From Stem Cells to Populations—Using hiPSC, Next-Generation Sequencing, and GWAS to Explore the Genetic and Molecular Mechanisms of Congenital Heart Defects
Abstract
1. Introduction
2. Identification of Causative Genes by Linkage Analysis and Chromosomal Microarray
3. Next-Generation Sequencing as a Tool for CHD
4. Genome-Wide Association Studies
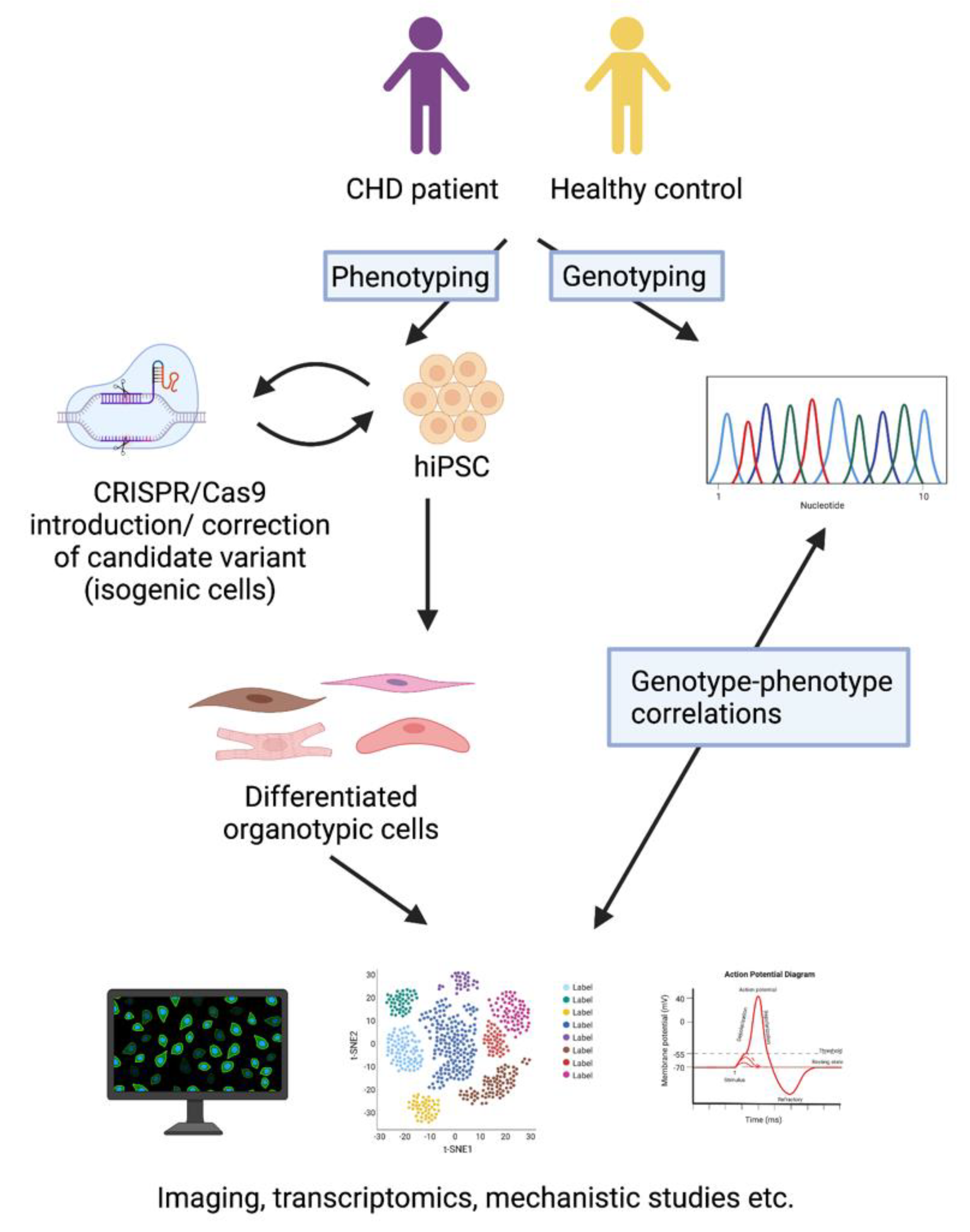
5. Human Induced Pluripotent Stem Cells
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Reller, M.D.; Strickland, M.J.; Riehle-Colarusso, T.; Mahle, W.T.; Correa, A. Prevalence of Congenital Heart Defects in Metropolitan Atlanta, 1998–2005. J. Pediatr. 2008, 153, 807–813. [Google Scholar] [CrossRef]
- Kerstjens-Frederikse, W.S.; van de Laar, I.M.B.H.; Vos, Y.J.; Verhagen, J.M.A.; Berger, R.M.F.; Lichtenbelt, K.D.; Klein Wassink-Ruiter, J.S.; van der Zwaag, P.A.; du Marchie Sarvaas, G.J.; Bergman, K.A.; et al. Cardiovascular Malformations Caused by NOTCH1 Mutations Do Not Keep Left: Data on 428 Probands with Left-Sided CHD and Their Families. Genet. Med. 2016, 18, 914–923. [Google Scholar] [CrossRef]
- Schott, J.J.; Benson, D.W.; Basson, C.T.; Pease, W.; Silberbach, G.M.; Moak, J.P.; Maron, B.J.; Seidman, C.E.; Seidman, J.G. Congenital Heart Disease Caused by Mutations in the Transcription Factor NKX2-5. Science 1998, 281, 108–111. [Google Scholar] [CrossRef]
- Blue, G.M.; Kirk, E.P.; Giannoulatou, E.; Dunwoodie, S.L.; Ho, J.W.K.; Hilton, D.C.K.; White, S.M.; Sholler, G.F.; Harvey, R.P.; Winlaw, D.S. Targeted next-Generation Sequencing Identifies Pathogenic Variants in Familial Congenital Heart Disease. J. Am. Coll. Cardiol. 2014, 64, 2498–2506. [Google Scholar] [CrossRef]
- Jia, Y.; Louw, J.J.; Breckpot, J.; Callewaert, B.; Barrea, C.; Sznajer, Y.; Gewillig, M.; Souche, E.; Dehaspe, L.; Vermeesch, J.R.; et al. The Diagnostic Value of next Generation Sequencing in Familial Nonsyndromic Congenital Heart Defects. Am. J. Med. Genet. A 2015, 167A, 1822–1829. [Google Scholar] [CrossRef]
- Szot, J.O.; Cuny, H.; Blue, G.M.; Humphreys, D.T.; Ip, E.; Harrison, K.; Sholler, G.F.; Giannoulatou, E.; Leo, P.; Duncan, E.L.; et al. A Screening Approach to Identify Clinically Actionable Variants Causing Congenital Heart Disease in Exome Data. Circ. Genom. Precis. Med. 2018, 11, e001978. [Google Scholar] [CrossRef]
- Zaidi, S.; Choi, M.; Wakimoto, H.; Ma, L.; Jiang, J.; Overton, J.D.; Romano-Adesman, A.; Bjornson, R.D.; Breitbart, R.E.; Brown, K.K.; et al. De Novo Mutations in Histone-Modifying Genes in Congenital Heart Disease. Nature 2013, 498, 220–223. [Google Scholar] [CrossRef]
- Blue, G.M.; Kirk, E.P.; Giannoulatou, E.; Sholler, G.F.; Dunwoodie, S.L.; Harvey, R.P.; Winlaw, D.S. Advances in the Genetics of Congenital Heart Disease: A Clinician’s Guide. J. Am. Coll. Cardiol. 2017, 69, 859–870. [Google Scholar] [CrossRef] [PubMed]
- Granados-Riveron, J.T.; Pope, M.; Bu’lock, F.A.; Thornborough, C.; Eason, J.; Setchfield, K.; Ketley, A.; Kirk, E.P.; Fatkin, D.; Feneley, M.P.; et al. Combined Mutation Screening of NKX2-5, GATA4, and TBX5 in Congenital Heart Disease: Multiple Heterozygosity and Novel Mutations: Multiple Heterozygosity in CHD. Congenit. Heart Dis. 2012, 7, 151–159. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, L.C.A.; Al Turki, S.; Manickaraj, A.K.; Manase, D.; Mulder, B.J.M.; Bergin, L.; Rosenberg, H.C.; Mondal, T.; Gordon, E.; Lougheed, J.; et al. Exome Sequencing Identifies Rare Variants in Multiple Genes in Atrioventricular Septal Defect. Genet. Med. 2016, 18, 189–198. [Google Scholar] [CrossRef] [PubMed]
- Tcheandjieu, C.; Zanetti, D.; Yu, M.; Priest, J.R. Inherited Extremes of Aortic Diameter Confer Risk for a Specific Class of Congenital Heart Disease. Circ. Genom. Precis. Med. 2020, 13, e003170. [Google Scholar] [CrossRef]
- Lim, T.B.; Foo, S.Y.R.; Chen, C.K. The Role of Epigenetics in Congenital Heart Disease. Genes 2021, 12, 390. [Google Scholar] [CrossRef] [PubMed]
- Helle, E.I.T.; Biegley, P.; Knowles, J.W.; Leader, J.B.; Pendergrass, S.; Yang, W.; Reaven, G.R.; Shaw, G.M.; Ritchie, M.; Priest, J.R. First Trimester Plasma Glucose Values in Women without Diabetes Are Associated with Risk for Congenital Heart Disease in Offspring. J. Pediatr. 2018, 195, 275–278. [Google Scholar] [CrossRef] [PubMed]
- Øyen, N.; Diaz, L.J.; Leirgul, E.; Boyd, H.A.; Priest, J.; Mathiesen, E.R.; Quertermous, T.; Wohlfahrt, J.; Melbye, M. Prepregnancy Diabetes and Offspring Risk of Congenital Heart Disease: A Nationwide Cohort Study. Circulation 2016, 133, 2243–2253. [Google Scholar] [CrossRef]
- Persson, M.; Razaz, N.; Edstedt Bonamy, A.-K.; Villamor, E.; Cnattingius, S. Maternal Overweight and Obesity and Risk of Congenital Heart Defects. J. Am. Coll. Cardiol. 2019, 73, 44–53. [Google Scholar] [CrossRef] [PubMed]
- Andersen, S.L.; Olsen, J.; Wu, C.S.; Laurberg, P. Birth Defects after Early Pregnancy Use of Antithyroid Drugs: A Danish Nationwide Study. J. Clin. Endocrinol. Metab. 2013, 98, 4373–4381. [Google Scholar] [CrossRef] [PubMed]
- Leatherbury, L.; Berul, C.I. Genetics of Congenital Heart Disease: Is the Glass Now Half-Full? Circ. Cardiovasc. Genet. 2017, 10, e001746. [Google Scholar] [CrossRef]
- Boskovski, M.T.; Homsy, J.; Nathan, M.; Sleeper, L.A.; Morton, S.; Manheimer, K.B.; Tai, A.; Gorham, J.; Lewis, M.; Swartz, M.; et al. De Novo Damaging Variants, Clinical Phenotypes, and Post-Operative Outcomes in Congenital Heart Disease. Circ. Genom. Precis. Med. 2020, 13, e002836. [Google Scholar] [CrossRef]
- Kim, D.S.; Kim, J.H.; Burt, A.A.; Crosslin, D.R.; Burnham, N.; Kim, C.E.; McDonald-McGinn, D.M.; Zackai, E.H.; Nicolson, S.C.; Spray, T.L.; et al. Burden of Potentially Pathologic Copy Number Variants Is Higher in Children with Isolated Congenital Heart Disease and Significantly Impairs Covariate-Adjusted Transplant-Free Survival. J. Thorac. Cardiovasc. Surg. 2016, 151, 1147–1151.e4. [Google Scholar] [CrossRef]
- Garg, V.; Kathiriya, I.S.; Barnes, R.; Schluterman, M.K.; King, I.N.; Butler, C.A.; Rothrock, C.R.; Eapen, R.S.; Hirayama-Yamada, K.; Joo, K.; et al. GATA4 Mutations Cause Human Congenital Heart Defects and Reveal an Interaction with TBX5. Nature 2003, 424, 443–447. [Google Scholar] [CrossRef] [PubMed]
- Garg, V.; Muth, A.N.; Ransom, J.F.; Schluterman, M.K.; Barnes, R.; King, I.N.; Grossfeld, P.D.; Srivastava, D. Mutations in NOTCH1 Cause Aortic Valve Disease. Nature 2005, 437, 270–274. [Google Scholar] [CrossRef]
- Basson, C.T.; Bachinsky, D.R.; Lin, R.C.; Levi, T.; Elkins, J.A.; Soults, J.; Grayzel, D.; Kroumpouzou, E.; Traill, T.A.; Leblanc-Straceski, J.; et al. Mutations in Human TBX5 [corrected] Cause Limb and Cardiac Malformation in Holt-Oram Syndrome. Nat. Genet. 1997, 15, 30–35. [Google Scholar] [CrossRef]
- Kirk, E.P.; Sunde, M.; Costa, M.W.; Rankin, S.A.; Wolstein, O.; Castro, M.L.; Butler, T.L.; Hyun, C.; Guo, G.; Otway, R.; et al. Mutations in Cardiac T-Box Factor Gene TBX20 Are Associated with Diverse Cardiac Pathologies, Including Defects of Septation and Valvulogenesis and Cardiomyopathy. Am. J. Hum. Genet. 2007, 81, 280–291. [Google Scholar] [CrossRef] [PubMed]
- Conrad, D.F.; Pinto, D.; Redon, R.; Feuk, L.; Gokcumen, O.; Zhang, Y.; Aerts, J.; Andrews, T.D.; Barnes, C.; Campbell, P.; et al. Origins and Functional Impact of Copy Number Variation in the Human Genome. Nature 2010, 464, 704–712. [Google Scholar] [CrossRef]
- Silversides, C.K.; Lionel, A.C.; Costain, G.; Merico, D.; Migita, O.; Liu, B.; Yuen, T.; Rickaby, J.; Thiruvahindrapuram, B.; Marshall, C.R.; et al. Rare Copy Number Variations in Adults with Tetralogy of Fallot Implicate Novel Risk Gene Pathways. PLoS Genet. 2012, 8, e1002843. [Google Scholar] [CrossRef]
- Soemedi, R.; Wilson, I.J.; Bentham, J.; Darlay, R.; Töpf, A.; Zelenika, D.; Cosgrove, C.; Setchfield, K.; Thornborough, C.; Granados-Riveron, J.; et al. Contribution of Global Rare Copy-Number Variants to the Risk of Sporadic Congenital Heart Disease. Am. J. Hum. Genet. 2012, 91, 489–501. [Google Scholar] [CrossRef]
- Costain, G.; Lionel, A.C.; Ogura, L.; Marshall, C.R.; Scherer, S.W.; Silversides, C.K.; Bassett, A.S. Genome-Wide Rare Copy Number Variations Contribute to Genetic Risk for Transposition of the Great Arteries. Int. J. Cardiol. 2016, 204, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Thienpont, B.; Mertens, L.; de Ravel, T.; Eyskens, B.; Boshoff, D.; Maas, N.; Fryns, J.-P.; Gewillig, M.; Vermeesch, J.R.; Devriendt, K. Submicroscopic Chromosomal Imbalances Detected by Array-CGH Are a Frequent Cause of Congenital Heart Defects in Selected Patients. Eur. Heart J. 2007, 28, 2778–2784. [Google Scholar] [CrossRef]
- Hitz, M.-P.; Lemieux-Perreault, L.-P.; Marshall, C.; Feroz-Zada, Y.; Davies, R.; Yang, S.W.; Lionel, A.C.; D’Amours, G.; Lemyre, E.; Cullum, R.; et al. Rare Copy Number Variants Contribute to Congenital Left-Sided Heart Disease. PLoS Genet. 2012, 8, e1002903. [Google Scholar] [CrossRef] [PubMed]
- Gamazon, E.R.; Stranger, B.E. The Impact of Human Copy Number Variation on Gene Expression. Brief. Funct. Genom. 2015, 14, 352–357. [Google Scholar] [CrossRef]
- Sifrim, A.; Hitz, M.-P.; Wilsdon, A.; Breckpot, J.; Turki, S.H.A.; Thienpont, B.; McRae, J.; Fitzgerald, T.W.; Singh, T.; Swaminathan, G.J.; et al. Distinct Genetic Architectures for Syndromic and Nonsyndromic Congenital Heart Defects Identified by Exome Sequencing. Nat. Genet. 2016, 48, 1060–1065. [Google Scholar] [CrossRef]
- Homsy, J.; Zaidi, S.; Shen, Y.; Ware, J.S.; Samocha, K.E.; Karczewski, K.J.; DePalma, S.R.; McKean, D.; Wakimoto, H.; Gorham, J.; et al. De Novo Mutations in Congenital Heart Disease with Neurodevelopmental and Other Congenital Anomalies. Science 2015, 350, 1262–1266. [Google Scholar] [CrossRef]
- Øyen, N.; Poulsen, G.; Boyd, H.A.; Wohlfahrt, J.; Jensen, P.K.A.; Melbye, M. Recurrence of Congenital Heart Defects in Families. Circulation 2009, 120, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.C.; Homsy, J.; Zaidi, S.; Lu, Q.; Morton, S.; DePalma, S.R.; Zeng, X.; Qi, H.; Chang, W.; Sierant, M.C.; et al. Contribution of Rare Inherited and de Novo Variants in 2,871 Congenital Heart Disease Probands. Nat. Genet. 2017, 49, 1593–1601. [Google Scholar] [CrossRef]
- Li, A.H.; Hanchard, N.A.; Furthner, D.; Fernbach, S.; Azamian, M.; Nicosia, A.; Rosenfeld, J.; Muzny, D.; D’Alessandro, L.C.A.; Morris, S.; et al. Whole Exome Sequencing in 342 Congenital Cardiac Left Sided Lesion Cases Reveals Extensive Genetic Heterogeneity and Complex Inheritance Patterns. Genome Med. 2017, 9, 95. [Google Scholar] [CrossRef]
- LaHaye, S.; Corsmeier, D.; Basu, M.; Bowman, J.L.; Fitzgerald-Butt, S.; Zender, G.; Bosse, K.; McBride, K.L.; White, P.; Garg, V. Utilization of Whole Exome Sequencing to Identify Causative Mutations in Familial Congenital Heart Disease. Circ. Cardiovasc. Genet. 2016, 9, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Reuter, M.S.; Chaturvedi, R.R.; Liston, E.; Manshaei, R.; Aul, R.B.; Bowdin, S.; Cohn, I.; Curtis, M.; Dhir, P.; Hayeems, R.Z.; et al. The Cardiac Genome Clinic: Implementing Genome Sequencing in Pediatric Heart Disease. Genet. Med. 2020, 22, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Sweeney, N.M.; Nahas, S.A.; Chowdhury, S.; Batalov, S.; Clark, M.; Caylor, S.; Cakici, J.; Nigro, J.J.; Ding, Y.; Veeraraghavan, N.; et al. Rapid Whole Genome Sequencing Impacts Care and Resource Utilization in Infants with Congenital Heart Disease. NPJ Genom. Med. 2021, 6, 1–10. [Google Scholar]
- Xu, J.; Wu, Q.; Wang, L.; Han, J.; Pei, Y.; Zhi, W.; Liu, Y.; Yin, C.; Jiang, Y. Next-Generation Sequencing Identified Genetic Variations in Families with Fetal Non-Syndromic Atrioventricular Septal Defects. Int. J. Clin. Exp. Pathol. 2018, 11, 3732. [Google Scholar]
- Øyen, N.; Poulsen, G.; Wohlfahrt, J.; Boyd, H.A.; Jensen, P.K.A.; Melbye, M. Recurrence of Discordant Congenital Heart Defects in Families. Circ. Cardiovasc. Genet. 2010, 2, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Helle, E.; Córdova-Palomera, A.; Ojala, T.; Saha, P.; Potiny, P.; Gustafsson, S.; Ingelsson, E.; Bamshad, M.; Nickerson, D.; Chong, J.X.; et al. Loss of Function, Missense, and Intronic Variants in NOTCH1 Confer Different Risks for Left Ventricular Outflow Tract Obstructive Heart Defects in Two European Cohorts. Genet. Epidemiol. 2019, 43, 215–226. [Google Scholar] [CrossRef]
- Reuter, M.S.; Jobling, R.; Chaturvedi, R.R.; Manshaei, R.; Costain, G.; Heung, T.; Curtis, M.; Hosseini, S.M.; Liston, E.; Lowther, C.; et al. Haploinsufficiency of Vascular Endothelial Growth Factor Related Signaling Genes Is Associated with Tetralogy of Fallot. Genet. Med. 2019, 21, 1001–1007. [Google Scholar] [CrossRef]
- Richter, F.; Morton, S.U.; Kim, S.W.; Kitaygorodsky, A.; Wasson, L.K.; Chen, K.M.; Zhou, J.; Qi, H.; Patel, N.; DePalma, S.R.; et al. Genomic Analyses Implicate Noncoding de Novo Variants in Congenital Heart Disease. Nat. Genet. 2020, 52, 769–777. [Google Scholar] [CrossRef]
- Lelieveld, S.H.; Spielmann, M.; Mundlos, S.; Veltman, J.A.; Gilissen, C. Comparison of Exome and Genome Sequencing Technologies for the Complete Capture of Protein-Coding Regions. Hum. Mutat. 2015, 36, 815–822. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and Guidelines for the Interpretation of Sequence Variants: A Joint Consensus Recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef] [PubMed]
- Paige, S.L.; Saha, P.; Priest, J.R. Beyond Gene Panels: Whole Exome Sequencing for Diagnosis of Congenital Heart Disease. Circ. Genom. Precis. Med. 2018, 11, e002097. [Google Scholar] [CrossRef] [PubMed]
- Hirschhorn, J.N.; Daly, M.J. Genome-Wide Association Studies for Common Diseases and Complex Traits. Nat. Rev. Genet. 2005, 6, 95–108. [Google Scholar] [CrossRef] [PubMed]
- Tang, N.K.Y.; McBeth, J.; Jordan, K.P.; Blagojevic-Bucknall, M.; Croft, P.; Wilkie, R. Impact of Musculoskeletal Pain on Insomnia Onset: A Prospective Cohort Study. Rheumatology 2015, 54, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Johnston, K.J.A.; Adams, M.J.; Nicholl, B.I.; Ward, J.; Strawbridge, R.J.; Ferguson, A.; McIntosh, A.M.; Bailey, M.E.S.; Smith, D.J. Genome-Wide Association Study of Multisite Chronic Pain in UK Biobank. PLoS Genet. 2019, 15, e1008164. [Google Scholar] [CrossRef]
- Hu, Z.; Shi, Y.; Mo, X.; Xu, J.; Zhao, B.; Lin, Y.; Yang, S.; Xu, Z.; Dai, J.; Pan, S.; et al. A Genome-Wide Association Study Identifies Two Risk Loci for Congenital Heart Malformations in Han Chinese Populations. Nat. Genet. 2013, 45, 818–821. [Google Scholar] [CrossRef] [PubMed]
- Cordell, H.J.; Bentham, J.; Topf, A.; Zelenika, D.; Heath, S.; Mamasoula, C.; Cosgrove, C.; Blue, G.; Granados-Riveron, J.; Setchfield, K.; et al. Genome-Wide Association Study of Multiple Congenital Heart Disease Phenotypes Identifies a Susceptibility Locus for Atrial Septal Defect at Chromosome 4p16. Nat. Genet. 2013, 45, 822–824. [Google Scholar] [CrossRef]
- Cordell, H.J.; Töpf, A.; Mamasoula, C.; Postma, A.V.; Bentham, J.; Zelenika, D.; Heath, S.; Blue, G.; Cosgrove, C.; Granados Riveron, J.; et al. Genome-Wide Association Study Identifies Loci on 12q24 and 13q32 Associated with Tetralogy of Fallot. Hum. Mol. Genet. 2013, 22, 1473–1481. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.; Guo, X.; Zhao, B.; Liu, J.; Da, M.; Wen, Y.; Hu, Y.; Ni, B.; Zhang, K.; Yang, S.; et al. Association Analysis Identifies New Risk Loci for Congenital Heart Disease in Chinese Populations. Nat. Commun. 2015, 6, 8082. [Google Scholar] [CrossRef]
- Agopian, A.J.; Goldmuntz, E.; Hakonarson, H.; Sewda, A.; Taylor, D.; Mitchell, L.E. Pediatric Cardiac Genomics Consortium* Genome-Wide Association Studies and Meta-Analyses for Congenital Heart Defects. Circ. Cardiovasc. Genet. 2017, 10, e001449. [Google Scholar] [CrossRef]
- Lahm, H.; Jia, M.; Dreßen, M.; Wirth, F.; Puluca, N.; Gilsbach, R.; Keavney, B.D.; Cleuziou, J.; Beck, N.; Bondareva, O.; et al. Congenital Heart Disease Risk Loci Identified by Genome-Wide Association Study in European Patients. J. Clin. Investig. 2021, 131. [Google Scholar] [CrossRef]
- Bjornsson, T.; Thorolfsdottir, R.B.; Sveinbjornsson, G.; Sulem, P.; Norddahl, G.L.; Helgadottir, A.; Gretarsdottir, S.; Magnusdottir, A.; Danielsen, R.; Sigurdsson, E.L.; et al. A Rare Missense Mutation in MYH6 Associates with Non-Syndromic Coarctation of the Aorta. Eur Heart J. 2018, 39. [Google Scholar] [CrossRef]
- Panagiotou, O.A.; Ioannidis, J.P.A. Genome-Wide Significance Project What Should the Genome-Wide Significance Threshold Be? Empirical Replication of Borderline Genetic Associations. Int. J. Epidemiol. 2012, 41, 273–286. [Google Scholar] [CrossRef] [PubMed]
- Begum, F.; Ghosh, D.; Tseng, G.C.; Feingold, E. Comprehensive Literature Review and Statistical Considerations for GWAS Meta-Analysis. Nucleic Acids Res. 2012, 40, 3777–3784. [Google Scholar] [CrossRef]
- Willer, C.J.; Li, Y.; Abecasis, G.R. METAL: Fast and Efficient Meta-Analysis of Genomewide Association Scans. Bioinformatics 2010, 26, 2190–2191. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- Magi, R.; Lindgren, C.M.; Morris, A.P. Meta-Analysis of Sex-Specific Genome-Wide Association Studies. Genet. Epidemiol. 2010, 34, 846–853. [Google Scholar] [CrossRef]
- Viechtbauer, W. Conducting Meta-Analyses inRwith themetaforPackage. J. Stat. Softw. 2010, 36, 1–48. [Google Scholar] [CrossRef]
- Liu, J.Z.; Tozzi, F.; Waterworth, D.M.; Pillai, S.G.; Muglia, P.; Middleton, L.; Berrettini, W.; Knouff, C.W.; Yuan, X.; Waeber, G.; et al. Meta-Analysis and Imputation Refines the Association of 15q25 with Smoking Quantity. Nat. Genet. 2010, 42, 436–440. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- Hamazaki, T.; El Rouby, N.; Fredette, N.C.; Santostefano, K.E.; Terada, N. Concise Review: Induced Pluripotent Stem Cell Research in the Era of Precision Medicine. Stem Cells 2017, 35, 545–550. [Google Scholar] [CrossRef]
- Theis, J.L.; Hrstka, S.C.; Evans, J.M.; O’Byrne, M.M.; de Andrade, M.; O’Leary, P.W.; Nelson, T.J.; Olson, T.M. Compound Heterozygous NOTCH1 Mutations Underlie Impaired Cardiogenesis in a Patient with Hypoplastic Left Heart Syndrome. Hum. Genet. 2015, 134, 1003–1011. [Google Scholar] [CrossRef]
- Yang, C.; Xu, Y.; Yu, M.; Lee, D.; Alharti, S.; Hellen, N.; Ahmad Shaik, N.; Banaganapalli, B.; Sheikh Ali Mohamoud, H.; Elango, R.; et al. Induced Pluripotent Stem Cell Modelling of HLHS Underlines the Contribution of Dysfunctional NOTCH Signalling to Impaired Cardiogenesis. Hum. Mol. Genet. 2017, 26, 3031–3045. [Google Scholar] [CrossRef] [PubMed]
- Hrstka, S.C.L.; Li, X.; Nelson, T.J. Wanek Program Genetics Pipeline Group NOTCH1-Dependent Nitric Oxide Signaling Deficiency in Hypoplastic Left Heart Syndrome Revealed Through Patient-Specific Phenotypes Detected in Bioengineered Cardiogenesis. Stem Cells 2017, 35, 1106–1119. [Google Scholar] [CrossRef]
- Miao, Y.; Tian, L.; Martin, M.; Paige, S.L.; Galdos, F.X.; Li, J.; Klein, A.; Zhang, H.; Ma, N.; Wei, Y.; et al. Intrinsic Endocardial Defects Contribute to Hypoplastic Left Heart Syndrome. Cell Stem Cell 2020, 27, 574–589. [Google Scholar] [CrossRef] [PubMed]
- Theodoris, C.V.; Li, M.; White, M.P.; Liu, L.; He, D.; Pollard, K.S.; Bruneau, B.G.; Srivastava, D. Human Disease Modeling Reveals Integrated Transcriptional and Epigenetic Mechanisms of NOTCH1 Haploinsufficiency. Cell 2015, 160, 1072–1086. [Google Scholar] [CrossRef]
- Ang, Y.-S.; Rivas, R.N.; Ribeiro, A.J.S.; Srivas, R.; Rivera, J.; Stone, N.R.; Pratt, K.; Mohamed, T.M.A.; Fu, J.-D.; Spencer, C.I.; et al. Disease Model of GATA4 Mutation Reveals Transcription Factor Cooperativity in Human Cardiogenesis. Cell 2016, 167, 1734–1749.e22. [Google Scholar] [CrossRef]
- Lam, Y.; Keung, W.; Chan, C.; Geng, L.; Wong, N.; Brenière-Letuffe, D.; Li, R.A.; Cheung, Y. Single-Cell Transcriptomics of Engineered Cardiac Tissues from Patient-Specific Induced Pluripotent Stem Cell–Derived Cardiomyocytes Reveals Abnormal Developmental Trajectory and Intrinsic Contractile Defects in Hypoplastic Right Heart Syndrome. J. Am. Heart Assoc. 2020, 9, e016528. [Google Scholar] [CrossRef]
- Ge, X.; Ren, Y.; Bartulos, O.; Lee, M.Y.; Yue, Z.; Kim, K.-Y.; Li, W.; Amos, P.J.; Bozkulak, E.C.; Iyer, A.; et al. Modeling Supravalvular Aortic Stenosis Syndrome with Human Induced Pluripotent Stem Cells. Circulation 2012, 126, 1695–1704. [Google Scholar] [CrossRef] [PubMed]
- Kinnear, C.; Chang, W.Y.; Khattak, S.; Hinek, A.; Thompson, T.; de Carvalho Rodrigues, D.; Kennedy, K.; Mahmut, N.; Pasceri, P.; Stanford, W.L.; et al. Modeling and Rescue of the Vascular Phenotype of Williams-Beuren Syndrome in Patient Induced Pluripotent Stem Cells. Stem Cells Transl. Med. 2013, 2, 2–15. [Google Scholar] [CrossRef]
- Jiang, Y.; Habibollah, S.; Tilgner, K.; Collin, J.; Barta, T.; Al-Aama, J.Y.; Tesarov, L.; Hussain, R.; Trafford, A.W.; Kirkwood, G.; et al. An Induced Pluripotent Stem Cell Model of Hypoplastic Left Heart Syndrome (HLHS) Reveals Multiple Expression and Functional Differences in HLHS-Derived Cardiac Myocytes. Stem Cells Transl. Med. 2014, 3, 416–423. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, J.; Yoshida, M.; Tarui, S.; Hirata, M.; Nagai, Y.; Kasahara, S.; Naruse, K.; Ito, H.; Sano, S.; Oh, H. Directed Differentiation of Patient-Specific Induced Pluripotent Stem Cells Identifies the Transcriptional Repression and Epigenetic Modification of NKX2-5, HAND1, and NOTCH1 in Hypoplastic Left Heart Syndrome. PLoS ONE 2014, 9, e102796. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.-S.; Fleres, B.; Lovett, J.; Anfinson, M.; Samudrala, S.S.K.; Kelly, L.J.; Teigen, L.E.; Cavanaugh, M.; Marquez, M.; Geurts, A.M.; et al. Contractility of Induced Pluripotent Stem Cell-Cardiomyocytes with an MYH6 Head Domain Variant Associated With Hypoplastic Left Heart Syndrome. Front. Cell Dev. Biol. 2020, 8, 440. [Google Scholar] [CrossRef]
- Paige, S.L.; Galdos, F.X.; Lee, S.; Chin, E.T.; Ranjbarvaziri, S.; Feyen, D.A.M.; Darsha, A.K.; Xu, S.; Ryan, J.A.; Beck, A.L.; et al. Patient-Specific Induced Pluripotent Stem Cells Implicate Intrinsic Impaired Contractility in Hypoplastic Left Heart Syndrome. Circulation 2020, 142, 1605–1608. [Google Scholar] [CrossRef]
- Paik, D.T.; Chandy, M.; Wu, J.C. Patient and Disease–Specific Induced Pluripotent Stem Cells for Discovery of Personalized Cardiovascular Drugs and Therapeutics. Pharmacol. Rev. 2020, 72, 320–342. [Google Scholar] [CrossRef]
- Yamanaka, S. Induced Pluripotent Stem Cells: Past, Present, and Future. Cell Stem Cell 2012, 10, 678–684. [Google Scholar] [CrossRef]
- Cyganek, L.; Tiburcy, M.; Sekeres, K.; Gerstenberg, K.; Bohnenberger, H.; Lenz, C.; Henze, S.; Stauske, M.; Salinas, G.; Zimmermann, W.-H.; et al. Deep Phenotyping of Human Induced Pluripotent Stem Cell–derived Atrial and Ventricular Cardiomyocytes. JCI Insight 2018, 3, e99941. [Google Scholar] [CrossRef] [PubMed]
- Ward, T.; Tai, W.; Morton, S.; Impens, F.; Van Damme, P.; Van Haver, D.; Timmerman, E.; Venturini, G.; Zhang, K.; Jang, M.Y.; et al. Mechanisms of Congenital Heart Disease Caused by NAA15 Haploinsufficiency. Circ. Res. 2021, 128, 1156–1169. [Google Scholar] [CrossRef]
- Theis, J.L.; Hu, J.J.; Sundsbak, R.S.; Evans, J.M.; Bamlet, W.R.; Qureshi, M.Y.; O’Leary, P.W.; Olson, T.M. Genetic Association Between Hypoplastic Left Heart Syndrome and Cardiomyopathies. Circ. Genom. Precis. Med. 2021, 14, e003126. [Google Scholar] [CrossRef]
- Tomita-Mitchell, A.; Stamm, K.D.; Mahnke, D.K.; Kim, M.-S.; Hidestrand, P.M.; Liang, H.L.; Goetsch, M.A.; Hidestrand, M.; Simpson, P.; Pelech, A.N.; et al. Impact of MYH6 Variants in Hypoplastic Left Heart Syndrome. Physiol. Genom. 2016, 48, 912–921. [Google Scholar] [CrossRef]
- Doyle, M.J.; Lohr, J.L.; Chapman, C.S.; Koyano-Nakagawa, N.; Garry, M.G.; Garry, D.J. Human Induced Pluripotent Stem Cell-Derived Cardiomyocytes as a Model for Heart Development and Congenital Heart Disease. Stem Cell Rev. Rep. 2015, 11, 710–727. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Helle, E.; Priest, J.R. Maternal Obesity and Diabetes Mellitus as Risk Factors for Congenital Heart Disease in the Offspring. J. Am. Heart Assoc. 2020, 9, e011541. [Google Scholar] [CrossRef] [PubMed]
- Stout, K.K.; Broberg, C.S.; Book, W.M.; Cecchin, F.; Chen, J.M.; Dimopoulos, K.; Everitt, M.D.; Gatzoulis, M.; Harris, L.; Hsu, D.T.; et al. Chronic Heart Failure in Congenital Heart Disease: A Scientific Statement from the American Heart Association. Circulation 2016, 133, 770–801. [Google Scholar] [CrossRef]
- Helle, E.; Pihkala, J.; Turunen, R.; Ruotsalainen, H.; Tuupanen, S.; Koskenvuo, J.; Ojala, T. Rare Variants in Genes Associated with Cardiomyopathy Are Not Common in Hypoplastic Left Heart Syndrome Patients with Myocardial Dysfunction. Front. Pediatr. 2020, 8, 596840. [Google Scholar] [CrossRef] [PubMed]
| Study | Cohorts and CHD Subtype | Nearest Genes | Significant Loci |
|---|---|---|---|
| Cordell et al., 2013 [52] 835 cases, 5159 controls | Tetralogy of Fallot | PTPN11 | 1 |
| Lin et al., 2015 [53] 945 cases, 1246 controls | Ventricular septal defect and/or atrial septal defect | EDNRA, SMARCA2, TBX3, PTPRT | 4 |
| Agopian et al., 2018 [54] 1940 cases and trios, 2976 controls | Conotruncal heart defect, left ventricular obstructive tract defect | OFCC1 | 1 |
| Bjornsson et al., 2018 [56] 120 cases, 355,166 controls | Coarctation of the aorta | MYH6 | 1 |
| Lahm et al., 2021 [55] 4034 cases, 8486 controls | All CHD phenotypes | KCNN2 | 1 |
| Transposition of the great arteries | MACROD2 | 6 | |
| Right heart lesions | SLC27A6 | 1 | |
| Left heart lesions | ARHGEF4 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Broberg, M.; Hästbacka, J.; Helle, E. From Stem Cells to Populations—Using hiPSC, Next-Generation Sequencing, and GWAS to Explore the Genetic and Molecular Mechanisms of Congenital Heart Defects. Genes 2021, 12, 921. https://doi.org/10.3390/genes12060921
Broberg M, Hästbacka J, Helle E. From Stem Cells to Populations—Using hiPSC, Next-Generation Sequencing, and GWAS to Explore the Genetic and Molecular Mechanisms of Congenital Heart Defects. Genes. 2021; 12(6):921. https://doi.org/10.3390/genes12060921
Chicago/Turabian StyleBroberg, Martin, Johanna Hästbacka, and Emmi Helle. 2021. "From Stem Cells to Populations—Using hiPSC, Next-Generation Sequencing, and GWAS to Explore the Genetic and Molecular Mechanisms of Congenital Heart Defects" Genes 12, no. 6: 921. https://doi.org/10.3390/genes12060921
APA StyleBroberg, M., Hästbacka, J., & Helle, E. (2021). From Stem Cells to Populations—Using hiPSC, Next-Generation Sequencing, and GWAS to Explore the Genetic and Molecular Mechanisms of Congenital Heart Defects. Genes, 12(6), 921. https://doi.org/10.3390/genes12060921






