Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient’s Information and Whole Genome Sequencing
2.2. Whole Genome Sequencing Data Processing
2.3. Variants Detection
2.4. Rare Recurrent Variants Selection and Enrichment Analysis
3. Results
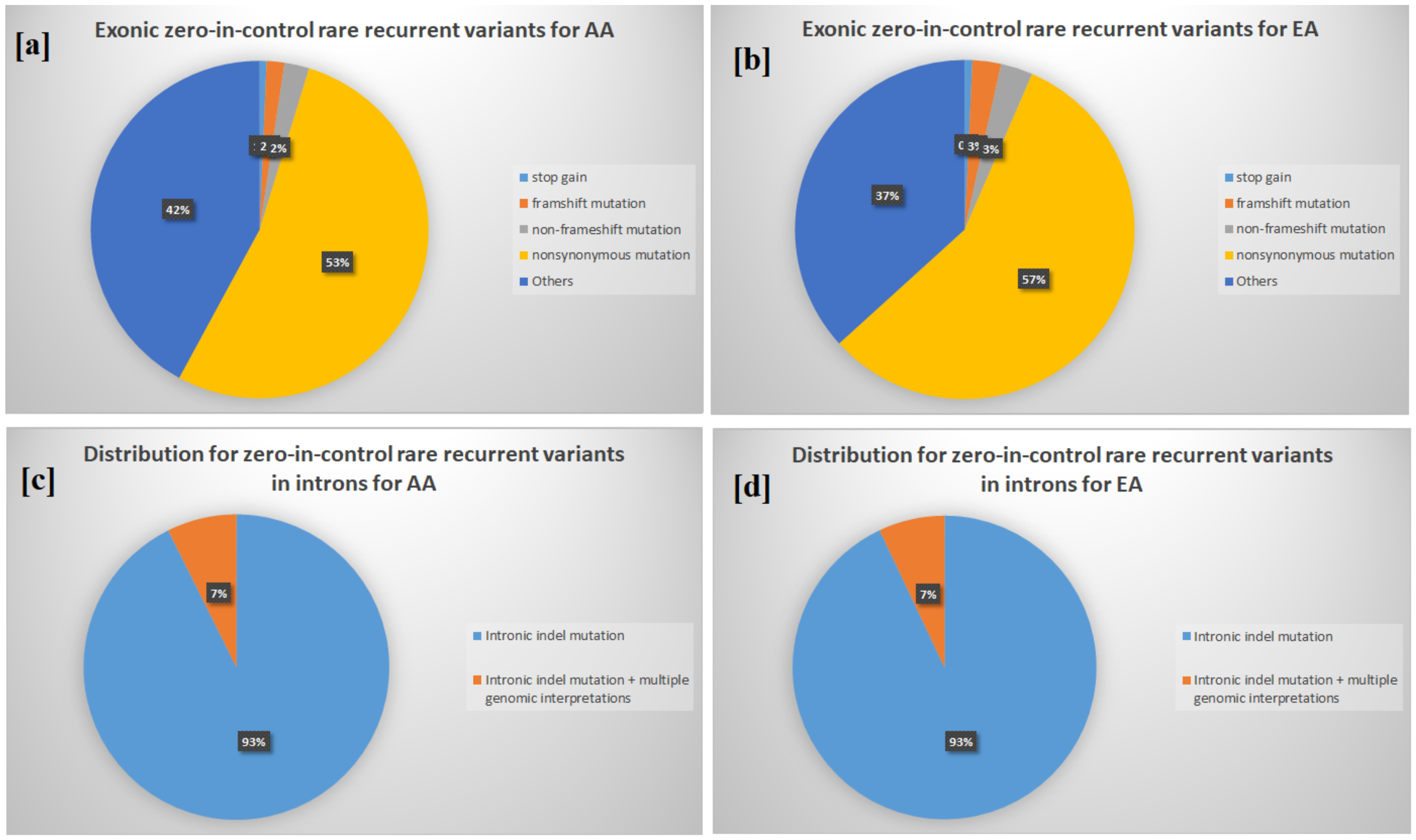
3.1. Novel Stop Codon/Frameshift Variants in Known ADHD Genes
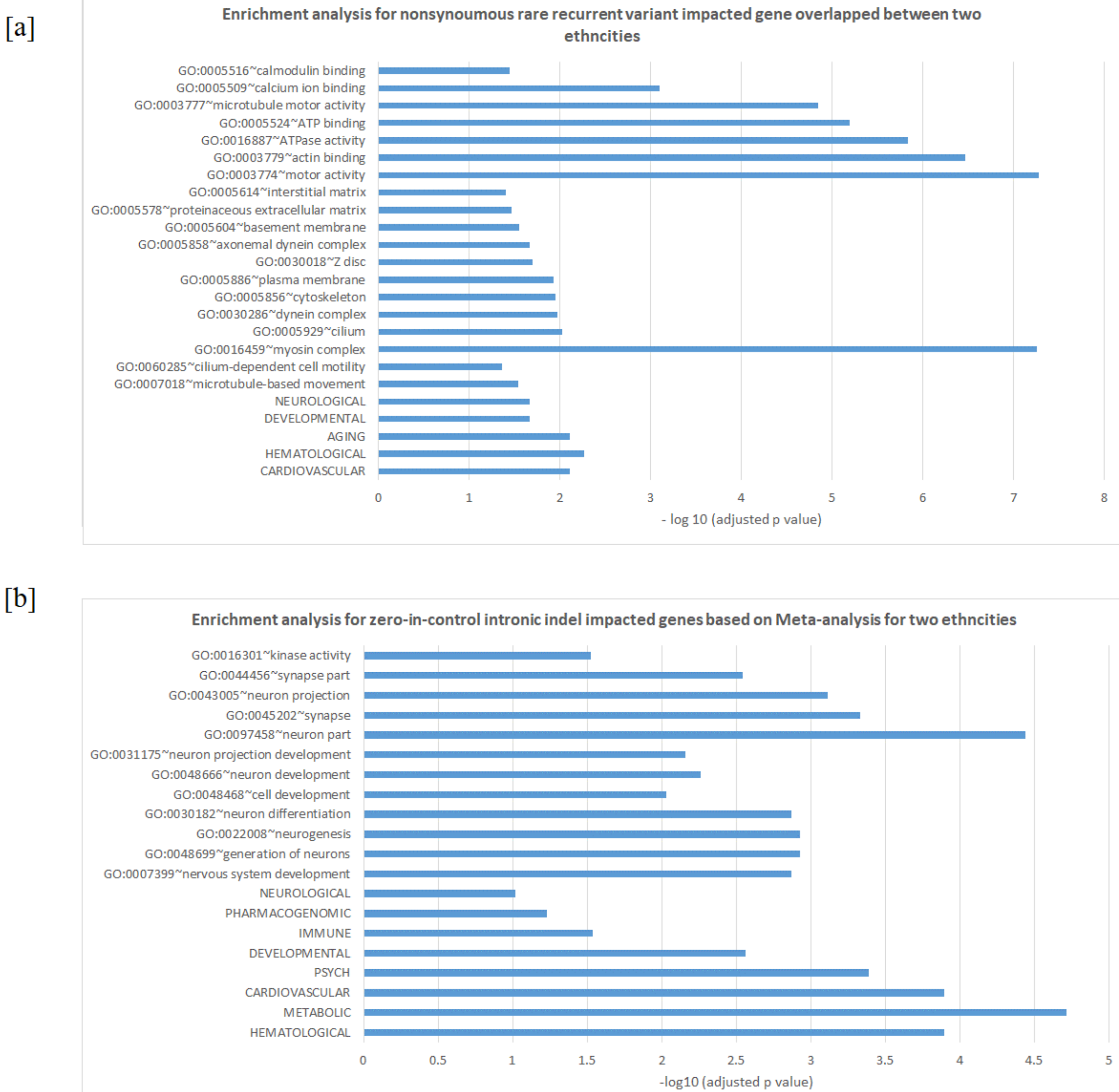
3.2. Enrichment in Homophilic Cell Adhesion of Rare Recurrent Variants in Noncoding RNA/Introns
3.3. Differences between Two Ethnicities and Meta-Analysis
3.4. Rare Recurrent Variants in Noncoding Regions of the Metabotropic Glutamate Receptor (Mglur) Pathway Genes
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Barbaresi, W.J.; Colligan, R.C.; Weaver, A.L.; Voigt, R.G.; Killian, J.M.; Katusic, S.K. Mortality, ADHD, and psychosocial adversity in adults with childhood ADHD: A prospective study. Pediatrics 2013, 131, 637–644. [Google Scholar] [CrossRef]
- Connolly, J.J.; Glessner, J.T.; Elia, J.; Hakonarson, H. ADHD & Pharmacotherapy: Past, Present and Future: A Review of the Changing Landscape of Drug Therapy for Attention Deficit Hyperactivity Disorder. Ther. Innov. Regul. Sci. 2015, 49, 632–642. [Google Scholar] [CrossRef]
- Lantieri, F.; Glessner, J.T.; Hakonarson, H.; Elia, J.; Devoto, M. Analysis of GWAS top hits in ADHD suggests association to two polymorphisms located in genes expressed in the cerebellum. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2010, 153B, 1127–1133. [Google Scholar] [CrossRef]
- Elia, J.; Glessner, J.T.; Wang, K.; Takahashi, N.; Shtir, C.J.; Hadley, D.; Sleiman, P.M.; Zhang, H.; Kim, C.E.; Robison, R.; et al. Genome-wide copy number variation study associates metabotropic glutamate receptor gene networks with attention deficit hyperactivity disorder. Nat. Genet. 2011, 44, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hakonarson, H. ANNOVAR: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic. Acids. Res. 2010, 38, e164. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.T.; Thorvaldsdóttir, H.; Wenger, A.M.; Zehir, A.; Mesirov, J.P. Variant Review with the Integrative Genomics Viewer. Cancer Res. 2017, 77, e31–e34. [Google Scholar] [CrossRef] [PubMed]
- Fu, W.; O’connor, T.D.; Jun, G.; Kang, H.M.; Abecasis, G.; Leal, S.M.; Gabriel, S.; Rieder, M.J.; Altshuler, D.; Shendure, J. Analysis of 6515 exomes reveals the recent origin of most human protein-coding variants. Nature 2013, 493, 216–220. [Google Scholar] [CrossRef] [PubMed]
- Siva, N. 1000 Genomes Project; Nature Publishing Group: Berlin, Germany, 2008. [Google Scholar]
- Karczewski, K.; Francioli, L. The Genome Aggregation Database (gnomAD). Available online: https://macarthurlab.org/2017/02/27/the-genome-aggregation-database-gnomad/ (accessed on 27 February 2017).
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alfoldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef]
- Ng, P.C.; Henikoff, S. SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res. 2003, 31, 3812–3814. [Google Scholar] [CrossRef] [PubMed]
- Chun, S.; Fay, J.C. Identification of deleterious mutations within three human genomes. Genome Res. 2009, 19, 1553–1561. [Google Scholar] [CrossRef] [PubMed]
- Adzhubei, I.A.; Schmidt, S.; Peshkin, L.; Ramensky, V.E.; Gerasimova, A.; Bork, P.; Kondrashov, A.S.; Sunyaev, S.R. A method and server for predicting damaging missense mutations. Nat. Methods 2010, 7, 248–249. [Google Scholar] [CrossRef]
- Schwarz, J.M.; Rodelsperger, C.; Schuelke, M.; Seelow, D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 2010, 7, 575–576. [Google Scholar] [CrossRef]
- Reva, B.; Antipin, Y.; Sander, C. Determinants of protein function revealed by combinatorial entropy optimization. Genome Biol. 2007, 8, R232. [Google Scholar] [CrossRef] [PubMed]
- Shihab, H.A.; Gough, J.; Cooper, D.N.; Stenson, P.D.; Barker, G.L.; Edwards, K.J.; Day, I.N.; Gaunt, T.R. Predicting the functional, molecular, and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum. Mutat. 2013, 34, 57–65. [Google Scholar] [CrossRef]
- Dong, C.; Wei, P.; Jian, X.; Gibbs, R.; Boerwinkle, E.; Wang, K.; Liu, X. Comparison and integration of deleteriousness prediction methods for nonsynonymous SNVs in whole exome sequencing studies. Hum. Mol. Genet. 2015, 24, 2125–2137. [Google Scholar] [CrossRef]
- Landrum, M.J.; Lee, J.M.; Riley, G.R.; Jang, W.; Rubinstein, W.S.; Church, D.M.; Maglott, D.R. ClinVar: Public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014, 42, D980–D985. [Google Scholar] [CrossRef]
- Zhang, L.; Chang, S.; Li, Z.; Zhang, K.; Du, Y.; Ott, J.; Wang, J. ADHDgene: A genetic database for attention deficit hyperactivity disorder. Nucleic Acids Res. 2012, 40, D1003–D1009. [Google Scholar] [CrossRef][Green Version]
- Koenekoop, R.K.; Wang, H.; Majewski, J.; Wang, X.; Lopez, I.; Ren, H.; Chen, Y.; Li, Y.; Fishman, G.A.; Genead, M.; et al. Mutations in NMNAT1 cause Leber congenital amaurosis and identify a new disease pathway for retinal degeneration. Nat. Genet. 2012, 44, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Bouziane, C.; Caan, M.W.A.; Tamminga, H.G.H.; Schrantee, A.; Bottelier, M.A.; de Ruiter, M.B.; Kooij, S.J.J.; Reneman, L. ADHD and maturation of brain white matter: A DTI study in medication naive children and adults. Neuroimage Clin. 2018, 17, 53–59. [Google Scholar] [CrossRef]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Stein, T.I.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards Suite: From Gene Data Mining to Disease Genome Sequence Analyses. Curr. Protoc. Bioinform. 2016, 54, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.; Cooper, M.; Hamshere, M.L.; Pocklington, A.; Scherer, S.W.; Kent, L.; Gill, M.; Owen, M.J.; Williams, N.; O’Donovan, M.C.; et al. Biological overlap of attention-deficit/hyperactivity disorder and autism spectrum disorder: Evidence from copy number variants. J. Am. Acad Child. Adolesc. Psychiatry 2014, 53, 761–770. [Google Scholar] [CrossRef]
- Sakane, F.; Mizuno, S.; Komenoi, S. Diacylglycerol Kinases as Emerging Potential Drug Targets for a Variety of Diseases: An Update. Front. Cell Dev. Biol. 2016, 4, 82. [Google Scholar] [CrossRef]
- Martins-de-Souza, D.; Guest, P.C.; Harris, L.W.; Vanattou-Saifoudine, N.; Webster, M.J.; Rahmoune, H.; Bahn, S. Identification of proteomic signatures associated with depression and psychotic depression in post-mortem brains from major depression patients. Transl. Psychiatry 2012, 2, e87. [Google Scholar] [CrossRef]
- Yamada, K.; Iwayama, Y.; Hattori, E.; Iwamoto, K.; Toyota, T.; Ohnishi, T.; Ohba, H.; Maekawa, M.; Kato, T.; Yoshikawa, T. Genome-wide association study of schizophrenia in Japanese population. PLoS ONE 2011, 6, e20468. [Google Scholar] [CrossRef] [PubMed]
- Lalani, S.R.; Hefner, M.A.; Belmont, J.W.; Davenport, S.L.H. CHARGE Syndrome. In GeneReviews®; Adam, M.P., Ardinger, H.H., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Stephens, K., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 1993. [Google Scholar]
- Kozlowska, A.; Wojtacha, P.; Rowniak, M.; Kolenkiewicz, M.; Huang, A.C.W. ADHD pathogenesis in the immune, endocrine and nervous systems of juvenile and maturating SHR and WKY rats. Psychopharmacology 2019. [Google Scholar] [CrossRef]
- Wong, M.L.; Dong, C.; Andreev, V.; Arcos-Burgos, M.; Licinio, J. Prediction of susceptibility to major depression by a model of interactions of multiple functional genetic variants and environmental factors. Mol. Psychiatry 2012, 17, 624–633. [Google Scholar] [CrossRef]
- Lesch, K.P.; Timmesfeld, N.; Renner, T.J.; Halperin, R.; Röser, C.; Nguyen, T.T.; Craig, D.W.; Romanos, J.; Heine, M.; Meyer, J.; et al. Molecular genetics of adult ADHD: Converging evidence from genome-wide association and extended pedigree linkage studies. J. Neural Transm. 2008, 115, 1573–1585. [Google Scholar] [CrossRef]
- El Hajj, N.; Dittrich, M.; Haaf, T. Epigenetic dysregulation of protocadherins in human disease. Semin. Cell Dev. Biol. 2017, 69, 172–182. [Google Scholar] [CrossRef]
- Franke, B.; Faraone, S.V.; Asherson, P.; Buitelaar, J.; Bau, C.H.; Ramos-Quiroga, J.A.; Mick, E.; Grevet, E.H.; Johansson, S.; Haavik, J.; et al. The genetics of attention deficit/hyperactivity disorder in adults, a review. Mol. Psychiatry 2012, 17, 960–987. [Google Scholar] [CrossRef]
- Rommelse, N.N.; Franke, B.; Geurts, H.M.; Hartman, C.A.; Buitelaar, J.K. Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur. Child. Adolesc. Psychiatry 2010, 19, 281–295. [Google Scholar] [CrossRef] [PubMed]
- Elia, J.; Ungal, G.; Kao, C.; Ambrosini, A.; De Jesus-Rosario, N.; Larsen, L.; Chiavacci, R.; Wang, T.; Kurian, C.; Titchen, K.; et al. Fasoracetam in adolescents with ADHD and glutamatergic gene network variants disrupting mGluR neurotransmitter signaling. Nat. Commun. 2018, 9, 4. [Google Scholar] [CrossRef]
- Acosta, M.T.; Arcos-Burgos, M.; Muenke, M. Attention deficit/hyperactivity disorder (ADHD): Complex phenotype, simple genotype? Genet. Med. 2004, 6, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chang, X.; Qu, H.; Glessner, J.; Tian, L.; Li, D.; Qiu, H.; Sleiman, P.M.; Hakonarson, H. Non-coding structural variation differentially impacts attention-deficit hyperactivity disorder (ADHD) gene networks in African American vs Caucasian children. Sci. Rep. 2020, 10, 1–8. [Google Scholar] [CrossRef]
- Zhang, F.; Lupski, J.R. Non-coding genetic variants in human disease. Hum. Mol. Genet. 2015, 24, R102–R110. [Google Scholar] [CrossRef] [PubMed]
- Stergiakouli, E.; Hamshere, M.; Holmans, P.; Langley, K.; Zaharieva, I.; Hawi, Z.; Kent, L.; Gill, M.; Williams, N.; Owen, M.J.; et al. Investigating the contribution of common genetic variants to the risk and pathogenesis of ADHD. Am. J. Psychiatry 2012, 169, 186–194. [Google Scholar] [CrossRef]
- Mullaney, J.M.; Mills, R.E.; Pittard, W.S.; Devine, S.E. Small insertions and deletions (INDELs) in human genomes. Hum. Mol. Genet. 2010, 19, R131–R136. [Google Scholar] [CrossRef]
- Bryen, S.J.; Joshi, H.; Evesson, F.J.; Girard, C.; Ghaoui, R.; Waddell, L.B.; Testa, A.C.; Cummings, B.; Arbuckle, S.; Graf, N.; et al. Pathogenic Abnormal Splicing Due to Intronic Deletions that Induce Biophysical Space Constraint for Spliceosome Assembly. Am. J. Hum. Genet. 2019, 105, 573–587. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Xiao, H.; Chiou, T.T.; Jin, H.; Bonhomme, B.; Miralles, C.P.; Pinal, N.; Ali, R.; Chen, W.V.; Maniatis, T.; et al. Molecular and functional interaction between protocadherin-gammaC5 and GABAA receptors. J. Neurosci. 2012, 32, 11780–11797. [Google Scholar] [CrossRef] [PubMed]
- Nagamitsu, S.; Yamashita, Y.; Tanigawa, H.; Chiba, H.; Kaida, H.; Ishibashi, M.; Kakuma, T.; Croarkin, P.E.; Matsuishi, T. Upregulated GABA Inhibitory Function in ADHD Children with Child Behavior Checklist-Dysregulation Profile: 123I-Iomazenil SPECT Study. Front. Psychiatry 2015, 6, 84. [Google Scholar] [CrossRef] [PubMed]

| Ethnicity | avsnp147 | chr | Locus | Ref | Alt | ADHD Occurrences | Genes | esp6500 Frequency | 1000Genome Frequency | gnomAD Frequency |
|---|---|---|---|---|---|---|---|---|---|---|
| rs369719214 | 19 | 33183093 | G | A | 2 | NUDT19 | 6.0 × 10−4 | 6.0 × 10−4 | 6.0 × 10−4 | |
| rs116059545 | 3 | 142537266 | G | A | 2 | PCOLCE2 | 2.0 × 10−4 | 2.0 × 10−4 | 2.0 × 10−4 | |
| rs146817618 | 11 | 6942649 | T | A | 2 | OR2D3 | 2.0 × 10−4 | . | 3.0 × 10−4 | |
| rs34789740 | 12 | 11091287 | G | A | 2 | TAS2R14 | 1.4 × 10−3 | 4.0 × 10−4 | 8.0 × 10−4 | |
| rs375920305 | 19 | 36259319 | C | T | 4 | PROSER3 | 7.0 × 10−4 | 8.0 × 10−4 | 3.0 × 10−4 | |
| AA | . | 19 | 52888074 | - | ATCATGAGGTCAGGAGATCGAGACCATCCTGGCTAACAAGGTGAAACC | 2 | ZNF880 | . | . | 1.8 × 10−3 |
| rs150768729 | 4 | 2073958 | C | T | 2 | POLN | 2.0 × 10−4 | . | 3.3E-05 | |
| rs370330395 | 19 | 52887191 | C | T | 3 | ZNF880 | 1.3 × 10−3 | 6.0 × 10−4 | 1.2 × 10−3 | |
| rs148053441 | 14 | 74361075 | G | A | 2 | ZNF410 | . | 4.0 × 10−4 | 1.0 × 10−3 | |
| rs181032032 | 19 | 12126454 | G | A | 2 | ZNF433 | 5.0 × 10−4 | 4.0 × 10−4 | 7.0 × 10−4 | |
| rs147487823 | 8 | 29202959 | G | A | 2 | DUSP4 | . | 2.0 × 10−4 | 3.2 × 10−5 | |
| rs138842904 | 16 | 74486025 | G | A | 2 | GLG1 | 9.0 × 10−4 | 4.0 × 10−4 | 8.0 × 10−4 | |
| rs147869298 | 4 | 140625183 | C | T | 2 | MGST2 | 1.5 × 10−3 | 6.0 × 10−4 | 5.0 × 10−4 | |
| . | 9 | 104239264 | - | ATTAAAAA | 2 | TMEM246 | . | . | 7.0 × 10−5 | |
| rs145322761 | 14 | 96730863 | C | T | 2 | BDKRB1 | 2.8 × 10−3 | 1.4 × 10−3 | 2.0 × 10−3 | |
| rs138652787 | 7 | 23871861 | C | G | 2 | STK31 | 1.8 × 10−3 | 6.0 × 10−4 | 1.6 × 10−3 | |
| EA | . | 2 | 11925167 | - | ATA | 3 | LPIN1 | . | . | 0.0E+00 |
| rs142358325 | 9 | 140139138 | G | A | 2 | FAM166A | 5.0 × 10−4 | 4.0 × 10−4 | 3.0 × 10−4 | |
| . | 13 | 32731436 | C | T | 2 | FRY | . | . | . | |
| rs370788593 | 10 | 82348410 | C | T | 2 | SH2D4B | 7.7 × 10−5 | . | 3.2 × 10−5 | |
| rs371526758 | 1 | 10042426 | G | A | 2 | NMNAT1 | 2.0 × 10−4 | . | 6.5 × 10−5 |
| Ethnicity | chr | Locus | Ref | Alt | ADHD Occurrences | Genes | esp6500 Frequency | 1000Genome Frequency | gnomAD Frequency |
|---|---|---|---|---|---|---|---|---|---|
| 7 | 48237838 | TTTG | - | 2 | ABCA13 | . | . | 3.38 × 10−5 | |
| 7 | 48237845 | - | GA | 2 | ABCA13 | . | . | 0 | |
| 7 | 48237846 | - | CA | 2 | ABCA13 | . | . | 0 | |
| AA | 12 | 112036782 | - | GCTGCTGCTGCTGC | 2 | ATXN2 | . | . | . |
| 4 | 962079 | - | TGCCTCTCCTGCCCCGCCCCCCCAACTCCTC | 3 | DGKQ | . | . | 0.0021 | |
| 4 | 962079 | - | TGCCTCTCCTGCCCCGCCC | 2 | DGKQ | . | . | 0.0014 | |
| 2 | 153417444 | - | GCCGT | 2 | FMNL2 | . | . | 0 | |
| 2 | 153417451 | GCCCTGG | - | 2 | FMNL2 | . | . | . | |
| 8 | 61732577 | GCTTT | - | 3 | CHD7 | . | . | 7.63 × 10−5 | |
| 8 | 61732592 | TT | - | 3 | CHD7 | . | . | 6.53 × 10−5 | |
| EA | 1 | 120056817 | - | G | 2 | HSD3B1 | . | . | 0.0005 |
| 1 | 120056818 | - | AA | 2 | HSD3B1 | 0.0007 | . | 0.0005 | |
| 10 | 73044507 | - | G | 2 | UNC5B | . | . | 0.0004 |
| Ethnicity | chr | Locus | Ref | Alt | avsnp147 | ADHD Occurrences | Genes | SIFT | Polyphen2 | LRT | Mutation Taster | Mutation Assessor | FATHMM | Radial SVM |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 8 | 26721808 | A | G | rs201951243 | 2 | ADRA1A | D | D | N | D | L | T | T | |
| 10 | 135139570 | C | T | rs149764782 | 2 | CALY | T | P | N | D | M | . | T | |
| 10 | 73501613 | C | T | rs200664666 | 2 | CDH23 | T | D | D | D | M | . | T | |
| 2 | 141459361 | G | A | rs147598746 | 2 | LRP1B | T | B | N | N | N | D | T | |
| 18 | 47429043 | C | A | rs114221227 | 2 | MYO5B | D | D | D | D | M | T | T | |
| 14 | 80327577 | A | G | . | 2 | NRXN3 | . | B | . | D | . | T | T | |
| 2 | 42991127 | G | A | rs146033252 | 2 | OXER1 | T | B | . | N | N | T | T | |
| EA | 17 | 71410819 | G | T | rs143251430 | 2 | SDK2 | D | P | D | D | L | T | T |
| 1 | 177930009 | C | T | rs185756583 | 2 | SEC16B | D | D | D | D | M | T | T | |
| 10 | 98945389 | G | A | rs375276519 | 2 | SLIT1 | D | D | N | N | N | T | T | |
| 6 | 24658948 | T | C | rs61757564 | 3 | TDP2 | T | B | N | N | L | T | T | |
| 11 | 78383209 | A | T | rs201179027 | 2 | TENM4 | . | D | D | D | L | T | D | |
| 11 | 78369726 | C | T | rs199594129 | 2 | TENM4 | . | D | N | D | N | D | T | |
| 3 | 36873195 | C | T | rs373979668 | 2 | TRANK1 | D | B | D | N | M | T | T | |
| 19 | 57764649 | G | A | rs377720732 | 2 | ZNF805 | D | B | . | N | L | T | T | |
| 11 | 74988453 | C | T | rs370469526 | 2 | ARRB1 | T | B | D | D | N | T | T | |
| 4 | 96025649 | C | G | rs145700191 | 2 | BMPR1B | T | B | N | D | L | T | D | |
| 9 | 90585782 | C | T | rs28364955 | 2 | CDK20 | T | P | D | N | L | T | T | |
| 7 | 50566868 | C | T | rs573103547 | 2 | DDC | D | D | D | D | M | T | T | |
| 9 | 1056887 | C | T | rs147461872 | 2 | DMRT2 | D | D | U | D | M | T | T | |
| 2 | 141250262 | T | C | rs140458851 | 2 | LRP1B | T | B | N | D | N | D | T | |
| 7 | 77648999 | C | A | rs773082728 | 2 | MAGI2 | T | B | . | D | N | T | T | |
| 15 | 91455385 | G | A | rs150171248 | 2 | MAN2A2 | T | B | D | D | M | D | T | |
| AA | 1 | 11856376 | C | T | rs150847674 | 2 | MTHFR | T | P | D | D | M | D | D |
| 12 | 117768154 | C | T | rs76090928 | 2 | NOS1 | D | B | D | D | L | T | T | |
| 14 | 79454391 | T | G | . | 2 | NRXN3 | . | D | D | D | . | T | T | |
| 20 | 4773214 | G | A | rs148303159 | 2 | RASSF2 | T | B | N | N | L | T | T | |
| 17 | 71397825 | T | C | rs138152327 | 2 | SDK2 | T | B | N | N | M | T | T | |
| 6 | 155577706 | C | A | rs114296676 | 2 | TIAM2 | T | D | D | N | M | T | T | |
| 5 | 14487903 | C | T | rs533386148 | 3 | TRIO | T | P | N | D | N | T | T | |
| 1 | 55571832 | C | T | . | 2 | USP24 | T | P | D | D | L | T | T | |
| 19 | 58772917 | A | C | rs148565349 | 2 | ZNF544 | T | B | . | N | L | T | T | |
| 5 | 123982926 | T | C | rs149444271 | 2 | ZNF608 | T | B | D | D | M | T | T | |
| 5 | 123980164 | T | C | rs113873110 | 2 | ZNF608 | . | B | N | D | L | T | T |
| ADHD Associated Genes | Gene | chr | Locus | Ref | Alt | avsnp147 | ADHD AA Occurrences | ADHD EA Occurrences | meta p-Value |
|---|---|---|---|---|---|---|---|---|---|
| Y | MAGI2 | 7 | 77687294 | - | TATA | . | 4 | 1 | 0.008 |
| - | ZMYND8 | 20 | 45929674 | - | TGTGTGTA | . | 4 | 1 | 0.008 |
| - | ERBB4 | 2 | 212606179 | - | ACACACAC | . | 4 | 1 | 0.008 |
| - | SOX6, | 11 | 16031671 | - | ACACACAC | . | 4 | 1 | 0.008 |
| - | ALCAM | 3 | 105241984 | - | AA | . | 3 | 2 | 0.009 |
| - | BRINP3 | 1 | 190425195 | A | - | . | 1 | 4 | 0.015 |
| Y | NRXN3 | 14 | 78954086 | ATAAATAAATAAATAA | - | . | 2 | 3 | 0.012 |
| - | COL25A1 | 4 | 110139513 | - | T | rs34056401 | 2 | 3 | 0.012 |
| - | CBFA2T2 | 20 | 32229919 | TTTTTGTGTGTGTG | - | . | 2 | 3 | 0.012 |
| Y | DCDC2 | 6 | 24280004 | T | - | rs563616388 | 1 | 4 | 0.015 |
| Y | DCDC2 | 6 | 24256576 | TA | - | rs556522905 | 1 | 4 | 0.015 |
| Y | NRXN1 | 2 | 50393425 | - | TG | . | 1 | 4 | 0.015 |
| Y | NRXN3 | 14 | 78937634 | - | AC | . | 3 | 1 | 0.038 |
| Y | CTNNA2 | 2 | 80029140 | T | - | . | 3 | 1 | 0.038 |
| - | RORA | 15 | 61494733 | T | - | . | 3 | 1 | 0.038 |
| - | ARHGEF2 | 1 | 155941370 | - | AAAAAAAAAAA | . | 3 | 1 | 0.038 |
| - | BICDL1 | 12 | 120451243 | - | GTGTGTGTGTGTGT | . | 3 | 1 | 0.038 |
| - | PARD3 | 10 | 34408932 | - | TTTTTTTTTTT | . | 3 | 1 | 0.038 |
| - | CNTN6 | 3 | 1136572 | AT | - | rs367911099 | 3 | 1 | 0.038 |
| - | DAB1 | 1 | 58589583 | AC | - | . | 3 | 1 | 0.038 |
| - | COL25A1 | 4 | 109743199 | - | TTTTTTTT | . | 3 | 1 | 0.038 |
| - | ANKS1A | 6 | 34891869 | - | GTGT | . | 3 | 1 | 0.038 |
| - | PTPRD | 9 | 9632617 | A | - | . | 3 | 1 | 0.038 |
| - | NPTN | 15 | 73897440 | - | GG | . | 3 | 1 | 0.038 |
| - | SLC4A7 | 3 | 27480018 | - | AAAAT | rs141000029 | 3 | 1 | 0.038 |
| - | LAMB1 | 7 | 107575792 | - | TTTTTTTTTTTTTT | . | 3 | 1 | 0.038 |
| - | DAB1 | 1 | 58222892 | - | TTTTTTTTTTTTTTTTTTTTTTTTTG | . | 3 | 1 | 0.038 |
| - | RORA | 15 | 61494730 | - | G | . | 3 | 1 | 0.038 |
| - | GNAQ | 9 | 80366732 | AAAA | - | . | 3 | 1 | 0.038 |
| - | MTOR | 1 | 11178676 | - | T | . | 3 | 1 | 0.038 |
| - | DAB1 | 1 | 58616263 | AA | - | . | 3 | 1 | 0.038 |
| - | LRP6 | 12 | 12392929 | - | A | . | 3 | 1 | 0.038 |
| - | NCAM2 | 21 | 22444188 | - | TATCTAT | . | 3 | 1 | 0.038 |
| - | SPTBN4 | 19 | 41005988 | - | AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAGAAAAGAAAAATCTTTGCTGAGCATGGTGGTACAC | . | 3 | 1 | 0.038 |
| - | RAB10 | 2 | 26331748 | - | TATT | . | 3 | 1 | 0.038 |
| Y | CLASP2 | 3 | 33577416 | A | - | . | 3 | 1 | 0.038 |
| Y | LINGO2 | 9 | 28664998 | - | ATATATAT | . | 2 | 2 | 0.046 |
| Y | PRKG1 | 10 | 52873716 | - | GT | . | 2 | 2 | 0.046 |
| - | TENM2 | 5 | 166755213 | - | T | . | 2 | 2 | 0.046 |
| - | DNER | 2 | 230419651 | GAGAGAAAAGGGAAGGG | - | . | 2 | 2 | 0.046 |
| - | EXT1 | 8 | 119053913 | GAAG | - | . | 2 | 2 | 0.046 |
| - | PTPRD | 9 | 8632122 | TG | - | rs755867249 | 2 | 2 | 0.046 |
| - | ABL2 | 1 | 179169177 | T | - | . | 2 | 2 | 0.046 |
| - | NOTCH3 | 19 | 15280017 | - | TCTCTCTC | . | 2 | 2 | 0.046 |
| Y | NRXN1 | 2 | 50932188 | ACAC | - | . | 2 | 2 | 0.046 |
| - | UST | 6 | 149379481 | ATGTGTGTGT | - | . | 2 | 2 | 0.046 |
| - | GCM1 | 6 | 53006798 | - | GAAA | . | 2 | 2 | 0.046 |
| Gene | chr | Locus | Ref | Alt | ADHD AA Occurrences | ADHD EA Occurrences | Meta-Analysis p-Value | esp6500 freq | 1000Genome freq | gnomAD freq |
|---|---|---|---|---|---|---|---|---|---|---|
| GRM7 | 3 | 7515182 | - | GAGAGAGAGA | 2 | 3 | 0.012 | . | . | . |
| DLGAP1 | 18 | 4330027 | AT | - | 3 | 1 | 0.038 | . | . | 0.0008 |
| GNG2 | 14 | 52368681 | - | A | 3 | 1 | 0.038 | . | . | . |
| GRIK2 | 6 | 102004196 | - | TTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTTT | 3 | 1 | 0.038 | . | . | . |
| PRKACB | 1 | 84664799 | - | ATAT | 3 | 1 | 0.038 | . | . | 0.0038 |
| GNAQ | 9 | 80366732 | AAAA | - | 3 | 1 | 0.038 | . | . | 0.0006 |
| PRKCG | 19 | 54406144 | - | AAAAAAAAAAAAAAAA | 3 | 1 | 0.038 | . | . | 0.0022 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Chang, X.; Qu, H.-Q.; Tian, L.; Glessner, J.; Qu, J.; Li, D.; Qiu, H.; Sleiman, P.; Hakonarson, H. Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry. Genes 2021, 12, 310. https://doi.org/10.3390/genes12020310
Liu Y, Chang X, Qu H-Q, Tian L, Glessner J, Qu J, Li D, Qiu H, Sleiman P, Hakonarson H. Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry. Genes. 2021; 12(2):310. https://doi.org/10.3390/genes12020310
Chicago/Turabian StyleLiu, Yichuan, Xiao Chang, Hui-Qi Qu, Lifeng Tian, Joseph Glessner, Jingchun Qu, Dong Li, Haijun Qiu, Patrick Sleiman, and Hakon Hakonarson. 2021. "Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry" Genes 12, no. 2: 310. https://doi.org/10.3390/genes12020310
APA StyleLiu, Y., Chang, X., Qu, H.-Q., Tian, L., Glessner, J., Qu, J., Li, D., Qiu, H., Sleiman, P., & Hakonarson, H. (2021). Rare Recurrent Variants in Noncoding Regions Impact Attention-Deficit Hyperactivity Disorder (ADHD) Gene Networks in Children of both African American and European American Ancestry. Genes, 12(2), 310. https://doi.org/10.3390/genes12020310






