Impacts of the Type I Toxin–Antitoxin System, SprG1/SprF1, on Staphylococcus aureus Gene Expression
Abstract
1. Introduction
2. Materials and Methods
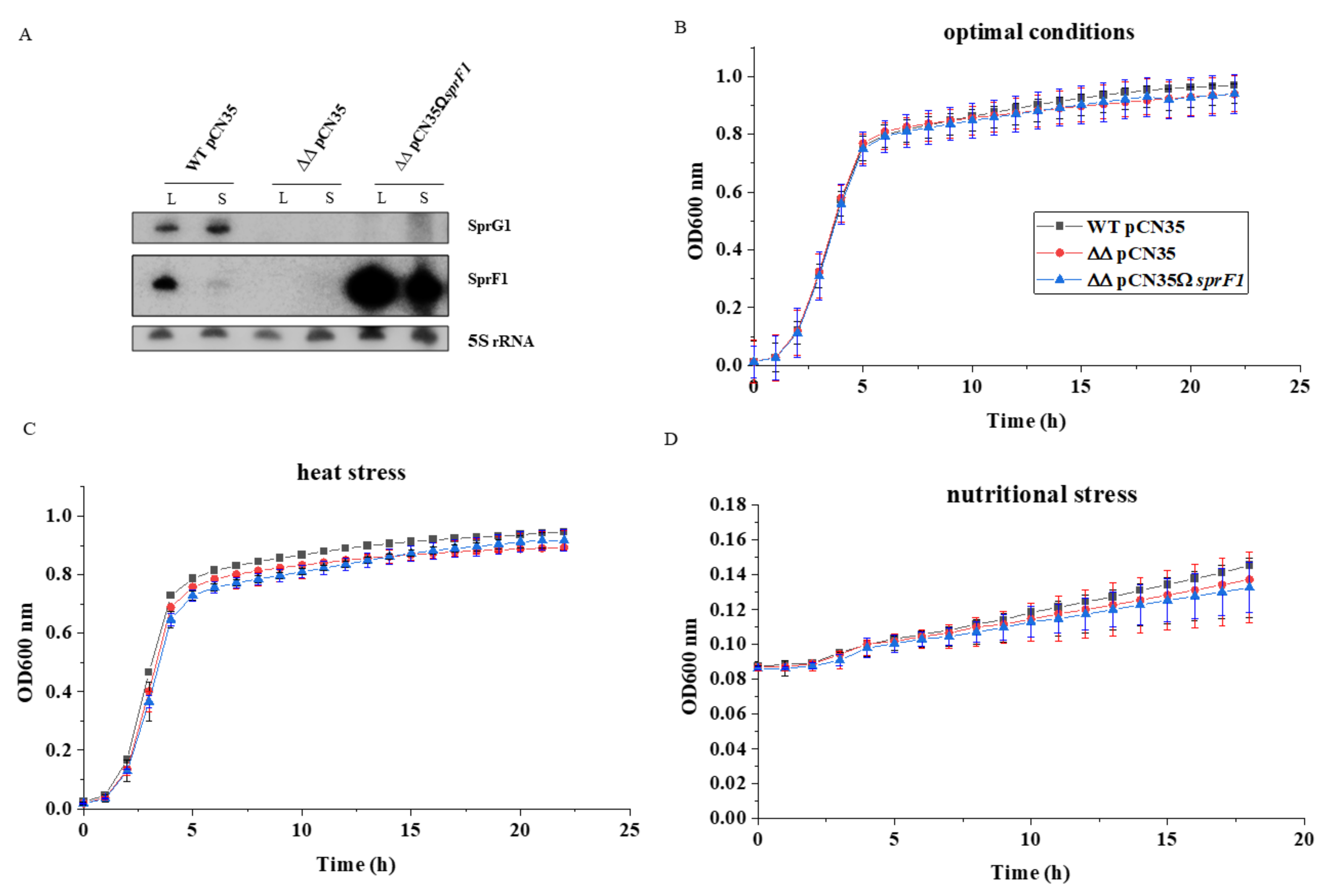
2.1. Bacterial Strains and Growth Conditions
2.2. Proteomic Analysis
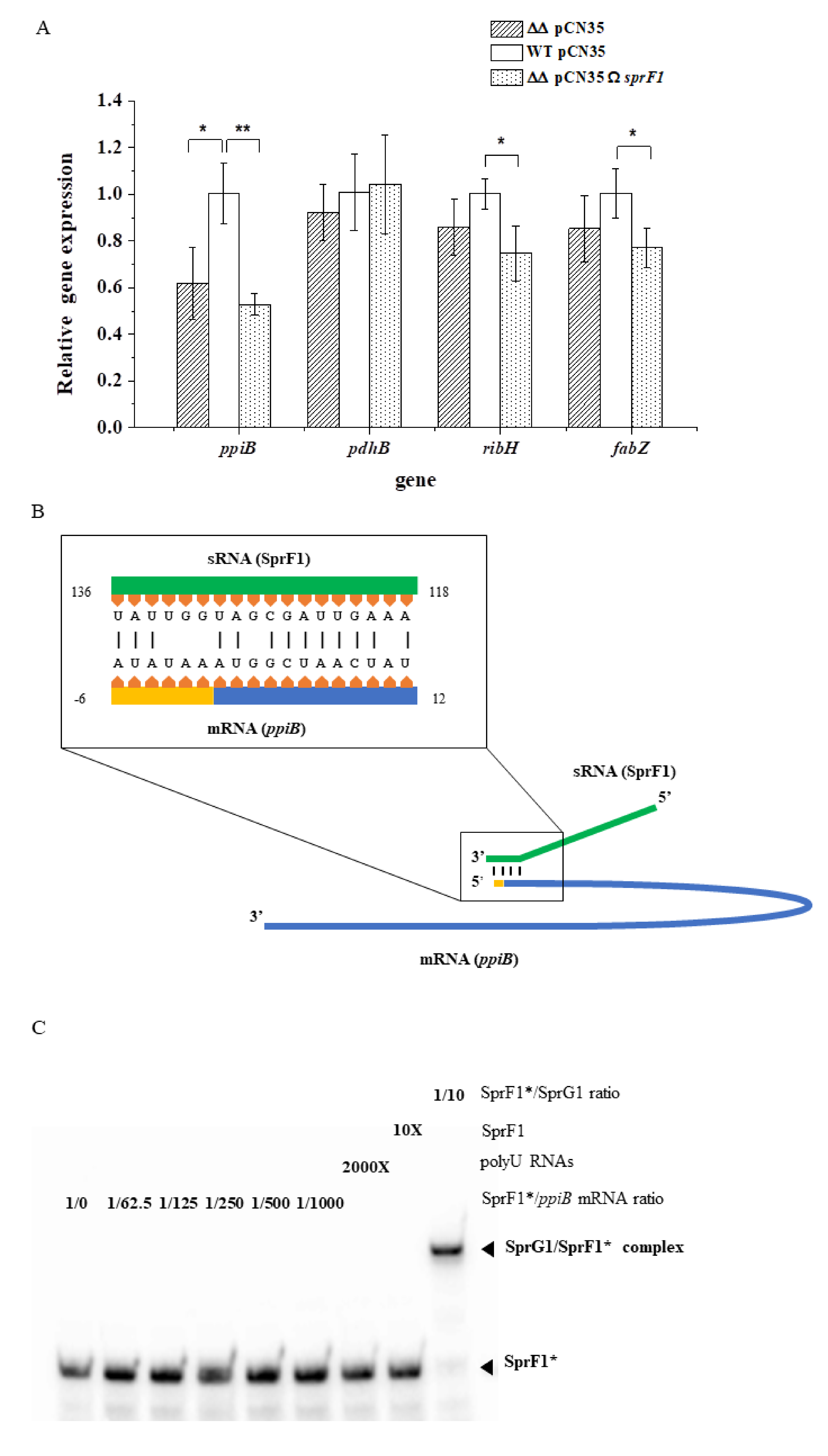
2.3. RNA Extraction, Northern Blots, and RT–qPCR Assays
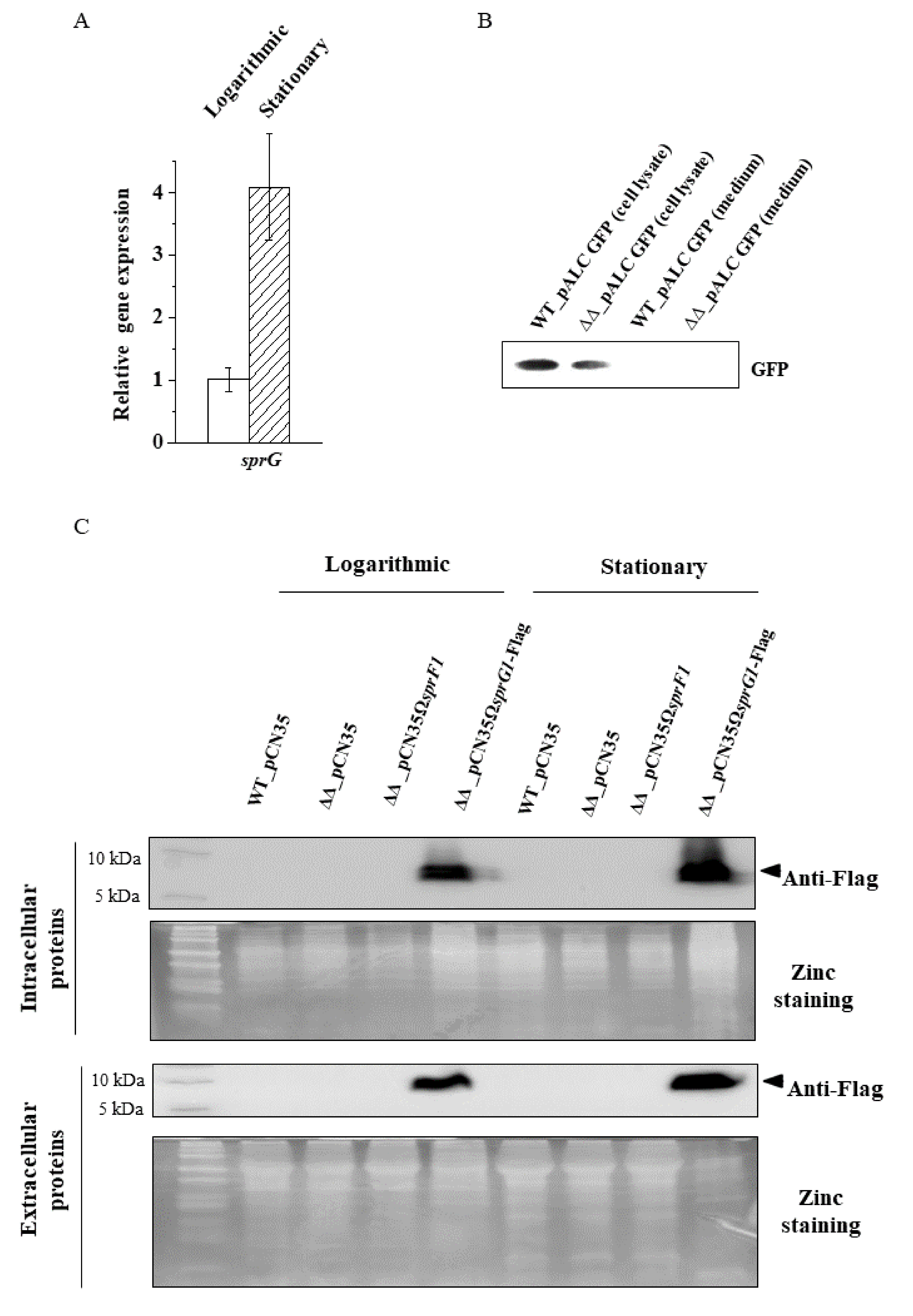
2.4. Protein Extraction and Western Blots
2.5. In Vitro Transcription, RNA Labeling, and Electrophoretic Mobility Shift Assays (EMSA) Assays
2.6. Statistics
3. Results
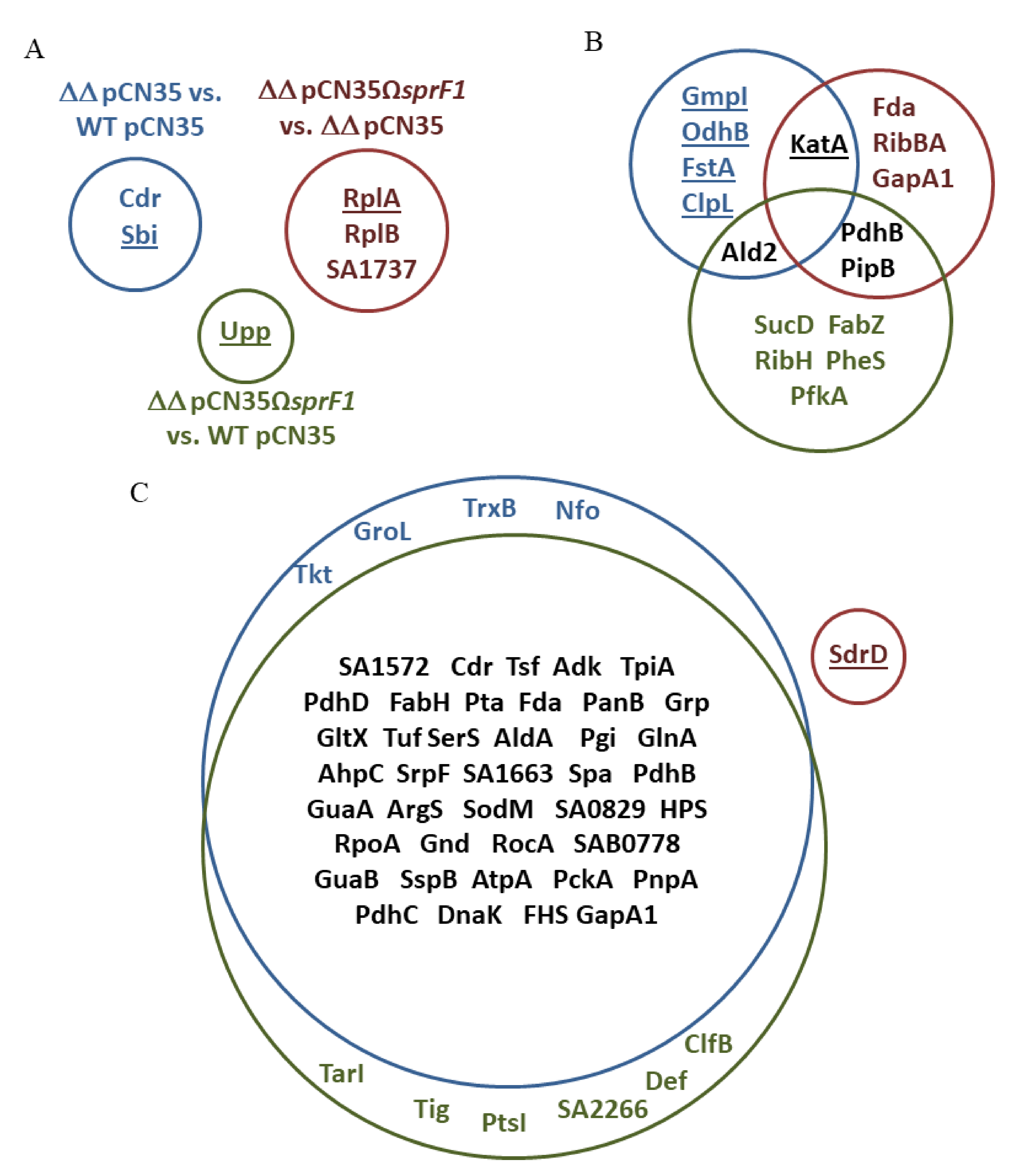
3.1. Deletion of the sprG1/sprF1 TA System Upregulates the Intracellular Proteome Whereas SprF1 Reverses It
3.2. The Lack of the SprG1 Toxin Correlates with Decreased Levels of Extracellular Proteins
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Harms, A.; Brodersen, D.E.; Mitarai, N.; Gerdes, K. Toxins, Targets, and Triggers: An Overview of Toxin-Antitoxin Biology. Mol. Cell 2018, 70, 768–784. [Google Scholar] [CrossRef] [PubMed]
- Helaine, S.; Cheverton, A.M.; Watson, K.G.; Faure, L.M.; Matthews, S.A.; Holden, D.W. Internalization of salmonella by macrophages induces formation of nonreplicating persisters. Science 2014, 343, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Lobato-Márquez, D.; Moreno-Córdoba, I.; Figueroa, V.; Diáz-Orejas, R.; Garciá-Del Portillo, F. Distinct type i and type II toxin-antitoxin modules control Salmonella lifestyle inside eukaryotic cells. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Ayrapetyan, M.; Williams, T.C.; Oliver, J.D. Bridging the gap between viable but non-culturable and antibiotic persistent bacteria. Trends Microbiol. 2015, 23, 7–13. [Google Scholar] [CrossRef]
- Zamakhaev, M.V.; Goncharenko, A.V.; Shumkov, M.S. Toxin-Antitoxin Systems and Bacterial Persistence (Review). Appl. Biochem. Microbiol. 2019, 55, 571–581. [Google Scholar] [CrossRef]
- Song, S.; Wood, T.K. Toxin/Antitoxin System Paradigms: Toxins Bound to Antitoxins Are Not Likely Activated by Preferential Antitoxin Degradation. Adv. Biosyst. 2020, 4, 1900290. [Google Scholar] [CrossRef]
- Page, R.; Peti, W. Toxin-antitoxin systems in bacterial growth arrest and persistence. Nat. Chem. Biol. 2016. [Google Scholar] [CrossRef]
- Brantl, S.; Jahn, N. sRNAs in bacterial type I and type III toxin-antitoxin systems. FEMS Microbiol. Rev. 2015, 39, 413–427. [Google Scholar] [CrossRef]
- Brielle, R.; Pinel-Marie, M.-L.; Felden, B. Linking bacterial type I toxins with their actions. Curr. Opin. Microbiol. 2016, 30, 114–121. [Google Scholar] [CrossRef]
- Maggi, S.; Yabre, K.; Ferrari, A.; Lazzi, C.; Kawano, M.; Rivetti, C.; Folli, C. Functional characterization of the type I toxin Lpt from Lactobacillus rhamnosus by fluorescence and atomic force microscopy. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef]
- Sayed, N.; Nonin-Lecomte, S.; Réty, S.; Felden, B. Functional and structural insights of a staphylococcus aureus apoptotic-like membrane peptide from a toxin-antitoxin module. J. Biol. Chem. 2012, 287, 43454–43463. [Google Scholar] [CrossRef]
- Pinel-Marie, M.L.; Brielle, R.; Felden, B. Dual Toxic-Peptide-Coding Staphylococcus aureus RNA under Antisense Regulation Targets Host Cells and Bacterial Rivals Unequally. Cell Rep. 2014, 7, 424–435. [Google Scholar] [CrossRef]
- Habib, G.; Zhu, Q.; Sun, B. Bioinformatics and functional assessment of toxin-antitoxin systems in staphylococcus aureus. Toxins 2018, 10, 473. [Google Scholar] [CrossRef]
- Germain-Amiot, N.; Augagneur, Y.; Camberlein, E.; Nicolas, I.; Lecureur, V.; Rouillon, A.; Felden, B. A novel Staphylococcus aureus cis-trans type I toxin-antitoxin module with dual effects on bacteria and host cells. Nucleic Acids Res. 2019, 47, 1759–1773. [Google Scholar] [CrossRef]
- Riffaud, C.; Pinel-Marie, M.L.; Pascreau, G.; Felden, B. Functionality and cross-regulation of the four SprG/SprF type I toxin-antitoxin systems in Staphylococcus aureus. Nucleic Acids Res. 2019, 47, 1740–1758. [Google Scholar] [CrossRef]
- Pinel-Marie, M.L.; Brielle, R.; Riffaud, C.; Germain-Amiot, N.; Polacek, N.; Felden, B. RNA antitoxin SprF1 binds ribosomes to attenuate translation and promote persister cell formation in Staphylococcus aureus. Nat. Microbiol. 2021, 6, 209–220. [Google Scholar] [CrossRef]
- Jenul, C.; Horswill, A.R. Regulation of Staphylococcus aureus Virulence. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef]
- Kuroda, M.; Ohta, T.; Uchiyama, I.; Baba, T.; Yuzawa, H.; Kobayashi, I.; Kobayashi, N.; Cui, L.; Oguchi, A.; Aoki, K.I.; et al. Whole genome sequencing of meticillin-resistant Staphylococcus aureus. Lancet 2001, 357, 1225–1240. [Google Scholar] [CrossRef]
- Bonar, E.; Chlebicka, K.; Dubin, G.; Wladyka, B. Application of two-dimensional difference gel electrophoresis in identification of factors responsible for virulence of staphylococcus aureus. In Methods in Molecular Biology; Humana: New York, NY, USA, 2020. [Google Scholar]
- Bodzon-Kulakowska, A.; Suder, P.; Mak, P.; Bierczynska-Krzysik, A.; Lubec, G.; Walczak, B.; Kotlinska, J.; Silberring, J. Proteomic analysis of striatal neuronal cell cultures after morphine administration. J. Sep. Sci. 2009, 32, 1200–1210. [Google Scholar] [CrossRef]
- Perez-Riverol, Y.; Csordas, A.; Bai, J.; Bernal-Llinares, M.; Hewapathirana, S.; Kundu, D.J.; Inuganti, A.; Griss, J.; Mayer, G.; Eisenacher, M.; et al. The PRIDE database and related tools and resources in 2019: Improving support for quantification data. Nucleic Acids Res. 2019, 47, D442–D450. [Google Scholar] [CrossRef]
- Schägger, H.; von Jagow, G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal. Biochem. 1987. [Google Scholar] [CrossRef]
- Bukowski, M.; Lyzen, R.; Helbin, W.M.; Bonar, E.; Szalewska-Palasz, A.; Wegrzyn, G.; Dubin, G.; Dubin, A.; Wladyka, B. A regulatory role for Staphylococcus aureus toxin-antitoxin system PemIK Sa. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef]
- Kery, M.B.; Feldman, M.; Livny, J.; Tjaden, B. TargetRNA2: Identifying targets of small regulatory RNAs in bacteria. Nucleic Acids Res. 2014, 42. [Google Scholar] [CrossRef]
- Lioliou, E.; Romilly, C.; Romby, P.; Fechter, P. RNA-mediated regulation in bacteria: From natural to artificial systems. N. Biotechnol. 2010, 27, 222–235. [Google Scholar] [CrossRef]
- Götz, F.; Yu, W.; Dube, L.; Prax, M.; Ebner, P. Excretion of cytosolic proteins (ECP) in bacteria. Int. J. Med. Microbiol. 2015, 305, 230–237. [Google Scholar] [CrossRef]
- Steinbrecher, T.; Prock, S.; Reichert, J.; Wadhwani, P.; Zimpfer, B.; Bürck, J.; Berditsch, M.; Elstner, M.; Ulrich, A.S. Peptide-lipid interactions of the stress-response peptide tisb that induces bacterial persistence. Biophys. J. 2012, 103, 1460–1469. [Google Scholar] [CrossRef]
- Wilmaerts, D.; Bayoumi, M.; Dewachter, L.; Knapen, W.; Mika, J.T.; Hofkens, J.; Dedecker, P.; Maglia, G.; Verstraeten, N.; Michiels, J. The persistence-inducing toxin hokb forms dynamic pores that cause ATP leakage. MBio 2018, 9. [Google Scholar] [CrossRef]
- Gurnev, P.A.; Ortenberg, R.; Dörr, T.; Lewis, K.; Bezrukov, S.M. Persister-promoting bacterial toxin TisB produces anion-selective pores in planar lipid bilayers. FEBS Lett. 2012, 586, 2529–2534. [Google Scholar] [CrossRef]
- Fraikin, N.; Goormaghtigh, F.; Van Melderen, L. Type II Toxin-Antitoxin Systems: Evolution and Revolutions. J. Bacteriol. 2020, 202. [Google Scholar] [CrossRef]
- Peyrusson, F.; Varet, H.; Nguyen, T.K.; Legendre, R.; Sismeiro, O.; Coppée, J.Y.; Wolz, C.; Tenson, T.; Van Bambeke, F. Intracellular Staphylococcus aureus persisters upon antibiotic exposure. Nat. Commun. 2020, 11. [Google Scholar] [CrossRef]
- Balaban, N.Q.; Helaine, S.; Lewis, K.; Ackermann, M.; Aldridge, B.; Andersson, D.I.; Brynildsen, M.P.; Bumann, D.; Camilli, A.; Collins, J.J.; et al. Definitions and guidelines for research on antibiotic persistence. Nat. Rev. Microbiol. 2019, 17, 441–448. [Google Scholar] [CrossRef]
- Riffaud, C.; Pinel-Marie, M.-L.; Felden, B. Cross-Regulations between Bacterial Toxin–Antitoxin Systems: Evidence of an Interconnected Regulatory Network? Trends Microbiol. 2020, 28, 851–866. [Google Scholar] [CrossRef] [PubMed]
- Alonzo, F.; Xayarath, B.; Whisstock, J.C.; Freitag, N.E. Functional analysis of the Listeria monocytogenes secretion chaperone PrsA2 and its multiple contributions to bacterial virulence. Mol. Microbiol. 2011, 80, 1530–1548. [Google Scholar] [CrossRef]
- Ünal, C.M.; Berges, M.; Smit, N.; Schiene-Fischer, C.; Priebe, C.; Strowig, T.; Jahn, D.; Steinert, M. PrsA2 (CD630_35000) of Clostridioides difficile is an active parvulin-type PPiase and a virulence modulator. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Xia, Y.; Song, X.; Ai, L. Common Non-classically Secreted Bacterial Proteins with Experimental Evidence. Curr. Microbiol. 2016, 72, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Dallo, S.F.; Kannan, T.R.; Blaylock, M.W.; Baseman, J.B. Elongation factor Tu and E1 β subunit of pyruvate dehydrogenase complex act as fibronectin binding proteins in Mycoplasma pneumoniae. Mol. Microbiol. 2002, 46, 1041–1051. [Google Scholar] [CrossRef] [PubMed]
- Wolf, C.; Kusch, H.; Monecke, S.; Albrecht, D.; Holtfreter, S.; von Eiff, C.; Petzl, W.; Rainard, P.; Bröker, B.M.; Engelmann, S. Genomic and proteomic characterization of Staphylococcus aureus mastitis isolates of bovine origin. Proteomics 2011, 11, 2491–2502. [Google Scholar] [CrossRef]
- Stach, N.; Karim, A.; Golik, P.; Kitel, R.; Pustelny, K.; Gruba, N.; Groborz, K.; Jankowska, U.; Kedracka-Krok, S.; Wladyka, B.; et al. Structural Determinants of Substrate Specificity of SplF Protease from Staphylococcus aureus. Int. J. Mol. Sci. 2021, 22, 2220. [Google Scholar] [CrossRef] [PubMed]
- Stach, N.; Kaszycki, P.; Wladyka, B.; Dubin, G. Extracellular Proteases of Staphylococcus spp. In Pet-to-Man Travelling Staphylococci: A World in Progress; Academic Press: Cambridge, MA, USA, 2018; ISBN 9780128135488. [Google Scholar]
- Kolar, S.L.; Antonio Ibarra, J.; Rivera, F.E.; Mootz, J.M.; Davenport, J.E.; Stevens, S.M.; Horswill, A.R.; Shaw, L.N. Extracellular proteases are key mediators of Staphylococcus aureus virulence via the global modulation of virulence-determinant stability. Microbiologyopen 2013, 2, 18–34. [Google Scholar] [CrossRef]
- Mootz, J.M.; Malone, C.L.; Shaw, L.N.; Horswill, A.R. Staphopains modulate staphylococcus aureus biofilm integrity. Infect. Immun. 2013. [Google Scholar] [CrossRef]
- Sonesson, A.; Przybyszewska, K.; Eriksson, S.; Mörgelin, M.; Kjellström, S.; Davies, J.; Potempa, J.; Schmidtchen, A. Identification of bacterial biofilm and the Staphylococcus aureus derived protease, staphopain, on the skin surface of patients with atopic dermatitis. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef]
- Lacey, K.A.; Mulcahy, M.E.; Towell, A.M.; Geoghegan, J.A.; McLoughlin, R.M. Clumping factor B is an important virulence factor during Staphylococcus aureus skin infection and a promising vaccine target. PLoS Pathog. 2019, 15, e1007713. [Google Scholar] [CrossRef]
- Ebner, P.; Rinker, J.; Nguyen, M.T.; Popella, P.; Nega, M.; Luqman, A.; Schittek, B.; Di Marco, M.; Stevanovic, S.; Götza, F. Excreted cytoplasmic proteins contribute to pathogenicity in Staphylococcus aureus. Infect. Immun. 2016, 84, 1672–1681. [Google Scholar] [CrossRef]
- Pasztor, L.; Ziebandt, A.K.; Nega, M.; Schlag, M.; Haase, S.; Franz-Wachtel, M.; Madlung, J.; Nordheim, A.; Heinrichs, D.E.; Götz, F. Staphylococcal major autolysin (Atl) is involved in excretion of cytoplasmic proteins. J. Biol. Chem. 2010, 285, 36794–36803. [Google Scholar] [CrossRef]
- Pichoff, S.; Lutkenhaus, J. Tethering the Z ring to the membrane through a conserved membrane targeting sequence in FtsA. Mol. Microbiol. 2005, 55, 1722–1734. [Google Scholar] [CrossRef]
- Masuda, H.; Tan, Q.; Awano, N.; Wu, K.P.; Inouye, M. YeeU enhances the bundling of cytoskeletal polymers of MreB and FtsZ, antagonizing the CbtA (YeeV) toxicity in Escherichia coli. Mol. Microbiol. 2012, 84, 979–989. [Google Scholar] [CrossRef]
- Hemmadi, V.; Biswas, M. An overview of moonlighting proteins in Staphylococcus aureus infection. Arch. Microbiol. 2021, 203, 481–498. [Google Scholar] [CrossRef]
| Name | Sequence (5′-3′) | Application |
|---|---|---|
| SprF1-NB | TAACTTTGGCTGGTTTCGATGGTT | Northern blot |
| SprG1-NB | ATGCCACCATAGGCACCACCTCCTT | Northern blot |
| 5S rRNA-NB | CGTAAGTTCGACTACCATCG | Northern blot |
| SprG1-F | AGTATACAAGCAGTAAAAAAAGTATATGTG | RT-qPCR |
| SprG1-R | ATTTCAGTAATGCCACCATAGGCA | RT-qPCR |
| gyrB-F | CAACAATGAACCCTGAGCACC | RT-qPCR |
| gyrB-R | CGGTTTTCTACAACGTCACCC | RT-qPCR |
| ribH-F | GTCGCGAAAGGTGTTTCTAAAGTA | RT-qPCR |
| ribH-R | CCAGCTTTCGTACCTGCTCT | RT-qPCR |
| fabZ-F | AACGTCAAGTAGTACCTGGTGATA | RT-qPCR |
| fabZ-R | CAAGCAAGTTGACCATCGACAG | RT-qPCR |
| ppiB-F | CATTGTTCAAATGAAAGAAGTACCTCA | RT-qPCR |
| ppiB-R | GTGTACCACCCTTTTCGCCATA | RT-qPCR |
| pdhB-F | GCTGAATCAGGTATTGGTGGTTTA | RT-qPCR |
| pdhB-R | TGTCCAGCAATCGCATCAAATACTT | RT-qPCR |
| SprF1-T7 | TAATACGACTCACTATAGGGATATATAGAAAAAGGGCAAC | In vitro transcription |
| SprF1-rev | AAAAAATAACCATCGCTAACTTTGGCT | In vitro transcription |
| ppiB-T7 | TAATACGACTCACTATAGGGTTTCCTCCCTTAAAAGTATGTTAATA | In vitro transcription |
| ppiB-rev | ATAACCACTTTAATTTCACCTTGTT | In vitro transcription |
| No. | Protein (Acronym) | Accession Number | N315-ΔΔ vs. N315 | N315-ΔΔSprF1 vs. N315 | N315-ΔΔSprF1 vs. N315-ΔΔ |
|---|---|---|---|---|---|
| Logarithmic phase | |||||
| 1 | Coenzyme A disulfide reductase (Cdr) | Q7A6H1 | 1.68 ↓ * | ||
| 2 | Immunoglobulin-binding protein (Sbi) | Q99RL2 | 1.72 ↑ | ||
| 3 | 50S ribosomal protein L1 (RplA) | Q99W68 | 1.82 ↑ | ||
| 4 | 50S ribosomal protein L2 (RplB) | P60432 | 1.51 ↓ | ||
| 5 | Uncharacterized protein (SA1737) | Q7A4P4 | 1.50 ↓ | ||
| 6 | Uracil phosphoribosyltransferase (Upp) | P67396 | 2.19 ↑ | ||
| Stationary phase | |||||
| 1 | Pyruvate dehydrogenase E1 component subunit β (PdhB) | P99063 | 2.19 ↓ | 1.82 ↓ | |
| 2 | Succinate--CoA ligase (ADP-forming) subunit α (SucD) | P99070 | 1.55 ↓ | ||
| 3 | Putative peptidyl-prolyl cis-trans isomerase (PpiB) | Q7A6I1 | 2.19 ↓ | 1.82 ↓ | |
| 4 | 3-hydroxyacyl-(acyl-carrier-protein) dehydrataseFabZ (FabZ) | P64108 | 2.36 ↓ | ||
| 5 | 6,7-dimethyl-8-ribityllumazine synthase (RibH) | P99141 | 1.58 ↓ | 2.36 ↓ | |
| 6 | Alanine dehydrogenase 2 (Ald2) | Q99TF4 | 1.86 ↓ | ||
| 7 | Phenylalanine--tRNA ligase α subunit (PheS) | P68848 | 1.55 ↓ | ||
| 8 | ATP-dependent 6-phosphofructokinase (PfkA) | P99165 | 1.55 ↓ | ||
| 9 | 2,3-bisphosphoglycerate-independent phosphoglycerate mutase (GpmI) | P64270 | 1.53 ↑ | ||
| 10 | Dihydrolipoyllysine-residue succinyltransferase component of 2-oxoglutarate dehydrogenase complex (OdhB) | Q7A5N4 | 1.53 ↑ | ||
| 11 | Catalase (KatA) | Q7A5T2 | 1.66 ↑ | 1.90 ↑ | |
| 12 | Cell division protein FtsA (FtsA) | P63765 | 1.53 ↑ | ||
| 13 | ATP-dependent Clp protease ATP-binding subunit ClpL (ClpL) | Q7A3F4 | 1.53 ↑ | ||
| 14 | Fructose-bisphosphate aldolase class 1 (Fda) | P99117 | 1.54 ↓ | ||
| 15 | Riboflavin biosynthesis protein RibBA (RibBA) | Q7A511 | 1.62 ↓ | ||
| 16 | Glyceraldehyde-3-phosphate dehydrogenase 1 (GapA1) | P99136 | 1.62 ↓ | ||
| No. | Protein (Acronym) | Accession Number | N315-ΔΔ vs. N315 | N315-ΔΔSprF1 vs. N315 | N315-ΔΔSprF1 vs. N315-ΔΔ |
|---|---|---|---|---|---|
| 1 | Glutamate----tRNA ligase (GltX) | P99170 | 1.74 ↓ | 1.72 ↓ | |
| 2 | 1-pyrroline-5-carboxylate dehydrogenase (RocA) | P99076 | 1.96 ↓ | 2.18 ↓ | |
| 3 | Glutamine synthetase (GlnA) | P99095 | 1.77 ↓ | 2.12 ↓ | |
| 4 | GMP synthase [glutamine-hydrolyzing] (GuaA) | P99105 | 1.87 ↓ | 1.80 ↓ | |
| 5 | Arginine--tRNA ligase (ArgS) | Q99W05 | 1.87 ↓ | 1.80 ↓ | |
| 6 | Transketolase (Tkt) | P99161 | 1.67 ↓ | ||
| 7 | 6-phosphogluconate dehydrogenase, decarboxylating (Gnd) | P63334 | 1.94 ↓ | 1.85 ↓ | |
| 8 | Phosphoenolpyruvate carboxykinase (ATP) (PckA) | P99128 | 2.57 ↓ | 2.42 ↓ | |
| 9 | Glucose-6-phosphate isomerase (Pgi) | P99078 | 1.77 ↓ | 1.86 ↓ | |
| 10 | Dihydrolipoyllysine-residue acetyltransferase component of pyruvate dehydrogenase complex (PdhC) | P65636 | 2.71 ↓ | 2.00 ↓ | |
| 11 | Pyruvate dehydrogenase E1 component subunit β (PdhB) | P99063 | 1.84 ↓ | 2.23 ↓ | |
| 12 | Dihydrolipoyl dehydrogenase (PdhD) | P99084 | 1.66 ↓ | 1.60 ↓ | |
| 13 | Fructose-bisphosphate aldolase class 1 (Fda) | P99117 | 1.68 ↓ | 1.56 ↓ | |
| 14 | Triosephosphate isomerase (TpiA) | P99133 | 1.59 ↓ | 1.92 ↓ | |
| 15 | Glyceraldehyde-3-phosphate dehydrogenase 1 (GapA1) | P99136 | 1.5 ↓ | ||
| 16 | 3-hexulose-6-phosphate synthase (HPS) | Q7A774 | 1.92 ↓ | 2.11 ↓ | |
| 17 | Formate--tetrahydrofolate ligase (FHS) | Q7A535 | 3.41 ↓ | 2.75 ↓ | |
| 18 | Phosphoenolpyruvate-protein phosphotransferase (PtsI) | Q99V14 | 1.63 ↓ | ||
| 19 | Elongation factor Ts (Tsf) | P99171 | 1.59 ↓ | 1.54 ↓ | |
| 20 | Elongation factor Tu (Tuf) | P99152 | 1.74 ↓ | 1.93 ↓ | |
| 21 | Serine--tRNA ligase (SerS) | P99178 | 1.74 ↓ | 2.12 ↓ | |
| 22 | Chaperone protein DnaK (DnaK) | P99110 | 1.82 ↓ | 1.64 ↓ | |
| 23 | Protein GrpE (GrpE) | P99086 | 1.73 ↓ | 1.50 ↓ | |
| 24 | Trigger factor (Tig) | P99080 | 1.54 ↓ | ||
| 25 | 60 kDa chaperonin (GroL) | P99083 | 1.51 ↓ | ||
| 26 | Thioredoxin reductase (TrxB) | Q6GIM7 | 2.02 ↓ | ||
| 27 | Alkyl hydroperoxide reductase C (AhpC) | P99074 | 1.82 ↓ | 1.82 ↓ | |
| 28 | Superoxide dismutase [Mn/Fe] 2 (SodM) | P66831 | 1.90 ↓ | 1.73 ↓ | |
| 29 | Coenzyme A disulfide reductase (Cdr) | Q7A6H1 | 1.54 ↓ | 1.60 ↓ | |
| 30 | Glutamyl endopeptidase (SspA) | Q7A6A6 | 1.80 ↓ | ||
| 31 | Immunoglobulin G-binding protein A (SpA) | P99134 | 1.83 ↓ | 1.84 ↓ | |
| 32 | Staphopain B (SspB) | Q7A6A7 | 2.27 ↓ | 1.81 ↓ | |
| 33 | Clumping factor B (ClfB) | Q7A382 | 1.63 ↓ | ||
| 34 | ATP synthase subunit α (AtpA) | P99111 | 2.42 ↓ | 2.41 ↓ | |
| 35 | Adenylate kinase (Adk) | P99062 | 1.59 ↓ | 1.59 ↓ | |
| 36 | Inosine-5’-monophosphate dehydrogenase (GuaB) | P99106 | 2.06 ↓ | 1.95 ↓ | |
| 37 | Polyribonucleotide nucleotidyltransferase (PnpA) | Q7A5 × 7 | 2.71 ↓ | 1.99 ↓ | |
| 38 | Probable endonuclease 4 (Nfo) | P63538 | 1.65 ↓ | ||
| 39 | DNA-directed RNA polymerase subunit α (RpoA) | P66706 | 1.93 ↓ | 1.81 ↓ | |
| 40 | Peptide deformylase (Def) | P99077 | 1.82 ↓ | ||
| 41 | Ribitol-5-phosphate cytidylyltransferase 1 (TarI) | Q7A7V0 | 2.01 ↓ | ||
| 42 | 3-oxoacyl-(acyl-carrier-protein) synthase 3 (FabH) | P99159 | 1.66 ↓ | 1.68 ↓ | |
| 43 | Phosphate acetyltransferase (Pta) | P99092 | 1.66 ↓ | 1.96 ↓ | |
| 44 | UPF0051 protein (SAB0778) | Q7A6L4 | 1.97 ↓ | 1.96 ↓ | |
| 45 | 3-methyl-2-oxobutanoate hydroxymethyltransferase (PanB) | P65656 | 1.71 ↓ | 1.89 ↓ | |
| 46 | DUF4242 domain-containing protein (SA0165) | A0A0H3JSJ2 | 1.58 ↓ | ||
| 47 | Putative aldehyde dehydrogenase (AldA) | Q7A825 | 1.74 ↓ | 2.12 ↓ | |
| 48 | Putative dipeptidase (SA1572) | Q7A522 | 1.51 ↓ | 2.32 ↓ | |
| 49 | Uncharacterized oxidoreductase (SA2266) | Q7A3L9 | 2.52 ↓ | ||
| 50 | Uncharacterized protein (SA0829) | Q7A6H3 | 1.90 ↓ | 1.79 ↓ | |
| 51 | UPF0342 protein (SA1663) | Q7A4V3 | 1.82 ↓ | 2.17 ↓ | |
| 52 | Serine-aspartate repeat-containing protein D (SdrD) | Q7A780 | 1.58 ↑ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chlebicka, K.; Bonar, E.; Suder, P.; Ostyn, E.; Felden, B.; Wladyka, B.; Pinel-Marie, M.-L. Impacts of the Type I Toxin–Antitoxin System, SprG1/SprF1, on Staphylococcus aureus Gene Expression. Genes 2021, 12, 770. https://doi.org/10.3390/genes12050770
Chlebicka K, Bonar E, Suder P, Ostyn E, Felden B, Wladyka B, Pinel-Marie M-L. Impacts of the Type I Toxin–Antitoxin System, SprG1/SprF1, on Staphylococcus aureus Gene Expression. Genes. 2021; 12(5):770. https://doi.org/10.3390/genes12050770
Chicago/Turabian StyleChlebicka, Kinga, Emilia Bonar, Piotr Suder, Emeline Ostyn, Brice Felden, Benedykt Wladyka, and Marie-Laure Pinel-Marie. 2021. "Impacts of the Type I Toxin–Antitoxin System, SprG1/SprF1, on Staphylococcus aureus Gene Expression" Genes 12, no. 5: 770. https://doi.org/10.3390/genes12050770
APA StyleChlebicka, K., Bonar, E., Suder, P., Ostyn, E., Felden, B., Wladyka, B., & Pinel-Marie, M.-L. (2021). Impacts of the Type I Toxin–Antitoxin System, SprG1/SprF1, on Staphylococcus aureus Gene Expression. Genes, 12(5), 770. https://doi.org/10.3390/genes12050770







