Clinical Relevance of VEGFA (rs3025039) +936 C>T Polymorphism in Primary Myelofibrosis: Susceptibility, Clinical Co-Variates, and Outcomes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. SNV Analyses
2.3. Data Analyses
2.4. Statistical Analyses
3. Results
3.1. Correlation between VEGFA rs3025039 Genotypes and PMF
3.2. VEGFA rs3025039 Genotypes and Somatic Driver Mutations
3.3. VEGFA rs3025039 Genotypes and Clinical Co-Variates at Diagnosis
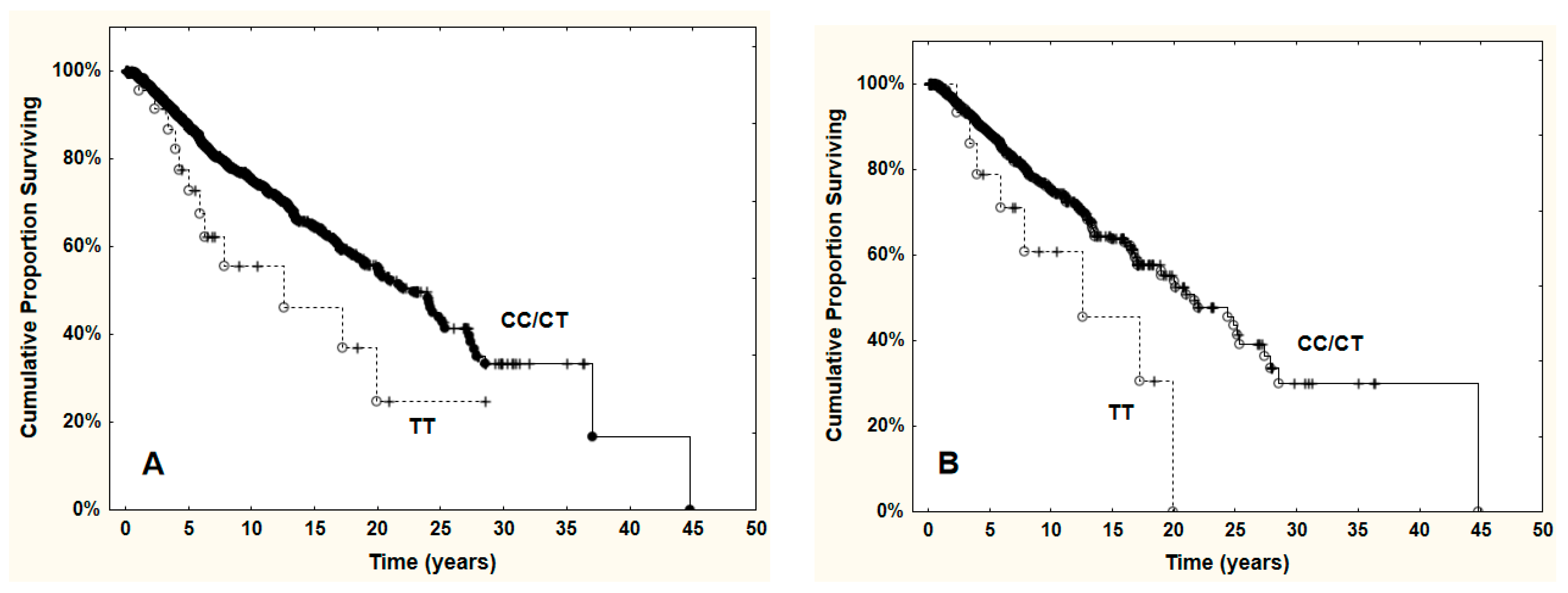
3.4. VEGFA rs3025039 Genotypes and Outcomes
3.5. VEGFA rs3025039 Genotypes and Thromboses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Grinfeld, J.; Nangalia, J.; Baxter, E.J.; Wedge, D.C.; Angelopoulos, N.; Cantrill, R.; Godfrey, A.L.; Papaemmanuil, E.; Gundem, G.; MacLean, C.; et al. Classification and Personalized Prognosis in Myeloproliferative Neoplasms. N. Engl. J. Med. 2018, 379, 1416–1430. [Google Scholar] [CrossRef] [PubMed]
- Gadomska, G.; Stankowska, K.; Boinska, J.; Ślusarz, R.; Tylicka, M.; Michalska, M.; Jachalska, A.; Rość, D. VEGF-A, sVEGFR-1, and sVEGFR-2 in BCR-ABL negative myeloproliferative neoplasms. Medicina 2017, 53, 34–39. [Google Scholar] [CrossRef] [PubMed]
- Medinger, M.; Passweg, J. Angiogenesis in myeloproliferative neoplasms, new markers and future directions. Memo-Mag. Eur. Med. Oncol. 2014, 7, 206–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boiocchi, L.; Vener, C.; Savi, F.; Bonoldi, E.; Moro, A.; Fracchiolla, N.S.; Iurlo, A.; Deliliers, G.L.; Coggi, G.; Bosari, S.; et al. Increased expression of vascular endothelial growth factor receptor 1 correlates with VEGF and microvessel density in Philadelphia chromosome-negative myeloproliferative neoplasms. J. Clin. Pathol. 2011, 64, 226–231. [Google Scholar] [CrossRef]
- Medinger, M.; Skoda, R.; Gratwohl, A.; Theocharides, A.; Buser, A.; Heim, D.; Dirnhofer, S.; Tichelli, A.; Tzankov, A. Angiogenesis and vascular endothelial growth factor-/receptor expression in myeloproliferative neoplasms: Correlation with clinical parameters and JAK2-V617F mutational status. Br. J. Haematol. 2009, 146, 150–157. [Google Scholar] [CrossRef]
- Alonci, A.; Allegra, A.; Bellomo, G.; Penna, G.; D’Angelo, A.; Quartarone, E.; Musolino, C. Evaluation of circulating endothelial cells, VEGF and VEGFR2 serum levels in patients with chronic myeloproliferative diseases. Hematol. Oncol. 2008, 26, 235–239. [Google Scholar] [CrossRef]
- Gianelli, U.; Vener, C.; Raviele, P.R.; Savi, F.; Somalvico, F.; Calori, R.; Iurlo, A.; Radaelli, F.; Fermo, E.; Bucciarelli, P.; et al. VEGF expression correlates with microvessel density in Philadelphia chromosome-negative chronic myeloproliferative disorders. Am. J. Clin. Pathol. 2007, 128, 966–973. [Google Scholar] [CrossRef] [Green Version]
- Panteli, K.; Bai, M.; Hatzimichael, E.; Zagorianakou, N.; Agnantis, N.J.; Bourantas, K. Serum levels, and bone marrow immunohistochemical expression of, vascular endothelial growth factor in patients with chronic myeloproliferative diseases. Hematology 2007, 12, 481–486. [Google Scholar] [CrossRef]
- Steurer, M.; Zoller, H.; Augustin, F.; Fong, D.; Heiss, S.; Strasser-Weippl, K.; Gastl, G.; Tzankov, A. Increased angiogenesis in chronic idiopathic myelofibrosis: Vascular endothelial growth factor as a prominent angiogenic factor. Hum. Pathol. 2007, 38, 1057–1064. [Google Scholar] [CrossRef]
- Ho, C.L.; Arora, B.; Hoyer, J.D.; Wellik, L.E.; Mesa, R.A.; Tefferi, A. Bone marrow expression of vascular endothelial growth factor in myelofibrosis with myeloid metaplasia. Eur. J. Haematol. 2005, 74, 35–39. [Google Scholar] [CrossRef]
- Di Raimondo, F.; Azzaro, M.P.; Palumbo, G.A.; Bagnato, S.; Stagno, F.; Giustolisi, G.M.; Cacciola, E.; Sortino, G.; Guglielmo, P.; Giustolisi, R. Elevated vascular endothelial growth factor (VEGF) serum levels in idiopathic myelofibrosis. Leukemia 2001, 15, 976–980. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grimm, C.; Watrowski, R.; Polterauer, S.; Baumühlner, K.; Natter, C.; Rahhal, J.; Heinze, G.; Schuster, E.; Hefler, L.; Reinthaller, A. Vascular endothelial growth factor gene polymorphisms and risk of cervical intraepithelial neoplasia. Int. J. Gynecol. Cancer 2011, 21, 597–601. [Google Scholar] [CrossRef] [PubMed]
- Hou, Q.; Li, M.-Y.; Huang, W.-T.; Wei, F.-F.; Peng, J.-P.; Lou, M.; Qiu, J.-G. Association between three VEGF polymorphisms and renal cell carcinoma susceptibility: A meta-analysis. Oncotarget 2017, 8, 50061–50070. [Google Scholar] [CrossRef]
- Kim, D.H.; Lee, N.Y.; Lee, M.-H.; Sohn, S.K.; Do, Y.R.; Park, J.Y. Vascular endothelial growth factor (VEGF) gene (VEGFA) polymorphism can predict the prognosis in acute myeloid leukaemia patients. Br. J. Haematol. 2007, 140, 71–79. [Google Scholar] [CrossRef] [Green Version]
- Mandal, R.K.; Yadav, S.S.; Panda, A.K.; Khattri, S. Vascular endothelial growth factor 936 c>T polymorphism increased oral cancer risk: Evidence from a meta-analysis. Genet. Test. Mol. Biomark. 2013, 17, 543–547. [Google Scholar] [CrossRef]
- Rodrigues, P.; Furriol, J.; Tormo, E.; Ballester, S.; Lluch, A.; Eroles, P. The single-nucleotide polymorphisms +936 C/T VEGF and 710 C/T VEGFR1 are associated with breast cancer protection in a Spanish population. Breast Cancer Res. Treat. 2012, 133, 769–778. [Google Scholar] [CrossRef]
- Zhang, J.; Yang, J.; Chen, Y.; Mao, Q.; Li, S.; Xiong, W.; Lin, Y.; Chen, J.; Ge, J. Genetic Variants of VEGF (rs201963 and rs3025039) and KDR (rs7667298, rs2305948, and rs1870377) Are Associated with Glioma Risk in a Han Chinese Population: A Case-Control Study. Mol. Neurobiol. 2015, 53, 2610–2618. [Google Scholar] [CrossRef]
- Swerdow, S.H.; Campo, E.; Harris, N.L.; Jaffe, E.S.; Pileri, S.A.; Stein, H.; Thiele, J. WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (Revised 4th Edition); IARC: Lyon, France, 2017. [Google Scholar]
- Barosi, G.; Viarengo, G.; Pecci, A.; Rosti, V.; Piaggio, G.; Marchetti, M.; Frassoni, F. Diagnostic and clinical relevance of the number of circulating CD34(+) cells in myelofibrosis with myeloid metaplasia. Blood J. Am. Soc. Hematol. 2001, 98, 3249–3255. [Google Scholar] [CrossRef]
- Barosi, G.; Rosti, V.; Catarsi, P.; Villani, L.; Abbà, C.; Carolei, A.; Magrini, U.; Gale, R.P.; Massa, M.; Campanelli, R. Reduced CXCR4-expression on CD34-positive blood cells predicts outcomes of persons with primary myelofibrosis. Leukemia 2021, 35, 468–475. [Google Scholar] [CrossRef] [PubMed]
- Barosi, G.; Massa, M.; Campanelli, R.; Fois, G.; Catarsi, P.; Viarengo, G.; Villani, L.; Poletto, V.; Bosoni, T.; Magrini, U.; et al. Primary myelofibrosis: Older age and high JAK2V617F allele burden are associated with elevated plasma high-sensitivity C-reactive protein levels and a phenotype of progressive disease. Leuk. Res. 2017, 60, 8–23. [Google Scholar] [CrossRef]
- Larson, D.P.; Akkari, Y.M.; Van Dyke, D.L.; Raca, G.; Gardner, J.A.; Rehder, C.W.; Kaiser-Rogers, K.A.; Eagle, P.; Yuhas, J.A.; Gu, J.; et al. Conventional Cytogenetic Analysis of Hematologic Neoplasms: A 20-Year Review of Proficiency Test Results From the College of American Pathologists/American College of Medical Genetics and Genomics Cytogenetics Committee. Arch. Pathol. Lab. Med. 2021, 145, 176–190. [Google Scholar] [CrossRef]
- Thiele, J.; Kvasnicka, H.M. Myelofibrosis—What’s in a name? Consensus on definition and EUMNET grading. Pathobiology 2007, 74, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Streiner, D.L.; Norman, G.R. Correction for multiple testing: Is there a resolution? Chest 2011, 140, 16–18. [Google Scholar] [CrossRef] [PubMed]
- Cervantes, F.; Dupriez, B.; Pereira, A.; Passamonti, F.; Reilly, J.T.; Morra, E.; Vannucchi, A.M.; Mesa, R.A.; Demory, J.L.; Barosi, G.; et al. New prognostic scoring system for primary myelofibrosis based on a study of the International Working Group for Myelofibrosis Research and Treatment. Blood J. Am. Soc. Hematol. 2009, 113, 2895–2901. [Google Scholar] [CrossRef] [PubMed]
- Jones, A.V.; Chase, A.; Silver, R.T.; Oscier, D.; Zoi, K.; Wang, Y.L.; Cario, H.; Pahl, H.L.; Collins, A.; Reiter, A.; et al. JAK2 haplotype is a major risk factor for the development of myeloproliferative neoplasms. Nat. Genet. 2009, 41, 446–449. [Google Scholar] [CrossRef] [Green Version]
- Olcaydu, D.; Harutyunyan, A.; Jäger, R.; Berg, T.; Gisslinger, B.; Pabinger, I.; Gisslinger, H.; Kralovics, R. A common JAK2 haplotype confers susceptibility to myeloproliferative neoplasms. Nat. Genet. 2009, 41, 450–454. [Google Scholar] [CrossRef] [PubMed]
- Kilpivaara, O.; Mukherjee, S.; Schram, A.M.; Wadleigh, M.; Mullally, A.; Ebert, B.L.; Bass, A.; Marubayashi, S.; Heguy, A.; Garcia-Manero, G.; et al. A germline JAK2 SNP is associated with predisposition to the development of JAK2(V617F)-positive myeloproliferative neoplasms. Nat. Genet. 2009, 41, 455–459. [Google Scholar] [CrossRef] [Green Version]
- Tapper, W.; Jones, A.V.; Kralovics, R.; Harutyunyan, A.S.; Zoi, K.; Leung, W.; Godfrey, A.L.; Guglielmelli, P.; Callaway, A.; Ward, D.; et al. Genetic variation at MECOM, TERT, JAK2 and HBS1L-MYB predisposes to myeloproliferative neoplasms. Nat. Commun. 2015, 6, 6691. [Google Scholar] [CrossRef] [Green Version]
- Trifa, A.P.; Bănescu, C.; Bojan, A.S.; Voina, C.M.; Popa, Ș.; Vișan, S.; Ciubean, A.D.; Tripon, F.; Dima, D.; Popov, V.M.; et al. MECOM, HBS1L-MYB, THRB-RARB, JAK2, and TERT polymorphisms defining the genetic predisposition to myeloproliferative neoplasms: A study on 939 patients. Am. J. Hematol. 2018, 93, 100–106. [Google Scholar] [CrossRef] [Green Version]
- Lighezan, D.L.; Bojan, A.S.; Iancu, M.; Pop, R.M.; Gligor-Popa, Ș.; Tripon, F.; Cosma, A.S.; Tomuleasa, C.; Dima, D.; Zdrenghea, M.; et al. TET2 rs1548483 SNP Associating with Susceptibility to Molecularly Annotated Polycythemia Vera and Primary Myelofibrosis. J. Pers. Med. 2020, 10, 259. [Google Scholar] [CrossRef]
- Tefferi, A.; Lasho, T.L.; Mudireddy, M.; Finke, C.M.; Hanson, C.A.; Ketterling, R.P.; Gangat, N.; Pardanani, A. The germline JAK2 GGCC (46/1) haplotype and survival among 414 molecularly-annotated patients with primary myelofibrosis. Am. J. Hematol. 2019, 94, 299–305. [Google Scholar] [CrossRef] [PubMed]
- Poletto, V.; Rosti, V.; Villani, L.; Catarsi, P.; Carolei, A.; Campanelli, R.; Massa, M.; Martinetti, M.; Viarengo, G.; Malovini, A.; et al. A3669G polymorphism of glucocorticoid receptor is a susceptibility allele for primary myelofibrosis and contributes to phenotypic diversity and blast transformation. Blood J. Am. Soc. Hematol. 2012, 120, 3112–3117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dong, P.P. Association of vascular endothelial growth factor expression and polymorphisms with the risk of gestational diabetes mellitus. J. Clin. Lab. Anal. 2019, 33, e22686. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhaskari, J.; Premalata, C.S.; Shilpa, V.; Rahul, B.; Pallavi, V.R.; Ramesh, G.; Krishnamoorthy, L. Vascular endothelial growth factor polymorphisms and a synchronized examination of plasma and tissue expression in epithelial ovarian cancers. Tumour. Biol. 2016, 37, 1017–1023. [Google Scholar] [CrossRef]
- Breunis, W.B.; Biezeveld, M.H.; Geissler, J.; Ottenkamp, J.; Kuipers, I.M.; Lam, J.; Hutchinson, A.; Welch, R.; Chanock, S.J.; Kuijpers, T.W. Vascular endothelial growth factor gene haplotypes in Kawasaki disease. Arthritis Rheum. Off. J. Am. Coll. Rheumatol. 2006, 54, 1588–1594. [Google Scholar] [CrossRef] [PubMed]
- Pasqualetti, G.; Danesi, R.; Del Tacca, M.; Bocci, G. Vascular endothelial growth factor pharmacogenetics: A new perspective for anti-angiogenic therapy. Pharmacogenomics 2007, 8, 49–66. [Google Scholar] [CrossRef]
- Sellami, N.; Lamine, L.B.; Turki, A.; Sarray, S.; Jailani, M.; Al-Ansari, A.K.; Ghorbel, M.; Mahjoub, T.; Almawi, W.Y. Association of VEGFA variants with altered VEGF secretion and type 2 diabetes: A case-control study. Cytokine 2018, 106, 29–34. [Google Scholar] [CrossRef]
- Al-Habboubi, H.H.; Sater, M.S.; Almawi, A.W.; Al-Khateeb, G.M.; Almawi, W.Y. Contribution of VEGF polymorphisms to variation in VEGF serum levels in a healthy population. Eur. Cytokine Netw. 2011, 22, 154–158. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mullaly, A. Both sides now: Losses and gains of mutant CALR. Blood J. Am. Soc. Hematol. 2020, 135, 82–83. [Google Scholar] [CrossRef]
- Balliu, M.; Calabresi, L.; Bartalucci, N.; Romagnoli, S.; Maggi, L.; Manfredini, R.; Lulli, M.; Guglielmelli, P.; Vannucchi, A.M. Activated IL-6 signaling contributes to the pathogenesis of, and is a novel therapeutic target for, CALR-mutated MPNs. Blood Adv. 2021, 5, 2184–2195. [Google Scholar] [CrossRef] [PubMed]
- Leone, P.; Shin, E.C.; Perosa, F.; Vacca, A.; Dammacco, F.; Racanelli, V. MHC class I antigen processing and presenting machinery: Organization, function, and defects in tumor cells. J. Natl. Cancer Inst. 2013, 105, 1172–1187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Beatty, G.L.; Gladney, W.L. Immune escape mechanisms as a guide for cancer immunotherapy. Clin. Cancer Res. 2015, 21, 687–692. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Barkal, A.A.; Weiskopf, K.; Kao, K.S.; Gordon, S.R.; Rosental, B.; Yiu, Y.Y.; George, B.M.; Markovic, M.; Ring, N.G.; Tsai, J.M.; et al. Engagement of MHC class I by the inhibitory receptor LILRB1 suppresses macrophages and is a target of cancer immunotherapy. Nat. Immunol. 2018, 19, 76–84. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, S.; Babiak, A.; Greiner, J. Immunotherapy for myeloproliferative neoplasms (MPN). Curr. Cancer Drug Targets 2011, 11, 72–84. [Google Scholar] [CrossRef] [PubMed]
- Barosi, G.; Campanelli, R.; Catarsi, P.; De Amici, M.; Abbà, C.; Viarengo, G.; Villani, L.; Gale, R.P.; Rosti, V.; Massa, M. Plasma sIL-2Rα levels are associated with disease progression in myelofibrosis with JAK2V617F but not CALR mutation. Leuk. Res. 2020, 90, 106319. [Google Scholar] [CrossRef] [PubMed]
- Nishi, A.; Milner, D.A., Jr.; Giovannucci, E.L.; Nishihara, R.; Tan, A.S.; Kawachi, I.; Ogino, S. Integration of molecular pathology, epidemiology and social science for global precision medicine. Expert Rev. Mol. Diagn. 2016, 16, 11–23. [Google Scholar] [CrossRef] [Green Version]
| Demographic Co-Variates | |
|---|---|
| Age, y, median (IQR) | 52 (46–61) |
| Male, N (%) | 503 (59) |
| Laboratory Co-Variates | |
| Hemoglobin, g/L, median (IQR) | 131 (109–148) |
| WBC × 109/L, median (IQR) | 8.5 (6.4–11.6) |
| Platelets × 109/L, median (IQR) | 467 (246–711) |
| Monocytes × 109/L, median (IQR) | 496 (332–688) |
| Spleen size, cm2, median (IQR) a | 120 (90–160) |
| Plasma LDH, × upper limit of normal (ULN), median (IQR) b | 1.29 (0.92–1.95) |
| Serum cholesterol, mg/dL, median (IQR) c | 158 (130–183) |
| Plasma high-sensitivity C-reactive protein, mg/dL, median (IQR) d | 10 (4–44) |
| Blood CD34-positive cells, × 106/L, median (IQR) e | 10 (4–43) |
| CXCR4/CD34, %, median (IQR) f | 41(21–63) |
| Molecular Co-Variates | |
| JAK2V617F-positive, N (%) | 544 (66) |
| JAK2V617F allele frequency, median (IQR) | 41 (22–68) |
| CALR mutation, N (%) | 171 (21) |
| MPL mutation, N (%) | 44 (5) |
| Triple negative, N (%) | 68 (8) |
| NGS detected mutations, N (%) g | 46 (20) |
| Cytogenetic abnormalities, N (%) h | 87 (30) |
| Bone Marrow Fibrosis Grade | |
| 0, N (%) | 258 (30) |
| 1, N (%) | 226 (27) |
| 2, N (%) | 245 (29) |
| 3, N (%) | 117 (14) |
| CC | CT | TT | CC/CT | CT/TT | T-Allele Frequency | ||
|---|---|---|---|---|---|---|---|
| PMF, N (%) | 849 | 601 (70.8) | 224 (26.4) | 24 (2.8) | 825 (97.2) | 248 (29.2) | 272/1698 (16) |
| Controls, N (%) | 250 | 165 (66) | 79 (31.6) | 6 (2.4) | 244 (97.6) | 85 (34) | 91/500 (18.2) |
| CT/TT (N = 248) vs. CC (N = 601) | TT (N = 24) vs. CC/CT (N = 825) | |||
|---|---|---|---|---|
| Outcome | HR (95% CI) | p-Value | HR (95% CI) | p-Value |
| Hemoglobin < 100 g/L | 1.16 (0.92, 1.45) | 0.20 | 1.41 (0.80, 2.44) | 0.23 |
| Spleen > 10 cm below left costal margin | 1.22 (0.97, 1.51) | 0.10 | 1.01 (0.51, 1.96) | 0.99 |
| WBC > 12 × 109/L | 1.10 (0.87, 1.37) | 0.45 | 1.02 (0.52, 1.98) | 0.94 |
| WBC < 4 × 109/L | 1.07 (0.73, 1.58) | 0.72 | 1.26 (0.46, 3.45) | 0.65 |
| Platelets < 150 × 109/L | 1.12 (0.93, 1.51) | 0.39 | 2,17 (1.25, 3.85) | 0.006 |
| Blood CD34-positive cells > 100 × 106/L | 1.25 (0.93, 1.51) | 0.11 | 1.11 (0.46, 2.70) | 0.81 |
| Transplant | 1.49 (0.98, 2.27) | 0.07 | 1.39 (0.44, 4.35) | 0.58 |
| Blast transformation | 1.15 (0.80, 1.65) | 0.43 | 1.22 (0.49, 2.94) | 0.67 |
| Death | 1.31 (1.00, 1.72) | 0.12 | 1.92 (1.06, 3.45) | 0.03 |
| VEGFA rs3025039 Genotype | CT/TT vs. CC OR (95% CI) | TT vs. CC/CT OR (95% CI) | ||||||
|---|---|---|---|---|---|---|---|---|
| All Subjects (N = 848) | CC (N = 600) | CT (N = 224) | TT (N = 24) | CC/CT (N = 824) | CT/TT (N = 248) | |||
| Thrombotic events, N (%) | 170 (20) | 124 (20.7) | 41 (18.3) | 5 (20.8) | 165 (20) | 46 (18.5) | OR = 0.87 (0.60, 1.27) p = 0.48 | OR = 1.05 (0.38, 2.85) p = 0.92 |
| Arterial thrombosis, N (% of PMF cases) | 49 (5.8) | 36 (6) | 12 (5.3) | 1 (4.2) | 48 (5.8) | 13 (5.2) | OR = 0.87 (0.45, 1.66) p = 0.67 | OR = 0.70 (0.09, 5.31) p = 0.73 |
| - In the year before diagnosis, N (% of thromboses) | 14 (28.6) | 10 (27.7) | 4 (33.3) | 0 (0) | 14 (29.2) | 4 (30.7) | ||
| - At diagnosis, N (% of thromboses) | 12 (24.5) | 9 (25) | 2 (16.7) | 1 (100) | 11 (22-9) | 3 (23.1) | ||
| - After diagnosis, N (% of thromboses) | 23 (46.9) | 17 (47.2) | 6 (50) | 0 (0) | 23 (48) | 6 (46.1) | ||
| Deep vein thrombosis in typical sites, N (% of PMF cases) | 24 (2.8) | 17 (2.8) | 4 (1.8) | 3 (12.5) | 21 (2.5) | 7 (2.8) | OR = 0.99 (0.41, 2.43) p = 0.99 | OR = 5.46 (1.51, 19.7) p = 0.0096 |
| - In the year before diagnosis, N (% of thromboses) | 3 (12.5) | 1 (5.9) | 1 (25) | 1 (33.3) | 2 (9.5) | 2 (28.6) | ||
| - At diagnosis, N (% of thromboses) | 5 (28.8) | 5 (29.4) | 0 (0) | 0 (0) | 5 (23.8) | 0 (0) | ||
| - After diagnosis, N (% of thromboses) | 16 (66.6) | 11 (64.7) | 3 (75) | 2 (66.6) | 14 (66.6) | 5 (71.4) | ||
| Vein thrombosis in atypical sites, N (% of PMF cases) | 97 (11.4) | 71 (11.8) | 25 (11.1) | 1 (4.2) | 96 (11.6) | 26 (9.7) | OR = 0.87 (0.54, 1.40) p = 0.57 | OR = 0.33 (0.04, 2.47) p = 0.28 |
| - In the year before diagnosis, N (% of thromboses) | 8 (8.2) | 6 (8.5) | 2 (8) | 0 (0) | 8 (8.3) | 2 (7.7) | ||
| - At diagnosis, N (% of thromboses) | 73 (75.2) | 53 (74.7) | 19 (76) | 1 (100) | 72 (75) | 20 (76.9) | ||
| - After diagnosis, N (% of thromboses) | 16 (16.5) | 12 (16.9) | 4 (16) | 0 (0) | 16 (16.7) | 4 (15.4) | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Villani, L.; Carolei, A.; Rosti, V.; Massa, M.; Campanelli, R.; Catarsi, P.; Abbà, C.; Gale, R.P.; Barosi, G. Clinical Relevance of VEGFA (rs3025039) +936 C>T Polymorphism in Primary Myelofibrosis: Susceptibility, Clinical Co-Variates, and Outcomes. Genes 2021, 12, 1271. https://doi.org/10.3390/genes12081271
Villani L, Carolei A, Rosti V, Massa M, Campanelli R, Catarsi P, Abbà C, Gale RP, Barosi G. Clinical Relevance of VEGFA (rs3025039) +936 C>T Polymorphism in Primary Myelofibrosis: Susceptibility, Clinical Co-Variates, and Outcomes. Genes. 2021; 12(8):1271. https://doi.org/10.3390/genes12081271
Chicago/Turabian StyleVillani, Laura, Adriana Carolei, Vittorio Rosti, Margherita Massa, Rita Campanelli, Paolo Catarsi, Carlotta Abbà, Robert Peter Gale, and Giovanni Barosi. 2021. "Clinical Relevance of VEGFA (rs3025039) +936 C>T Polymorphism in Primary Myelofibrosis: Susceptibility, Clinical Co-Variates, and Outcomes" Genes 12, no. 8: 1271. https://doi.org/10.3390/genes12081271
APA StyleVillani, L., Carolei, A., Rosti, V., Massa, M., Campanelli, R., Catarsi, P., Abbà, C., Gale, R. P., & Barosi, G. (2021). Clinical Relevance of VEGFA (rs3025039) +936 C>T Polymorphism in Primary Myelofibrosis: Susceptibility, Clinical Co-Variates, and Outcomes. Genes, 12(8), 1271. https://doi.org/10.3390/genes12081271






