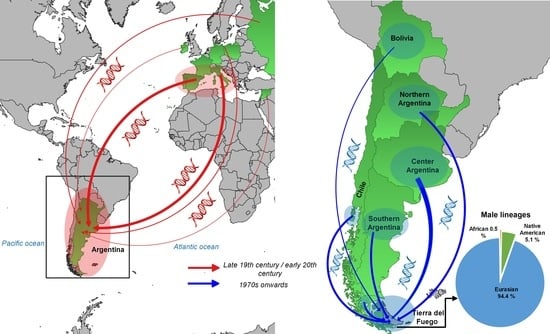
Tierra Del Fuego: What Is Left from the Precolonial Male Lineages?
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling
2.2. Y Chromosome STR Typing
2.3. Y Chromosome SNP Typing
2.4. Data Analyses
3. Results and Discussion
3.1. Tierra Del Fuego Genetic Diversity
3.2. Tierra Del Fuego Haplogroups
3.3. Tierra Del Fuego Continental Ancestry
3.4. Comparisons with Latin American Populations
3.5. Comparisons with Eurasian Populations
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Steele, J.; Politis, G. AMS 14C dating of early human occupation of southern South America. J. Archaeol. Sci. 2009, 36, 419–429. [Google Scholar] [CrossRef]
- Borrero, L. The origins of ethnographic subsistence patterns in Fuego Patagonia. In Patagonia: Natural History, Prehistory and Ethnography at the Uttermost End of the Earth; McEwan, C., Borrero, L., Prieto, A., Eds.; British Museum Press: London, UK, 1997; pp. 60–81. [Google Scholar]
- Piana, E.L.; Orquera, L.A. The southern top of the world: The first peopling of Patagonia and Tierra del Fuego and the cultural endurance of the Fuegian Sea-Nomads. Arct. Anthropol. 2009, 46, 103–117. [Google Scholar] [CrossRef]
- Rodríguez, M.E.; Horlent, L. Tehuelches y Selk’nam (Santa Cruz y Tierra del Fuego): No Desaparecimos, 1st ed.; Ministerio de Educación y Deportes: Ciudad Autónoma de Buenos Aires, Argentina, 2016. [Google Scholar]
- Van Aert, P. Tierra del Fuego. In The Political Economy of Divided Islands, 1st ed.; Baldacchino, G., Ed.; Palgrave Macmillan: London, UK, 2013; pp. 195–211. [Google Scholar]
- Devoto, F. Historia de la Inmigración en la Argentina, 1st ed.; Editorial Sudamericana: Buenos Aires, Argentina, 2002. [Google Scholar]
- Hermida, M.; Malizia, M.; Van Aert, P. Ser fueguino. Un estudio sobre migración y construcción de pertenencia. In X Jornadas de Sociología; Facultad de Ciencias Sociales, Universidad de Buenos Aires: Buenos Aires, Argentina, 2013. [Google Scholar]
- Lattes, A.E. Esplendor y ocaso de las migraciones internas. In Población y Bienestar en la Argentina del Primero al Segundo Centenario, 1st ed.; Torrado, S., Ed.; Edhasa: Buenos Aires, Argentina, 2007. [Google Scholar]
- INDEC. Provincia de Tierra del Fuego. Población Total Nacida en el Extranjero por Lugar de Nacimiento, Según Sexo y Grupos de Edad. 2010. Available online: https://www.indec.gob.ar/ (accessed on 24 January 2022).
- Cardoso, S.; Palencia-Madrid, L.; Valverde, L.; Alfonso-Sánchez, M.A.; Gómez-Pérez, L.; Alfaro, E.; Bravi, C.M.; Dipierri, J.E.; Peña, J.A.; de Pancorbo, M.M. Mitochondrial DNA control region data reveal high prevalence of Native American lineages in Jujuy province, NW Argentina. Forensic Sci. Int. Genet. 2013, 7, e52–e55. [Google Scholar] [CrossRef] [PubMed]
- García, A.; Garita-onandía, Y.; Demarchi, D.A.; Altuna, M.E. Patrones de migración y niveles de diversidad genética de linajes maternos en la población de La Esperanza, provincia de Jujuy. Rev. Argent. Antropol. Biol. 2021, 23, 033. [Google Scholar] [CrossRef]
- Motti, J.M.B.; Muzzio, M.; Ramallo, V.; Rodenak Kladniew, B.E.; Alfaro Gómez, E.L.; Dipierri, J.E.; Bailliet, G.; Bravi, C.M. Origen y distribución espacial de linajes maternos nativos en el noroeste y centro oeste argentinos. Rev. Argent. Antropol. Biol. 2013, 15, 3–14. [Google Scholar]
- Pauro, M.; García, A.; Nores, R.; Demarchi, D.A. Analysis of uniparental lineages in two villages of Santiago del Estero, Argentina, seat of “Pueblos de Indios” in colonial times. Hum. Biol. 2013, 85, 699–719. [Google Scholar] [CrossRef]
- Corach, D.; Lao, O.; Bobillo, C.; van Der Gaag, K.; Zuniga, S.; Vermeulen, M.; van Duijn, K.; Goedbloed, M.; Vallone, P.M.; Parson, W.; et al. Inferring continental ancestry of argentineans from Autosomal, Y-chromosomal and mitochondrial DNA. Ann. Hum. Genet. 2010, 74, 65–76. [Google Scholar] [CrossRef]
- Castagnola, J.; Cano, H.; Hulaniuk, M.L.; Trinks, J.; Corach, D.; Caputo, M. Inferring the genetic structure of Northwestern Argentina by uniparental SNP typing. Forensic Sci. Int. Genet. Suppl. Ser. 2019, 7, 306–309. [Google Scholar] [CrossRef]
- Salas, A.; Jaime, J.C.; Álvarez-Iglesias, V.; Carracedo, A. Gender bias in the multiethnic genetic composition of central Argentina. Eur. J. Hum. Genet. 2008, 53, 662–674. [Google Scholar] [CrossRef][Green Version]
- Bailliet, G.; Ramallo, V.; Muzzio, M.; Santos, M.R.; Motti, J.M.B.; Bianchi, N.O.; Bravi, C.M. Antecedentes y nuevos aportes en el estudio del Cromosoma Y en poblaciones humanas sudamericanas. BAG J. Basic Appl. Genet. 2011, 22, 1–9. [Google Scholar]
- Sala, C.; Paz Sepulveda, P.B.; Cuello, M.; Schwab, M.; Jurado Medina, L.S.; Motti, J.M.B.; Santos, M.R.; Aquilano, E.; Martín Alva, E.A.; Mejia Porturas, M.; et al. Análisis de los linajes paternos en la ciudad de Trujillo, Perú. Runa 2022, 43, 173–190. [Google Scholar] [CrossRef]
- Bobillo, M.C.; Zimmermann, B.; Sala, A.; Huber, G.; Röck, A.; Bandelt, H.-J.; Corach, D.; Parson, W. Amerindian mitochondrial DNA haplogroups predominate in the population of Argentina: Towards a first nationwide forensic mitochondrial DNA sequence database. Int. J. Legal Med. 2010, 124, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Avena, S.A.; Parolin, M.L.; Dejean, C.B.; Ríos Part, M.C.; Fabrikant, G.; Goicoechea, A.S.; Dugoujon, J.M.; Carnese, F.R. Mezcla génica y linajes uniparentales en Comodoro Rivadavia (provincia de Chubut, Argentina). Rev. Arg. Antrop. Biol. 2009, 11, 25–41. [Google Scholar]
- Avena, S.A.; Parolin, M.L.; Boquet, M.; Dejean, C.B.; Postillone, M.B.; Alvarez Trentini, Y.; Di Fabio Rocca, F.; Mansilla, F.; Jones, L.; Dugoujon, J.M.; et al. Mezcla génica y linajes uniparentales en Esquel (Pcia. de Chubut): Su comparación con otras muestras poblacionales argentinas. BAG J. Basic Appl. Genet 2010, 21, 1–14. [Google Scholar]
- Parolin, M.L.; Avena, S.A.; Fleischer, S.; Pretell, M.; Rocca, F.D.F.; Rodríguez, D.A.; Dejean, C.B.; Postillone, M.B.; Vaccaro, M.S.; Dahinten, S.L.; et al. Análisis de la diversidad biológica y mestizaje en la ciudad de Puerto Madryn (Prov. De Chubut, Argentina). Rev. Arg. Antrop. Biol. 2013, 15, 61–75. [Google Scholar]
- Parolin, M.L.; Baillet, G.; Bravi, C.M.; Sala, C.; Schwab, M.; Paz, P. Variabilidad genética y autopercepción del mestizaje en la población Cordillerana de San Carlos De Bariloche (Río Negro, Arg). In Proceedings of the XIV Congreso de la Asociación Latinoamericana de Antropología Biológica, Tacuarembó, Uruguay, 18–21 October 2016; pp. 130–131. [Google Scholar]
- Tamburrini, C.; de Saint Pierre, M.; Bravi, C.M.; Bailliet, G.; Jurado Medina, L.; Velázquez, I.F.; Real, L.E.; Holley, A.; Tedeschi, C.M.; Basso, N.G.; et al. Uniparental origins of the admixed Argentine Patagonia. Am. J. Hum. Biol. 2021, 34, e23682. [Google Scholar] [CrossRef]
- Jurado Medina, L.S.; Parolin, M.L.; Ramallo, V.; Sepúlveda, P.B.; Real, L.E.; Avena, S.A.; Carnese, F.R.; Basso, N.G.; Baillet, G. En búsqueda de la herencia americana en poblaciones mestizas análisis comparativo entre muestras masculinas de Patagonia, Noroeste y Cuyo. In Proceedings of the XII Jornadas Nacionales de Antropología Biológica, Corrientes, Argentina, 22–25 September 2015; p. 82. [Google Scholar]
- Brion, M.; Sobrino, B.; Blanco-Verea, A.; Lareu, M.V.; Carracedo, A. Hierarchical analysis of 30 Y-chromosome SNPs in European populations. Int. J. Legal Med. 2005, 119, 10–15. [Google Scholar] [CrossRef]
- Gomes, V.; Sánchez-Diaz, P.; Amorim, A.; Carracedo, A.; Gusmão, L. Digging deeper into East African human Y chromosome lineages. Hum. Genet. 2010, 127, 603–613. [Google Scholar] [CrossRef]
- Campos, A. Genetic Characterization of the Male Lineages Present in Nigeria Population. Master’s Thesis, Faculty of Sciences of the University of Porto, Porto, Portugal, 2018. [Google Scholar]
- Resque, R.; Gusmão, L.; Geppert, M.; Roewer, L.; Palha, T.; Alvarez, L.; Ribeiro-dos-Santos, Â.; Santos, S. Male lineages in Brazil: Intercontinental admixture and stratification of the European background. PLoS ONE 2016, 11, e0152573. [Google Scholar] [CrossRef]
- Aragão, G. Characterization of Male Lineages in the Ashaninka from Peru. Master’s Thesis, Faculty of Sciences of the University of Porto, Porto, Portugal, 2018. [Google Scholar]
- González, M.; Gomes, V.; López-Parra, A.M.; Amorim, A.; Carracedo, A.; Sánchez-Diaz, P.; Arroyo-Pardo, E.; Gusmão, L. The genetic landscape of Equatorial Guinea and the origin and migration routes of the Y chromosome haplogroup RV88. Eur. J. Hum. Genet. 2013, 21, 324–331. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.J.; Forster, P.; Sykes, B.C.; Richards, M.B. Mitochondrial portraits of human populations using median networks. Genetics 1995, 141, 743–753. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.J.; Forster, P.; Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Qamar, R.; Ayub, Q.; Mohyuddin, A.; Helgason, A.; Mazhar, K.; Mansoor, A.; Zerjal, T.; Tyler-Smith, C.; Mehdi, S.Q. Y-chromosomal DNA variation in Pakistan. Am. J. Hum. Genet. 2002, 70, 1107–1124. [Google Scholar] [CrossRef]
- Dupanloup, I.; Bertorelle, G. Inferring admixture proportions from molecular data: Extension to any number of parental populations. Mol. Biol. Evol. 2001, 18, 672–675. [Google Scholar] [CrossRef]
- Bertorelle, G.; Excoffier, L. Inferring admixture proportions from molecular data. Mol. Biol. Evol. 1998, 15, 1298–1311. [Google Scholar] [CrossRef] [PubMed]
- Helgason, A.; Einarsson, A.W.; Guðmundsdóttir, V.B.; Sigurðsson, Á.; Gunnarsdóttir, E.D.; Jagadeesan, A.; Ebenesersdóttir, S.S.; Kong, A.; Stefánsson, K. The Y-chromosome point mutation rate in humans. Nat. Genet. 2015, 47, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Alonso Morales, L.A.; Casas-Vargas, A.; Rojas Castro, M.; Resque, R.; Ribeiro-Dos-Santos, Â.K.; Santos, S.; Gusmão, L.; Usaquén, W. Paternal portrait of populations of the middle Magdalena River region (Tolima and Huila, Colombia): New insights on the peopling of Central America and northernmost South America. PLoS ONE 2018, 13, e0207130. [Google Scholar] [CrossRef]
- Cárdenas, J.M.; Heinz, T.; Pardo-Seco, J.; Álvarez-Iglesias, V.; Taboada-Echalar, P.; Sánchez-Diz, P.; Carracedo, Á.; Salas, A. The multiethnic ancestry of Bolivians as revealed by the analysis of Y-chromosome markers. Forensic Sci. Int. Genet. 2015, 14, 210–218. [Google Scholar] [CrossRef]
- Núñez, C.; Geppert, M.; Baeta, M.; Roewer, L.; Martínez-Jarreta, B. Y chromosome haplogroup diversity in a Mestizo population of Nicaragua. Forensic Sci. Int. Genet. 2012, 6, e192–e195. [Google Scholar] [CrossRef]
- Parolin, M.L.; Toscanini, U.F.; Velázquez, I.F.; Llull, C.; Berardi, G.L.; Holley, A.; Tamburrini, C.; Avena, S.; Carnese, F.R.; Lanata, J.L.; et al. Genetic admixture patterns in Argentinian Patagonia. PLoS ONE 2019, 14, e0214830. [Google Scholar] [CrossRef] [PubMed]
- Purps, J.; Siegert, S.; Willuweit, S.; Nagy, M.; Alves, C.; Salazar, R.; Angustia, S.M.; Santos, L.H.; Anslinger, K.; Bayer, B.; et al. A global analysis of Y-chromosomal haplotype diversity for 23 STR loci. Forensic Sci. Int. Genet. 2014, 12, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, J.; Romero, M.; Simão, F.; Ferreira Almeida, A.P.; Quiroz, A.; Machado, P.; Velázquez, V.; de Carvalho, E.F.; Vullo, C.; Gusmão, L. Analysis of 23 Y-STRs in a population sample from eastern Paraguay. Forensic Sci. Int. Genet. 2018, 37, e20–e22. [Google Scholar] [CrossRef] [PubMed]
- Simão, F.; Ribeiro, J.; Vullo, C.; Catelli, L.; Gomes, V.; Xavier, C.; Huber, G.; Bodner, M.; Quiroz, A.; Ferreira, A.P.; et al. The Ancestry of Eastern Paraguay: A Typical South American Profile with a Unique Pattern of Admixture. Genes 2021, 12, 1788. [Google Scholar] [CrossRef] [PubMed]
- Toscanini, U.; Vullo, C.; Berardi, G.; Llull, C.; Borosky, A.; Gómez, A.; Pardo-Seco, J.; Salas, A. A comprehensive Y-STR portrait of Argentinean populations. Forensic Sci. Int. Genet. 2016, 20, 1–5. [Google Scholar] [CrossRef]
- Borucki, A. The slave trade to the Rio de La Plata, 1777–1812: Trans-imperial networks and Atlantic warfare. Colon. Lat. Am. Rev. 2011, 20, 81–107. [Google Scholar] [CrossRef]
- Borucki, A.; Eltis, D.; Wheat, D. Atlantic history and the slave trade to Spanish America. Am. Hist. Rev. 2015, 120, 433–461. [Google Scholar] [CrossRef]
- Cruciani, F.; La Fratta, R.; Santolamazza, P.; Sellitto, D.; Pascone, R.; Moral, P.; Watson, E.; Guida, V.; Colomb, E.B.; Zaharova, B.; et al. Phylogeographic analysis of haplogroup E3b (E-M215) y chromosomes reveals multiple migratory events within and out of Africa. Am. J. Hum. Genet. 2004, 74, 1014–1022. [Google Scholar] [CrossRef]
- Henn, B.M.; Gignoux, C.; Lin, A.A.; Oefner, P.J.; Shen, P.; Scozzari, R.; Cruciani, F.; Tishkoff, S.A.; Mountain, J.L.; Underhill, P.A. Y-chromosomal evidence of a pastoralist migration through Tanzania to southern Africa. Proc. Natl. Acad. Sci. USA 2008, 105, 10693–10698. [Google Scholar] [CrossRef]
- Semino, O.; Magri, C.; Benuzzi, G.; Lin, A.A.; Al-Zahery, N.; Battaglia, V.; Maccioni, L.; Triantaphyllidis, C.; Shen, P.; Oefner, P.J.; et al. Origin, diffusion, and differentiation of Y-chromosome haplogroups E and J: Inferences on the neolithization of Europe and later migratory events in the Mediterranean area. Am. J. Hum. Genet. 2004, 74, 1023–1034. [Google Scholar] [CrossRef]
- De Filippo, C.; Barbieri, C.; Whitten, M.; Mpoloka, S.W.; Gunnarsdóttir, E.D.; Bostoen, K.; Nyambe, T.; Beyer, K.; Schreiber, H.; de Knijff, P.; et al. Y-chromosomal variation in sub-Saharan Africa: Insights into the history of Niger-Congo groups. Mol. Biol. Evol. 2011, 28, 1255–1269. [Google Scholar] [CrossRef] [PubMed]
- Larmuseau, M.H.; Vessi, A.; Jobling, M.A.; van Geystelen, A.; Primativo, G.; Biondi, G.; Martínez-Labarga, C.; Ottoni, C.; Decorte, R.; Rickards, O. The Paternal Landscape along the Bight of Benin—Testing Regional Representativeness of West-African Population Samples Using Y-Chromosomal Markers. PLoS ONE 2015, 10, e0141510. [Google Scholar] [CrossRef] [PubMed]
- Rosa, A.; Ornelas, C.; Jobling, M.A.; Brehm, A.; Villems, R. Y-chromosomal diversity in the population of Guinea-Bissau: A multiethnic perspective. BMC Evol. Biol. 2007, 7, 124. [Google Scholar] [CrossRef]
- Adams, S.M.; Bosch, E.; Balaresque, P.L.; Ballereau, S.J.; Lee, A.C.; Arroyo, E.; López-Parra, A.M.; Aler, M.; Grifo, M.S.; Brion, M.; et al. The genetic legacy of religious diversity and intolerance: Paternal lineages of Christians, Jews, and Muslims in the Iberian Peninsula. Am. J. Hum. Genet. 2008, 83, 725–736. [Google Scholar] [CrossRef]
- Boattini, A.; Martinez-Cruz, B.; Sarno, S.; Harmant, C.; Useli, A.; Sanz, P.; Yang-Yao, D.; Manry, J.; Ciani, G.; Luiselli, D.; et al. Uniparental markers in Italy reveal a sex-biased genetic structure and different historical strata. PLoS ONE 2013, 8, e65441. [Google Scholar] [CrossRef]
- Rębała, K.; Martínez-Cruz, B.; Tönjes, A.; Kovacs, P.; Stumvoll, M.; Lindner, I.; Büttner, A.; Wichmann, H.E.; Siváková, D.; Soták, M.; et al. Contemporary paternal genetic landscape of Polish and German populations: From early medieval Slavic expansion to post-World War II resettlements. Eur. J. Hum. Genet. 2013, 21, 415–422. [Google Scholar] [CrossRef] [PubMed]
- Zalloua, P.A.; Xue, Y.; Khalife, J.; Makhoul, N.; Debiane, L.; Platt, D.E.; Royyuru, A.K.; Herrera, R.J.; Hernanz, D.F.; Blue-Smith, J.; et al. Y-chromosomal diversity in Lebanon is structured by recent historical events. Am. J. Hum. Genet. 2008, 82, 873–882. [Google Scholar] [CrossRef]
- Berniell-Lee, G.; Calafell, F.; Bosch, E.; Heyer, E.; Sica, L.; Mouguiama-Daouda, P.; van der Veen, L.; Hombert, J.M.; Quintana-Murci, L.; Comas, D. Genetic and demographic implications of the Bantu expansion: Insights from human paternal lineages. Mol. Biol. Evol. 2009, 26, 1581–1589. [Google Scholar] [CrossRef]
- Cruciani, F.; Trombetta, B.; Sellitto, D.; Massaia, A.; Destro-Bisol, G.; Watson, E.; Beraud Colomb, E.; Dugoujon, J.M.; Moral, P.; Scozzari, R. Human Y chromosome haplogroup R-V88: A paternal genetic record of early mid Holocene trans-Saharan connections and the spread of Chadic languages. Eur. J. Hum. Genet. 2010, 18, 800–807. [Google Scholar] [CrossRef][Green Version]
- D’Atanasio, E.; Trombetta, B.; Bonito, M.; Finocchio, A.; Di Vito, G.; Seghizzi, M.; Romano, R.; Russo, G.; Paganotti, G.M.; Watson, E.; et al. The peopling of the last Green Sahara revealed by high-coverage resequencing of trans-Saharan patrilineages. Genome Biol. 2018, 19, 20. [Google Scholar] [CrossRef]
- Shriner, D.; Rotimi, C.N. Genetic history of Chad. Am. J. Phys. Anthropol. 2018, 167, 804–812. [Google Scholar] [CrossRef] [PubMed]
- Marcus, J.H.; Posth, C.; Ringbauer, H.; Lai, L.; Skeates, R.; Sidore, C.; Beckett, J.; Furtwängler, A.; Olivieri, A.; Chiang, C.; et al. Genetic history from the Middle Neolithic to present on the Mediterranean island of Sardinia. Nat. Commun. 2020, 11, 939. [Google Scholar] [CrossRef] [PubMed]
- Contu, D.; Morelli, L.; Santoni, F.; Foster, J.W.; Francalacci, P.; Cucca, F. Y-chromosome based evidence for pre-neolithic origin of the genetically homogeneous but diverse Sardinian population: Inference for association scans. PLoS ONE 2008, 3, e1430. [Google Scholar] [CrossRef]
- Morelli, L.; Contu, D.; Santoni, F.; Whalen, M.B.; Francalacci, P.; Cucca, F. A comparison of Y-chromosome variation in Sardinia and Anatolia is more consistent with cultural rather than demic diffusion of agriculture. PLoS ONE 2010, 5, e10419. [Google Scholar] [CrossRef] [PubMed]
- Underhill, P.A.; Shen, P.; Lin, A.A.; Jin, L.; Passarino, G.; Yang, W.H.; Kauffman, E.; Bonné-Tamir, B.; Bertranpetit, J.; Francalacci, P.; et al. Y chromosome sequence variation and the history of human populations. Nat. Genet. 2000, 26, 358–361. [Google Scholar] [CrossRef]
- Di Cristofaro, J.; Mazières, S.; Tous, A.; Di Gaetano, C.; Lin, A.A.; Nebbia, P.; Piazza, A.; King, R.J.; Underhill, P.; Chiaroni, J. Prehistoric migrations through the Mediterranean basin shaped Corsican Y-chromosome diversity. PLoS ONE 2018, 13, e0200641. [Google Scholar] [CrossRef] [PubMed]
- Fortes-Lima, C.; Brucato, N.; Croze, M.; Bellis, G.; Schiavinato, S.; Massougbodji, A.; Migot-Nabias, F.; Dugoujon, J.M. Genetic population study of Y-chromosome markers in Benin and Ivory Coast ethnic groups. Forensic Sci. Int. Genet. 2015, 19, 232–237. [Google Scholar] [CrossRef]
- Battaglia, V.; Grugni, V.; Perego, U.A.; Angerhofer, N.; Gomez-Palmieri, J.E.; Woodward, S.R.; Achilli, A.; Myres, N.; Torroni, A.; Semino, O. The first peopling of South America: New evidence from Y-chromosome haplogroup Q. PLoS ONE 2013, 8, e71390. [Google Scholar] [CrossRef]
- Grugni, V.; Raveane, A.; Ongaro, L.; Battaglia, V.; Trombetta, B.; Colombo, G.; Capodiferro, M.R.; Olivieri, A.; Achilli, A.; Perego, U.A.; et al. Analysis of the human Y-chromosome haplogroup Q characterizes ancient population movements in Eurasia and the Americas. BMC Biol. 2019, 17, 1–14. [Google Scholar] [CrossRef]
- Pinotti, T.; Bergström, A.; Geppert, M.; Bawn, M.; Ohasi, D.; Shi, W.; Lacerda, D.R.; Solli, A.; Norstedt, J.; Reed, K.; et al. Y chromosome sequences reveal a short Beringian Standstill, rapid expansion, and early population structure of Native American founders. Curr. Biol. 2019, 29, 149–157. [Google Scholar] [CrossRef]
- Zegura, S.L.; Karafet, T.M.; Zhivotovsky, L.A.; Hammer, M.F. High-resolution SNPs and microsatellite haplotypes point to a single, recent entry of Native American Y chromosomes into the Americas. Mol. Biol. Evol. 2004, 21, 164–175. [Google Scholar] [CrossRef] [PubMed]
- Jota, M.S.; Lacerda, D.R.; Sandoval, J.R.; Vieira, P.P.; Ohasi, D.; Santos-Júnior, J.E.; Acosta, O.; Cuellar, C.; Revollo, S.; Paz-Y-Miño, C.; et al. New native South American Y chromosome lineages. J. Hum. Genet. 2016, 61, 593–603. [Google Scholar] [CrossRef]
- Sandoval, J.R.; Lacerda, D.R.; Jota, M.S.; Elward, R.; Acosta, O.; Pinedo, D.; Danos, P.; Cuellar, C.; Revollo, S.; Santos, F.R.; et al. Genetic ancestry of families of putative Inka descent. Mol. Genet. Genom. 2018, 293, 873–881. [Google Scholar] [CrossRef] [PubMed]
- Gayà-Vidal, M.; Moral, P.; Saenz-Ruales, N.; Gerbault, P.; Tonasso, L.; Villena, M.; Vasquez, R.; Bravi, C.M.; Dugoujon, J.M. mtDNA and Y-chromosome diversity in Aymaras and Quechuas from Bolivia: Different stories and special genetic traits of the Andean Altiplano populations. Am. J. Phys. Anthropol. 2011, 145, 215–230. [Google Scholar] [CrossRef]
- Barbieri, C.; Sandoval, J.R.; Valqui, J.; Shimelman, A.; Ziemendorff, S.; Schröder, R.; Geppert, M.; Roewer, L.; Gray, R.; Stoneking, M.; et al. Enclaves of genetic diversity resisted Inca impacts on population history. Sci. Rep. 2017, 7, 17411. [Google Scholar] [CrossRef] [PubMed]
- Roewer, L.; Nothnagel, M.; Gusmão, L.; Gomes, V.; González, M.; Corach, D.; Sala, A.; Alechine, E.; Palha, T.; Santos, N.; et al. Continent-wide decoupling of Y-chromosomal genetic variation from language and geography in native South Americans. PLoS Genet. 2013, 9, e1003460. [Google Scholar] [CrossRef]
- Ansari-Pour, N.; Plaster, C.A.; Bradman, N. Evidence from Y-chromosome analysis for a late exclusively eastern expansion of the Bantu-speaking people. Eur. J. Hum. Genet. 2013, 21, 423–429. [Google Scholar] [CrossRef]
- Montano, V.; Ferri, G.; Marcari, V.; Batini, C.; Anyaele, O.; Destro-Bisol, G.; Comas, D. The Bantu expansion revisited: A new analysis of Y chromosome variation in Central Western Africa. Mol. Ecol. 2011, 20, 2693–2708. [Google Scholar] [CrossRef]
- Ansari-Pour, N.; Moñino, Y.; Duque, C.; Gallego, N.; Bedoya, G.; Thomas, M.G.; Bradman, N. Palenque de San Basilio in Colombia: Genetic data support an oral history of a paternal ancestry in Congo. Proc. Biol. Sci. 2016, 283, 20152980. [Google Scholar] [CrossRef]
- Sims, L.M.; Garvey, D.; Ballantyne, J. Sub-populations within the major European and African derived haplogroups R1b3 and E3a are differentiated by previously phylogenetically undefined Y-SNPs. Hum. Mutat. 2007, 28, 97. [Google Scholar] [CrossRef]
- Jannuzzi, J.; Ribeiro, J.; Alho, C.; de Oliveira Lázaro E Arão, G.; Cicarelli, R.; Simões Dutra Corrêa, H.; Ferreira, S.; Fridman, C.; Gomes, V.; Loiola, S.; et al. Male lineages in Brazilian populations and performance of haplogroup prediction tools. Forensic Sci. Int. Genet. 2020, 44, 102163. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, A.M.; Domingues, P.M.; Gomes, V.; Amorim, A.; Jannuzzi, J.; Carvalho, E.F.; Gusmão, L. Male lineage strata of Brazilian population disclosed by the simultaneous analysis of STRs and SNPs. Forensic Sci. Int. Genet. 2014, 13, 264–268. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, M.; Brito, P.; Lopes, V.; Andrade, L.; Anjos, M.J.; Real, F.C.; Gusmão, L. Analysis of paternal lineages in Brazilian and African populations. Genet. Mol. Biol. 2010, 33, 422–427. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Schaan, A.P.; Gusmão, L.; Jannuzzi, J.; Modesto, A.; Amador, M.; Marques, D.; Rabenhorst, S.H.; Montenegro, R.; Lopes, T.; Yoshioka, F.K.; et al. New insights on intercontinental origins of paternal lineages in Northeast Brazil. BMC Evol. Biol. 2020, 20, 15. [Google Scholar] [CrossRef]
- de Figueiredo, R.; Ambrosio, I.B.; Braganholi, D.F.; Chemale, G.; Martins, J.A.; Gomes, V.; Gusmão, L.; Cicarelli, R. Male-specific contributions to the Brazilian population of Espirito Santo. Int. J. Leg. Med. 2016, 130, 679–681. [Google Scholar] [CrossRef]
- de Azevedo, D.A.; da Silva, L.A.F.; Gusmão, L.; de Carvalho, E.F. Analysis of Y chromosome SNPs in Alagoas, northeastern Brazil. Forensic Sci. Int. Genet. Suppl. Ser. 2009, 2, 421–422. [Google Scholar] [CrossRef]
- Rojas, W.; Parra, M.V.; Campo, O.; Caro, M.A.; Lopera, J.G.; Arias, W.; Duque, C.; Naranjo, A.; García, J.; Vergara, C.; et al. Genetic make up and structure of Colombian populations by means of uniparental and biparental DNA markers. Am. J. Phys. Anthropol. 2010, 143, 13–20. [Google Scholar] [CrossRef]
- Acosta, M.A.; Blanco-Verea, A.; Lareu, M.V.; Brion, M.; Carracedo, A. The genetic male component of two South-Western Colombian populations. Forensic Sci. Int. Genet. 2009, 3, e59–e61. [Google Scholar] [CrossRef]
- Toscanini, U.; Brisighelli, F.; Moreno, F.; Pantoja-Astudillo, J.A.; Morales, E.A.; Bustos, P.; Pardo-Seco, J.; Salas, A. Analysis of Y-chromosome STRs in Chile confirms an extensive introgression of European male lineages in urban populations. Forensic Sci. Int. Genet. 2016, 21, 76–80. [Google Scholar] [CrossRef]
- Toscanini, U.; Gaviria, A.; Pardo-Seco, J.; Gómez-Carballa, A.; Moscoso, F.; Vela, M.; Cobos, S.; Lupero, A.; Zambrano, A.K.; Martinón-Torres, F.; et al. The geographic mosaic of Ecuadorian Y-chromosome ancestry. Forensic Sci. Int. Genet. 2018, 33, 59–65. [Google Scholar] [CrossRef]
- Villaescusa, P.; Seidel, M.; Nothnagel, M.; Pinotti, T.; González-Andrade, F.; Alvarez-Gila, O.; de Pancorbo, M.M.; Roewer, L. A Y-chromosomal survey of Ecuador’s multi-ethnic population reveals new insights into the tri-partite population structure and supports an early Holocene age of the rare Native American founder lineage C3-MPB373. Forensic Sci. Int. Genet. 2021, 51, 102427. [Google Scholar] [CrossRef] [PubMed]
- Vullo, C.; Gomes, V.; Romanini, C.; Oliveira, A.M.; Rocabado, O.; Aquino, J.; Amorim, A.; Gusmão, L. Association between Y haplogroups and autosomal AIMs reveals intra-population substructure in Bolivian populations. Int. J. Leg. Med. 2015, 129, 673–680. [Google Scholar] [CrossRef] [PubMed]
- Marcheco-Teruel, B.; Parra, E.J.; Fuentes-Smith, E.; Salas, A.; Buttenschøn, H.N.; Demontis, D.; Torres-Español, M.; Marín-Padrón, L.C.; Gómez-Cabezas, E.J.; Alvarez-Iglesias, V.; et al. Cuba: Exploring the history of admixture and the genetic basis of pigmentation using autosomal and uniparental markers. PLoS Genet. 2014, 10, e1004488. [Google Scholar] [CrossRef] [PubMed]
- Edwards, E.D. An African Tree Produces White Flowers: The Disappearance of the Black Community in Buenos Aires, Argentina, 1850–1890. McNair Sch. J. 2002, 6, 48–55. [Google Scholar]
- Edwards, E.D. The making of a White nation: The disappearance of the Black population in Argentina. Hist. Compass 2018, 16, e12456. [Google Scholar] [CrossRef]
- López-Ramírez, Y.L.; Aguilar-Velázquez, J.A.; López-Armenta, M.; Ruiz-Hernández, M.; Rangel-Villalobos, H. Paternal lineages and forensic parameters based on 23 Y-STRs (Powerplex® Y23) in Mestizo males from Mexico City. Int. J. Leg. Med. 2020, 134, 199–202. [Google Scholar] [CrossRef]
- Martínez-Cortés, G.; Salazar-Flores, J.; Fernández-Rodríguez, L.G.; Rubi-Castellanos, R.; Rodríguez-Loya, C.; Velarde-Félix, J.S.; Muñoz-Valle, J.F.; Parra-Rojas, I.; Rangel-Villalobos, H. Admixture and population structure in Mexican-Mestizos based on paternal lineages. J. Hum. Genet. 2012, 57, 568–574. [Google Scholar] [CrossRef]
- Núñez, C.; Baeta, M.; Fernández, M.; Zarrabeitia, M.; Martinez-Jarreta, B.; de Pancorbo, M.M. Highly discriminatory capacity of the PowerPlex(®) Y23 System for the study of isolated populations. Forensic Sci. Int. Genet. 2015, 17, 104–107. [Google Scholar] [CrossRef]
- Toscanini, U.; Gusmão, L.; Berardi, G.; Gomes, V.; Amorim, A.; Salas, A.; Raimondi, E. Male lineages in South American native groups: Evidence of M19 traveling south. Am. J. Phys. Anthropol. 2011, 146, 188–196. [Google Scholar] [CrossRef]
- Baily, S.L. Immigrants in the Lands of Promise: Italians in Buenos Aires and New York City, 1870–1914; Cornell University Press: Ithaca, NY, USA, 1999. [Google Scholar]
- Klein, H.S. A integração dos imigrantes italianos no Brasil, na Argentina e Estados Unidos. Novos Estudos CEBRAP 1989, 25, 95–117. [Google Scholar]
- Marquiegui, D. La inmigración española en la Argentina: Los gallegos de Luján, 1880–1920. Ciclos Hist. Econ. Soc. 1993, 3, 133–154. [Google Scholar]
- Solberg, C. Immigration and Nationalism: Argentina and Chile, 1890–1914; University of Texas Press: Austin, TX, USA, 1969. [Google Scholar]
- Yáñez Gallardo, C.R. Argentina como país de destino. La emigración española entre 1860–1930. Estud. Migr. Latinoam. 1989, 13, 467–498. [Google Scholar]
- Robino, C.; Ralf, A.; Pasino, S.; De Marchi, M.R.; Ballantyne, K.N.; Barbaro, A.; Bini, C.; Carnevali, E.; Casarino, L.; Di Gaetano, C.; et al. Development of an Italian RM Y-STR haplotype database: Results of the 2013 GEFI collaborative exercise. Forensic Sci. Int. Genet. 2015, 15, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Beleza, S.; Gusmão, L.; Lopes, A.; Alves, C.; Gomes, I.; Giouzeli, M.; Calafell, F.; Carracedo, A.; Amorim, A. Micro-phylogeographic and demographic history of Portuguese male lineages. Ann. Hum. Genet. 2006, 70, 181–194. [Google Scholar] [CrossRef] [PubMed]
- Cinnioğlu, C.; King, R.; Kivisild, T.; Kalfoğlu, E.; Atasoy, S.; Cavalleri, G.L.; Lillie, A.S.; Roseman, C.C.; Lin, A.A.; Prince, K.; et al. Excavating Y-chromosome haplotype strata in Anatolia. Hum. Genet. 2004, 114, 127–148. [Google Scholar] [CrossRef] [PubMed]
- Peričić, M.; Barać Lauc, L.; Martinović Klarić, I.; Janićijević, B.; Rudan, P. Review of Croatian genetic heritage as revealed by mitochondrial DNA and Y chromosomal lineages. Croat. Med. J. 2005, 46, 502–513. [Google Scholar]
- Ramos-Luis, E.; Blanco-Verea, A.; Brión, M.; van Huffel, V.; Sánchez-Diz, P.; Carracedo, A. Y-chromosomal DNA analysis in French male lineages. Forensic Sci. Int. Genet. 2014, 9, 162–168. [Google Scholar] [CrossRef]
- Sanchez, J.J.; Børsting, C.; Hernandez, A.; Mengel-Jørgensen, J.; Morling, N. Y chromosome SNP haplogroups in Danes, Greenlanders and Somalis. Int. Congr. Ser. 2004, 1261, 347–349. [Google Scholar] [CrossRef]
- Myres, N.M.; Rootsi, S.; Lin, A.A.; Järve, M.; King, R.J.; Kutuev, I.; Cabrera, V.M.; Khusnutdinova, E.K.; Pshenichnov, A.; Yunusbayev, B.; et al. A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe. Eur. J. Hum. Genet. 2011, 19, 95–101. [Google Scholar] [CrossRef]
- Busby, G.B.; Brisighelli, F.; Sánchez-Diz, P.; Ramos-Luis, E.; Martinez-Cadenas, C.; Thomas, M.G.; Bradley, D.G.; Gusmão, L.; Winney, B.; Bodmer, W.; et al. The peopling of Europe and the cautionary tale of Y chromosome lineage R-M269. Proc. Biol. Sci. 2012, 279, 884–892. [Google Scholar] [CrossRef]
- Cruciani, F.; Trombetta, B.; Antonelli, C.; Pascone, R.; Valesini, G.; Scalzi, V.; Vona, G.; Melegh, B.; Zagradisnik, B.; Assum, G.; et al. Strong intra- and inter-continental differentiation revealed by Y chromosome SNPs M269, U106 and U152. Forensic Sci. Int. Genet. 2011, 5, e49–e52. [Google Scholar] [CrossRef] [PubMed]
- Lucotte, G. The major Y-chromosome haplogroup R1b-M269 in West Europe, subdivided by the three SNPs S21/U106, S145/L21 and S28/U152, shows a clear pattern of geographic differentiation. Adv. Anthropol. 2015, 5, 22. [Google Scholar] [CrossRef]
- Zieger, M.; Utz, S. The Y-chromosomal haplotype and haplogroup distribution of modern Switzerland still reflects the alpine divide as a geographical barrier for human migration. Forensic Sci. Int. Genet. 2020, 48, 102345. [Google Scholar] [CrossRef] [PubMed]
- Valverde, L.; Illescas, M.J.; Villaescusa, P.; Gotor, A.M.; García, A.; Cardoso, S.; Algorta, J.; Catarino, S.; Rouault, K.; Férec, C.; et al. New clues to the evolutionary history of the main European paternal lineage M269: Dissection of the Y-SNP S116 in Atlantic Europe and Iberia. Eur. J. Hum. Genet. 2016, 24, 437–441. [Google Scholar] [CrossRef] [PubMed]
- Solé-Morata, N.; García-Fernández, C.; Urasin, V.; Bekada, A.; Fadhlaoui-Zid, K.; Zalloua, P.; Comas, D.; Calafell, F. Whole Y-chromosome sequences reveal an extremely recent origin of the most common North African paternal lineage E-M183 (M81). Sci. Rep. 2017, 7, 15941. [Google Scholar] [CrossRef]
- Altena, E.; Smeding, R.; van der Gaag, K.J.; Larmuseau, M.; Decorte, R.; Lao, O.; Kayser, M.; Kraaijenbrink, T.; de Knijff, P. The Dutch Y-chromosomal landscape. Eur. J. Hum. Genet. 2020, 28, 287–299. [Google Scholar] [CrossRef]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodrigues, P.; Velázquez, I.F.; Ribeiro, J.; Simão, F.; Amorim, A.; Carvalho, E.F.; Bravi, C.M.; Basso, N.G.; Real, L.E.; Galli, C.; et al. Tierra Del Fuego: What Is Left from the Precolonial Male Lineages? Genes 2022, 13, 1712. https://doi.org/10.3390/genes13101712
Rodrigues P, Velázquez IF, Ribeiro J, Simão F, Amorim A, Carvalho EF, Bravi CM, Basso NG, Real LE, Galli C, et al. Tierra Del Fuego: What Is Left from the Precolonial Male Lineages? Genes. 2022; 13(10):1712. https://doi.org/10.3390/genes13101712
Chicago/Turabian StyleRodrigues, Pedro, Irina Florencia Velázquez, Julyana Ribeiro, Filipa Simão, António Amorim, Elizeu F. Carvalho, Claudio Marcelo Bravi, Néstor Guillermo Basso, Luciano Esteban Real, Claudio Galli, and et al. 2022. "Tierra Del Fuego: What Is Left from the Precolonial Male Lineages?" Genes 13, no. 10: 1712. https://doi.org/10.3390/genes13101712
APA StyleRodrigues, P., Velázquez, I. F., Ribeiro, J., Simão, F., Amorim, A., Carvalho, E. F., Bravi, C. M., Basso, N. G., Real, L. E., Galli, C., González, A. d. C., Gamulin, A., Saldutti, R., Parolin, M. L., Gomes, V., & Gusmão, L. (2022). Tierra Del Fuego: What Is Left from the Precolonial Male Lineages? Genes, 13(10), 1712. https://doi.org/10.3390/genes13101712