Causal Association between Periodontitis and Parkinson’s Disease: A Bidirectional Mendelian Randomization Study
Abstract
1. Introduction
2. Materials and Methods
2.1. MR of Periodontitis on Risk of PD
2.2. MR of PD on Risk of Periodontitis
2.3. MR Analysis
3. Results
Mendelian Randomization
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Tysnes, O.B.; Storstein, A. Epidemiology of Parkinson’s disease. J. Neural Transm. 2017, 124, 901–905. [Google Scholar] [CrossRef]
- Parnetti, L.; Gaetani, L.; Eusebi, P.; Paciotti, S.; Hansson, O.; El-Agnaf, O.; Mollenhauer, B.; Blennow, K.; Calabresi, P. CSF and blood biomarkers for Parkinson’s disease. Lancet Neurol. 2019, 18, 573–586. [Google Scholar] [CrossRef]
- Noyce, A.J.; Bandres-Ciga, S.; Kim, J.; Heilbron, K.; Kia, D.; Hemani, G.; Xue, A.; Lawlor, D.A.; Smith, G.D.; Duran, R.; et al. The Parkinson’s Disease Mendelian Randomization Research Portal. Mov. Disord. 2019, 34, 1864–1872. [Google Scholar] [CrossRef] [PubMed]
- Qiu, X.; Xiao, Y.; Wu, J.; Gan, L.; Huang, Y.; Wang, J. C-Reactive Protein and Risk of Parkinson’s Disease: A Systematic Review and Meta-Analysis. Front. Neurol. 2019, 10, 384. [Google Scholar] [CrossRef]
- Botelho, J.; Lyra, P.; Proença, L.; Godinho, C.; Mendes, J.J.; Machado, V. Relationship between blood and standard biochemistry levels with Periodontitis in Parkinson’s Disease patients: Data from the NHANES 2011-2012. J. Pers. Med. 2020, 10, 69. [Google Scholar] [CrossRef] [PubMed]
- Tonetti, M.S.; Jepsen, S.; Jin, L.; Otomo-Corgel, J. Impact of the global burden of periodontal diseases on health, nutrition and wellbeing of mankind: A call for global action. J. Clin. Periodontol. 2017, 44, 456–462. [Google Scholar] [CrossRef] [PubMed]
- Caton, J.G.; Armitage, G.; Berglundh, T.; Chapple, I.L.C.; Jepsen, S.; Kornman, K.S.; Mealey, B.L.; Papapanou, P.N.; Sanz, M.; Tonetti, M.S. A new classification scheme for periodontal and peri-implant diseases and conditions—Introduction and key changes from the 1999 classification. J. Clin. Periodontol. 2018, 45, S1–S8. [Google Scholar] [CrossRef]
- Kaur, T.; Uppoor, A.; Naik, D. Parkinson’s disease and periodontitis—The missing link? A review. Gerodontology 2016, 33, 434–438. [Google Scholar] [CrossRef]
- Lyra, P.; Machado, V.; Proença, L.; Domingos, J.; Godinho, C.; Mendes, J.J.; Botelho, J. Parkinson’s Disease, Periodontitis and patient-related outcomes: A cross-sectional study. Medicina 2020, 56, 383. [Google Scholar] [CrossRef]
- Schwarz, J.; Heimhilger, E.; Storch, A. Increased periodontal pathology in Parkinson’s disease. J. Neurol. 2006, 253, 608–611. [Google Scholar] [CrossRef]
- Einarsdóttir, E.R.; Gunnsteinsdóttir, H.; Hallsdóttir, M.H.; Sveinsson, S.; Jónsdóttir, S.R.; Olafsson, V.G.; Bragason, T.H.; Saemundsson, S.R.; Holbrook, W.P. Dental health of patients with Parkinson’s disease in Iceland. Spec. Care Dent. 2009, 29, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Hanaoka, A.; Kashihara, K. Increased frequencies of caries, periodontal disease and tooth loss in patients with Parkinson’s disease. J. Clin. Neurosci. 2009, 16, 1279–1282. [Google Scholar] [CrossRef]
- Nakayama, Y.; Washio, M.; Mori, M. Oral health conditions in patients with Parkinson’s disease. J. Epidemiol. 2004, 14, 143–150. [Google Scholar] [CrossRef]
- Van Stiphout, M.A.E.; Marinus, J.; Van Hilten, J.J.; Lobbezoo, F.; De Baat, C. Oral Health of Parkinson’s Disease Patients: A Case-Control Study. Parkinsons. Dis. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Burgess, S.; Labrecque, J.A. Mendelian randomization with a binary exposure variable: Interpretation and presentation of causal estimates. Eur. J. Epidemiol. 2018, 33, 947–952. [Google Scholar] [CrossRef]
- Burgess, S.; Butterworth, A.; Thompson, S.G. Mendelian Randomization Analysis With Multiple Genetic Variants Using Summarized Data. Genet. Epidemiol. 2013, 37, 658–665. [Google Scholar] [CrossRef]
- Bowden, J.; Davey Smith, G.; Haycock, P.C.; Burgess, S. Consistent Estimation in Mendelian Randomization with Some Invalid Instruments Using a Weighted Median Estimator. Genet. Epidemiol. 2016, 40, 304–314. [Google Scholar] [CrossRef] [PubMed]
- Teumer, A.; Holtfreter, B.; Völker, U.; Petersmann, A.; Nauck, M.; Biffar, R.; Völzke, H.; Kroemer, H.K.; Meisel, P.; Homuth, G.; et al. Genome-wide association study of chronic periodontitis in a general German population. J. Clin. Periodontol. 2013, 40, 977–985. [Google Scholar] [CrossRef]
- Munz, M.; Richter, G.M.; Loos, B.G.; Jepsen, S.; Divaris, K.; Offenbacher, S.; Teumer, A.; Holtfreter, B.; Kocher, T.; Bruckmann, C.; et al. Meta-analysis of genome-wide association studies of aggressive and chronic periodontitis identifies two novel risk loci. Eur. J. Hum. Genet. 2019, 27, 102–113. [Google Scholar] [CrossRef]
- Shungin, D.; Haworth, S.; Divaris, K.; Agler, C.S.; Kamatani, Y.; Keun Lee, M.; Grinde, K.; Hindy, G.; Alaraudanjoki, V.; Pesonen, P.; et al. Genome-wide analysis of dental caries and periodontitis combining clinical and self-reported data. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef]
- Nalls, M.A.; Blauwendraat, C.; Vallerga, C.L.; Heilbron, K.; Bandres-Ciga, S.; Chang, D.; Tan, M.; Kia, D.A.; Noyce, A.J.; Xue, A.; et al. Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: A meta-analysis of genome-wide association studies. Lancet Neurol. 2019, 18, 1091–1102. [Google Scholar] [CrossRef]
- Chang, D.; Nalls, M.A.; Hallgrímsdóttir, I.B.; Hunkapiller, J.; van der Brug, M.; Cai, F.; Kerchner, G.A.; Ayalon, G.; Bingol, B.; Sheng, M.; et al. A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci. Nat. Genet. 2017, 49, 1511–1516. [Google Scholar] [CrossRef]
- Offenbacher, S.; Divaris, K.; Barros, S.P.; Moss, K.L.; Marchesan, J.T.; Morelli, T.; Zhang, S.; Kim, S.; Sun, L.; Beck, J.D.; et al. Genome-wide association study of biologically informed periodontal complex traits offers novel insights into the genetic basis of periodontal disease. Hum. Mol. Genet. 2016, 25, 2113–2129. [Google Scholar] [CrossRef] [PubMed]
- Verbanck, M.; Chen, C.-Y.; Neale, B.; Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 2018, 50, 693–698. [Google Scholar] [CrossRef]
- Bowden, J.; Davey Smith, G.; Burgess, S. Mendelian randomization with invalid instruments: Effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 2015, 44, 512–525. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Wang, J.; Hemani, G.; Bowden, J.; Small, D.S. Statistical inference in two-sample summary-data Mendelian randomization using robust adjusted profile score. Ann. Statist. 2020, 48, 1742–1769. [Google Scholar] [CrossRef]
- Welter, D.; MacArthur, J.; Morales, J.; Burdett, T.; Hall, P.; Junkins, H.; Klemm, A.; Flicek, P.; Manolio, T.; Hindorff, L.; et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014, 42, D1001–D1006. [Google Scholar] [CrossRef]
- Leira, Y.; Domínguez, C.; Seoane, J.; Seoane-Romero, J.; Pías-Peleteiro, J.M.; Takkouche, B.; Blanco, J.; Aldrey, J.M. Is Periodontal Disease Associated with Alzheimer’s Disease? A Systematic Review with Meta-Analysis. Neuroepidemiology 2017, 48, 21–31. [Google Scholar] [CrossRef]
- Nadim, R.; Tang, J.; Dilmohamed, A.; Yuan, S.; Wu, C.; Bakre, A.T.; Partridge, M.; Ni, J.; Copeland, J.R.; Anstey, K.J.; et al. Influence of periodontal disease on risk of dementia: A systematic literature review and a meta-analysis. Eur. J. Epidemiol. 2020, 35, 821–833. [Google Scholar] [CrossRef]
- Ide, M.; Harris, M.; Stevens, A.; Sussams, R.; Hopkins, V.; Culliford, D.; Fuller, J.; Ibbett, P.; Raybould, R.; Thomas, R.; et al. Periodontitis and cognitive decline in Alzheimer’s disease. PLoS ONE 2016, 11, e0151081. [Google Scholar] [CrossRef]
- Dominy, S.S.; Lynch, C.; Ermini, F.; Benedyk, M.; Marczyk, A.; Konradi, A.; Nguyen, M.; Haditsch, U.; Raha, D.; Griffin, C.; et al. Porphyromonas gingivalis in Alzheimer’s disease brains: Evidence for disease causation and treatment with small-molecule inhibitors. Sci. Adv. 2019, 5, eaau3333. [Google Scholar] [CrossRef] [PubMed]
- Noble, J.M.; Borrell, L.N.; Papapanou, P.N.; Elkind, M.S.V.; Scarmeas, N.; Wright, C.B. Periodontitis is associated with cognitive impairment among older adults: Analysis of NHANES-III. J. Neurol. Neurosurg. Psychiatry 2009, 80, 1206–1211. [Google Scholar] [CrossRef] [PubMed]
- Ilievski, V.; Zuchowska, P.K.; Green, S.J.; Toth, P.T.; Ragozzino, M.E.; Le, K.; Aljewari, H.W.; O’Brien-Simpson, N.M.; Reynolds, E.C.; Watanabe, K. Chronic oral application of a periodontal pathogen results in brain inflammation, neurodegeneration and amyloid beta production in wild type mice. PLoS ONE 2018, 13, e0204941. [Google Scholar] [CrossRef] [PubMed]
- Do Lee, C.; Folsom, A.R.; Nieto, F.J.; Chambless, L.E.; Shahar, E.; Wolfe, D.A. White blood cell count and incidence of coronary heart disease and ischemic stroke and mortality from cardiovascular disease in African-American and White men and women: Atherosclerosis Risk in Communities Study. Am. J. Epidemiol. 2001, 154, 758–764. [Google Scholar] [CrossRef] [PubMed]
- Hegde, R.; Awan, K.H. Effects of periodontal disease on systemic health. Disease-a-Month 2019, 65, 185–192. [Google Scholar] [CrossRef] [PubMed]
- Tong, P.C.; Lee, K.F.; So, W.Y.; Ng, M.H.; Chan, W.B.; Lo, M.K.; Chan, N.N.; Chan, J.C. White Blood Cell Count Is Associated with Macro- and Microvascular Complications in Chinese Patients with Type 2 Diabetes. Diabetes Care 2004, 27, 216–222. [Google Scholar] [CrossRef]
- Divaris, K.; Monda, K.L.; North, K.E.; Olshan, A.F.; Reynolds, L.M.; Hsueh, W.C.; Lange, E.M.; Moss, K.; Barros, S.P.; Weyant, R.J.; et al. Exploring the genetic basis of chronic periodontitis: A genome-wide association study. Hum. Mol. Genet. 2013, 22, 2312–2324. [Google Scholar] [CrossRef]
- Schaefer, A.S.; Richter, G.M.; Nothnagel, M.; Manke, T.; Dommisch, H.; Jacobs, G.; Arlt, A.; Rosenstiel, P.; Noack, B.; Groessner-Schreiber, B.; et al. A genome-wide association study identifies GLT6D1 as a susceptibility locus for periodontitis. Hum. Mol. Genet. 2009, 19, 553–562. [Google Scholar] [CrossRef]
- Tonetti, M.S.; Greenwell, H.; Kornman, K.S. Staging and grading of periodontitis: Framework and proposal of a new classification and case definition. J. Periodontol. 2018, 45, S149–S161. [Google Scholar] [CrossRef] [PubMed]
- Botelho, J.; Machado, V.; Proença, L.; Mendes, J.J. The 2018 periodontitis case definition improves accuracy performance of full-mouth partial diagnostic protocols. Sci. Rep. 2020, 10, 7093. [Google Scholar] [CrossRef]
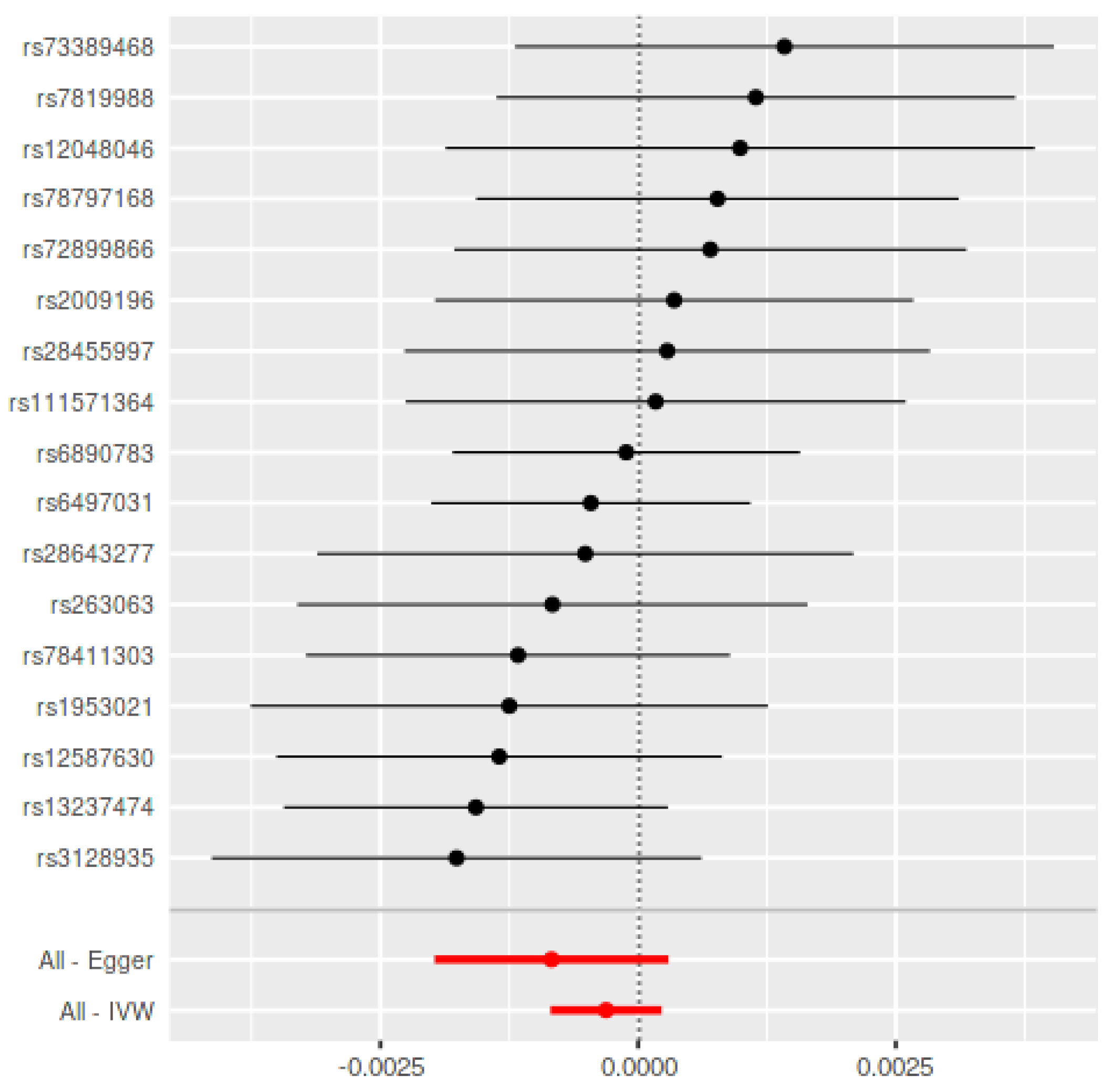
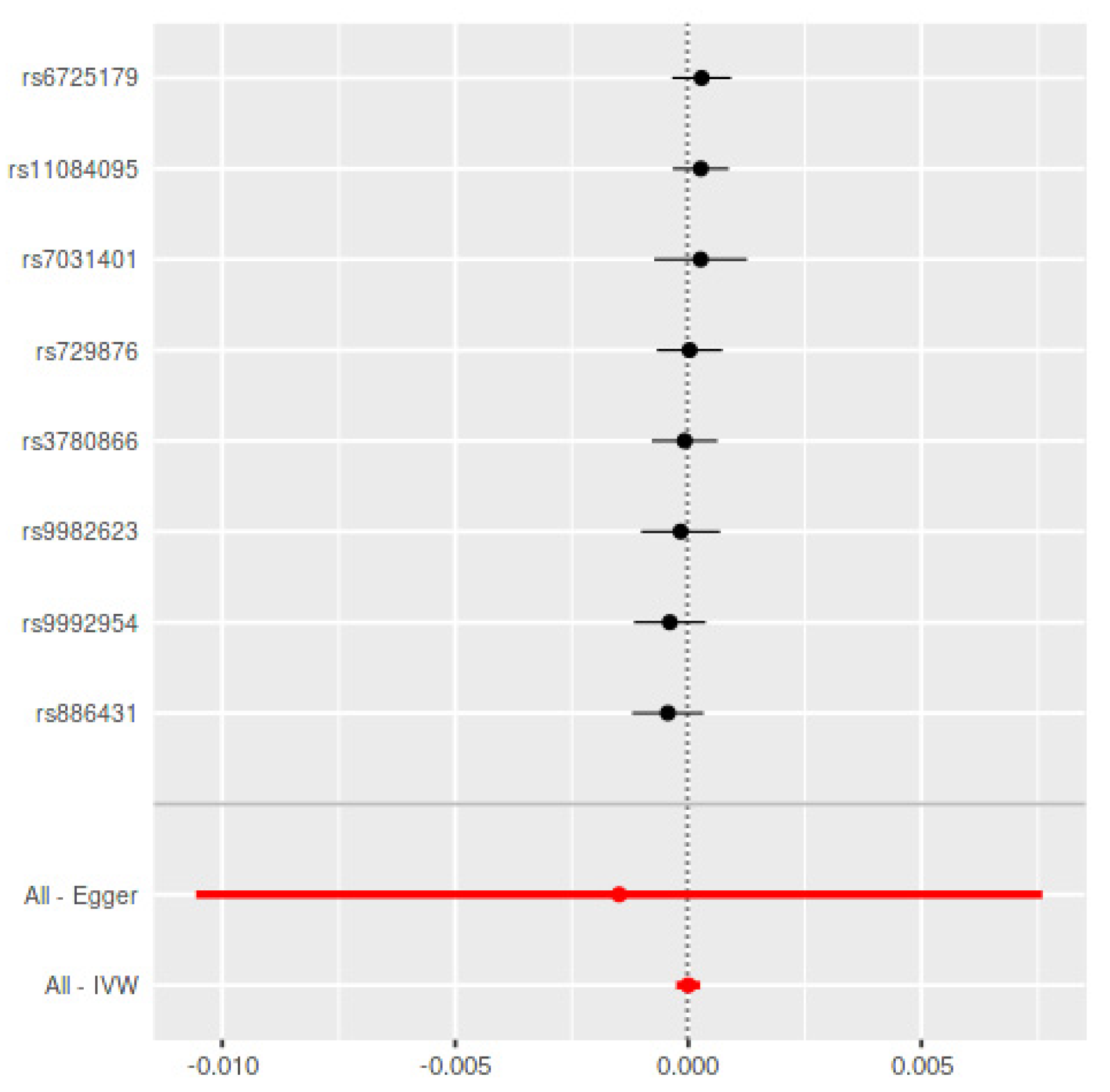
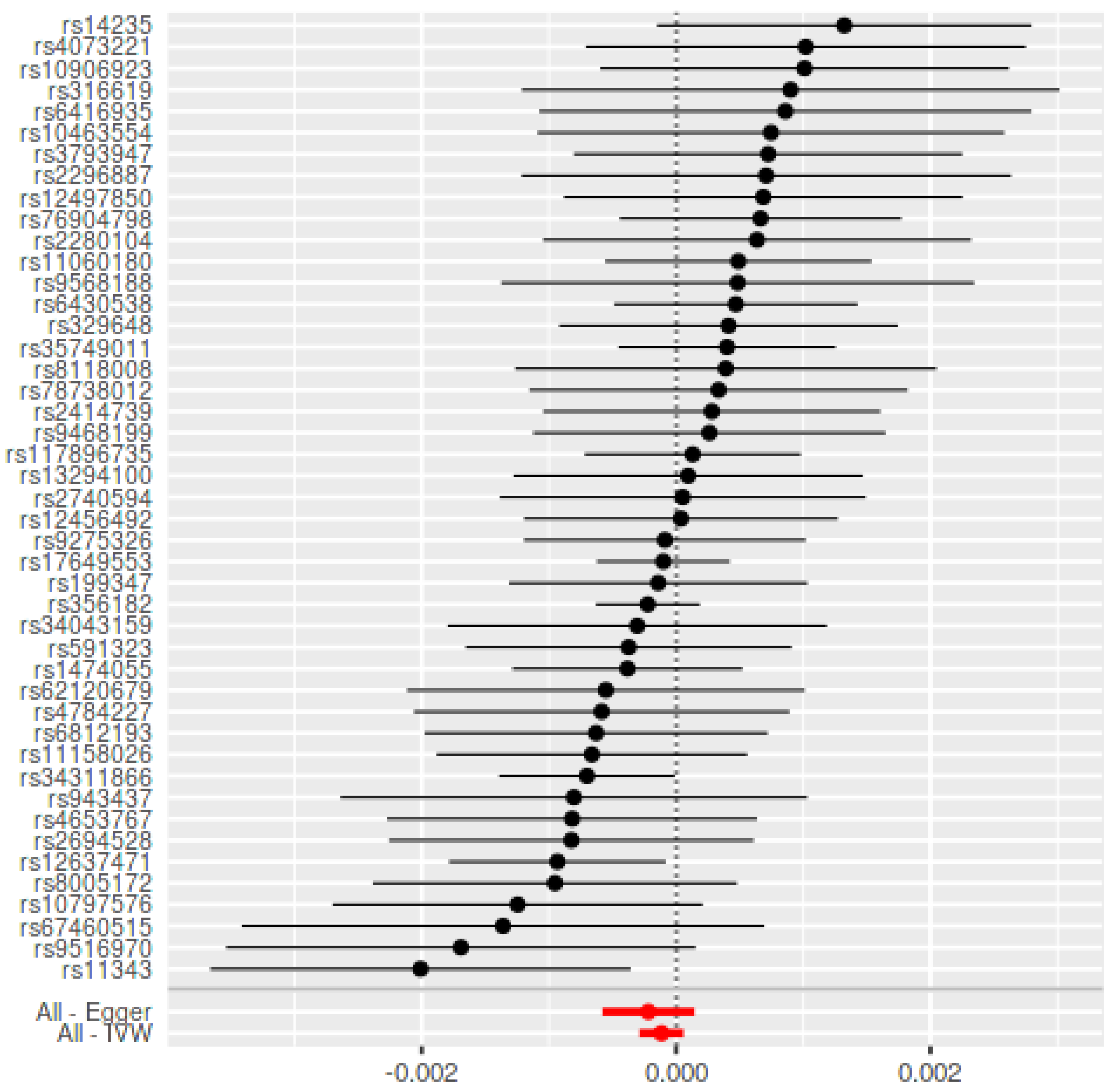
| Method | Periodontitis on PD | Parkinson’s Disease on Periodontitis Using Chang et al., 2017 ± | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Instrumental SNPs from Teumer et al. * | Instrumental SNPs from Munz et al. # | ||||||||
| B (SE) | p-Value | Q Statistic/p Value | B (Se) | p-Value | Q Statistic/p Value | B (Se) | p-Value | Q Statistic/p Value | |
| IVW | −0.0003 (0.0003) | 0.26 | 11.45/0.78 | −0.0000 (0.0001) | 0.99 | 4.30/0.75 | −0.0001 (0.0001) | 0.19 | 46.83/0.36 |
| Weighten median | −0.0003 (0.0003) | 0.37 | - | 0.0000 (0.0002) | 0.93 | - | −0.0001 (0.0001) | 0.31 | - |
| MR-Egger | −0.0008 (0.0006) | 0.17 | 10.37/0.80 | −0.0014 (0.0046) | 0.76 | 4.19/0.65 | −0.0002 (0.0002) | 0.24 | 46.39/0.33 |
| MR-RAPS | −0.0003 (0.0003) | 0.27 | - | −0.0000 (0.0001) | 0.99 | - | −0.0001 (0.0001) | 0.31 | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Botelho, J.; Machado, V.; Mendes, J.J.; Mascarenhas, P. Causal Association between Periodontitis and Parkinson’s Disease: A Bidirectional Mendelian Randomization Study. Genes 2021, 12, 772. https://doi.org/10.3390/genes12050772
Botelho J, Machado V, Mendes JJ, Mascarenhas P. Causal Association between Periodontitis and Parkinson’s Disease: A Bidirectional Mendelian Randomization Study. Genes. 2021; 12(5):772. https://doi.org/10.3390/genes12050772
Chicago/Turabian StyleBotelho, João, Vanessa Machado, José João Mendes, and Paulo Mascarenhas. 2021. "Causal Association between Periodontitis and Parkinson’s Disease: A Bidirectional Mendelian Randomization Study" Genes 12, no. 5: 772. https://doi.org/10.3390/genes12050772
APA StyleBotelho, J., Machado, V., Mendes, J. J., & Mascarenhas, P. (2021). Causal Association between Periodontitis and Parkinson’s Disease: A Bidirectional Mendelian Randomization Study. Genes, 12(5), 772. https://doi.org/10.3390/genes12050772









