Arabidopsis NPF4.6 and NPF5.1 Control Leaf Stomatal Aperture by Regulating Abscisic Acid Transport
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Growth Conditions
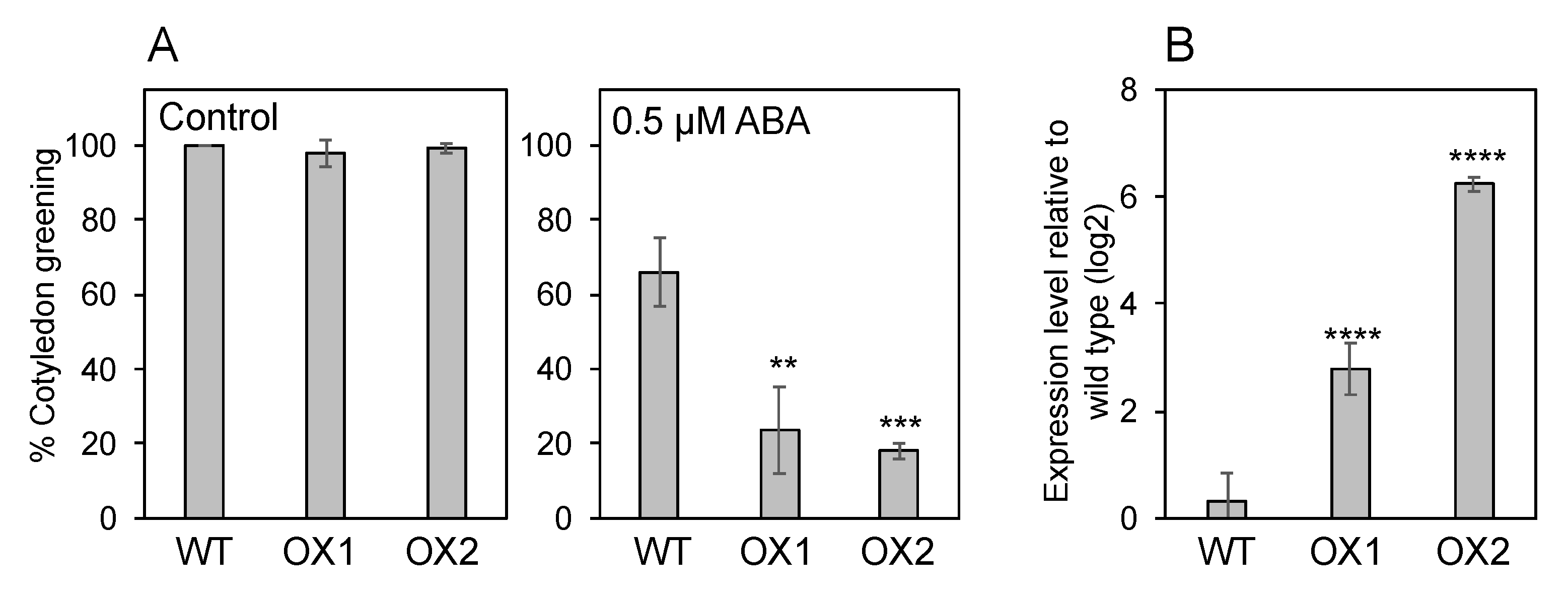
2.2. Germination Assays
2.3. Vector Construction
2.4. Thermal Imaging
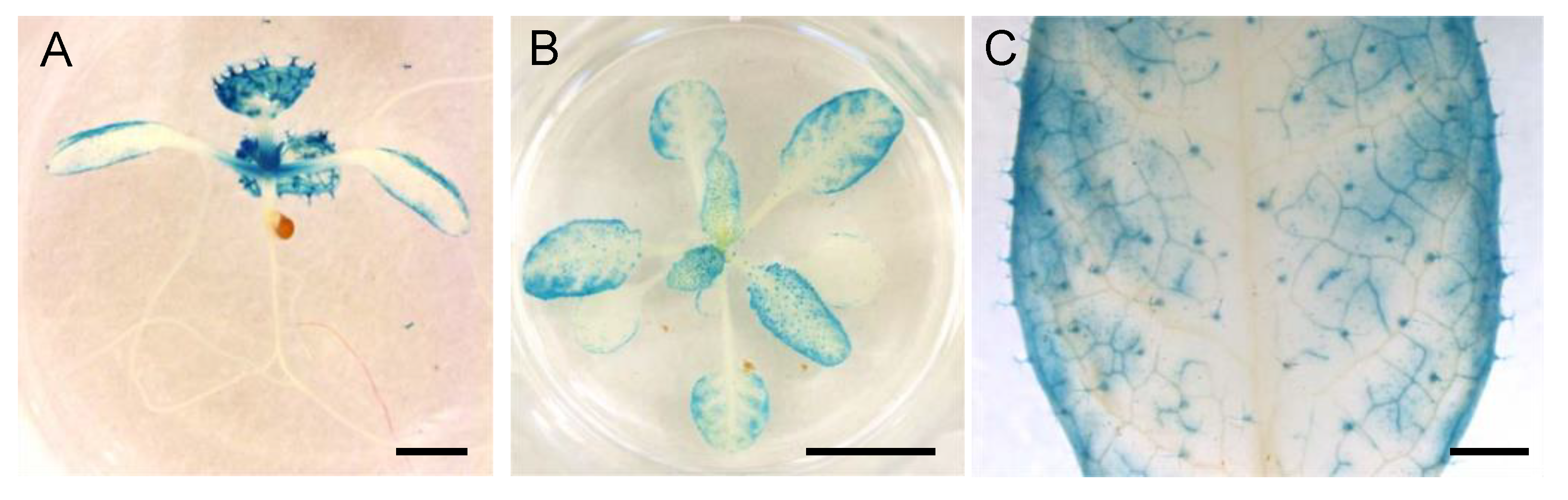
2.5. GUS Staining
2.6. Transport Assays
2.7. ABA Measurements
2.8. Chemicals
2.9. Quantitative Reverse Transcription-PCR
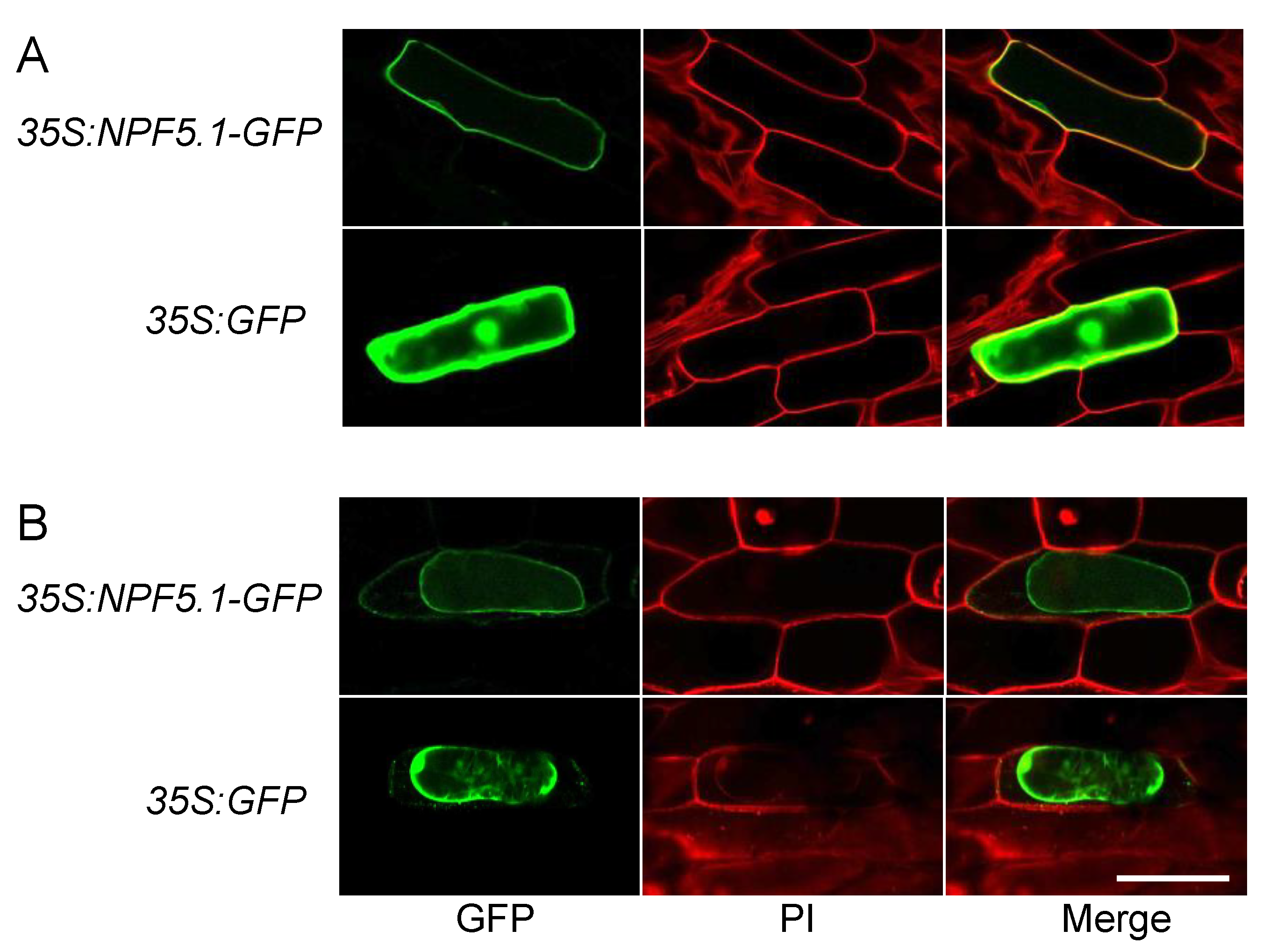
2.10. Transient Expression of GPF Fused NPF5.1 in Onion Epidermal Cells
3. Results
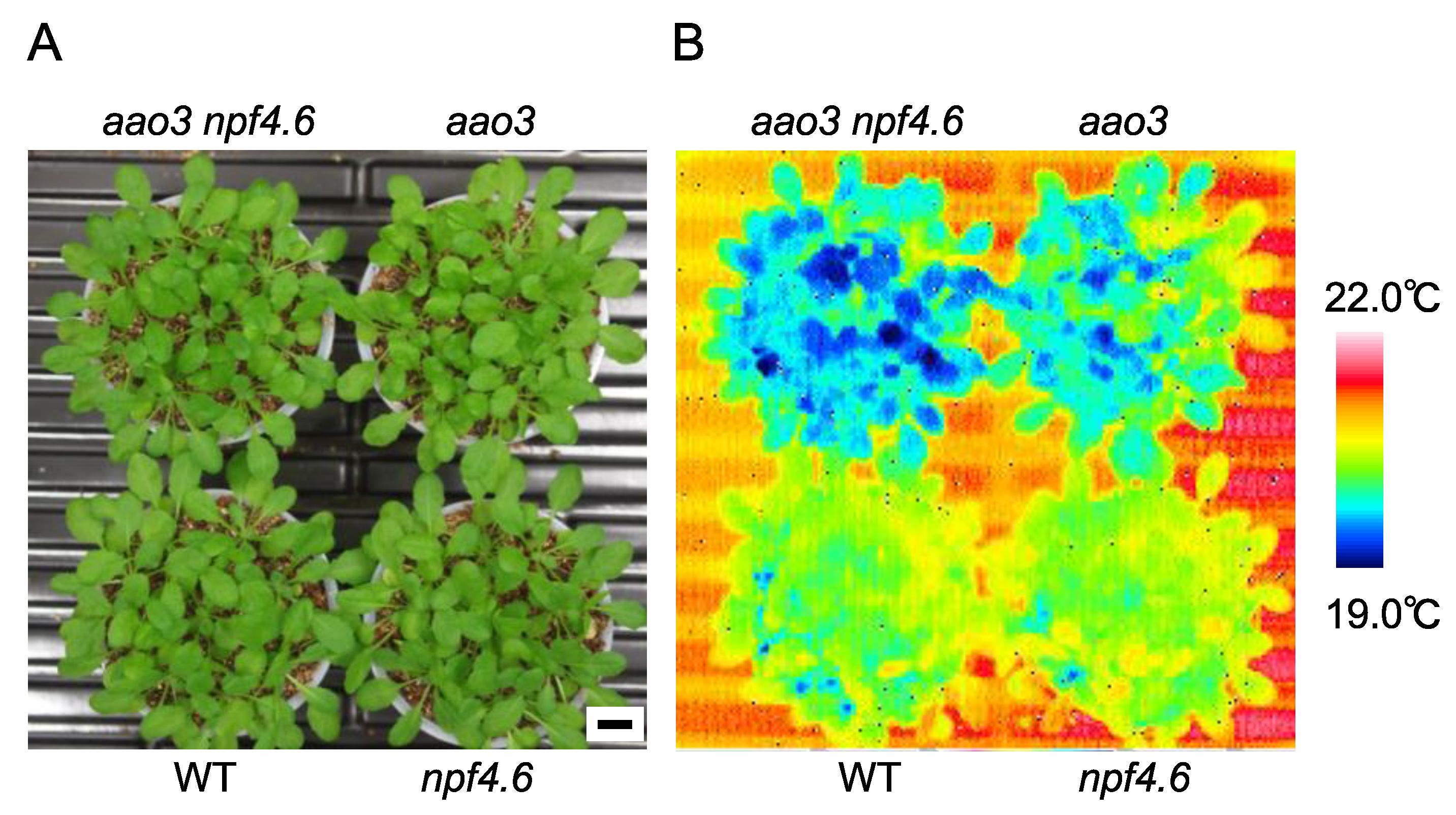
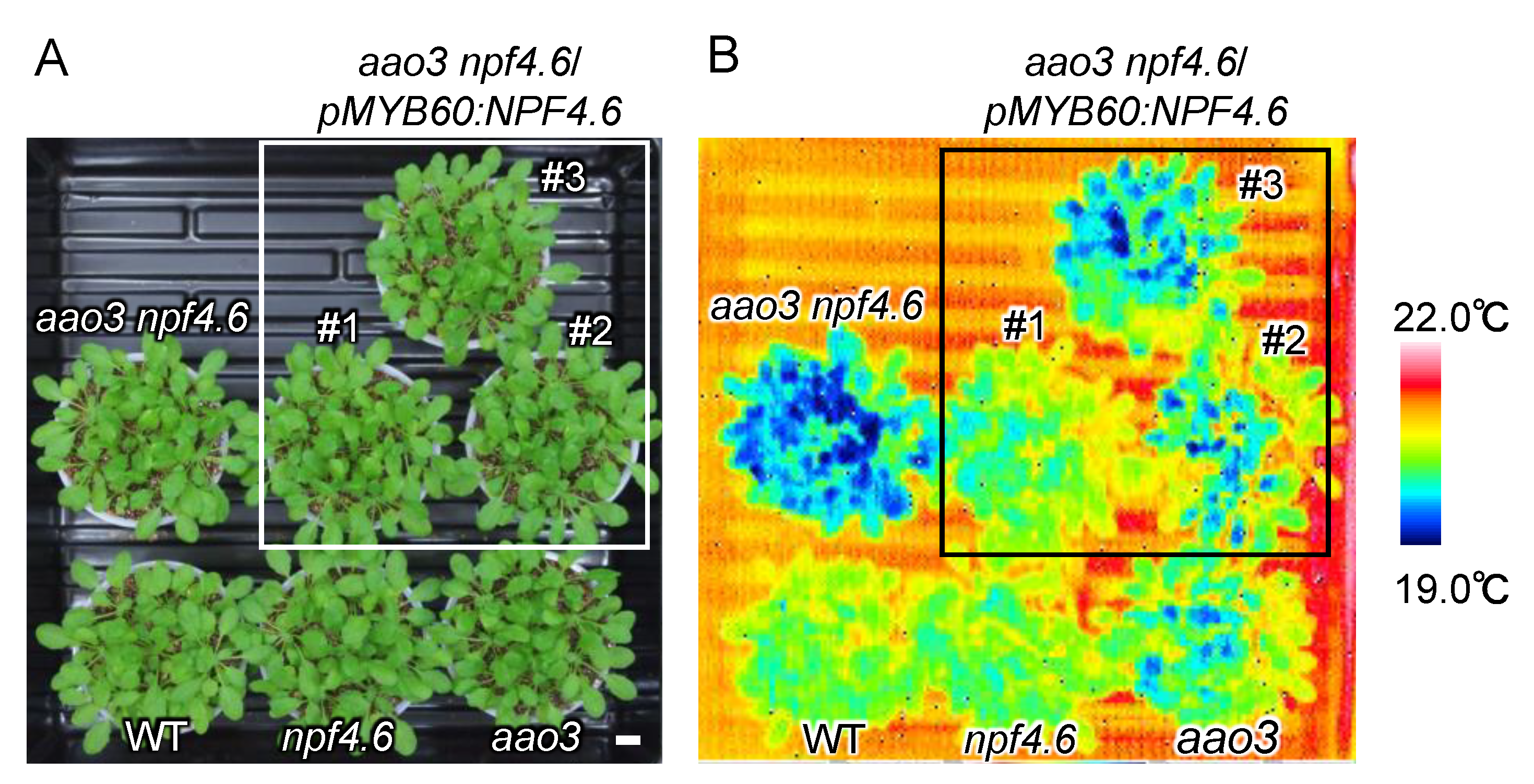
3.1. NPF4.6 Mediates ABA Uptake into Guard Cells
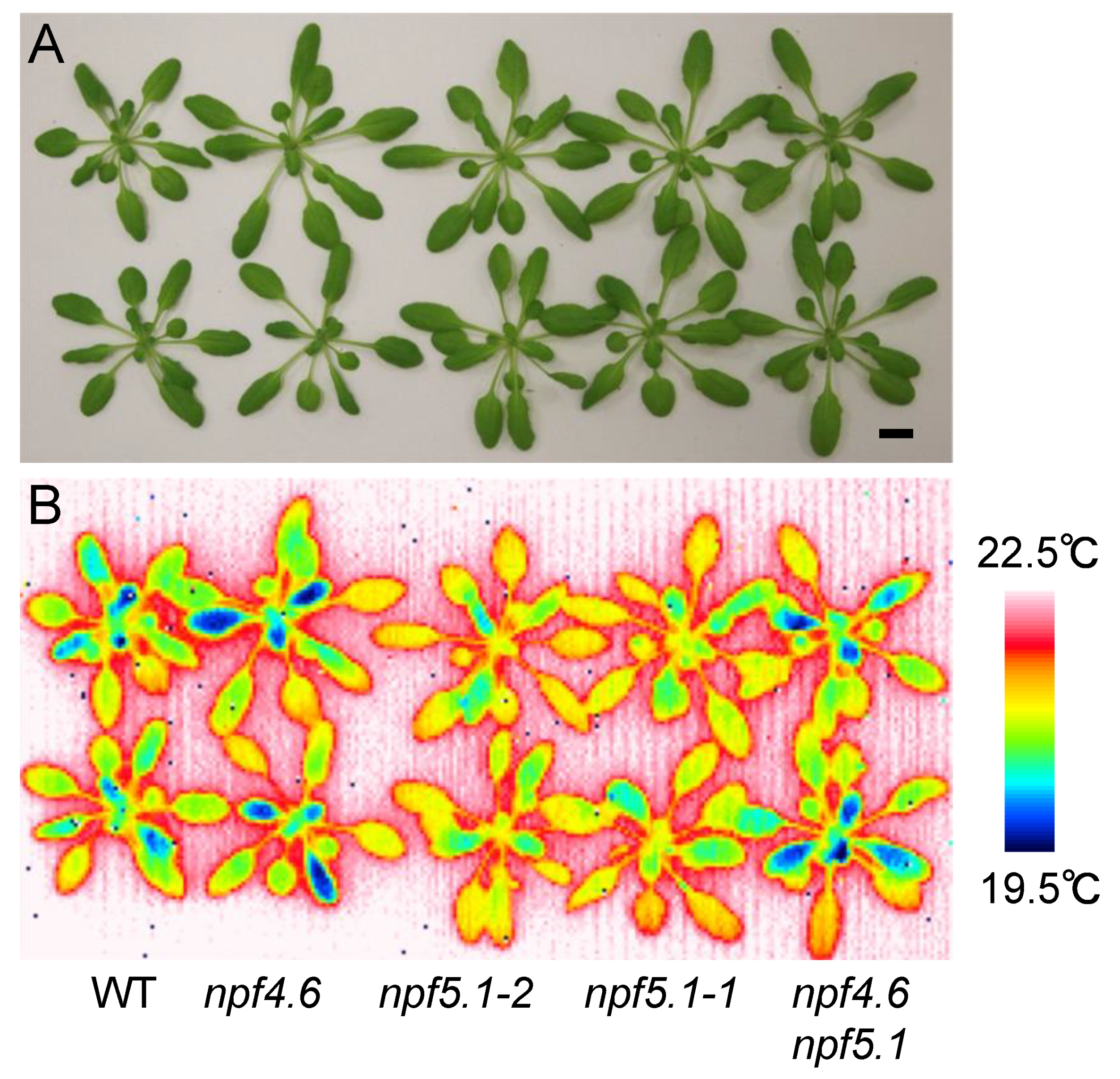
3.2. Identification of Another NPF That Regulates Stomatal Aperture
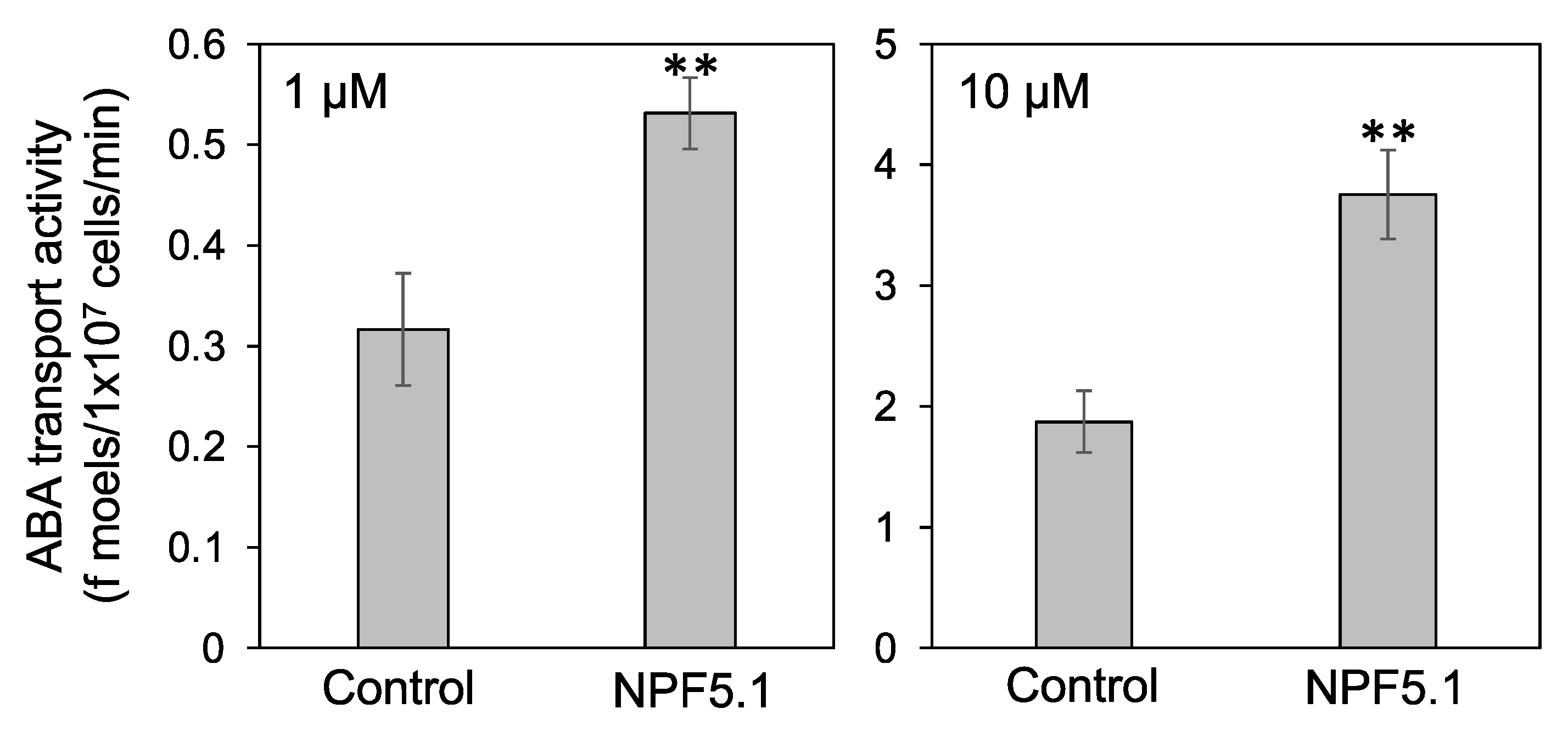
3.3. NPF5.1 Has an ABA Uptake Activity
3.4. Endogenous ABA Levels in NPF5.1
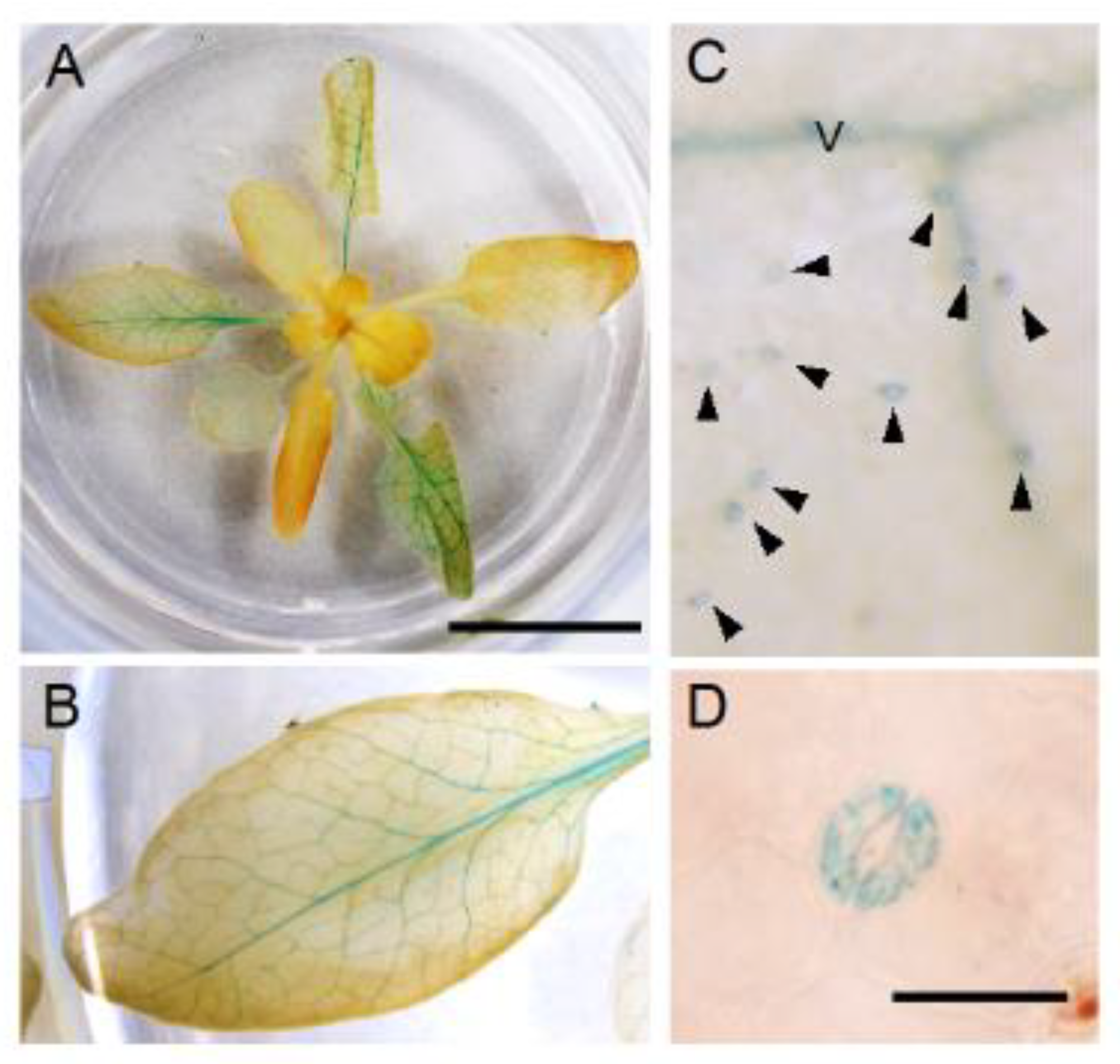
3.5. Spatial Expression Patterns of NPF5.1
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zeevaart, J.A.D.; Creelman, R.A. Metabolism and physiology of abscisic acid. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1988, 39, 439–473. [Google Scholar] [CrossRef]
- Nambara, E.; Marion-Poll, A. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 2005, 56, 165–185. [Google Scholar] [CrossRef]
- Finkelstein, R. Abscisic acid synthesis and response. Arabidopsis Book 2013, 11, e0166. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Zeevaart, J.A. The 9-cis-epoxycarotenoid cleavage reaction is the key regulatory step of abscisic acid biosynthesis in water-stressed bean. Proc. Natl. Acad. Sci. USA 1999, 96, 15354–15361. [Google Scholar] [CrossRef] [PubMed]
- Endo, A.; Sawada, Y.; Takahashi, H.; Okamoto, M.; Ikegami, K.; Koiwai, H.; Seo, M.; Toyomasu, T.; Mitsuhashi, W.; Shinozaki, K.; et al. Drought induction of Arabidopsis 9-cis-epoxycarotenoid dioxygenase occurs in vascular parenchyma cells. Plant Physiol. 2008, 147, 1984–1993. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Fujita, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. ABA-mediated transcriptional regulation in response to osmotic stress in plants. J. Plant Res. 2011, 124, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Karssen, C.M.; Brinkhorstvanderswan, D.L.C.; Breekland, A.E.; Koornneef, M. Induction of dormancy during seed development by endogenous abscisic acid: Studies on abscisic acid deficient genotypes of Arabidopsis thaliana (L) Heynh. Planta 1983, 157, 158–165. [Google Scholar] [CrossRef]
- Kanno, Y.; Jikumaru, Y.; Hanada, A.; Nambara, E.; Abrams, S.R.; Kamiya, Y.; Seo, M. Comprehensive hormone profiling in developing Arabidopsis seeds: Examination of the site of ABA biosynthesis, ABA transport and hormone interactions. Plant Cell Physiol. 2010, 51, 1988–2001. [Google Scholar] [CrossRef]
- Nambara, E.; Okamoto, M.; Tatematsu, K.; Yano, R.; Seo, M.; Kamiya, Y. Abscisic acid and the control of seed dormancy and germination. Seed Sci. Res. 2010, 20, 55–67. [Google Scholar] [CrossRef]
- Seo, M.; Marion-Poll, A. Abscisic acid metabolism and transport. Adv. Bot. Res. 2019, 92, 1–49. [Google Scholar]
- Iuchi, S.; Kobayashi, M.; Taji, T.; Naramoto, M.; Seki, M.; Kato, T.; Tabata, S.; Kakubari, Y.; Yamaguchi-Shinozaki, K.; Shinozaki, K. Regulation of drought tolerance by gene manipulation of 9-cis-epoxycarotenoid dioxygenase, a key enzyme in abscisic acid biosynthesis in Arabidopsis. Plant J. 2001, 27, 325–333. [Google Scholar] [CrossRef]
- Frey, A.; Effroy, D.; Lefebvre, V.; Seo, M.; Perreau, F.; Berger, A.; Sechet, J.; To, A.; North, H.M.; Marion-Poll, A. Epoxycarotenoid cleavage by NCED5 fine-tunes ABA accumulation and affects seed dormancy and drought tolerance with other NCED family members. Plant J. 2012, 70, 501–512. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, V.; North, H.; Frey, A.; Sotta, B.; Seo, M.; Okamoto, M.; Nambara, E.; Marion-Poll, A. Functional analysis of Arabidopsis NCED6 and NCED9 genes indicates that ABA synthesized in the endosperm is involved in the induction of seed dormancy. Plant J. 2006, 45, 309–319. [Google Scholar] [CrossRef]
- Tan, B.C.; Joseph, L.M.; Deng, W.T.; Liu, L.; Li, Q.B.; Cline, K.; McCarty, D.R. Molecular characterization of the Arabidopsis 9-cis epoxycarotenoid dioxygenase gene family. Plant J. 2003, 35, 44–56. [Google Scholar] [CrossRef] [PubMed]
- Seo, M.; Hanada, A.; Kuwahara, A.; Endo, A.; Okamoto, M.; Yamauchi, Y.; North, H.; Marion-Poll, A.; Sun, T.P.; Koshiba, T.; et al. Regulation of hormone metabolism in Arabidopsis seeds: Phytochrome regulation of abscisic acid metabolism and abscisic acid regulation of gibberellin metabolism. Plant J. 2006, 48, 354–366. [Google Scholar] [CrossRef]
- Toh, S.; Imamura, A.; Watanabe, A.; Nakabayashi, K.; Okamoto, M.; Jikumaru, Y.; Hanada, A.; Aso, Y.; Ishiyama, K.; Tamura, N.; et al. High temperature-induced abscisic acid biosynthesis and its role in the inhibition of gibberellin action in Arabidopsis seeds. Plant Physiol. 2008, 146, 1368–1385. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, F.; Suzuki, T.; Osakabe, Y.; Betsuyaku, S.; Kondo, Y.; Dohmae, N.; Fukuda, H.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A small peptide modulates stomatal control via abscisic acid in long-distance signalling. Nature 2018, 556, 235–238. [Google Scholar] [CrossRef]
- Sato, H.; Takasaki, H.; Takahashi, F.; Suzuki, T.; Iuchi, S.; Mitsuda, N.; Ohme-Takagi, M.; Ikeda, M.; Seo, M.; Yamaguchi-Shinozaki, K.; et al. Arabidopsis thaliana NGATHA1 transcription factor induces ABA biosynthesis by activating NCED3 gene during dehydration stress. Proc. Natl. Acad. Sci. USA 2018, 115, E11178–E11187. [Google Scholar] [CrossRef]
- Iuchi, S.; Kobayashi, M.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A stress-inducible gene for 9-cis-epoxycarotenoid dioxygenase involved in abscisic acid biosynthesis under water stress in drought-tolerant cowpea. Plant Physiol. 2000, 123, 553–562. [Google Scholar] [CrossRef]
- Chernys, J.T.; Zeevaart, J.A. Characterization of the 9-cis-epoxycarotenoid dioxygenase gene family and the regulation of abscisic acid biosynthesis in avocado. Plant Physiol. 2000, 124, 343–353. [Google Scholar] [CrossRef]
- Thompson, A.J.; Jackson, A.C.; Parker, R.A.; Morpeth, D.R.; Burbidge, A.; Taylor, I.B. Abscisic acid biosynthesis in tomato: Regulation of zeaxanthin epoxidase and 9-cis-epoxycarotenoid dioxygenase mRNAs by light/dark cycles, water stress and abscisic acid. Plant Mol. Biol. 2000, 42, 833–845. [Google Scholar] [CrossRef] [PubMed]
- Sawada, Y.; Aoki, M.; Nakaminami, K.; Mitsuhashi, W.; Tatematsu, K.; Kushiro, T.; Koshiba, T.; Kamiya, Y.; Inoue, Y.; Nambara, E.; et al. Phytochrome- and gibberellin-mediated regulation of abscisic acid metabolism during germination of photoblastic lettuce seeds. Plant Physiol. 2008, 146, 1386–1396. [Google Scholar] [CrossRef]
- Endo, A.; Okamoto, M.; Koshiba, T. ABA biosynthesis and catabolism pathways. In Abscisic Acid: Metabolism, Transport and Signaling; Zhang, D.P., Ed.; Springer: Berlin/Heidelberg, Germany, 2014; pp. 21–46. [Google Scholar]
- Kushiro, T.; Okamoto, M.; Nakabayashi, K.; Yamagishi, K.; Kitamura, S.; Asami, T.; Hirai, N.; Koshiba, T.; Kamiya, Y.; Nambara, E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8′-hydroxylases: Key enzymes in ABA catabolism. EMBO J. 2004, 23, 1647–1656. [Google Scholar] [CrossRef] [PubMed]
- Saito, S.; Hirai, N.; Matsumoto, C.; Ohigashi, H.; Ohta, D.; Sakata, K.; Mizutani, M. Arabidopsis CYP707As encode (+)-abscisic acid 8′-hydroxylase, a key enzyme in the oxidative catabolism of abscisic acid. Plant Physiol. 2004, 134, 1439–1449. [Google Scholar] [CrossRef] [PubMed]
- Umezawa, T.; Okamoto, M.; Kushiro, T.; Nambara, E.; Oono, Y.; Seki, M.; Kobayashi, M.; Koshiba, T.; Kamiya, Y.; Shinozaki, K. CYP707A3, a major ABA 8′-hydroxylase involved in dehydration and rehydration response in Arabidopsis thaliana. Plant J. 2006, 46, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Okamoto, M.; Kuwahara, A.; Seo, M.; Kushiro, T.; Asami, T.; Hirai, N.; Kamiya, Y.; Koshiba, T.; Nambara, E. CYP707A1 and CYP707A2, which encode abscisic acid 8′-hydroxylases, are indispensable for proper control of seed dormancy and germination in Arabidopsis. Plant Physiol. 2006, 141, 97–107. [Google Scholar] [CrossRef]
- Yang, S.H.; Zeevaart, J.A. Expression of ABA 8′-hydroxylases in relation to leaf water relations and seed development in bean. Plant J. 2006, 47, 675–686. [Google Scholar] [CrossRef]
- Chono, M.; Honda, I.; Shinoda, S.; Kushiro, T.; Kamiya, Y.; Nambara, E.; Kawakami, N.; Kaneko, S.; Watanabe, Y. Field studies on the regulation of abscisic acid content and germinability during grain development of barley: Molecular and chemical analysis of pre-harvest sprouting. J. Exp. Bot. 2006, 57, 2421–2434. [Google Scholar] [CrossRef]
- Saika, H.; Okamoto, M.; Miyoshi, K.; Kushiro, T.; Shinoda, S.; Jikumaru, Y.; Fujimoto, M.; Arikawa, T.; Takahashi, H.; Ando, M.; et al. Ethylene promotes submergence-induced expression of OsABA8ox1, a gene that encodes ABA 8′-hydroxylase in rice. Plant Cell Physiol. 2007, 48, 287–298. [Google Scholar] [CrossRef]
- Kuromori, T.; Seo, M.; Shinozaki, K. ABA transport and plant water stress responses. Trends Plant Sci. 2018, 23, 513–522. [Google Scholar] [CrossRef]
- Koiwai, H.; Nakaminami, K.; Seo, M.; Mitsuhashi, W.; Toyomasu, T.; Koshiba, T. Tissue-specific localization of an abscisic acid biosynthetic enzyme, AAO3, in Arabidopsis. Plant Physiol. 2004, 134, 1697–1707. [Google Scholar] [CrossRef] [PubMed]
- Kuromori, T.; Sugimoto, E.; Shinozaki, K. Intertissue signal transfer of abscisic acid from vascular cells to guard cells. Plant Physiol. 2014, 164, 1587–1592. [Google Scholar] [CrossRef]
- Kuromori, T.; Miyaji, T.; Yabuuchi, H.; Shimizu, H.; Sugimoto, E.; Kamiya, A.; Moriyama, Y.; Shinozaki, K. ABC transporter AtABCG25 is involved in abscisic acid transport and responses. Proc. Natl. Acad. Sci. USA 2010, 107, 2361–2366. [Google Scholar] [CrossRef]
- Kanno, Y.; Hanada, A.; Chiba, Y.; Ichikawa, T.; Nakazawa, M.; Matsui, M.; Koshiba, T.; Kamiya, Y.; Seo, M. Identification of an abscisic acid transporter by functional screening using the receptor complex as a sensor. Proc. Natl. Acad. Sci. USA 2012, 109, 9653–9658. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Hwang, J.U.; Lee, M.; Kim, Y.Y.; Assmann, S.M.; Martinoia, E.; Lee, Y. PDR-type ABC transporter mediates cellular uptake of the phytohormone abscisic acid. Proc. Natl. Acad. Sci. USA 2010, 107, 2355–2360. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, H.; Pan, Y.; Yu, Y.; Luan, S.; Li, L. A DTX/MATE-type transporter facilitates abscisic acid efflux and modulates ABA sensitivity and drought tolerance in Arabidopsis. Mol. Plant 2014, 7, 1522–1532. [Google Scholar] [CrossRef]
- Kang, J.; Yim, S.; Choi, H.; Kim, A.; Lee, K.P.; Lopez-Molina, L.; Martinoia, E.; Lee, Y. Abscisic acid transporters cooperate to control seed germination. Nat. Commun. 2015, 6, 8113. [Google Scholar] [CrossRef] [PubMed]
- Yao, L.; Cheng, X.; Gu, Z.; Huang, W.; Li, S.; Wang, L.; Wang, Y.F.; Xu, P.; Ma, H.; Ge, X. The AWPM-19 family protein OsPM1 mediates abscisic acid influx and drought response in rice. Plant Cell 2018, 30, 1258–1276. [Google Scholar] [CrossRef] [PubMed]
- Pawela, A.; Banasiak, J.; Biala, W.; Martinoia, E.; Jasinski, M. MtABCG20 is an ABA exporter influencing root morphology and seed germination of Medicago truncatula. Plant J. 2019, 98, 511–523. [Google Scholar] [CrossRef] [PubMed]
- Krattinger, S.G.; Kang, J.; Braunlich, S.; Boni, R.; Chauhan, H.; Selter, L.L.; Robinson, M.D.; Schmid, M.W.; Wiederhold, E.; Hensel, G.; et al. Abscisic acid is a substrate of the ABC transporter encoded by the durable wheat disease resistance gene Lr34. New Phytol. 2019, 223, 853–866. [Google Scholar] [CrossRef] [PubMed]
- Fu, X.; Liu, H.; Hassani, D.; Peng, B.; Yan, X.; Wang, Y.; Wang, C.; Li, L.; Liu, P.; Pan, Q.; et al. AaABCG40 enhances artemisinin content and modulates rrought tolerance in Artemisia annua. Front. Plant Sci. 2020, 11, 950. [Google Scholar] [CrossRef]
- Kuromori, T.; Sugimoto, E.; Shinozaki, K. Brachypodium BdABCG25 is a homolog of Arabidopsis AtABCG25 involved in the transport of abscisic acid. FEBS Lett. 2021, 595, 954–959. [Google Scholar] [CrossRef] [PubMed]
- Shohat, H.; Illouz-Eliaz, N.; Kanno, Y.; Seo, M.; Weiss, D. The tomato DELLA protein PROCERA promotes abscisic acid responses in guard cells by upregulating an abscisic acid transporter. Plant Physiol. 2020, 184, 518–528. [Google Scholar] [CrossRef]
- Seo, M.; Aoki, H.; Koiwai, H.; Kamiya, Y.; Nambara, E.; Koshiba, T. Comparative studies on the Arabidopsis aldehyde oxidase (AAO) gene family revealed a major role of AAO3 in ABA biosynthesis in seeds. Plant Cell Physiol. 2004, 45, 1694–1703. [Google Scholar] [CrossRef] [PubMed]
- Chiba, Y.; Shimizu, T.; Miyakawa, S.; Kanno, Y.; Koshiba, T.; Kamiya, Y.; Seo, M. Identification of Arabidopsis thaliana NRT1/PTR FAMILY (NPF) proteins capable of transporting plant hormones. J. Plant Res. 2015, 128, 679–686. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, T.; Kurose, T.; Hino, T.; Tanaka, K.; Kawamukai, M.; Niwa, Y.; Toyooka, K.; Matsuoka, K.; Jinbo, T.; Kimura, T. Development of series of gateway binary vectors, pGWBs, for realizing efficient construction of fusion genes for plant transformation. J. Biosci. Bioeng. 2007, 104, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Rusconi, F.; Simeoni, F.; Francia, P.; Cominelli, E.; Conti, L.; Riboni, M.; Simoni, L.; Martin, C.R.; Tonelli, C.; Galbiati, M. The Arabidopsis thaliana MYB60 promoter provides a tool for the spatio-temporal control of gene expression in stomatal guard cells. J. Exp. Bot. 2013, 64, 3361–3371. [Google Scholar] [CrossRef]
- Becker, D. Binary vectors which allow the exchange of plant selectable markers and reporter genes. Nucleic Acids Res. 1990, 18, 203. [Google Scholar] [CrossRef]
- Nakagawa, T.; Suzuki, T.; Murata, S.; Nakamura, S.; Hino, T.; Maeo, K.; Tabata, R.; Kawai, T.; Tanaka, K.; Niwa, Y.; et al. Improved Gateway binary vectors: High-performance vectors for creation of fusion constructs in transgenic analysis of plants. Biosci. Biotechnol. Biochem. 2007, 71, 2095–2100. [Google Scholar] [CrossRef]
- Kanno, Y.; Oikawa, T.; Chiba, Y.; Ishimaru, Y.; Shimizu, T.; Sano, N.; Koshiba, T.; Kamiya, Y.; Ueda, M.; Seo, M. AtSWEET13 and AtSWEET14 regulate gibberellin-mediated physiological processes. Nat. Commun. 2016, 7, 13245. [Google Scholar] [CrossRef]
- Watanabe, S.; Takahashi, N.; Kanno, Y.; Suzuki, H.; Aoi, Y.; Takeda-Kamiya, N.; Toyooka, K.; Kasahara, H.; Hayashi, K.I.; Umeda, M.; et al. The Arabidopsis NRT1/PTR FAMILY protein NPF7.3/NRT1.5 is an indole-3-butyric acid transporter involved in root gravitropism. Proc. Natl. Acad. Sci. USA 2020, 117, 31500–31509. [Google Scholar] [CrossRef] [PubMed]
- Seo, M.; Koiwai, H.; Akaba, S.; Komano, T.; Oritani, T.; Kamiya, Y.; Koshiba, T. Abscisic aldehyde oxidase in leaves of Arabidopsis thaliana. Plant J. 2000, 23, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Seo, M.; Peeters, A.J.; Koiwai, H.; Oritani, T.; Marion-Poll, A.; Zeevaart, J.A.; Koornneef, M.; Kamiya, Y.; Koshiba, T. The Arabidopsis aldehyde oxidase 3 (AAO3) gene product catalyzes the final step in abscisic acid biosynthesis in leaves. Proc. Natl. Acad. Sci. USA 2000, 97, 12908–12913. [Google Scholar] [CrossRef]
- Bauer, H.; Ache, P.; Lautner, S.; Fromm, J.; Hartung, W.; Al-Rasheid, K.A.; Sonnewald, S.; Sonnewald, U.; Kneitz, S.; Lachmann, N.; et al. The stomatal response to reduced relative humidity requires guard cell-autonomous ABA synthesis. Curr. Biol. 2013, 23, 53–57. [Google Scholar] [CrossRef] [PubMed]
- Leran, S.; Varala, K.; Boyer, J.C.; Chiurazzi, M.; Crawford, N.; Daniel-Vedele, F.; David, L.; Dickstein, R.; Fernandez, E.; Forde, B.; et al. A unified nomenclature of NITRATE TRANSPORTER 1/PEPTIDE TRANSPORTER family members in plants. Trends Plant Sci. 2014, 19, 5–9. [Google Scholar] [CrossRef] [PubMed]
- Krouk, G.; Lacombe, B.; Bielach, A.; Perrine-Walker, F.; Malinska, K.; Mounier, E.; Hoyerova, K.; Tillard, P.; Leon, S.; Ljung, K.; et al. Nitrate-regulated auxin transport by NRT1.1 defines a mechanism for nutrient sensing in plants. Dev. Cell 2010, 18, 927–937. [Google Scholar] [CrossRef]
- Saito, H.; Oikawa, T.; Hamamoto, S.; Ishimaru, Y.; Kanamori-Sato, M.; Sasaki-Sekimoto, Y.; Utsumi, T.; Chen, J.; Kanno, Y.; Masuda, S.; et al. The jasmonate-responsive GTR1 transporter is required for gibberellin-mediated stamen development in Arabidopsis. Nat. Commun. 2015, 6, 6095. [Google Scholar] [CrossRef]
- David, L.C.; Berquin, P.; Kanno, Y.; Seo, M.; Daniel-Vedele, F.; Ferrario-Mery, S. N availability modulates the role of NPF3.1, a gibberellin transporter, in GA-mediated phenotypes in Arabidopsis. Planta 2016, 244, 1315–1328. [Google Scholar] [CrossRef]
- Tal, I.; Zhang, Y.; Jorgensen, M.E.; Pisanty, O.; Barbosa, I.C.; Zourelidou, M.; Regnault, T.; Crocoll, C.; Erik Olsen, C.; Weinstain, R.; et al. The Arabidopsis NPF3 protein is a GA transporter. Nat. Commun. 2016, 7, 1486. [Google Scholar] [CrossRef]
- Ishimaru, Y.; Oikawa, T.; Suzuki, T.; Takeishi, S.; Matsuura, H.; Takahashi, K.; Hamamoto, S.; Uozumi, N.; Shimizu, T.; Seo, M.; et al. GTR1 is a jasmonic acid and jasmonoyl-l-isoleucine transporter in Arabidopsis thaliana. Biosci. Biotechnol. Biochem. 2017, 81, 249–255. [Google Scholar] [CrossRef]
- Nour-Eldin, H.H.; Andersen, T.G.; Burow, M.; Madsen, S.R.; Jorgensen, M.E.; Olsen, C.E.; Dreyer, I.; Hedrich, R.; Geiger, D.; Halkier, B.A. NRT/PTR transporters are essential for translocation of glucosinolate defence compounds to seeds. Nature 2012, 488, 531–534. [Google Scholar] [CrossRef]
- Larsen, B.; Fuller, V.L.; Pollier, J.; Van Moerkercke, A.; Schweizer, F.; Payne, R.; Colinas, M.; O’Connor, S.E.; Goossens, A.; Halkier, B.A. Identification of iridoid glucoside transporters in catharanthus roseus. Plant Cell Physiol. 2017, 58, 1507–1518. [Google Scholar] [CrossRef] [PubMed]
- Payne, R.M.; Xu, D.; Foureau, E.; Teto Carqueijeiro, M.I.; Oudin, A.; Bernonville, T.D.; Novak, V.; Burow, M.; Olsen, C.E.; Jones, D.M.; et al. An NPF transporter exports a central monoterpene indole alkaloid intermediate from the vacuole. Nat. Plants 2017, 3, 16208. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Qiu, J.; Jayakannan, M.; Xu, B.; Li, Y.; Mayo, G.M.; Tester, M.; Gilliham, M.; Roy, S.J. AtNPF2.5 modulates chloride (Cl-) efflux from roots of Arabidopsis thaliana. Front. Plant Sci. 2016, 7, 2013. [Google Scholar] [CrossRef]
- Li, B.; Byrt, C.; Qiu, J.; Baumann, U.; Hrmova, M.; Evrard, A.; Johnson, A.A.; Birnbaum, K.D.; Mayo, G.M.; Jha, D.; et al. Identification of a stelar-localized transport protein that facilitates root-to-shoot transfer of chloride in Arabidopsis. Plant Physiol. 2016, 170, 1014–1029. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Yu, M.; Du, X.Q.; Wang, Z.F.; Wu, W.H.; Quintero, F.J.; Jin, X.H.; Li, H.D.; Wang, Y. NRT1.5/NPF7.3 functions as a proton-coupled H+/K+ antiporter for K+ loading into the xylem in Arabidopsis. Plant Cell 2017, 29, 2016–2026. [Google Scholar] [CrossRef]
- Corratge-Faillie, C.; Lacombe, B. Substrate (un)specificity of Arabidopsis NRT1/PTR FAMILY (NPF) proteins. J. Exp. Bot. 2017, 68, 3107–3113. [Google Scholar] [CrossRef] [PubMed]
- Huang, N.C.; Liu, K.H.; Lo, H.J.; Tsay, Y.F. Cloning and functional characterization of an Arabidopsis nitrate transporter gene that encodes a constitutive component of low-affinity uptake. Plant Cell 1999, 11, 1381–1392. [Google Scholar] [CrossRef]
- Lin, S.-H.; Kuo, H.-F.; Canivenc, G.; Lin, C.-S.; Lepetit, M.; Hsu, P.-K.; Tillard, P.; Lin, H.-L.; Wang, Y.-Y.; Tsai, C.-B.; et al. Mutation of the Arabidopsis NRT1.5 nitrate transporter causes defective root-to-shoot nitrate transport. Plant Cell 2008, 20, 2514–2528. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shimizu, T.; Kanno, Y.; Suzuki, H.; Watanabe, S.; Seo, M. Arabidopsis NPF4.6 and NPF5.1 Control Leaf Stomatal Aperture by Regulating Abscisic Acid Transport. Genes 2021, 12, 885. https://doi.org/10.3390/genes12060885
Shimizu T, Kanno Y, Suzuki H, Watanabe S, Seo M. Arabidopsis NPF4.6 and NPF5.1 Control Leaf Stomatal Aperture by Regulating Abscisic Acid Transport. Genes. 2021; 12(6):885. https://doi.org/10.3390/genes12060885
Chicago/Turabian StyleShimizu, Takafumi, Yuri Kanno, Hiromi Suzuki, Shunsuke Watanabe, and Mitsunori Seo. 2021. "Arabidopsis NPF4.6 and NPF5.1 Control Leaf Stomatal Aperture by Regulating Abscisic Acid Transport" Genes 12, no. 6: 885. https://doi.org/10.3390/genes12060885
APA StyleShimizu, T., Kanno, Y., Suzuki, H., Watanabe, S., & Seo, M. (2021). Arabidopsis NPF4.6 and NPF5.1 Control Leaf Stomatal Aperture by Regulating Abscisic Acid Transport. Genes, 12(6), 885. https://doi.org/10.3390/genes12060885





