Investigation of Isocitrate Dehydrogenase 1 and 2 Mutations in Acute Leukemia Patients in Saudi Arabia
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients Data
2.2. Cytogenetic and Molecular Data
2.3. DNA Extraction and PCR Amplification
2.4. DNA Sequencing
3. Results
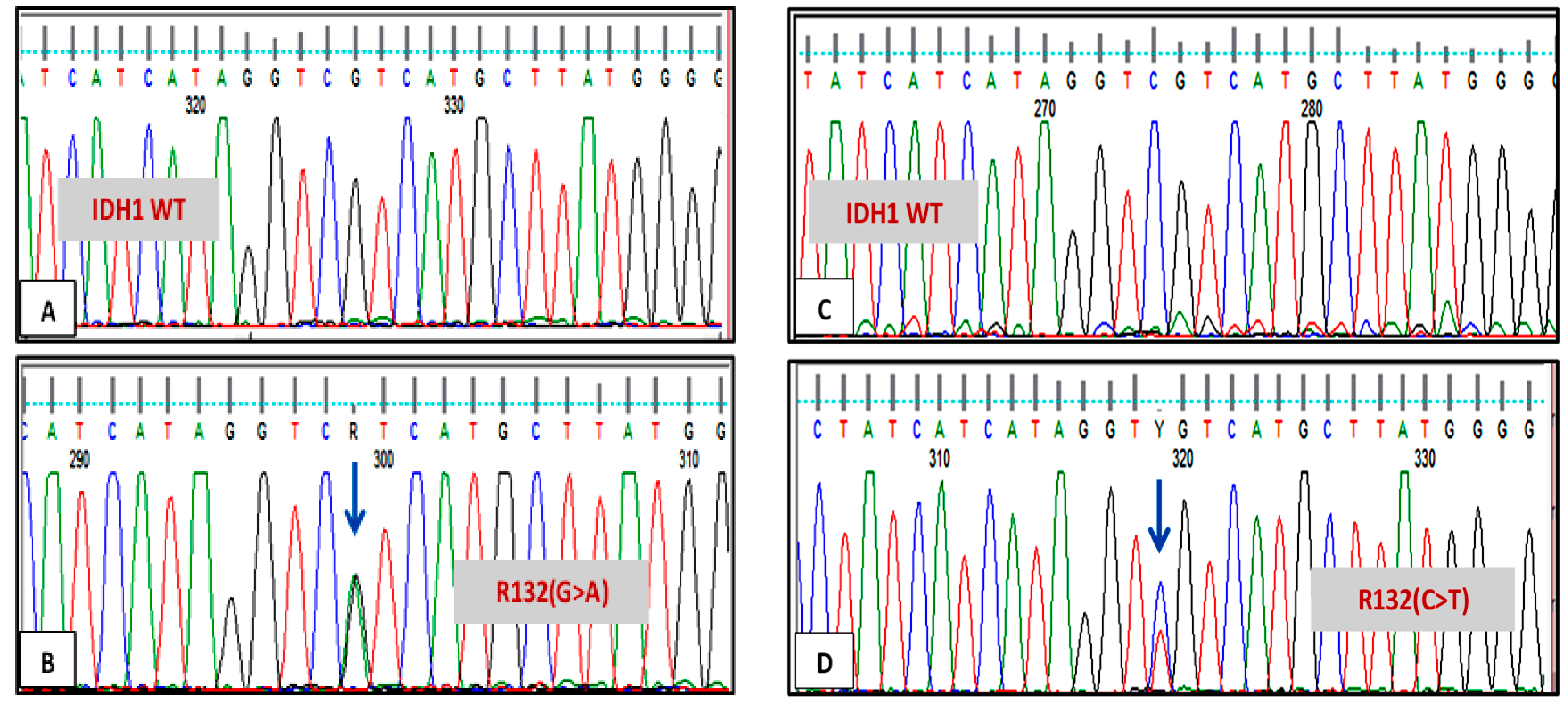
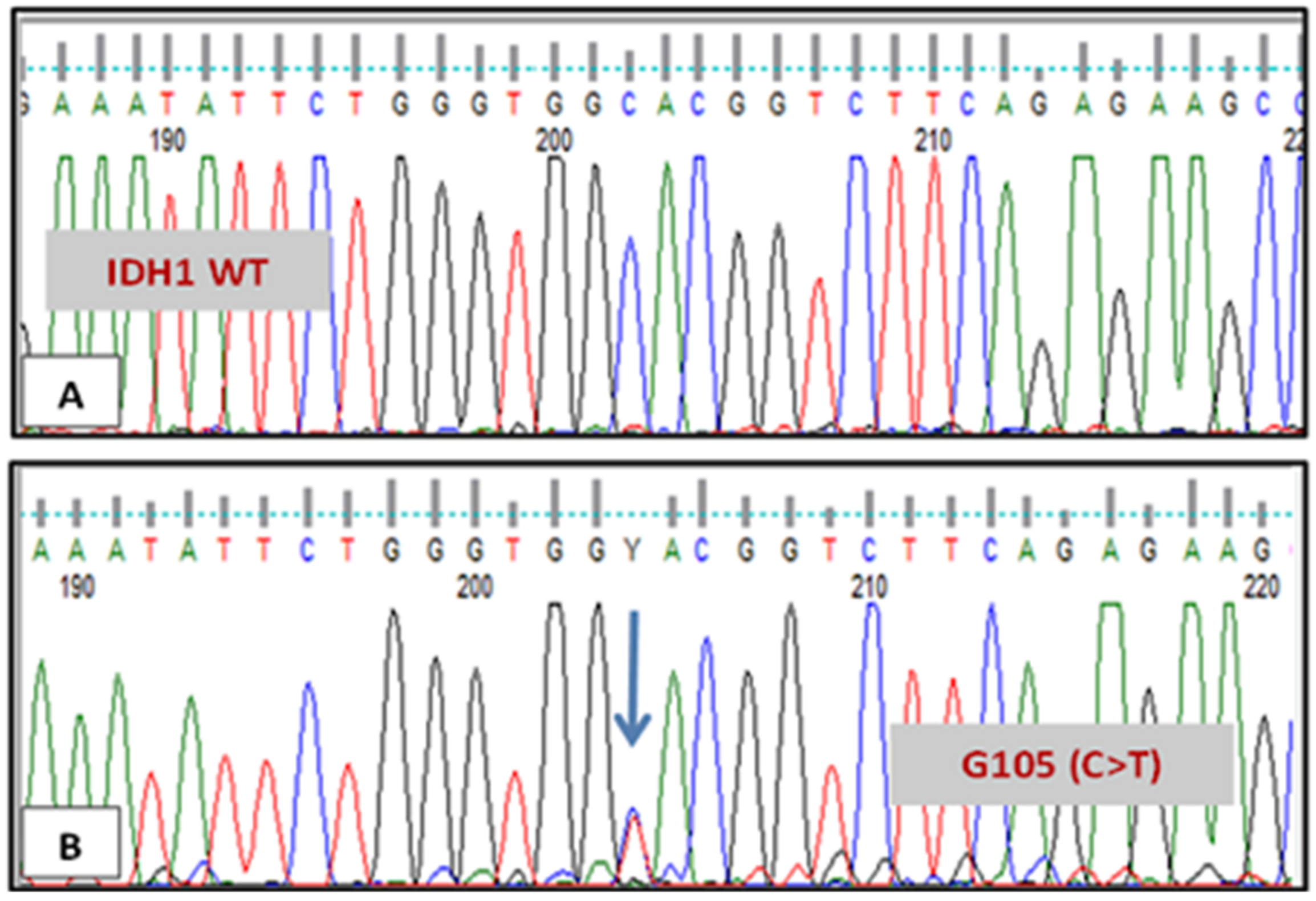
3.1. IDH1 and IDH2 Mutations Analysis
3.2. Correlation of IDH1 and IDH2 with the Cytogenetic Aberrations
3.3. Correlation of IDH1/2 with Molecular Results
3.4. IDH1 and IDH2 Mutations Association with the Treatment Response
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yang, H.; Ye, D.; Guan, K.-L.; Xiong, Y. IDH1 and IDH2 mutations in tumorigenesis: Mechanistic insights and clinical perspectives. Clin. Cancer Res. 2012, 18, 5562–5571. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yen, K.; Bittinger, M.; Su, S.; Fantin, V. Cancer-associated IDH mutations: Biomarker and therapeutic opportunities. Oncogene 2010, 29, 6409–6417. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balss, J.; Meyer, J.; Mueller, W.; Korshunov, A.; Hartmann, C.; von Deimling, A. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol. 2008, 116, 597–602. [Google Scholar] [CrossRef] [PubMed]
- Dang, L.; White, D.W.; Gross, S.; Bennett, B.D.; Bittinger, M.A.; Driggers, E.M.; Fantin, V.R.; Jang, H.G.; Jin, S.; Keenan, M.C.; et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 2009, 462, 739–744. [Google Scholar] [CrossRef] [Green Version]
- Ahmad, F.; Mohota, R.; Sanap, S.; Mandava, S.; Das, B.R. Molecular evaluation of DNMT3A and IDH1/2 gene mutation: Frequency, distribution pattern and associations with additional molecular markers in normal karyotype Indian acute myeloid leukemia patients. Asian Pac. J. Cancer Prev. 2014, 15, 1247–1253. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Horbinski, C. What do we know about IDH1/2 mutations so far, and how do we use it? Acta Neuropathol. 2013, 125, 621–636. [Google Scholar] [CrossRef] [Green Version]
- Seltzer, M.J.; Bennett, B.D.; Joshi, A.D.; Gao, P.; Thomas, A.G.; Ferraris, D.V.; Tsukamoto, T.; Rojas, C.J.; Slusher, B.S.; Rabinowitz, J.D.; et al. Inhibition of glutaminase preferentially slows growth of glioma cells with mutant IDH1. Cancer Res. 2010, 70, 8981–8987. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mondesir, J.; Willekens, C.; Touat, M.; de Botton, S. IDH1 and IDH2 mutations as novel therapeutic targets: Current perspectives. J. Blood Med. 2016, 7, 171. [Google Scholar]
- Hillis, L.D.; Smith, P.K.; Anderson, J.L.; Bittl, J.A.; Bridges, C.R.; Byrne, J.G.; Cigarroa, J.E.; DiSesa, V.J.; Hiratzka, L.F.; Hutter, A.M.; et al. 2011 ACCF/AHA guideline for coronary artery bypass graft surgery: A report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines developed in collaboration with the American Association for Thoracic Surgery, Society of Cardiovascular Anesthesiologists, and Society of Thoracic Surgeons. J. Am. Coll. Cardiol. 2011, 58, e123–e210. [Google Scholar]
- AlYami, M.S.; Watson, R. An overview of nursing in Saudi Arabia. J. Health Spec. 2014, 2, 10–12. [Google Scholar] [CrossRef]
- Bleeker, F.E.; Lamba, S.; Leenstra, S.; Troost, D.; Hulsebos, T.; Vandertop, W.P.; Frattini, M.; Molinari, F.; Knowles, M.; Cerrato, A.; et al. IDH1 mutations at residue p. R132 (IDH1R132) occur frequently in high-grade gliomas but not in other solid tumors. Hum. Mutat. 2009, 30, 7–11. [Google Scholar] [CrossRef]
- Emadi, A.; Faramand, R.; Carter-Cooper, B.; Tolu, S.; Ford, L.A.; Lapidus, R.G.; Wetzler, M.; Wang, E.S.; Etemadi, A.; Griffiths, E.A. Presence of isocitrate dehydrogenase mutations may predict clinical response to hypomethylating agents in patients with acute myeloid leukemia. Am. J. Hematol. 2015, 90, E77–E79. [Google Scholar] [CrossRef]
- Phoenix, C.; Smith, B. Telling a (good?) counterstory of aging: Natural bodybuilding meets the narrative of decline. J. Gerontol. Ser. B Psychol. Sci. Soc. Sci. 2011, 66, 628–639. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cocciardi, S.; Dolnik, A.; Kapp-Schwoerer, S.; Rücker, F.G.; Lux, S.; Blätte, T.J.; Skambraks, S.; Krönke, J.; Heidel, F.H.; Schnöder, T.M.; et al. Clonal evolution patterns in acute myeloid leukemia with NPM1 mutation. Nat. Commun. 2019, 10, 2031. [Google Scholar] [CrossRef] [Green Version]
- Patel, K.P.; Ravandi, F.; Ma, D.; Paladugu, A.; Barkoh, B.A.; Medeiros, L.J.; Luthra, R. Acute myeloid leukemia with IDH1 or IDH2 mutation: Frequency and clinicopathologic features. Am. J. Clin. Pathol. 2011, 135, 35–45. [Google Scholar] [CrossRef] [Green Version]
- Ward, P.S.; Patel, J.; Wise, D.R.; Abdel-Wahab, O.; Bennett, B.D.; Coller, H.A.; Cross, J.R.; Fantin, V.R.; Hedvat, C.V.; Perl, A.E.; et al. The common feature of leukemia-associated IDH1 and IDH2 mutations is a neomorphic enzyme activity converting α-ketoglutarate to 2-hydroxyglutarate. Cancer Cell 2010, 17, 225–234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wagner, K.; Damm, F.; Göhring, G.; Görlich, K.; Heuser, M.; Schäfer, I.; Ottmann, O.; Lübbert, M.; Heit, W.; Kanz, L.; et al. Impact of IDH1 R132 mutations and an IDH1 single nucleotide polymorphism in cytogenetically normal acute myeloid leukemia: SNP rs11554137 is an adverse prognostic factor. J. Clin. Oncol. 2010, 28, 2356–2364. [Google Scholar] [CrossRef]
- Paschka, P.; Schlenk, R.F.; Gaidzik, V.I.; Habdank, M.; Krönke, J.; Bullinger, L.; Späth, D.; Kayser, S.; Zucknick, M.; Götze, K.; et al. IDH1 and IDH2 mutations are frequent genetic alterations in acute myeloid leukemia and confer adverse prognosis in cytogenetically normal acute myeloid leukemia with NPM1 mutation without FLT3 internal tandem duplication. J. Clin. Oncol. 2010, 28, 3636–3643. [Google Scholar] [CrossRef] [PubMed]
- Andersson, A.K.; Miller, D.W.; Lynch, J.A.; Lemoff, A.S.; Cai, Z.; Pounds, S.B.; Radtke, I.; Yan, B.; Schuetz, J.D.; Rubnitz, J.E.; et al. IDH1 and IDH2 mutations in pediatric acute leukemia. Leukemia 2011, 25, 1570–1577. [Google Scholar] [CrossRef] [PubMed]
- Simonin, M.; Schmidt, A.; Bontoux, C.; Dourthe, M.-É.; Lengliné, E.; Andrieu, G.P.; Lhermitte, L.; Graux, C.; Grardel, N.; Cayuela, J.-M.; et al. Oncogenetic landscape and clinical impact of IDH1 and IDH2 mutations in T-ALL. J. Hematol. Oncol. 2021, 14, 74. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, S. Downstream molecular pathways of FLT3 in the pathogenesis of acute myeloid leukemia: Biology and therapeutic implications. J. Hematol. Oncol. 2011, 4, 13. [Google Scholar] [CrossRef] [Green Version]
- Takahashi, S. Current findings for recurring mutations in acute myeloid leukemia. J. Hematol. Oncol. 2011, 4, 36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kaminska, B.; Czapski, B.; Guzik, R.; Król, S.K.; Gielniewski, B. Consequences of IDH1/2 mutations in gliomas and an assessment of inhibitors targeting mutated IDH proteins. Molecules 2019, 24, 968. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Age, years, median (range) | (18–76) 34 |
| Gender (female/male) | 18/25 |
| Cytogenetic, n (%) | |
| normal | 24 (56%) |
| single abnormality | 16 (37%) |
| complex | 2 (5%) |
| unknown | 1 (2%) |
| Molecular Changes (%) | |
| FLT3/ITD | 2 (5%) |
| FLT3/TKD | 2 (5%) |
| NPM1 | 5 (12%) |
| BCR/ABL | 2 (5%) |
| PML/RARA | 2 (5%) |
| None | 31 (72%) |
| Outcome Following Treatment | |
| In remission | 28 (65%) |
| Relapse | 15 (35%) |
| Age, years, median (range) | (1–14) 8 |
| Gender (female/male) | 16/14 |
| Cytogenetic, n (%) | |
| normal | 15 (50%) |
| single abnormality | 1 (3%) |
| complex | 3 (10%) |
| unknown | 11 (37%) |
| Outcome Following Treatment | |
| In remission | 19 (63%) |
| Relapse | 11 (37%) |
| Patient No. | Age/Sex | Diagnosis | Karyotype | Type of IDH Mutation | Presence of Other Mutations | Treatment Response |
|---|---|---|---|---|---|---|
| 1 | 31/F | AML | trisomy 4,8 and 21 | IDH1 codon R132 c.395G>A, pArg13His | none | Relapse |
| 2 | 17/F | AML | cytogenetically normal | IDH1 codon R132, c.394C>T, p.Arg131Cys | NPM1 | In remission |
| 3 | 21/M | AML | t(8:21)(q22:q22), -2q deletion | IDH1 codon R132 c.395G>A, pArg13His | none | In remission |
| 4 | 29/F | AML | Complex karyotype: translocation (9;11), trisomy 13,12 and 22 | IDH1 codon G105, c.315C>T, pGly105Gly | none | Relapse |
| 5 | 18/F | AML | cytogenetically normal | IDH1 codon R132, c.394C>T, p.Arg131Cys | none | In remission |
| 6 | 36/F | AML | cytogenetically normal | IDH1 codon R135 c. T>A, | none | In remission |
| 7 | 22/M | AML | cytogenetically normal | IDH1 codon R132, c.394C>T, p.Arg131Cys + codon G105, c.315C>T, pGly105Gly | none | Relapse |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alkhatabi, H.; Bin Saddeq, H.A.; Alyamani, L.; Shinawi, T.; Yasin, E.B.; Alserihi, R.; Felimban, R.; Tayeb, H.H.; Mimani, R.; Alalla, Z.; et al. Investigation of Isocitrate Dehydrogenase 1 and 2 Mutations in Acute Leukemia Patients in Saudi Arabia. Genes 2021, 12, 1963. https://doi.org/10.3390/genes12121963
Alkhatabi H, Bin Saddeq HA, Alyamani L, Shinawi T, Yasin EB, Alserihi R, Felimban R, Tayeb HH, Mimani R, Alalla Z, et al. Investigation of Isocitrate Dehydrogenase 1 and 2 Mutations in Acute Leukemia Patients in Saudi Arabia. Genes. 2021; 12(12):1963. https://doi.org/10.3390/genes12121963
Chicago/Turabian StyleAlkhatabi, Heba, Haneen Abdulfattah Bin Saddeq, Luay Alyamani, Thoraia Shinawi, Elrashed B. Yasin, Raed Alserihi, Raed Felimban, Hossam H. Tayeb, Rawan Mimani, Zainab Alalla, and et al. 2021. "Investigation of Isocitrate Dehydrogenase 1 and 2 Mutations in Acute Leukemia Patients in Saudi Arabia" Genes 12, no. 12: 1963. https://doi.org/10.3390/genes12121963
APA StyleAlkhatabi, H., Bin Saddeq, H. A., Alyamani, L., Shinawi, T., Yasin, E. B., Alserihi, R., Felimban, R., Tayeb, H. H., Mimani, R., Alalla, Z., Abu-Elmagd, M., & Abuzenadah, A. (2021). Investigation of Isocitrate Dehydrogenase 1 and 2 Mutations in Acute Leukemia Patients in Saudi Arabia. Genes, 12(12), 1963. https://doi.org/10.3390/genes12121963







