ALDH2 p.E504K Variation and Sex Are Major Factors Associated with Current and Quitting Alcohol Drinking in Japanese Oldest Old
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Baseline Characteristics
2.3. Genotyping
2.4. Pre-Phasing and Imputation
2.5. Genome-Wide Association Study
2.6. Statistical Analysis
3. Results
3.1. Baseline Characteristics of the Kawasaki Aging and Wellbeing Project Cohort
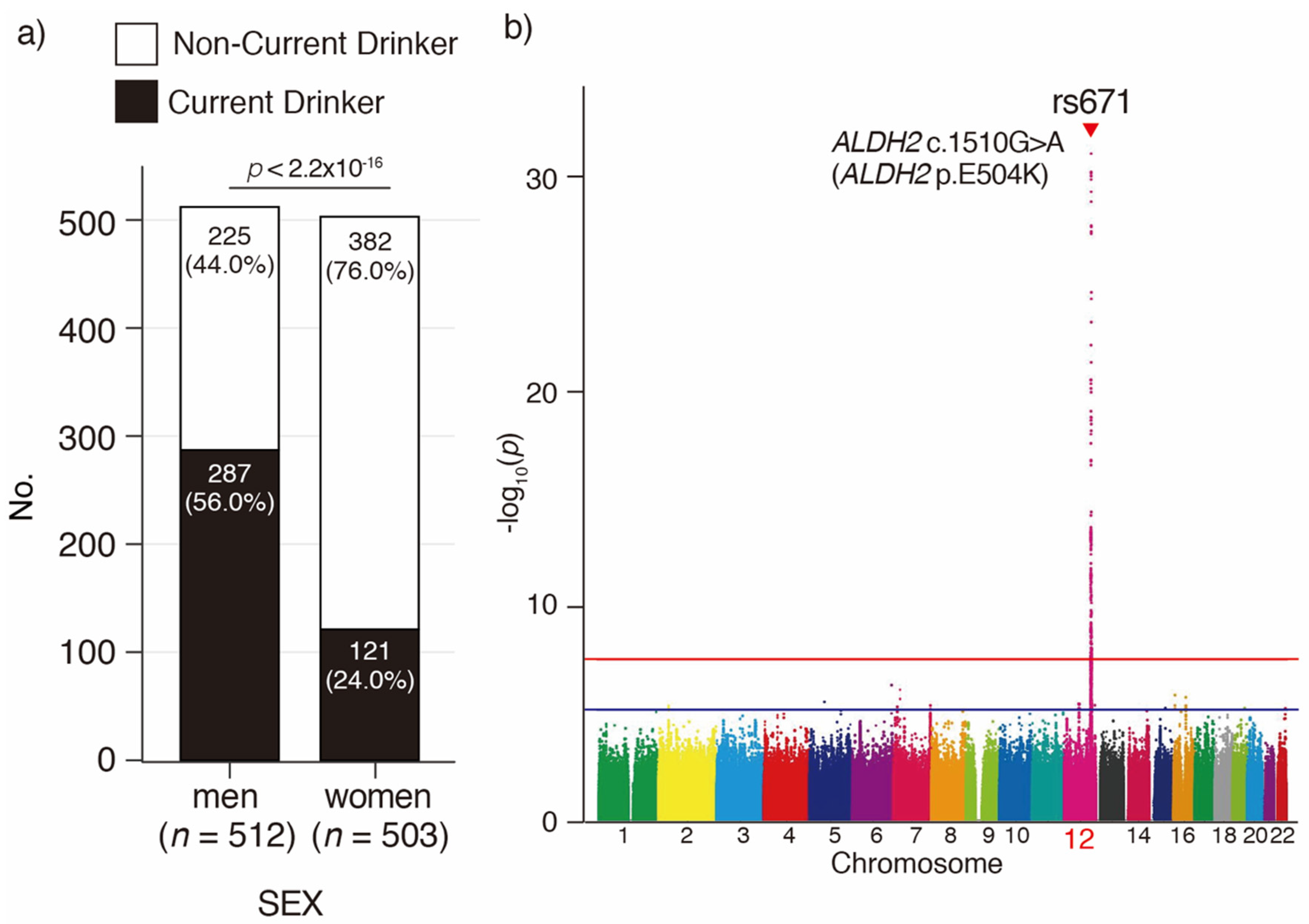
3.2. Imputation and GWAS for Current Drinking in Japanese Oldest Old
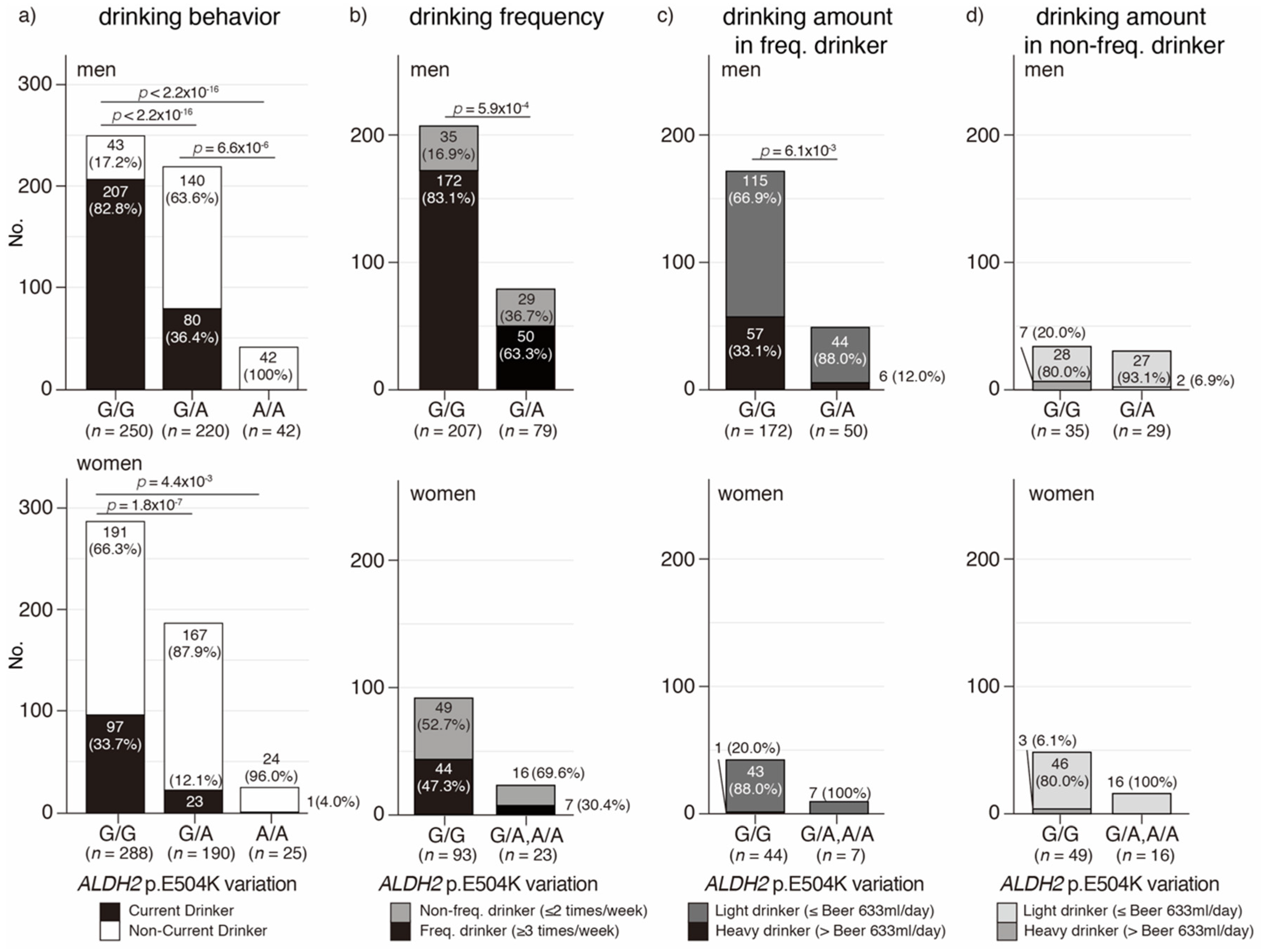
3.3. ALDH2 p.E504K Variation Is Associated with Drinking Behavior, Frequency, and Amount in the Japanese Oldest Old
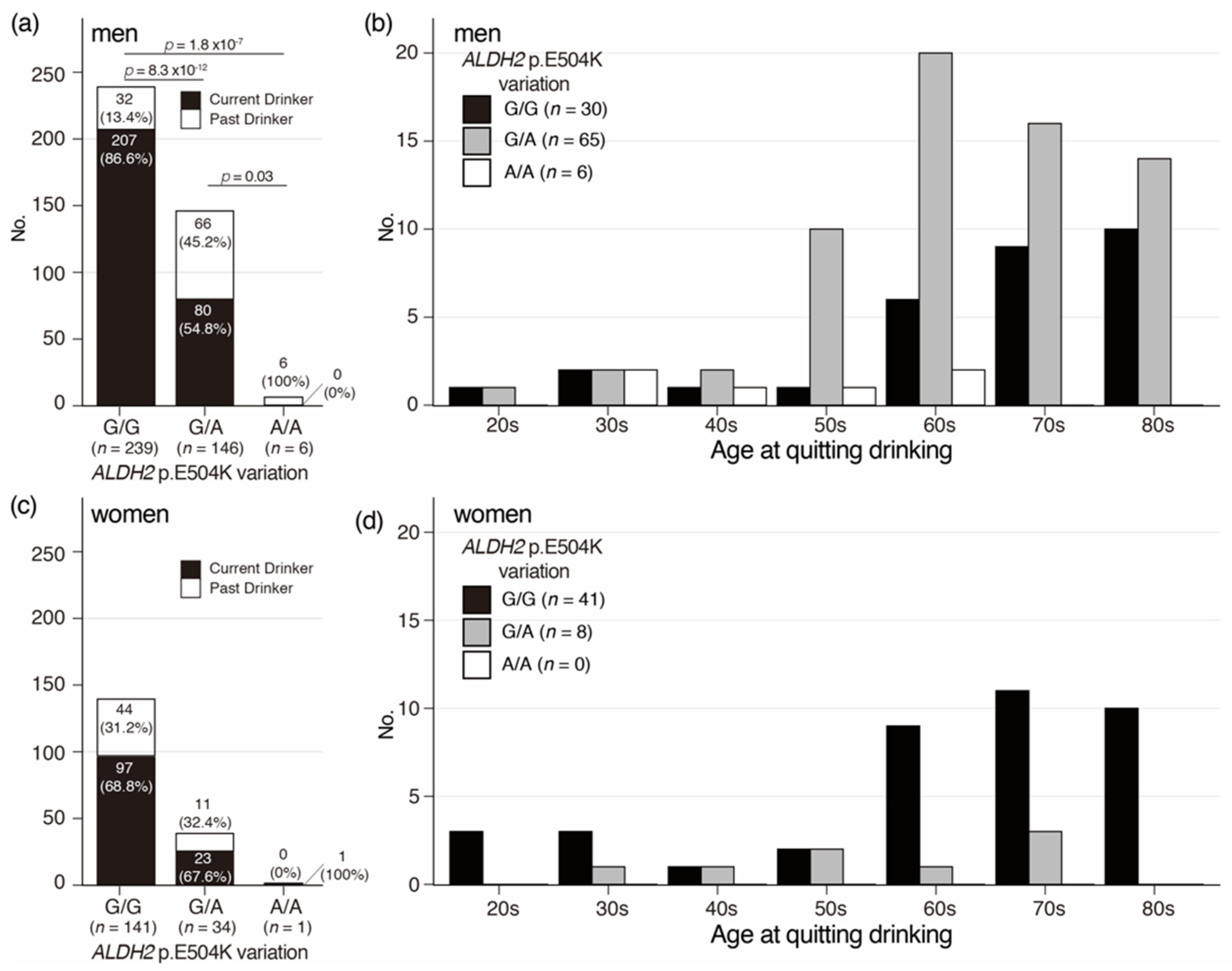
3.4. Distribution of Age at Quitting Drinking in Oldest Old Men and Women with the ALDH2 p.E504K Variation
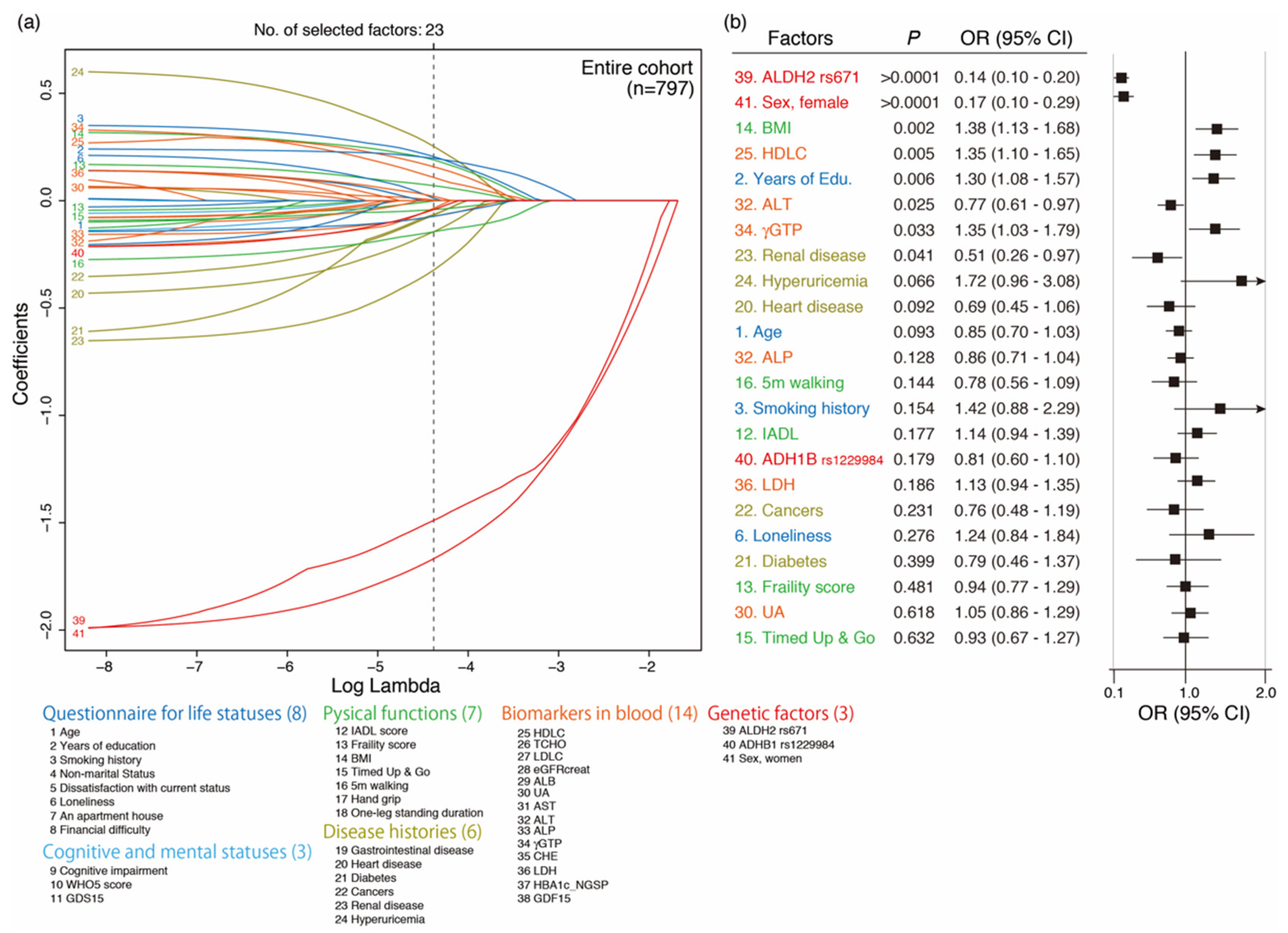
3.5. Variable Selection Associated with Current Drinking by LASSO and Its Multivariate Regression Logistic Analysis
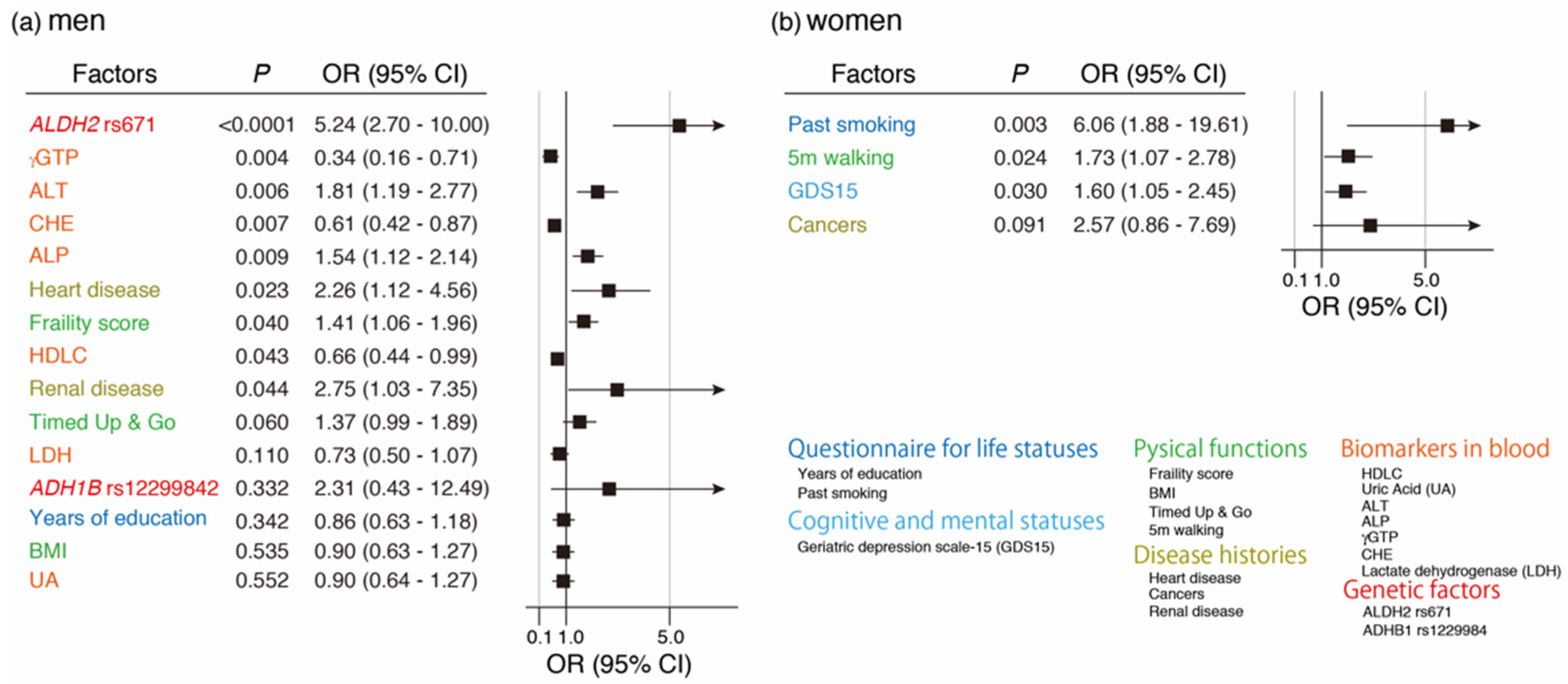
3.6. Variable Selection Associated with Quitting Drinking by LASSO and Its Multivariate Regression Logistic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Status Report on Alcohol and Health 2018; WHO: Genova, Switzerland, 2018. [Google Scholar]
- Hannah, R.; Max, R. Alcohol Consumption. Available online: OurWorldInData.org (accessed on 1 January 2018).
- Spear, L.P. Effects of adolescent alcohol consumption on the brain and behaviour. Nat. Rev. Neurosci. 2018, 19, 197–214. [Google Scholar] [CrossRef] [PubMed]
- Mende, M.A. Alcohol in the aging brain-the interplay between alcohol consumption, cognitive decline and the cardiovascular system. Front. Neurosci. 2019, 13, 713. [Google Scholar] [CrossRef] [PubMed]
- Public_Health_Scotland. Alcohol and Ageing; Public_Health_Scotland: Edinburgh, UK, 2006. [Google Scholar]
- Higuchi, S.; Ozaki, Y.; Shirasaka, T.; Hiro, H.; Matsushita, S. Research on the Actual Drinking Situation of Adults and the Prevention of Related Problems.; Health Labour Sciences Research: Tokyo, Japan, 2003. [Google Scholar]
- Edenberg, H.J. The genetics of alcohol metabolism: Role of alcohol dehydrogenase and aldehyde dehydrogenase variants. Alcohol. Res. Health 2007, 30, 5–13. [Google Scholar] [PubMed]
- Lai, C.L.; Yao, C.T.; Chau, G.Y.; Yang, L.F.; Kuo, T.Y.; Chiang, C.P.; Yin, S.J. Dominance of the inactive Asian variant over activity and protein contents of mitochondrial aldehyde dehydrogenase 2 in human liver. Alcohol. Clin. Exp. Res. 2014, 38, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Kitagawa, K.; Kawamoto, T.; Kunugita, N.; Tsukiyama, T.; Okamoto, K.; Yoshida, A.; Nakayama, K.; Nakayama, K. Aldehyde dehydrogenase (ALDH) 2 associates with oxidation of methoxyacetaldehyde; in vitro analysis with liver subcellular fraction derived from human and Aldh2 gene targeting mouse. FEBS Lett. 2000, 476, 306–311. [Google Scholar] [CrossRef]
- Bosron, W.F.; Li, T.K. Genetic polymorphism of human liver alcohol and aldehyde dehydrogenases, and their relationship to alcohol metabolism and alcoholism. Hepatology 1986, 6, 502–510. [Google Scholar] [CrossRef]
- Mulligan, C.J.; Robin, R.W.; Osier, M.V.; Sambuughin, N.; Goldfarb, L.G.; Kittles, R.A.; Hesselbrock, D.; Goldman, D.; Long, J.C. Allelic variation at alcohol metabolism genes (ADH1B, ADH1C, ALDH2) and alcohol dependence in an American Indian population. Hum. Genet. 2003, 113, 325–336. [Google Scholar] [CrossRef]
- Takeuchi, F.; Isono, M.; Nabika, T.; Katsuya, T.; Sugiyama, T.; Yamaguchi, S.; Kobayashi, S.; Ogihara, T.; Yamori, Y.; Fujioka, A.; et al. Confirmation of ALDH2 as a Major locus of drinking behavior and of its variants regulating multiple metabolic phenotypes in a Japanese population. Circ. J. 2011, 75, 911–918. [Google Scholar] [CrossRef]
- Okamoto, K.; Murawaki, Y.; Yuasa, I.; Kawasaki, H. Effect of ALDH2 and CYP2E1 gene polymorphisms on drinking behavior and alcoholic liver disease in Japanese male workers. Alcohol. Clin. Exp. Res. 2001, 25, 19S–23S. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Chang, S.; Wang, F.; Sun, H.; Ni, Z.; Yue, W.; Zhou, H.; Gelernter, J.; Malison, R.T.; Kalayasiri, R.; et al. Genome-wide association study of alcohol dependence in male Han Chinese and cross-ethnic polygenic risk score comparison. Transl. Psychiatry 2019, 9, 249. [Google Scholar] [CrossRef] [PubMed]
- Arai, Y.; Oguma, Y.; Abe, Y.; Takayama, M.; Hara, A.; Urushihara, H.; Takebayashi, T. Behavioral changes and hygiene practices of older adults in Japan during the first wave of COVID-19 emergency. BMC Geriatr. 2021, 21, 137. [Google Scholar] [CrossRef] [PubMed]
- Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Statist. Soc. B 1996, 58, 267–288. [Google Scholar] [CrossRef]
- Arai, Y.; Iinuma, T.; Takayama, M.; Takayama, M.; Abe, Y.; Fukuda, R.; Ando, J.; Ohta, K.; Hanabusa, H.; Asakura, K.; et al. The Tokyo Oldest Old survey on Total Health (TOOTH): A longitudinal cohort study of multidimensional components of health and well-being. BMC Geriatr. 2010, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Hirata, T.; Arai, Y.; Yuasa, S.; Abe, Y.; Takayama, M.; Sasaki, T.; Kunitomi, A.; Inagaki, H.; Endo, M.; Morinaga, J.; et al. Associations of cardiovascular biomarkers and plasma albumin with exceptional survival to the highest ages. Nat. Commun. 2020, 11, 3820. [Google Scholar] [CrossRef] [PubMed]
- Delaneau, O.; Zagury, J.F.; Robinson, M.R.; Marchini, J.L.; Dermitzakis, E.T. Accurate, scalable and integrative haplotype estimation. Nat. Commun. 2019, 10, 5436. [Google Scholar] [CrossRef] [PubMed]
- Rubinacci, S.; Delaneau, O.; Marchini, J. Genotype imputation using the Positional Burrows Wheeler Transform. PLoS Genet. 2020, 16, e1009049. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- Cui, R.; Kamatani, Y.; Takahashi, A.; Usami, M.; Hosono, N.; Kawaguchi, T.; Tsunoda, T.; Kamatani, N.; Kubo, M.; Nakamura, Y.; et al. Functional variants in ADH1B and ALDH2 coupled with alcohol and smoking synergistically enhance esophageal cancer risk. Gastroenterology 2009, 137, 1768–1775. [Google Scholar] [CrossRef] [PubMed]
| Oldest Old Men | Oldest Old Women | Oldest Old Men vs. Oldest Old Women | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| N | Current Drinker | Non-Current Drinker | p | Effect Size 4 | N | Current Drinker | Non-Current Drinker | p | Effect Size 4 | Men | Women | p | ||
| No of participants, no | 512 | 287 | 225 | - | - | 503 | 121 | 382 | - | - | 512 | 503 | - | |
| No of frequent drinker (≥3 times/week) | 286 | 222 | - | - | - | 116 | 51 | - | - | - | 222 | 51 | <0.001 | |
| No of higher amount drinker (>Beer 633ml/time) | 286 | 72 | - | - | - | 116 | 4 | - | - | - | 72 | 4 | <0.001 | |
| No of quitted drinker | 512 | - | 96 | - | - | 503 | - | 55 | - | - | 96 | 55 | <0.001 | |
| Questionnaire for life statuses (8) | ||||||||||||||
| Age (years), mean (SD) 1 | 512 | 87.0 (1.3) | 87.0 (1.3) | 0.645 | −0.04 | 503 | 86.9 (1.3) | 87.1 (1.3) | 0.060 | −0.20 | 87.0 (1.4) | 87.1 (1.4) | 0.330 | |
| Educated years (year), mean (SD) 2 | 511 | 12.3 (3.7) | 11.8 (3.5) | 0.122 | 0.07 | 503 | 11.0 (2.8) | 10.5 (2.4) | 0.114 | 0.07 | 12.1 (3.7) | 10.6 (2.6) | <0.001 | |
| Smoking history, no (%) 3 | 512 | 222 (77.3) | 166 (73.7) | 0.404 | 0.04 | 500 | 10 (8.3) | 29 (7.6) | 0.956 | 0.01 | 388 (75.8) | 39 (7.8) | <0.001 | |
| Non-marital Status, no (%) 3 | 509 | 63 (22.0) | 48 (21.6) | 0.999 | 0.01 | 500 | 95 (79.8) | 295 (77.4) | 0.670 | 0.03 | 111 (21.9) | 390 (78.0) | <0.001 | |
| Dissatisfaction with current status, no (%) 3 | 510 | 45 (15.7) | 32 (14.4) | 0.787 | 0.02 | 501 | 13 (10.8) | 46 (12.1) | 0.722 | 0.02 | 77 (15.1) | 59 (11.8) | 0.149 | |
| Loneliness, no (%) 3 | 474 | 91 (33.8) | 67 (32.7) | 0.870 | 0.01 | 470 | 40 (35.1) | 93 (26.1) | 0.084 | 0.09 | 158 (33.3) | 133 (28.3) | 0.109 | |
| An apartment house, no (%) 3 | 499 | 86 (30.8) | 66 (29.6) | 0.986 | 0.01 | 483 | 49 (40.8) | 136 (35.6) | 0.354 | 0.05 | 152 (29.9) | 185 (36.9) | 0.022 | |
| Financial difficulty, no (%) 3 | 503 | 59 (20.9) | 40 (18.1) | 0.498 | 0.04 | 481 | 17 (14.8) | 72 (19.7) | 0.298 | 0.05 | 99 (19.7) | 89 (18.5) | 0.697 | |
| Cognitive and mental statuses (3) | ||||||||||||||
| Cognitive impairment (MMSE ≤ 23), no (%) 3 | 511 | 49 (17.1) | 42 (18.7) | 0.739 | 0.02 | 500 | 14 (11.6) | 57 (15.0) | 0.422 | 0.04 | 91 (17.8) | 71 (14.2) | 0.139 | |
| WHO5 score, mean (SD) 2 | 504 | 18.2 (5.3) | 18.2 (5.1) | 0.900 | 0.00 | 491 | 18.5 (4.9) | 18.5 (5.1) | 0.951 | 0.00 | 18.2 (5.2) | 18.5 (5.1) | 0.395 | |
| GDS15, mean (SD) 2 | 504 | 3.3 (2.8) | 3.6 (3.0) | 0.377 | 0.04 | 494 | 2.9 (2.5) | 3.2 (2.6) | 0.221 | 0.06 | 3.5 (2.9) | 3.2 (2.6) | 0.249 | |
| Pysical functions (7) | ||||||||||||||
| IADL score, mean (SD) 2 | 512 | 4.8 (0.4) | 4.8 (0.5) | 0.929 | 0.00 | 502 | 4.9 (0.4) | 4.8 (0.5) | 0.122 | 0.07 | 4.8 (0.4) | 4.8 (0.5) | 0.483 | |
| Frailty score, mean (SD) 2 | 502 | 1.4 (0.9) | 1.5 (1.1) | 0.353 | 0.04 | 494 | 1.4 (1.0) | 1.4 (0.9) | 0.583 | 0.02 | 1.5 (1.0) | 1.4 (0.9) | 0.797 | |
| BMI, mean (SD) 1 | 512 | 23.8 (2.6) | 23.3 (3.3) | 0.098 | 0.15 | 503 | 23.1 (3.0) | 22.8 (3.2) | 0.242 | 0.12 | 23.6 (2.9) | 22.9 (3.2) | <0.001 | |
| Timed Up & Go (sec), mean (SD) 2 | 505 | 11.4 (2.7) | 11.5 (2.6) | 0.570 | 0.03 | 497 | 11.2 (2.7) | 11.8 (3.3) | 0.134 | 0.07 | 11.4 (2.7) | 11.6 (3.2) | 0.854 | |
| 5m walking (sec), mean (SD) 2 | 510 | 5.4 (1.1) | 5.6 (1.7) | 0.878 | 0.01 | 499 | 5.3 (1.1) | 5.7 (1.7) | 0.012 | 0.11 | 5.5 (1.4) | 5.6 (1.6) | 0.735 | |
| Hand grip (kg), mean (SD) 2 | 506 | 27.2 (4.9) | 27.4 (5.0) | 0.609 | 0.02 | 497 | 18.5 (3.3) | 18.0 (3.3) | 0.092 | 0.08 | 27.3 (4.9) | 18.1 (3.3) | <0.001 | |
| One-leg standing duration (sec), mean (SD) 2 | 501 | 23.2 (29.2) | 21.5 (27.7) | 0.834 | 0.01 | 493 | 21.5 (23.6) | 20.2 (24.7) | 0.278 | 0.05 | 22.4 (28.5) | 20.5 (24.4) | 0.842 | |
| Disease histories (6) | ||||||||||||||
| Gastrointestinal disease, no (%) 3 | 511 | 171 (59.8) | 137 (60.9) | 0.872 | 0.01 | 502 | 72 (60.0) | 227 (59.4) | 0.996 | 0.01 | 308 (60.3) | 299 (59.6) | 0.867 | |
| Heart disease, no (%) 3 | 511 | 82 (28.7) | 79 (35.1) | 0.144 | 0.07 | 499 | 26 (22.0) | 78 (20.5) | 0.814 | 0.02 | 161 (31.5) | 104 (20.8) | <0.001 | |
| Diabetes, no (%) 3 | 507 | 38 (13.4) | 44 (19.6) | 0.077 | 0.08 | 494 | 11 (9.2) | 45 (12.0) | 0.509 | 0.04 | 82 (16.2) | 56 (11.3) | 0.034 | |
| Cancers, no (%) 3 | 508 | 70 (24.5) | 66 (29.7) | 0.220 | 0.06 | 497 | 13 (11.0) | 60 (15.8) | 0.254 | 0.06 | 136 (26.8) | 73 (14.7) | <0.001 | |
| Renal disease, no (%) 3 | 510 | 25 (8.7) | 31 (13.8) | 0.092 | 0.08 | 497 | 7 (5.8) | 35 (9.3) | 0.320 | 0.05 | 56 (11.0) | 42 (8.5) | 0.212 | |
| Hyperuricemia, no (%) 3 | 501 | 53 (18.9) | 27 (12.3) | 0.061 | 0.09 | 487 | 7 (6.2) | 18 (4.8) | 0.734 | 0.03 | 80 (16.0) | 25 (5.1) | <0.001 | |
| Biomarkers in plasma (14) | ||||||||||||||
| HDLC (mg/dL), mean (SD) 2 | 512 | 58.8 (14.7) | 51.9 (13.6) | <0.001 | 0.24 | 503 | 66.9 (15.3) | 65.3 (15.6) | 0.367 | 0.04 | 55.8 (14.6) | 65.7 (15.6) | <0.001 | |
| TCHO (mg/dL), mean (SD) 2 | 512 | 192.4 (31.1) | 184.9 (31.7) | 0.007 | 0.12 | 503 | 210.5 (30.5) | 211.2 (34.1) | 0.934 | 0.00 | 189.1 (31.5) | 211.0 (33.3) | <0.001 | |
| LDLC (mg/dL), mean (SD) 2 | 512 | 106.4 (26.3) | 105.6 (26.9) | 0.735 | 0.01 | 503 | 114.8 (27.1) | 117.1 (28.7) | 0.494 | 0.03 | 106.0 (26.6) | 116.5 (28.3) | <0.001 | |
| eGFRcreat (mL/min/1.73m2), mean (SD) 2 | 512 | 58.8 (14.3) | 56.8 (15.1) | 0.091 | 0.07 | 503 | 61.1 (15.6) | 59.7 (13.8) | 0.495 | 0.03 | 57.9 (14.7) | 60.0 (14.3) | 0.025 | |
| ALB (g/dL), mean (SD) 2 | 512 | 4.1 (0.3) | 4.2 (0.3) | 0.624 | 0.02 | 503 | 4.2 (0.3) | 4.2 (0.3) | 0.842 | 0.01 | 4.2 (0.3) | 4.2 (0.3) | 0.202 | |
| UA (mg/dL), mean (SD) 2 | 512 | 6.0 (1.3) | 5.8 (1.2) | 0.145 | 0.06 | 503 | 5.1 (1.2) | 5.1 (1.2) | 0.542 | 0.03 | 5.9 (1.3) | 5.1 (1.2) | <0.001 | |
| AST (U/L), mean (SD) 2 | 512 | 24.4 (8.9) | 23.9 (9.9) | 0.260 | 0.05 | 503 | 24.0 (6.3) | 25.8 (27.7) | 0.973 | 0.00 | 24.2 (9.3) | 25.4 (24.3) | 0.192 | |
| ALT (U/L), mean (SD) 2 | 512 | 17.6 (8.8) | 17.8 (10.9) | 0.746 | 0.01 | 503 | 16.4 (5.4) | 18.6 (38.1) | 0.476 | 0.03 | 17.7 (9.8) | 18.1 (33.3) | 0.148 | |
| ALP (U/L), mean (SD) 2 | 512 | 229.3 (66.0) | 245.0 (75.3) | 0.012 | 0.11 | 503 | 233.7 (72.6) | 239.9 (75.7) | 0.447 | 0.03 | 236.2 (70.6) | 238.4 (75.0) | 0.762 | |
| gGTP (U/L), mean (SD) 2 | 512 | 35.9 (41.5) | 25.1 (22.3) | <0.001 | 0.24 | 503 | 23.7 (40.9) | 22.9 (16.7) | 0.741 | 0.01 | 31.1 (34.8) | 23.0 (24.7) | <0.001 | |
| CHE (U/L), mean (SD) 2 | 512 | 283.4 (59.7) | 278.4 (59.8) | 0.461 | 0.03 | 503 | 305.8 (57.0) | 312.0 (66.8) | 0.570 | 0.03 | 281.2 (59.8) | 310.5 (64.6) | <0.001 | |
| LDH (U/L), mean (SD) 2 | 512 | 194.4 (37.2) | 191.8 (40.1) | 0.388 | 0.04 | 503 | 211.1 (38.1) | 207.1 (41.2) | 0.244 | 0.05 | 193.3 (38.5) | 208.1 (40.5) | <0.001 | |
| HBA1c_NGSP (%), mean (SD) 2 | 512 | 5.9 (0.6) | 6.0 (0.7) | 0.047 | 0.09 | 503 | 5.9 (0.4) | 6.0 (0.6) | 0.826 | 0.01 | 6.0 (0.6) | 6.0 (0.6) | 0.635 | |
| GDF15, mean (SD) 2 | 512 | 1854.4 (859.0) | 1900.0 (909.0) | 0.715 | 0.02 | 503 | 1431.8 (497.9) | 1530.2 (521.5) | 0.064 | 0.08 | 1874.6 (880.7) | 1506.7 (517.2) | <0.001 | |
| Genetic factors (3) | ||||||||||||||
| ALDH2 rs671 (p.E504K ), minor allele frequency 3 | 512 | 0.139 | 0.497 | <0.001 | 0.39 | 503 | 0.103 | 0.281 | <0.001 | 0.18 | 0.297 | 0.239 | 0.004 | |
| ADH1B rs1229984 (p.H48R), major allele frequency 3 | 512 | 0.777 | 0.782 | 0.460 | 0.03 | 503 | 0.747 | 0.797 | 0.125 | 0.05 | 0.780 | 0.785 | 0.632 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sasaki, T.; Nishimoto, Y.; Hirata, T.; Abe, Y.; Takebayashi, T.; Arai, Y. ALDH2 p.E504K Variation and Sex Are Major Factors Associated with Current and Quitting Alcohol Drinking in Japanese Oldest Old. Genes 2021, 12, 799. https://doi.org/10.3390/genes12060799
Sasaki T, Nishimoto Y, Hirata T, Abe Y, Takebayashi T, Arai Y. ALDH2 p.E504K Variation and Sex Are Major Factors Associated with Current and Quitting Alcohol Drinking in Japanese Oldest Old. Genes. 2021; 12(6):799. https://doi.org/10.3390/genes12060799
Chicago/Turabian StyleSasaki, Takashi, Yoshinori Nishimoto, Takumi Hirata, Yukiko Abe, Toru Takebayashi, and Yasumichi Arai. 2021. "ALDH2 p.E504K Variation and Sex Are Major Factors Associated with Current and Quitting Alcohol Drinking in Japanese Oldest Old" Genes 12, no. 6: 799. https://doi.org/10.3390/genes12060799
APA StyleSasaki, T., Nishimoto, Y., Hirata, T., Abe, Y., Takebayashi, T., & Arai, Y. (2021). ALDH2 p.E504K Variation and Sex Are Major Factors Associated with Current and Quitting Alcohol Drinking in Japanese Oldest Old. Genes, 12(6), 799. https://doi.org/10.3390/genes12060799







