A Genome-Wide Analysis of Pathogenesis-Related Protein-1 (PR-1) Genes from Piper nigrum Reveals Its Critical Role during Phytophthora capsici Infection
Abstract
:1. Introduction
2. Materials and Methods
2.1. Identification and Analysis of PR-1 Genes from the P. nigrum Genome
2.2. GO and KEGG Analysis
2.3. Secondary and Tertiary Structure Prediction
2.4. Role of PnPR-1 in P. capsici-Infected P. nigrum
2.5. Plant-Pathogen Infections, Staining and RT-qPCR
2.6. Prediction of cis-Acting Regulatory Elements in Promoter Regions
3. Results
3.1. Genome-Wide Identification and Analysis of P. nigrum PR-1 Genes
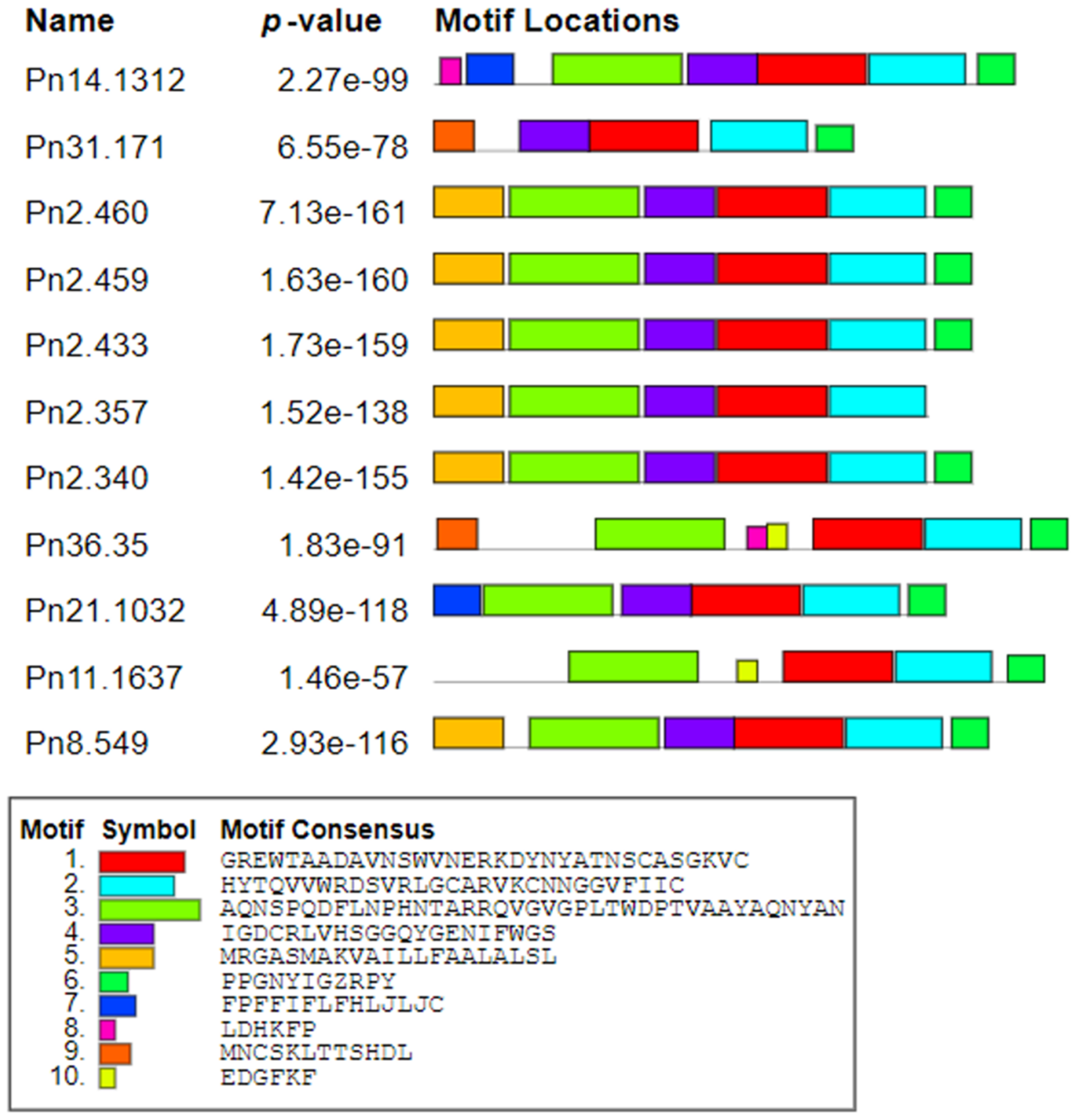
3.2. Sequence Conservation of PnPR-1 Genes
3.3. GO and KEGG Pathway Analysis of PnPR-1 Genes
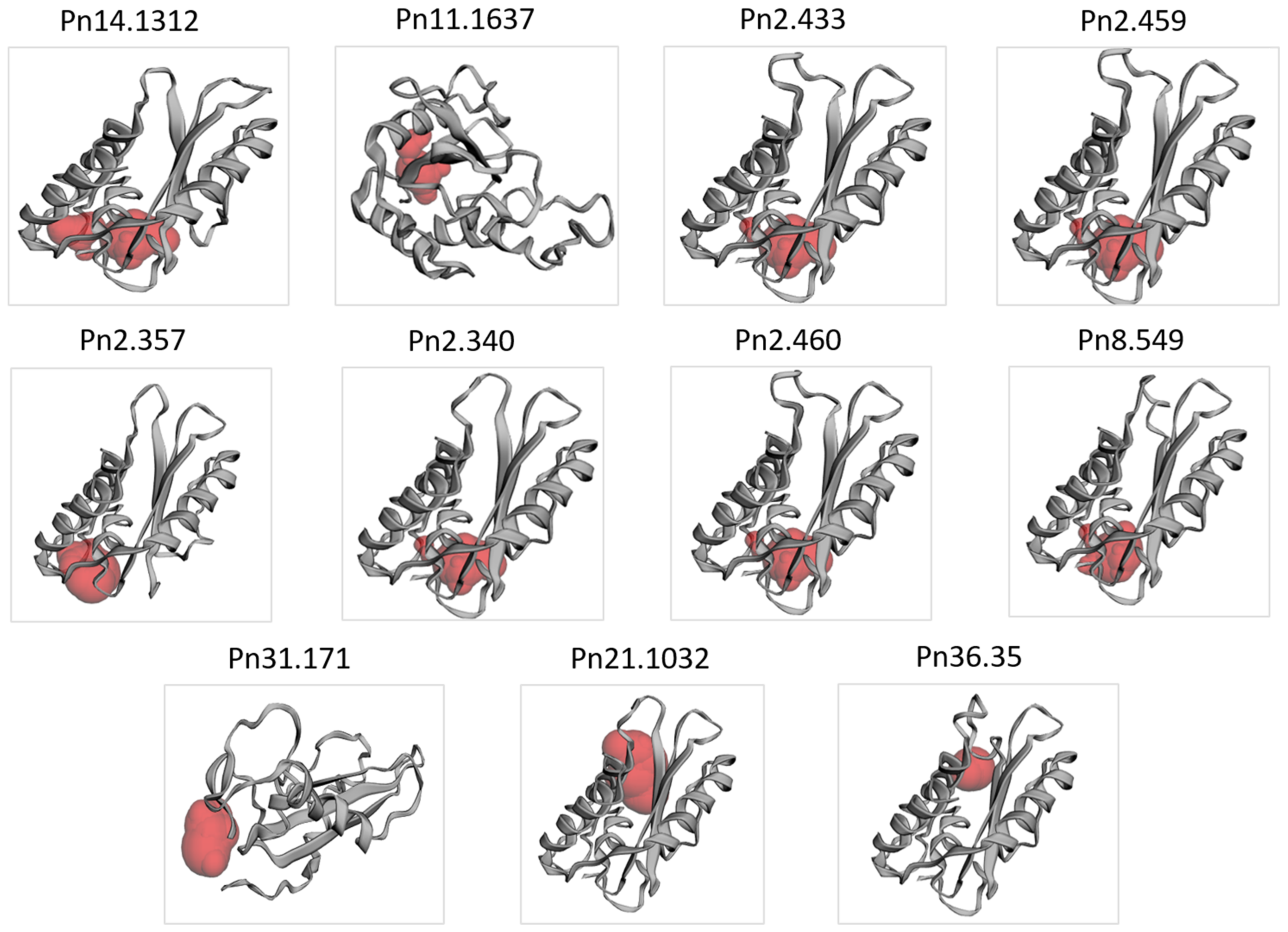
3.4. Secondary and 3D Structure of the PnPR-1 Protein
3.5. Cis-Regulatory Elements of the PnPR-1 Genes
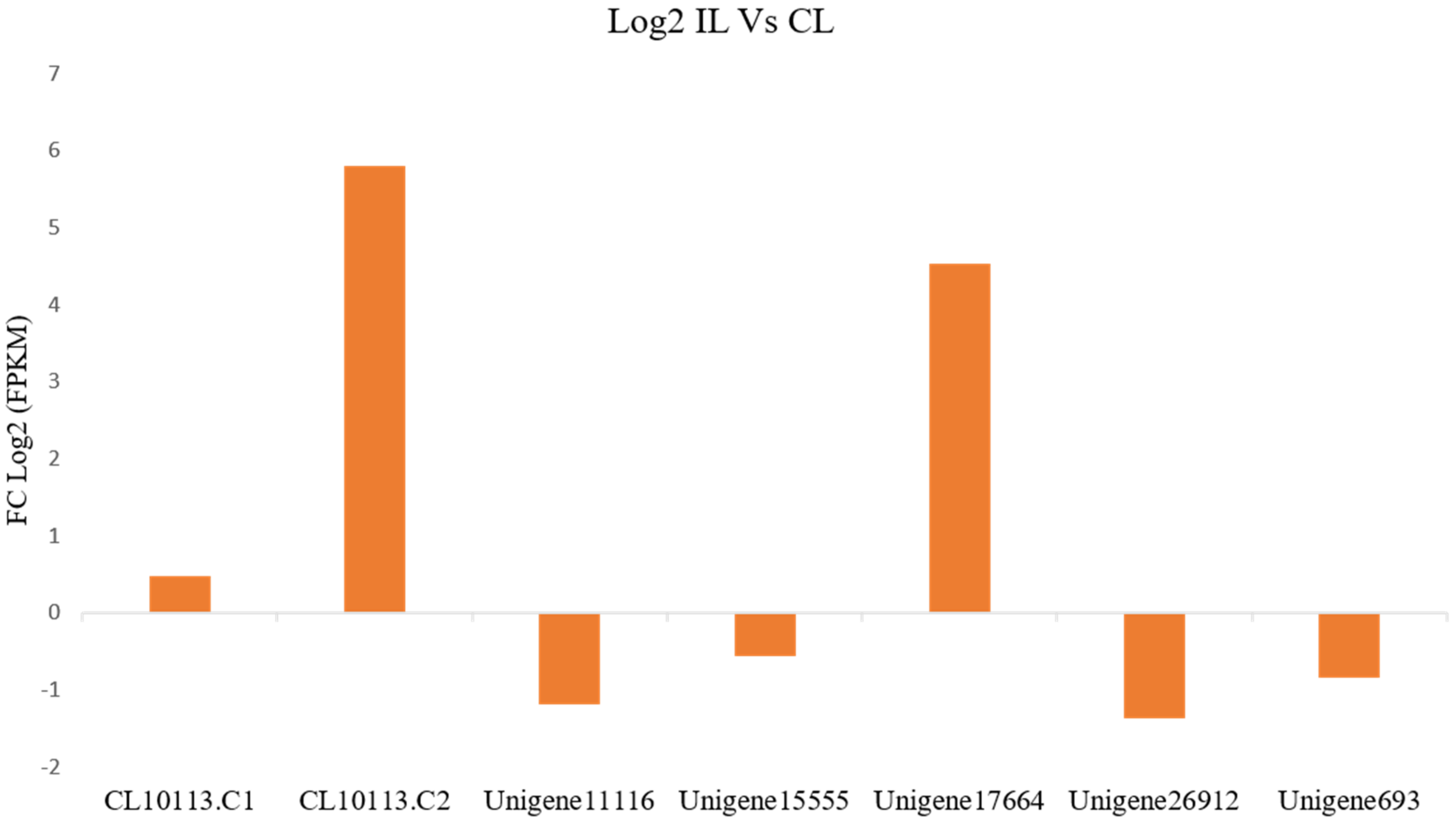
3.6. Expression of PR-1 Genes during P. capsici Infection in P. nigrum
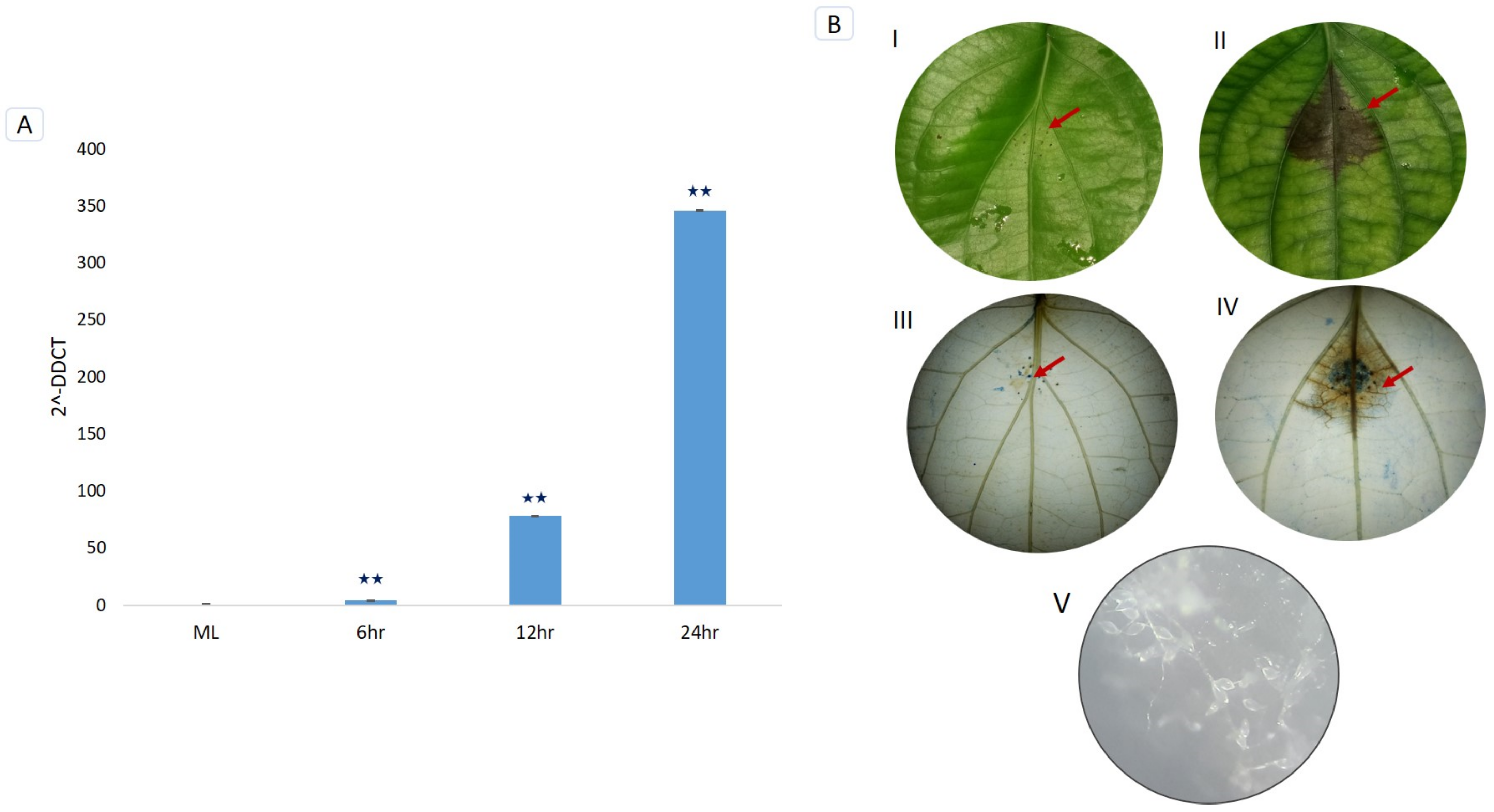
3.7. Trypan Blue Staining and Microscopic Detection of P. capsici Infection
3.8. RT-qPCR Validation of PnPR-1 Genes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dangl, J.L.; Jones, J.D.G. Plant pathogens and integrated defence responses to infection. Nature 2001, 411, 826–833. [Google Scholar] [CrossRef] [PubMed]
- Van Baarlen, P.; Van Belkum, A.; Summerbell, R.C.; Crous, P.W.; Thomma, B.P. Molecular mechanisms of pathogenicity: How do pathogenic microorganisms develop cross-kingdom host jumps? FEMS Microbiol. Rev. 2007, 31, 239–277. [Google Scholar] [CrossRef] [Green Version]
- Chassot, C.; Nawrath, C.; Métraux, J.-P. Cuticular defects lead to full immunity to a major plant pathogen. Plant J. 2007, 49, 972–980. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Boccardo, N.A.; Segretin, M.E.; Hernandez, I.; Mirkin, F.G.; Chacón, O.; Lopez, Y.; Borras-Hidalgo, O.; Bravo-Almonacid, F.F. Expression of pathogenesis-related proteins in transplastomic tobacco plants confers resistance to filamentous pathogens under field trials. Sci. Rep. 2019, 9, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Le Henanff, G.; Farine, S.; Kieffer-Mazet, F.; Miclot, A.-S.; Heitz, T.; Mestre, P.; Bertsch, C.; Chong, J. Vitis vinifera VvNPR1.1 is the functional ortholog of AtNPR1 and its overexpression in grapevine triggers constitutive activation of PR genes and enhanced resistance to powdery mildew. Planta 2011, 234, 405–417. [Google Scholar] [CrossRef]
- Molla, K.A.; Karmakar, S.; Chanda, P.; Sarkar, S.N.; Datta, S.K.; Datta, K. Tissue-specific expression of Arabidopsis NPR1 gene in rice for sheath blight resistance without compromising phenotypic cost. Plant Sci. 2016, 250, 105–114. [Google Scholar] [CrossRef] [PubMed]
- Malnoy, M.; Jin, Q.; Borejsza-Wysocka, E.E.; He, S.Y.; Aldwinckle, H.S. Overexpression of the apple MpNPR1 gene confers increased disease resistance in Malus x domestica. Mol. Plant Microbe Interact. 2007, 20, 1568–1580. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Christensen, A.B.; Cho, B.H.O.; Næsby, M.; Gregersen, P.L.; Brandt, J.; Madriz-Ordeñana, K.; Collinge, D.B.; Thordal-Christensen, H. The molecular characterization of two barley proteins establishes the novel PR-17 family of pathogenesis-related proteins. Mol. Plant Pathol. 2002, 3, 135–144. [Google Scholar] [CrossRef]
- Loake, G.; Grant, M. Salicylic acid in plant defence—The players and protagonists. Curr. Opin. Plant Biol. 2007, 10, 466–472. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of Inducible Defense-related Proteins in Infected Plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.-E.; Li, D.-W.; Zhang, Y.-L.; Zhao, Q.; He, Y.-M.; Gong, Z.-H. Defence responses of pepper (Capsicum annuum L.) infected with incompatible and compatible strains of Phytophthora capsici. Eur. J. Plant Pathol. 2013, 136, 625–638. [Google Scholar] [CrossRef]
- Akbudak, M.A.; Yildiz, S.; Filiz, E. Pathogenesis related protein-1 (PR-1) genes in tomato (Solanum lycopersicum L.): Bioinformatics analyses and expression profiles in response to drought stress. Genomics 2020, 112, 4089–4099. [Google Scholar] [CrossRef]
- Breen, S.; Williams, S.; Outram, M.; Kobe, B.; Solomon, P.S. Emerging Insights into the Functions of Pathogenesis-Related Protein 1. Trends Plant Sci. 2017, 22, 871–879. [Google Scholar] [CrossRef] [PubMed]
- Kiba, A.; Nishihara, M.; Nakatsuka, T.; Yamamura, S. Pathogenesis-related protein 1 homologue is an antifungal protein in Wasabia japonica leaves and confers resistance to Botrytis cinerea in transgenic tobacco. Plant Biotechnol. 2007, 24, 247–253. [Google Scholar] [CrossRef]
- Sarowar, S.; Young, J.K.; Eui, N.K.; Ki, D.K.; Byung, K.H.; Islam, R.; Jeong, S.S. Overexpression of a pepper basic pathogenesis-related protein 1 gene in tobacco plants enhances resistance to heavy metal and pathogen stresses. Plant Cell Rep. 2005, 24, 216–224. [Google Scholar] [CrossRef]
- Shin, S.H.; Pak, J.-H.; Kim, M.J.; Kim, H.J.; Oh, J.S.; Choi, H.K.; Jung-Hun, P.; Chung, Y.S. An Acidic PATHOGENESIS-RELATED1 Gene of Oryza grandiglumis is Involved in Disease Resistance Response Against Bacterial Infection. Plant Pathol. J. 2014, 30, 208–214. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cooper, B.; Clarke, J.D.; Budworth, P.; Kreps, J.; Hutchison, D.; Park, S.; Guimil, S.; Dunn, M.; Luginbühl, P.; Ellero, C.; et al. A network of rice genes associated with stress response and seed development. Proc. Natl. Acad. Sci. USA 2003, 100, 4945–4950. [Google Scholar] [CrossRef] [Green Version]
- Lotan, T.; Ori, N.; Fluhr, R. Pathogenesis-related proteins are developmentally regulated in tobacco flowers. Plant Cell 1989, 1, 881–887. [Google Scholar]
- Chen, Y.L.; Lee, C.Y.; Cheng, K.T.; Chang, W.H.; Huang, R.N.; Nam, H.G.; Chen, Y.R. Quantitative peptidomics study reveals that a wound-induced peptide from PR-1 regulates immune signaling in tomato. Plant Cell 2014, 26, 4135–4148. [Google Scholar] [CrossRef] [Green Version]
- Gamir, J.; Darwiche, R.; Hof, P.V.; Choudhary, V.; Stumpe, M.; Schneiter, R.; Mauch, F. The sterol-binding activity of PATHOGENESIS-RELATED PROTEIN 1 reveals the mode of action of an antimicrobial protein. Plant J. 2017, 89, 502–509. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Asha, S.; Sreekumar, S.; Soniya, E.V. Unravelling the complexity of microRNA-mediated gene regulation in black pepper (Piper nigrum L.) using high-throughput small RNA profiling. Plant Cell Rep. 2016, 35, 53–63. [Google Scholar] [CrossRef]
- Hu, L.; Xu, Z.; Wang, M.; Fan, R.; Yuan, D.; Wu, B.; Wu, H.; Qin, X.; Yan, L.; Tan, L.; et al. The chromosome-scale reference genome of black pepper provides insight into piperine biosynthesis. Nat. Commun. 2019, 10, 1–11. [Google Scholar] [CrossRef]
- Hall, T.A. BIOEDIT: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/ NT. In Nucleic Acids Symposium Series; Information Retrieval Ltd.: London, UK, 1999. [Google Scholar]
- Marchler-Bauer, A.; Derbyshire, M.K.; Gonzales, N.R.; Lu, S.; Chitsaz, F.; Geer, L.Y.; Geer, R.C.; He, J.; Gwadz, M.; Hurwitz, D.I.; et al. CDD: NCBI’s conserved domain database. Nucleic Acids Res. 2015, 43, D222–D226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef] [Green Version]
- Törönen, P.; Medlar, A.; Holm, L. PANNZER2: A rapid functional annotation web server. Nucleic Acids Res. 2018, 46, W84–W88. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Sato, Y.; Morishima, K. BlastKOALA and GhostKOALA: KEGG Tools for Functional Characterization of Genome and Metagenome Sequences. J. Mol. Biol. 2016, 428, 726–731. [Google Scholar] [CrossRef] [Green Version]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.; Sternberg, M.J.E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef] [Green Version]
- Dundas, J.; Ouyang, Z.; Tseng, J.; Binkowski, A.; Turpaz, Y.; Liang, J. CASTp: Computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res. 2006, 34, W116–W118. [Google Scholar] [CrossRef] [Green Version]
- Fernández-Bautista, N.; Domínguez-Núñez, J.; Moreno, M.M.; Berrocal-Lobo, M. Plant Tissue Trypan Blue Staining during Phytopathogen Infection. Bio-Protocol 2016, 6. [Google Scholar] [CrossRef] [Green Version]
- Asha, S.; Soniya, E.V. Transfer RNA Derived Small RNAs Targeting Defense Responsive Genes Are Induced during Phytophthora capsici Infection in Black Pepper (Piper nigrum L.). Front. Plant Sci. 2016, 7, 1–16. [Google Scholar] [CrossRef] [Green Version]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-DeltaDeltaC(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Higo, K.; Ugawa, Y.; Iwamoto, M.; Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 1999, 27, 297–300. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaur, A.; Pati, P.K.; Pati, A.M.; Nagpal, A.K. In-silico analysis of cis-acting regulatory elements of pathogenesis-related proteins of Arabidopsis thaliana and Oryza sativa. PLoS ONE 2017, 12, e0184523. [Google Scholar] [CrossRef] [Green Version]
- Agrios, G.N. How plants defend themselves against pathogens. In Plant Pathology, 5th ed.; Elsevier Academic Press: Amsterdam, The Netherlands, 2005. [Google Scholar]
- Van Loon, L.C.; Gerritsen, Y.A.M.; Ritter, C.E. Identification, purification, and characterization of pathogenesis-related proteins from virus-infected Samsun NN tobacco leaves. Plant Mol. Biol. 1987, 9, 593–609. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Pierpoint, W.S.; Boller, T.; Conejero, V. Recommendations for naming plant pathogenesis-related proteins. Plant Mol. Biol. Rep. 1994, 12, 245–264. [Google Scholar] [CrossRef]
- Memelink, J.; Linthorst, H.J.M.; Schilperoort, R.A.; Hoge, J.H.C. Tobacco genes encoding acidic and basic isoforms of pathogenesis-related proteins display different expression patterns. Plant Mol. Biol. 1990, 14, 119–126. [Google Scholar] [CrossRef] [PubMed]
- Mitsuhara, I.; Iwai, T.; Seo, S.; Yanagawa, Y.; Kawahigasi, H.; Hirose, S.; Ohkawa, Y.; Ohashi, Y. Characteristic expression of twelve rice PR1 family genes in response to pathogen infection, wounding, and defense-related signal compounds (121/180). Mol. Genet. Genom. 2008, 279, 415–427. [Google Scholar] [CrossRef] [Green Version]
- Almeida-Silva, F.; Venancio, T.M. Pathogenesis-related protein 1 (PR-1) genes in soybean: Genome-wide identification, structural analysis and expression profiling under multiple biotic and abiotic stresses. bioRxiv 2021. [Google Scholar] [CrossRef]
- Choudhary, V.; Darwiche, R.; Gfeller, D.; Zoete, V.; Michielin, O.; Schneiter, R. The caveolin-binding motif of the pathogen-related yeast protein Pry1, a member of the CAP protein superfamily, is required for in vivo export of cholesteryl acetate. J. Lipid Res. 2014, 55, 883–894. [Google Scholar] [CrossRef] [Green Version]
- Lincoln, J.E.; Sanchez, J.P.; Zumstein, K.; Gilchrist, D.G. Plant and animal PR1 family members inhibit programmed cell death and suppress bacterial pathogens in plant tissues. Mol. Plant Pathol. 2018, 19, 2111–2123. [Google Scholar] [CrossRef] [Green Version]
- Breen, S.; Williams, S.J.; Winterberg, B.; Kobe, B.; Solomon, P.S. Wheat PR-1 proteins are targeted by necrotrophic pathogen effector proteins. Plant J. 2016, 88, 13–25. [Google Scholar] [CrossRef] [Green Version]
- Lu, S.; Faris, J.; Sherwood, R.; Friesen, T.L.; Edwards, M.C. A dimeric PR-1-type pathogenesis-related protein interacts with ToxA and potentially mediates ToxA-induced necrosis in sensitive wheat. Mol. Plant Pathol. 2014, 15, 650–663. [Google Scholar] [CrossRef]
- Danino, Y.M.; Even, D.; Ideses, D.; Juven-Gershon, T. The core promoter: At the heart of gene expression. Biochim. Biophys. Acta BBA Bioenerg. 2015, 1849, 1116–1131. [Google Scholar] [CrossRef]
- Kaur, G.; Pati, P.K. Analysis of cis-acting regulatory elements of Respiratory burst oxidase homolog ( Rboh ) gene families in Arabidopsis and rice provides clues for their diverse functions. Comput. Biol. Chem. 2016, 62, 104–118. [Google Scholar] [CrossRef]
- Ross, C.; Shen, Q.J. Computational Prediction and Experimental Verification of HVA1-like Abscisic Acid Responsive Promoters in Rice (Oryza sativa). Plant Mol. Biol. 2006, 62, 233–246. [Google Scholar] [CrossRef]
- Kesarwani, M.; Yoo, J.; Dong, X. Genetic Interactions of TGA Transcription Factors in the Regulation of Pathogenesis-Related Genes and Disease Resistance in Arabidopsis. Plant Physiol. 2007, 144, 336–346. [Google Scholar] [CrossRef] [Green Version]
- Fujimoto, S.Y.; Ohta, M.; Usui, A.; Shinshi, H.; Ohme-Takagi, M. Arabidopsis ethylene-responsive element binding factors act as transcriptional activators or repressors of GCC box-mediated gene expression. Plant Cell 2000, 12, 393–404. [Google Scholar] [PubMed] [Green Version]
- Foyer, C.H.; Kerchev, P.I.; Hancock, R.D. The ABA-INSENSITIVE-4 (ABI4) transcription factor links redox, hormone and sugar signaling pathways. Plant Signal. Behav. 2012, 7, 276–281. [Google Scholar] [CrossRef] [Green Version]
- Akagi, T.; Ikegami, A.; Yonemori, K. DkMyb2 wound-induced transcription factor of persimmon (Diospyros kaki Thunb.), contributes to proanthocyanidin regulation. Planta 2010, 232, 1045–1059. [Google Scholar] [CrossRef] [PubMed]
- Ryu, K.H.; Kang, Y.H.; Park, Y.H.; Hwang, I.; Schiefelbein, J.; Lee, M.M. The WEREWOLF MYB protein directly regulates CAPRICE transcription during cell fate specification in the Arabidopsis root epidermis. Development 2005, 132, 4765–4775. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Taoka, K.-I.; Ohtsubo, N.; Fujimoto, Y.; Mikami, K.; Meshi, T.; Iwabuchi, M. The Modular Structure and Function of the Wheat H1 Promoter with S Phase-Specific Activity. Plant Cell Physiol. 1998, 39, 294–306. [Google Scholar] [CrossRef] [Green Version]
- Ali, S.; Mir, Z.A.; Bhat, J.A.; Tyagi, A.; Chandrashekar, N.; Yadav, P.; Rawat, S.; Sultana, M.; Grover, A. Isolation and characterization of systemic acquired resistance marker gene PR1 and its promoter from Brassica juncea. 3 Biotech 2018, 8, 10. [Google Scholar] [CrossRef] [PubMed]
- Tunsagool, P.; Jutidamrongphan, W.; Phaonakrop, N.; Jaresitthikunchai, J.; Roytrakul, S.; Leelasuphakul, W. Insights into stress responses in mandarins triggered by Bacillus subtilis cyclic lipopeptides and exogenous plant hormones upon Penicillium digitatum infection. Plant Cell Rep. 2019, 38, 559–575. [Google Scholar] [CrossRef]
- Ali, S.; Mir, Z.A.; Tyagi, A.; Bhat, J.A.; Chandrashekar, N.; Papolu, P.K.; Rawat, S.; Grover, A. Identification and comparative analysis of Brassica juncea pathogenesis-related genes in response to hormonal, biotic and abiotic stresses. Acta Physiol. Plant. 2017, 39, 268. [Google Scholar] [CrossRef]
- Seo, P.J.; Kim, M.J.; Park, J.-Y.; Kim, S.-Y.; Jeon, J.; Lee, Y.-H.; Kim, J.; Park, C.-M. Cold activation of a plasma membrane-tethered NAC transcription factor induces a pathogen resistance response in Arabidopsis. Plant J. 2010, 61, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Kothari, K.S.; Dansana, P.K.; Giri, J.; Tyagi, A.K. Rice Stress Associated Protein 1 (OsSAP1) Interacts with Aminotransferase (OsAMTR1) and Pathogenesis-Related 1a Protein (OsSCP) and Regulates Abiotic Stress Responses. Front. Plant Sci. 2016, 7, 1057. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, W.-X.; Zhang, F.-C.; Zhang, W.-Z.; Song, L.-F.; Wu, W.-H.; Chen, Y.-F. Arabidopsis Di19 Functions as a Transcription Factor and Modulates PR1, PR2, and PR5 Expression in Response to Drought Stress. Mol. Plant 2013, 6, 1487–1502. [Google Scholar] [CrossRef] [Green Version]
- Nazeem, P.A.; Achuthan, C.R.; Babu, T.D.; Parab, G.V.; Girija, D.; Keshavachandran, R.; Samiyappan, R. Expression of pathogenesis related proteins in black pepper (Piper nigrum L.) in relation to Phytophthora foot rot disease. J. Trop. Agric. 2008, 46, 45–51. [Google Scholar]
- Trang Anh, T.; Bao Linh, T.; Vu Phong, N.; Lan Thanh Bien, T.; Thi Nha Tram, T.; Dinh Don, L. Expression of Proteins Related to Phytophthora capsici Tolerance in Black Pepper (Piper nigrum L.). Int. J. Agric. Innov. Res. 2018, 6, 2319-1473. [Google Scholar]
- Vijesh Kumar, I.P.; Johnson, G.K.; Anandaraj, M. Real-Time Quantitative RT-PCR of Some Defense Inoculated with Phytophthora Capsici. Int. J. Agric. Sci. Res. 2016, 6, 69–78. [Google Scholar]


| Genome CDS Id | Genome Scaffold No. | Exon No. | Start | Stop | Strand | Protein Length (AA) | Molecular Weight (kDa) | Theoretical pI | |
|---|---|---|---|---|---|---|---|---|---|
| Pn2.460 | Pn23 | 1 | 1315655 | 1316146 | − | 155 | 17.09488 | 4.82 | Acidic |
| Pn2.459 | Pn23 | 1 | 1320928 | 1321419 | − | 192 | 21.91361 | 5.8 | |
| Pn2.433 | Pn23 | 1 | 1493563 | 1494054 | − | 168 | 18.35074 | 7.58 | Basic |
| Pn2.357 | Pn23 | 2 | 2226595 | 2229193 | − | 127 | 14.37881 | 7.62 | |
| Pn2.340 | Pn23 | 1 | 2402850 | 2403341 | − | 185 | 19.80756 | 9.1 | |
| Pn21.1032 | Pn3 | 1 | 3202437 | 3202904 | + | 163 | 17.85002 | 9.15 | |
| Pn31.171 | Pn15 | 1 | 26854949 | 26855332 | − | 163 | 17.87807 | 9.3 | |
| Pn36.35 | Pn25 | 1 | 5754975 | 5755553 | − | 163 | 17.88005 | 9.3 | |
| Pn11.1637 | Pn4 | 1 | 33322945 | 33323502 | − | 176 | 19.63337 | 9.37 | |
| Pn8.549 | Pn8 | 1 | 25779763 | 25780269 | − | 163 | 17.96421 | 9.44 | |
| Pn14.1312 | Pn14 | 1 | 5932334 | 5932864 | + | 357 | 38.49596 | 11.3 | |
| Genome Id | Cleavage Site Position | Sequence Position | Probability | Protein Type | Signal Peptide (Sec/SPI) | Other |
|---|---|---|---|---|---|---|
| Pn21.1032 | 16 and 17 | CNA-QN | 0.9541 | Likelihood | 0.9981 | 0.0019 |
| Pn2.340 | 24 and 25 | AQA-QN | 0.8849 | Likelihood | 0.9984 | 0.0016 |
| Pn2.357 | 24 and 25 | AQA-QN | 0.8898 | Likelihood | 0.9986 | 0.0014 |
| Pn2.433 | 24 and 25 | AQA-QN | 0.8892 | Likelihood | 0.9987 | 0.0013 |
| Pn2.459 | 24 and 25 | AQA-QN | 0.8898 | Likelihood | 0.9986 | 0.0014 |
| Pn2.460 | 24 and 25 | AQA-QN | 0.8892 | Likelihood | 0.9987 | 0.0013 |
| Pn36.35 | 29 and 30 | ASS-SP | 0.6292 | Likelihood | 0.9813 | 0.0187 |
| Pn14.1312 | 31 and 32 | TNA-AL | 0.4399 | Likelihood | 0.7324 | 0.2676 |
| Pn8.549 | 30 and 31 | TLA-QN | 0.3454 | Likelihood | 0.5106 | 0.4894 |
| Pn31.171 | - | - | - | Likelihood | 0.0029 | 0.9971 |
| Pn11.1637 | - | - | - | Likelihood | 0.2691 | 0.7309 |
| GO ID | GO Domain | Function Description |
|---|---|---|
| GO:0006952 | Biological process | Defense response |
| GO:0009607 | Biological process | Response to biotic stimulus |
| GO:0048544 | Biological process | Recognition of pollen |
| GO:0006468 | Biological process | Protein phosphorylation |
| GO:0010274 | Biological process | Hydrotropism |
| GO:0004672 | Molecular function | Protein kinase activity |
| GO:0030554 | Molecular function | Adenyl nucleotide binding |
| GO:0035639 | Molecular function | Purine ribonucleoside triphosphate binding |
| GO:0032555 | Molecular function | Purine ribonucleotide binding |
| GO:0005576 | Cellular component | Extracellular region |
| GO:0016020 | Cellular component | Membrane |
| GO:0031224 | Cellular component | Intrinsic component of membrane |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kattupalli, D.; Srinivasan, A.; Soniya, E.V. A Genome-Wide Analysis of Pathogenesis-Related Protein-1 (PR-1) Genes from Piper nigrum Reveals Its Critical Role during Phytophthora capsici Infection. Genes 2021, 12, 1007. https://doi.org/10.3390/genes12071007
Kattupalli D, Srinivasan A, Soniya EV. A Genome-Wide Analysis of Pathogenesis-Related Protein-1 (PR-1) Genes from Piper nigrum Reveals Its Critical Role during Phytophthora capsici Infection. Genes. 2021; 12(7):1007. https://doi.org/10.3390/genes12071007
Chicago/Turabian StyleKattupalli, Divya, Asha Srinivasan, and Eppurath Vasudevan Soniya. 2021. "A Genome-Wide Analysis of Pathogenesis-Related Protein-1 (PR-1) Genes from Piper nigrum Reveals Its Critical Role during Phytophthora capsici Infection" Genes 12, no. 7: 1007. https://doi.org/10.3390/genes12071007
APA StyleKattupalli, D., Srinivasan, A., & Soniya, E. V. (2021). A Genome-Wide Analysis of Pathogenesis-Related Protein-1 (PR-1) Genes from Piper nigrum Reveals Its Critical Role during Phytophthora capsici Infection. Genes, 12(7), 1007. https://doi.org/10.3390/genes12071007





