Investigating Olfactory Gene Variation and Odour Identification in Older Adults
Abstract
1. Introduction
2. Materials and Methods
2.1. Participants
2.2. Olfaction
2.3. Lifestyle and Health Variables
2.4. Selection of Olfactory Receptor Genes and Single Nucleotide Polymorphisms (SNPs)
2.5. Genotyping and Calculation of Polygenic Risk Scores (PRS)
2.6. Statistical Analyses
3. Results
3.1. Sample Characteristics
3.2. BSIT Item Identification
3.3. Potential Covariates Influencing Olfaction
3.4. Genetic Analyses
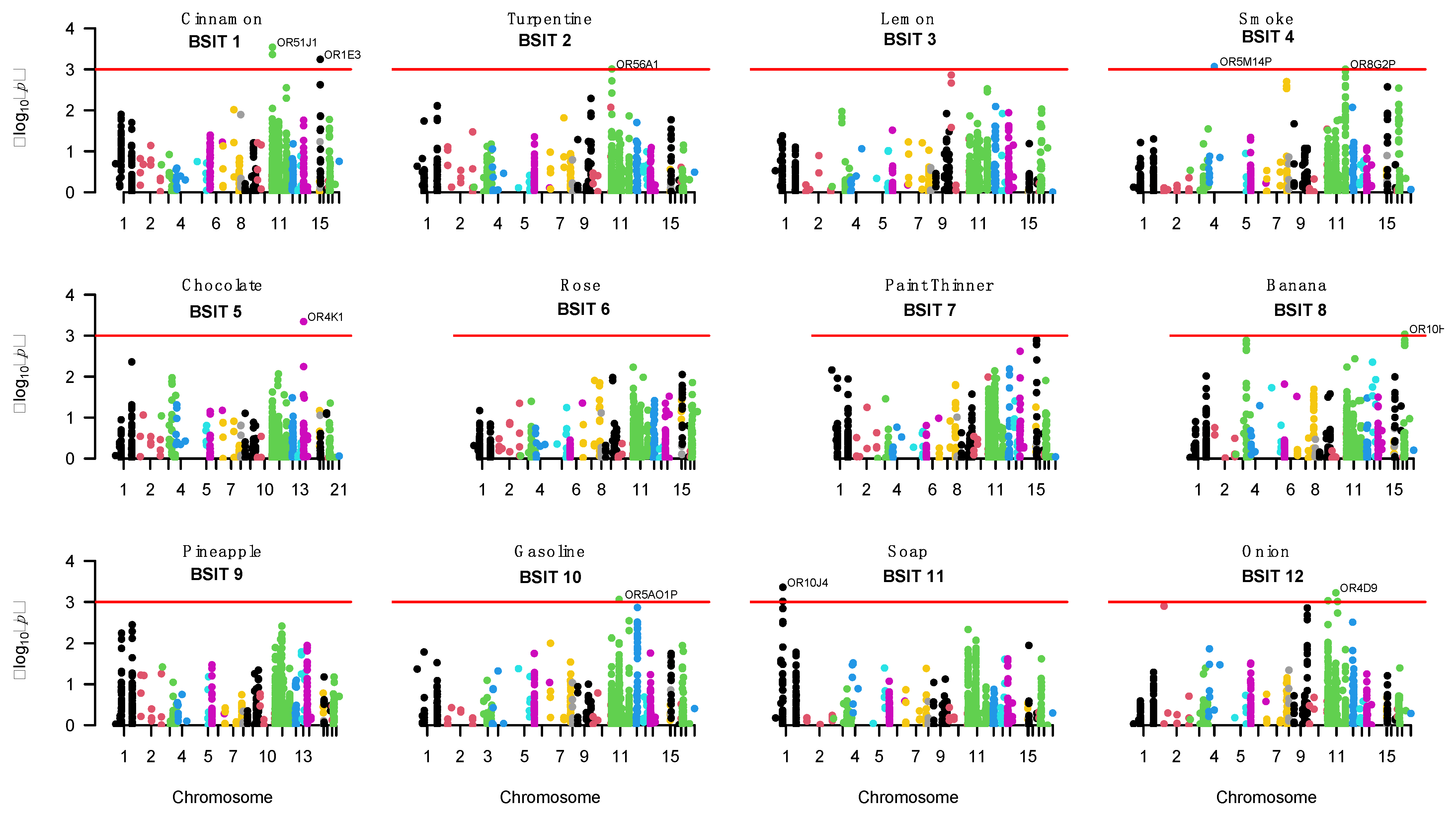
3.4.1. OR-Related Genes and Single SNP Analysis
3.4.2. Replication SNP Analysis
3.5. Polygenic Risk Scores
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dintica, C.S.; Marseglia, A.; Rizzuto, D.; Wang, R.; Seubert, J.; Arfanakis, K.; Bennett, D.A.; Xu, W. Impaired olfaction is associated with cognitive decline and neurodegeneration in the Brain. Neurology 2019, 92, e700–e709. [Google Scholar] [CrossRef]
- Doty, R.L.; Petersen, I.; Mensah, N.; Christensen, K. Genetic and environmental influences on odor identification ability in the very old. Psychol. Aging 2011, 26, 864–871. [Google Scholar] [CrossRef]
- Murphy, C.; Schubert, C.R.; Cruickshanks, K.J.; Klein, B.E.; Klein, R.; Nondahl, D.M. Prevalence of olfactory impairment in older adults. JAMA 2002, 288, 2307–2312. [Google Scholar] [CrossRef] [PubMed]
- Heinrich, J.; Vidal, J.S.; Simon, A.; Rigaud, A.S.; Hanon, O.; Epelbaum, J.; Viollet, C.; Duron, E. Relationships between lower olfaction and brain white matter lesions in elderly subjects with mild cognitive impairment. J. Alzheimers Dis. 2018, 61, 1133–1141. [Google Scholar] [CrossRef]
- Liu, B.; Luo, Z.; Pinto, J.M.; Shiroma, E.J.; Tranah, G.J.; Wirdefeldt, K.; Fang, F.; Harris, T.B.; Chen, H. Relationship between poor olfaction and mortality among community-dwelling older adults: A cohort study. Ann. Intern. Med. 2019, 170, 673–681. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, K.P.; Kennedy, R.C.; Baron, G.; Greenberg, R.A. Number of mitral cells and the bulb volume in the aging human olfactory bulb: A quantitative morphological study. Anat. Rec. 1987, 218, 73–87. [Google Scholar] [CrossRef] [PubMed]
- Doty, R.L.; Kamath, V. The influences of age on olfaction: A review. Front. Psychol. 2014, 5, 20. [Google Scholar] [CrossRef]
- Bathini, P.; Brai, E.; Auber, L.A. Olfactory dysfunction in the pathophysiological continuum of dementia. Ageing Res. Rev. 2019, 55, 100956. [Google Scholar] [CrossRef]
- Hawkes, C.; Doty, R.L. The Neurology of Olfaction; Cambridge University Press: Cambridge, UK, 2009. [Google Scholar]
- Segal, N.L.; Topolski, T.D.; Wilson, S.M.; Brown, K.W.; Araki, L. Twin analysis of odor identification and perception. Physiol. Behav. 1995, 57, 605–609. [Google Scholar] [CrossRef]
- Finkel, D.; Pedersen, N.L.; Larsson, M. Olfactory functioning and cognitive abilities: A twin study. J. Gerontol. B Psychol. Sci. Soc. Sci. 2001, 56, P226–P233. [Google Scholar] [CrossRef]
- Knaapila, A.; Keskitalo, K.; Kallela, M.; Wessman, M.; Sammalisto, S.; Hiekkalinna, T.; Palotie, A.; Peltonen, L.; Tuorila, H.; Perola, M. Genetic component of identification, intensity and pleasantness of odours: A finnish family study. Eur. J. Hum. Genet. 2007, 15, 596–602. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Yang, J.; Tranah, G.; Franceschini, N.; Parimi, N.; Alkorta-Aranburu, G.; Xu, Z.; Alonso, A.; Cummings, S.R.; Fornage, M.; et al. Genome-wide meta-analysis on the sense of smell among us older adults. Medicine 2015, 94, e1892. [Google Scholar] [CrossRef]
- Dong, J.; Wyss, A.; Yang, J.; Price, T.R.; Nicolas, A.; Nalls, M.; Tranah, G.; Franceschini, N.; Xu, Z.; Schulte, C.; et al. Genome-wide association analysis of the sense of smell in u.s. older adults: Identification of novel risk loci in African-Americans and European-Americans. Mol. Neurobiol. 2017, 54, 8021–8032. [Google Scholar] [CrossRef]
- Trimmer, C.; Keller, A.; Murphy, N.R.; Snyder, L.L.; Willer, J.R.; Nagai, M.H.; Katsanis, N.; Vosshall, L.B.; Matsunami, H.; Mainland, J.D. Genetic variation across the human olfactory receptor repertoire alters odor perception. Proc. Natl. Acad. Sci. USA 2019, 116, 9475–9480. [Google Scholar] [CrossRef] [PubMed]
- Menashe, I.; Man, O.; Lancet, D.; Gilad, Y. Different noses for different people. Nat. Genet. 2003, 34, 143–144. [Google Scholar] [CrossRef]
- Malnic, B.; Godfrey, P.A.; Buck, L.B. The human olfactory receptor gene family. Proc. Natl. Acad. Sci. USA 2004, 101, 2584–2589. [Google Scholar] [CrossRef]
- Sachdev, P.S.; Brodaty, H.; Reppermund, S.; Kochan, N.A.; Trollor, J.N.; Draper, B.; Slavin, M.J.; Crawford, J.; Kang, K.; Broe, G.A.; et al. The sydney memory and ageing study (mas): Methodology and baseline medical and neuropsychiatric characteristics of an elderly epidemiological non-demented cohort of australians aged 70–90 years. Int. Psychogeriatr. 2010, 22, 1248–1264. [Google Scholar] [CrossRef] [PubMed]
- Sachdev, P.S.; Lammel, A.; Trollor, J.N.; Lee, T.; Wright, M.J.; Ames, D.; Wen, W.; Martin, N.G.; Brodaty, H.; Schofield, P.R. A comprehensive neuropsychiatric study of elderly twins: The older australian twins study. Twin Res. Hum. Genet. 2009, 12, 573–582. [Google Scholar] [CrossRef]
- Menon, C.; Westervelt, H.J.; Jahn, D.R.; Dressel, J.A.; O’Bryant, S.E. Normative performance on the brief smell identification test (bsit) in a multi-ethnic bilingual cohort: A project frontier study. Clin. Neuropsychol. 2013, 27, 946–961. [Google Scholar] [CrossRef]
- Doty, R.L.; Marcus, A.; Lee, W.W. Development of the 12-item cross-cultural smell identification test (cc-sit). Laryngoscope 1996, 1, 353–356. [Google Scholar] [CrossRef]
- Wilson, R.S.; Schneider, J.A.; Arnold, S.E.; Tang, Y.; Boyle, P.A.; Bennett, D.A. Olfactory identification and incidence of mild cognitive impairment in older age. Arch. Gen. Psychiatry 2007, 64, 802–808. [Google Scholar] [CrossRef] [PubMed]
- Crasto, C.; Marenco, L.; Miller, P.; Shepherd, G. Olfactory receptor database: A metadata-driven automated population from sources of gene and protein sequences. Nucleic Acids Res. 2002, 30, 354–360. [Google Scholar] [CrossRef] [PubMed]
- Nyholt, D.R. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am. J. Hum. Genet. 2004, 74, 765–769. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, S.R.; McRae, J.F.; Bava, C.M.; Beresford, M.K.; Hunter, D.; Jia, Y.; Chheang, S.L.; Jin, D.; Peng, M.; Gamble, J.C.; et al. A mendelian trait for olfactory sensitivity affects odor experience and food selection. Curr. Biol. 2013, 23, 1601–1605. [Google Scholar] [CrossRef]
- Jaeger, S.R.; McRae, J.F.; Salzman, Y.; Williams, L.; Newcomb, R.D. A preliminary investigation into a genetic basis for cis-3-hexen-1-ol odour perception: A genome-wide association approach. Food Qual. Prefer. 2010, 21, 121–131. [Google Scholar] [CrossRef]
- Keller, A.; Zhuang, H.; Chi, Q.; Vosshall, L.B.; Matsunami, H. Genetic variation in a human odorant receptor alters odour perception. Nature 2007, 449, 468–472. [Google Scholar] [CrossRef]
- Poljak, A.; Crawford, J.D.; Smythe, G.A.; Brodaty, H.; Slavin, M.J.; Kochan, N.A.; Trollor, J.N.; Wen, W.; Mather, K.A.; Assareh, A.A.; et al. The relationship between plasma abeta levels, cognitive function and brain volumetrics: Sydney memory and ageing study. Curr. Alzheimer Res. 2016, 13, 243–255. [Google Scholar] [CrossRef] [PubMed]
- Price, A.L.; Patterson, N.J.; Plenge, R.M.; Weinblatt, M.E.; Shadick, N.A.; Reich, D. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 2006, 38, 904–909. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Forer, L.; Schonherr, S.; Sidore, C.; Locke, A.E.; Kwong, A.; Vrieze, S.I.; Chew, E.Y.; Levy, S.; McGue, M.; et al. Next-generation genotype imputation service and methods. Nat. Genet. 2016, 48, 1284–1287. [Google Scholar] [CrossRef]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-generation plink: Rising to the challenge of larger and richer datasets. Gigascience 2015, 4, 7. [Google Scholar] [CrossRef]
- Euesden, J.; Lewis, C.M.; O’Reilly, P.F. Prsice: Polygenic risk score software. Bioinformatics 2015, 31, 1466–1468. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Wang, C.; Conomos, M.P.; Stilp, A.M.; Li, Z.; Sofer, T.; Szpiro, A.A.; Chen, W.; Brehm, J.M.; Celedon, J.C.; et al. Control for population structure and relatedness for binary traits in genetic association studies via logistic mixed models. Am. J. Hum. Genet. 2016, 98, 653–666. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Huang, J.; Liu, Y.; Ma, L.; Potash, J.B.; Han, S. Combat: A combined association test for genes using summary statistics. Genetics 2017, 207, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Gogarten, S.M.; Bhangale, T.; Conomos, M.P.; Laurie, C.A.; McHugh, C.P.; Painter, I.; Zheng, X.; Crosslin, D.R.; Levine, D.; Lumley, T.; et al. Gwastools: An r/bioconductor package for quality control and analysis of genome-wide association studies. Bioinformatics 2012, 28, 3329–3331. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Olender, T.; Waszak, S.M.; Viavant, M.; Khen, M.; Ben-Asher, E.; Reyes, A.; Nativ, N.; Wysocki, C.J.; Ge, D.; Lancet, D. Personal receptor repertoires: Olfaction as a model. BMC Genomics 2012, 13, 414. [Google Scholar] [CrossRef] [PubMed]
- Glusman, G.; Yanai, I.; Rubin, I.; Lancet, D. The complete human olfactory subgenome. Genome Res. 2001, 11, 685–702. [Google Scholar] [CrossRef]
| Characteristic | Sydney MAS | OATS | Total |
|---|---|---|---|
| N | 881 | 514 | 1395 |
| Age at assessment (years), mean ± SD | 78.7 ± 4.8 | 70.7 ± 5.5 | 75.5 ± 6.5 |
| Men n, (%) | 393 (44.4) | 207 (35.1) | 600 (41.0) |
| Brief Smell identification Test (BSIT) score, mean ± SD | 9.3 ± 2.2 | 9.7 ± 1.7 | 9.4 ± 2.0 |
| SMOKING STATUS n, (%) | |||
| Past/Never Smoked | 840 (96.4) | 558 (94.3) | 1398 (95.5) |
| Current Smoker | 31 (3.6) | 34 (5.7) | 65 (4.4) |
| BSIT Item | Odour | Sydney MAS n, (%) | OATS n, (%) | Total n, (%) |
|---|---|---|---|---|
| 1 | Cinnamon | 767, (87.1) | 455, (88.6) | 1222, (87.6) |
| 2 | Turpentine | 195, (22.1) | 121, (23.5) | 316, (22.7) |
| 3 | Lemon | 540, (61.3) | 374, (72.8) | 914, (65.5) |
| 4 | Smoke | 658, (74.7) | 413, (80.4) | 1071, (76.8) |
| 5 | Chocolate | 749, (85.0) | 381, (74.1) | 1130, (81.6) |
| 6 | Rose | 629, (71.4) | 408, (79.4) | 1037, (74.3) |
| 7 | Paint Thinner | 785, (89.1) | 478, (93.0) | 1263, (90.5) |
| 8 | Banana | 700, (79.5) | 421, (81.9) | 1121, (80.4) |
| 9 | Pineapple | 743, (84.4) | 460, (89.5) | 1203, (86.2) |
| 10 | Gasoline | 787, (89.3) | 487, (94.8) | 1274, (91.3) |
| 11 | Soap | 800, (90.1) | 483, (94.0) | 1283, (92.0) |
| 12 | Onion | 785, (89.1) | 488, (95.0) | 1273, (91.3) |
| BSIT Item 1 | Odour | Chromosome | Gene 2 | Gene Based p-Value | Sentinel SNP | Sentinel SNP Base Position 3 | Sentinel SNP p-Value | SNP Function/Position |
|---|---|---|---|---|---|---|---|---|
| 1 | Cinnamon | 11 | OR51J1 | 2.90 × 10−4 | rs6578634 | 5420909 | 4.03 × 10−5 | Intronic |
| 11 | OR51Q1 | 4.35 × 10−4 | rs61894107 | 5442376 | 3.24 × 10−5 | Intronic | ||
| 17 | OR1E3 | 5.73 × 10−4 | rs769433 | 3019790 | 8.12 × 10−5 | 98 bp upstream variant | ||
| 2 | Turpentine | 11 | OR56A1 | 9.81 × 10−4 | rs117555183 | 6049351 | 5.32 × 10−5 | 380 bp upstream variant |
| 4 | OR5M14P | 8.59 × 10−4 | rs1985012 | 41719745 | 1.09 × 10−4 | Intergenic | ||
| 4 | Smoke | 11 | OR8G2P | 9.92 × 10−4 | rs10893174 | 124090872 | 5.38 × 10−4 | Intergenic |
| 11 | OR8B2 | 9.99 × 10−4 | rs530704 | 124247361 | 1.57 × 10−4 | Exonic (synonymous) | ||
| 5 | Chocolate | 14 | OR4K1 | 4.57 × 10−4 | rs9323231 | 20408911 | 1.99 × 10−3 | Intergenic |
| 8 | Banana | 19 | OR10H3 | 9.22 × 10−4 | rs79876008 | 15851350 | 1.38 × 10−4 | 853 bp upstream variant |
| 10 | Gasoline | 11 | OR5AO1P | 8.74 × 10−4 | rs11228886 | 56815339 | 4.86 × 10−5 | Intergenic |
| 11 | Soap | 1 | OR10J4 | 4.41 × 10−4 | rs78689883 | 159406445 | 1.99 × 10−4 | Intergenic |
| 1 | OR10J1 | 9.66 × 10−4 | rs4128726 | 159405844 | 2.08 × 10−4 | Intronic | ||
| 12 | Onion | 11 | OR52D1 | 9.30 × 10−4 | rs4638331 | 5511431 | 6.81 × 10−5 | Intronic |
| 11 | OR4D9 | 6.01 × 10−4 | rs116937381 | 59281930 | 5.72 × 10−5 | 456 bp upstream variant | ||
| 11 | OR7E4P | 9.80 × 10−4 | rs61128173 | 71334302 | 3.74 × 10−5 | Intergenic |
| Chromosome | SNP 1 | SNP BP | β ± S.E. | p-Value | Association with BSIT Variable (p-Value < 0.05) |
|---|---|---|---|---|---|
| 4 | rs72679931a | 162245253 | 0.045 ± 0.023 | 5.04 × 10−2 | BSIT Total |
| 0.490 ± 0.200 | 1.41 × 10−2 | Rose (item 6), | |||
| 0.659 ± 0.275 | 1.65 × 10−2 | Paint thinner (item 7) | |||
| 0.644 ± 0.285 | 2.35 × 10−2 | Gasoline (item 10) | |||
| 8 | rs34276508b | 1328679 | 0.008 ± 0.028 | 7.77 × 10−1 | BSIT Total |
| 0.550 ± 0.221 | 1.30 × 10−2 | Lemon (item 3) | |||
| 8 | rs2730141 a | 40282221 | 0.013 ± 0.012 | 3.02 × 10−1 | BSIT Total |
| 0.375 ± 0.177 | 3.43 × 10−2 | Onion (item 12) | |||
| 9 | rs4442206 a | 73747369 | −0.002 ± 0.010 | 8.35 × 10−1 | BSIT Total |
| −0.207 ± 0.099 | 3.64 × 10−2 | Turpentine (item 2) | |||
| 9 | rs6560178 a | 73748538 | −0.002 ± 0.010 | 8.50 × 10−1 | BSIT Total |
| −0.207 ± 0.099 | 3.55 × 10−2 | Turpentine (item 2) | |||
| 9 | rs2251885 b | 103793544 | −0.002 ± 0.010 | 8.52 × 10−1 | BSIT Total |
| −0.253 ± 0.124 | 4.05 × 10−2 | Pineapple (item 9), | |||
| −0.394 ± 0.159 | 1.29 × 10−2 | Soap (item 11) | |||
| 9 | rs193020892 a | 128840966 | 0.051 ± 0.039 | 1.91 × 10−1 | BSIT Total |
| 1.083 ± 0.445 | 1.49 × 10−2 | Gasoline (item 10) | |||
| 11 | rs7938698 c | 19709674 | 0.025 ± 0.016 | 1.15 × 10−1 | BSIT Total |
| 0.338 ± 0.158 | 3.22 × 10−2 | Chocolate (item 5), | |||
| 0.590 ± 0.204 | 3.78 × 10−3 | Gasoline (item 10 | |||
| 11 | rs6591536 d | 59211188 | 0.004 ± 0.011 | 7.08 × 10−1 | BSIT Total |
| 0.359 ± 0.121 | 2.95 × 10−3 | Pineapple (item 9 | |||
| 15 | rs78633367 a | 38816290 | 0.044 ± 0.037 | 2.64 × 10−1 | BSIT Total |
| 0.866 ± 0.435 | 4.67 × 10−2 | Onion (item 12) | |||
| 16 | rs964745 c | 82540806 | −0.022 ± 0.021 | 3.11 × 10−1 | BSIT Total |
| −0.541 ± 0.240 | 2.40 × 10−2 | Smoke (item 4) | |||
| 19 | rs5020278 e | 9325116 | −0.001 ± 0.013 | 9.38 × 10−1 | BSIT Total |
| −0.442 ± 0.164 | 9.00 × 10−3 | Cinnamon (item 1) | |||
| 0.234 ± 0.106 | 3.04 × 10−2 | Lemon (item 3) | |||
| 20 | rs6052484 b | 4260610 | 0.014 ± 0.011 | 1.96 × 10−1 | BSIT Total |
| 0.235 ± 0.109 | 3.05 × 10−2 | Turpentine (item 2) |
| BSIT Odour (Item) | Polygenic Risk Score (PRS) | β Value (PRS) | S.E. (PRS) | p Value (PRS) | FDR |
|---|---|---|---|---|---|
| Cinnamon (1) | AD PRS | 0.234 | 0.080 | 3.31 × 10−3 | 0.982 |
| Chocolate (5) | WMH PRS | −0.352 | 0.081 | 1.70 × 10−5 | 0.001 |
| Chocolate (5) | Smoking PRS | −0.451 | 0.084 | 8.87 × 10−8 | 1.15 × 10−5 |
| Chocolate (5) | PD PRS | −0.338 | 0.081 | 2.67 × 10−5 | 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raj, S.; Thalamuthu, A.; Armstrong, N.J.; Wright, M.J.; Kwok, J.B.; Trollor, J.N.; Ames, D.; Schofield, P.R.; Brodaty, H.; Sachdev, P.S.; et al. Investigating Olfactory Gene Variation and Odour Identification in Older Adults. Genes 2021, 12, 669. https://doi.org/10.3390/genes12050669
Raj S, Thalamuthu A, Armstrong NJ, Wright MJ, Kwok JB, Trollor JN, Ames D, Schofield PR, Brodaty H, Sachdev PS, et al. Investigating Olfactory Gene Variation and Odour Identification in Older Adults. Genes. 2021; 12(5):669. https://doi.org/10.3390/genes12050669
Chicago/Turabian StyleRaj, Siddharth, Anbupalam Thalamuthu, Nicola J Armstrong, Margaret J Wright, John B Kwok, Julian N Trollor, David Ames, Peter R Schofield, Henry Brodaty, Perminder S Sachdev, and et al. 2021. "Investigating Olfactory Gene Variation and Odour Identification in Older Adults" Genes 12, no. 5: 669. https://doi.org/10.3390/genes12050669
APA StyleRaj, S., Thalamuthu, A., Armstrong, N. J., Wright, M. J., Kwok, J. B., Trollor, J. N., Ames, D., Schofield, P. R., Brodaty, H., Sachdev, P. S., & Mather, K. A. (2021). Investigating Olfactory Gene Variation and Odour Identification in Older Adults. Genes, 12(5), 669. https://doi.org/10.3390/genes12050669








