10.3390/genes13040644
APA StyleDisciglio, V., Sanese, P., Fasano, C., Lotesoriere, C., Valentini, A. M., Forte, G., Lepore Signorile, M., De Marco, K., Grossi, V., Lolli, I., Cariola, F., & Simone, C. (2022). Identification and Somatic Characterization of the Germline PTEN Promoter Variant rs34149102 in a Family with Gastrointestinal and Breast Tumors. Genes, 13(4), 644. https://doi.org/10.3390/genes13040644






