ABA and Bud Dormancy in Perennials: Current Knowledge and Future Perspective
Abstract
:1. Introduction
1.1. ABA Metabolism and Signaling
1.2. Bud Dormancy
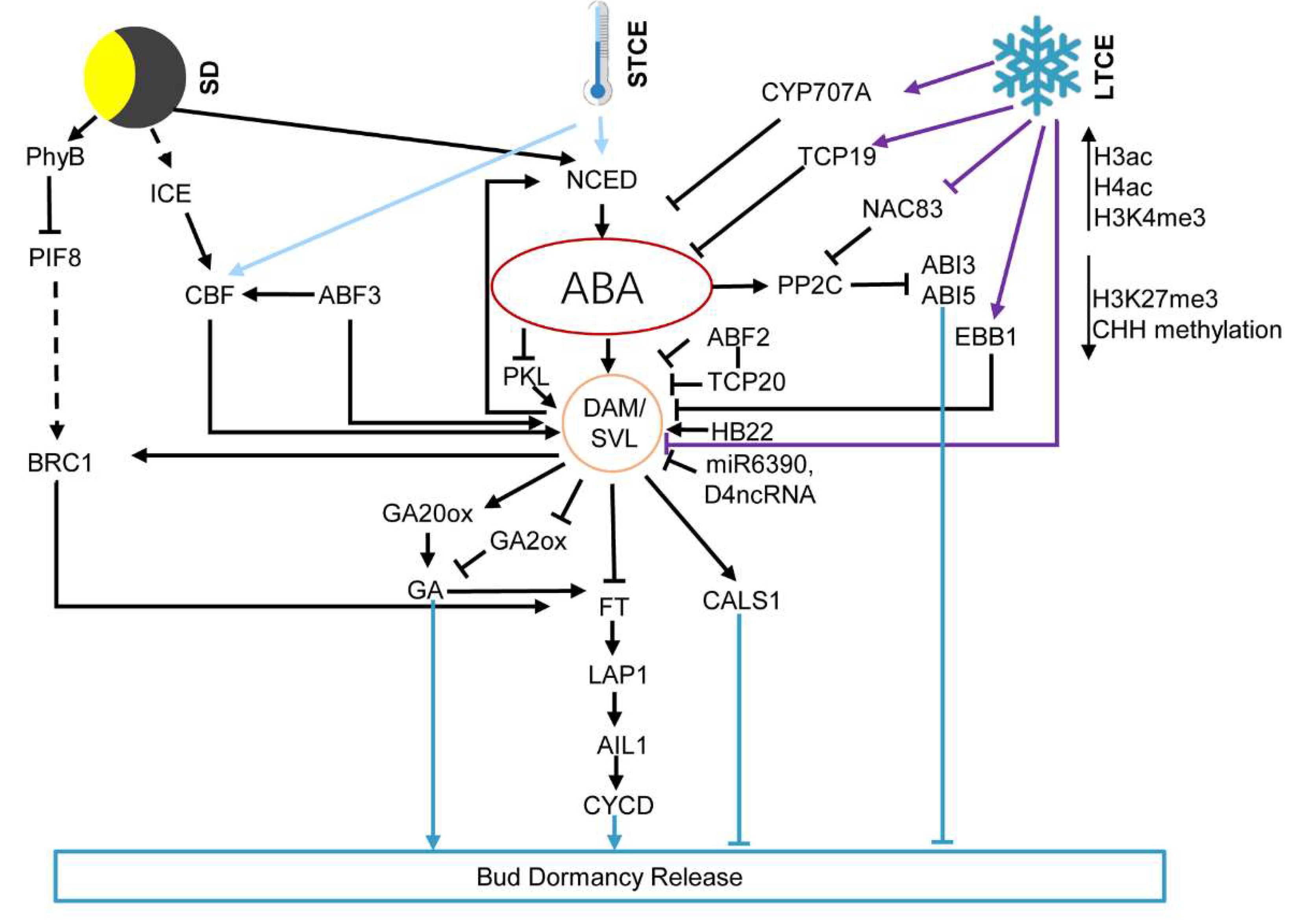
2. ABA Integrates Environments Signaling in Regulating Bud Dormancy
2.1. ABA Mediates Photoperiod Response of the Bud Dormancy
2.2. ABA Mediates Temperature Signals Regulated Bud Dormancy
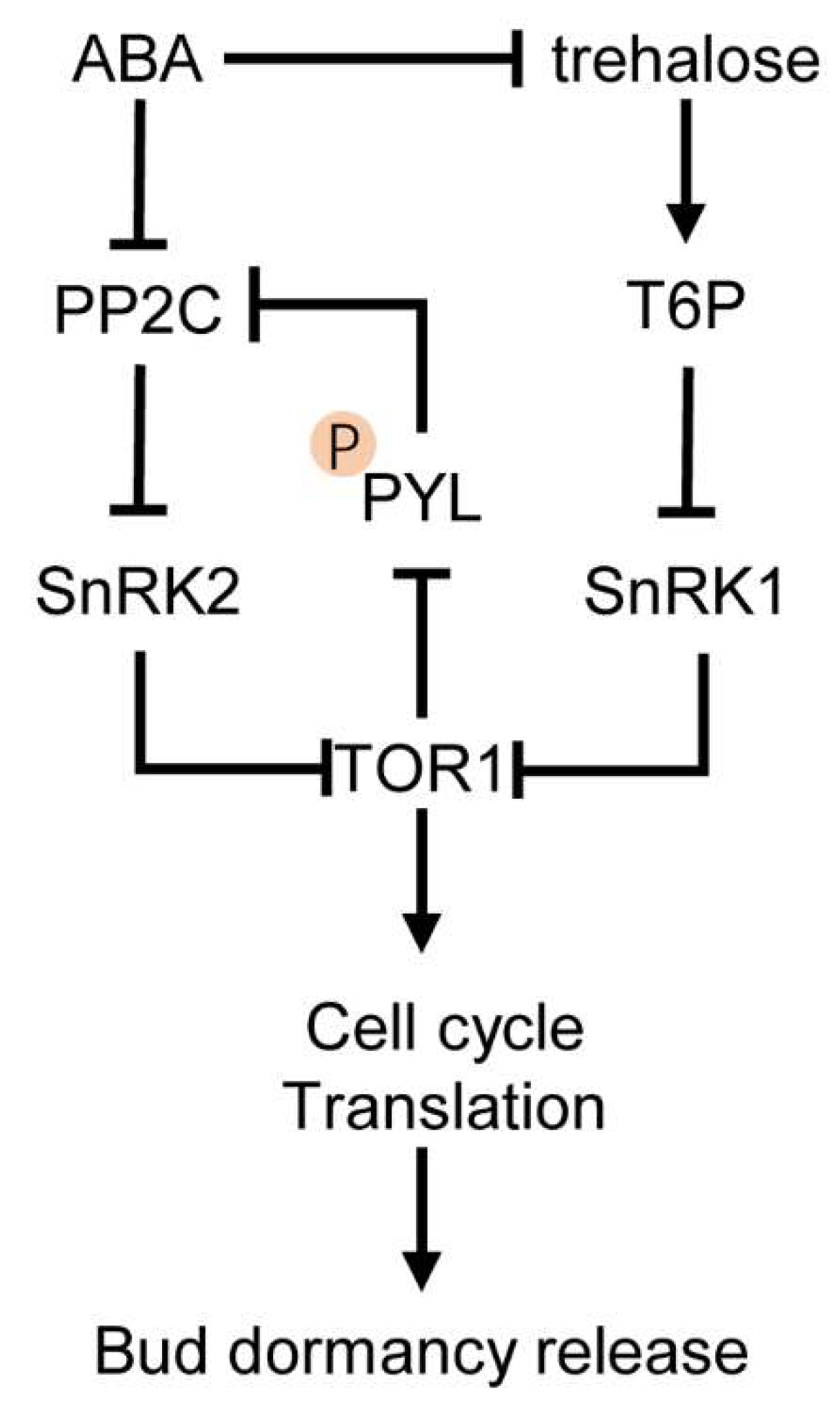
3. Cross-Talk between ABA and Sugars in Bud Dormancy
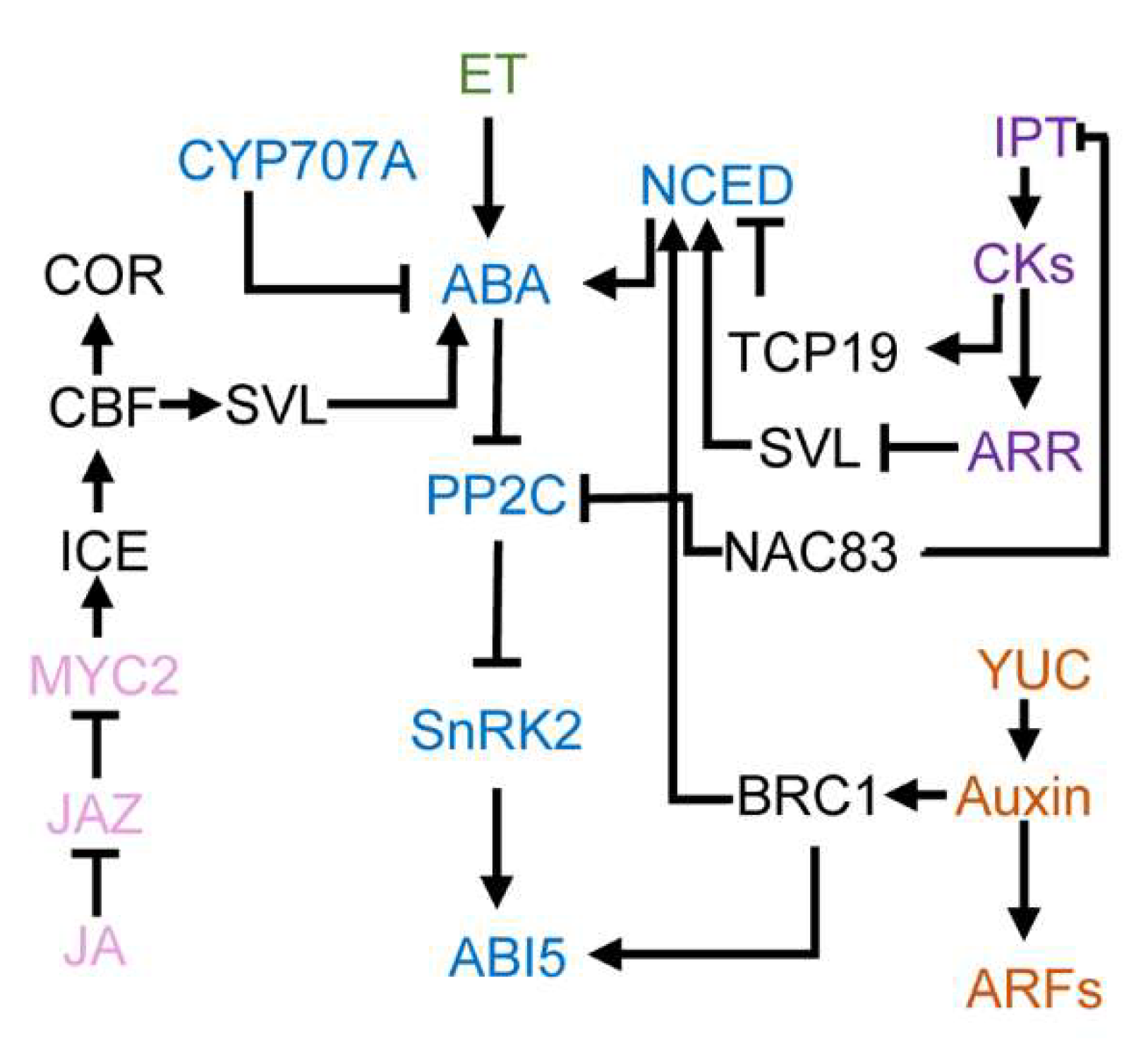
4. Cross-Talks between ABA and Other Hormones in Regulating Bud Dormancy
4.1. ABA and Gibberellins
4.2. ABA and Cytokinins
4.3. ABA and Auxin
4.4. ABA and Ethylene
4.5. ABA and Jasmonic Acid
5. Epigenetic Regulation of Bud Dormancy Mediated by ABA
6. Conclusions and Outlook
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kermode, A.R. Role of Abscisic Acid in Seed Dormancy. J. Plant Growth Regul. 2005, 24, 319–344. [Google Scholar] [CrossRef] [Green Version]
- Zheng, C.; Halaly, T.; Acheampong, A.K.; Takebayashi, Y.; Jikumaru, Y.; Kamiya, Y.; Or, E. Abscisic acid (ABA) regulates grape bud dormancy, and dormancy release stimuli may act through modification of ABA metabolism. J. Exp. Bot. 2015, 66, 1527–1542. [Google Scholar] [CrossRef] [PubMed]
- Walker, M.; Pérez, M.; Steinbrecher, T.; Gawthrop, F.; Pavlović, I.; Novák, O.; Tarkowská, D.; Strnad, M.; Marone, F.; Nakabayashi, K.; et al. Molecular mechanisms and hormonal regulation underpinning morphological dormancy: A case study using Apium graveolens (Apiaceae). Plant J. 2021. [Google Scholar] [CrossRef]
- Graeber, K.; Nakabayashi, K.; Miatton, E.; Leubner, G.; Soppe, W.J.J. Molecular mechanisms of seed dormancy. Plant Cell Environ. 2012, 35, 1769–1786. [Google Scholar] [CrossRef] [PubMed]
- Nambara, E.; Marion-Poll, A. Abscisic acid biosynthesis and catabolism. Annu. Rev. Plant Biol. 2005, 56, 165–185. [Google Scholar] [CrossRef] [Green Version]
- Chen, K.; Li, G.J.; Bressan, R.A.; Song, C.P.; Zhu, J.K.; Zhao, Y. Abscisic acid dynamics, signaling, and functions in plants. J. Integr. Plant Biol. 2000, 62, 25–54. [Google Scholar] [CrossRef] [Green Version]
- Koornneef, M.; Jorna, M.L.; Der Swan, D.L.C.B.-V.; Karssen, C.M. The isolation of abscisic acid (ABA) deficient mutants by selection of induced revertants in non-germinating gibberellin sensitive lines of Arabidopsis thaliana (L.) heynh. Theor. Appl. Genet. 1982, 61, 385–393. [Google Scholar] [CrossRef]
- Marin, E.; Nussaume, L.; Quesada, A.; Gonneau, M.; Sotta, B.; Hugueney, P.; Frey, A.; Marion-Poll, A. Molecular identification of zeaxanthin epoxidase of Nicotiana plumbaginifolia, a gene involved in abscisic acid biosynthesis and corresponding to the ABA locus of Arabidopsis thaliana. EMBO J. 1996, 15, 2331–2342. [Google Scholar] [CrossRef]
- Schwartz, S.H.; Qin, X.; Zeevaart, J.A. Elucidation of the Indirect Pathway of Abscisic Acid Biosynthesis by Mutants, Genes, and Enzymes. Plant Physiol. 2003, 131, 1591–1601. [Google Scholar] [CrossRef] [Green Version]
- Kushiro, T.; Okamoto, M.; Nakabayashi, K.; Yamagishi, K.; Kitamura, S.; Asami, T.; Hirai, N.; Koshiba, T.; Kamiya, Y.; Nambara, E. The Arabidopsis cytochrome P450 CYP707A encodes ABA 8′-hydroxylases: Key enzymes in ABA catabolism. EMBO J. 2004, 23, 1647–1656. [Google Scholar] [CrossRef] [Green Version]
- Saito, S.; Hirai, N.; Matsumoto, C.; Ohigashi, H.; Ohta, D.; Sakata, K.; Mizutani, M. Arabidopsis CYP707As Encode (+)-Abscisic Acid 8′-Hydroxylase, a Key Enzyme in the Oxidative Catabolism of Abscisic Acid. Plant Physiol. 2004, 134, 1439–1449. [Google Scholar] [CrossRef] [Green Version]
- Lee, K.H.; Piao, H.L.; Kim, H.Y.; Choi, S.M.; Jiang, F.; Hartung, W.; Hwang, I.; Kwak, J.M.; Lee, I.J.; Hwang, I. Activation of glucosidase via stress-induced polymerization rapidly increases active pools of abscisic acid. Cell 2006, 126, 1109–1120. [Google Scholar] [CrossRef] [Green Version]
- Xu, Z.-Y.; Lee, K.H.; Dong, T.; Jeong, J.C.; Jin, J.B.; Kanno, Y.; Kim, D.H.; Kim, S.Y.; Seo, M.; Bressan, R.A.; et al. A Vacuolar β-Glucosidase Homolog That Possesses Glucose-Conjugated Abscisic Acid Hydrolyzing Activity Plays an Important Role in Osmotic Stress Responses in Arabidopsis. Plant Cell 2012, 24, 2184–2199. [Google Scholar] [CrossRef] [Green Version]
- Dietz, K.J.; Sauter, A.; Wichert, K.; Messdaghi, D.; Hartung, W. Extracellular β-glucosidase activity in barley involved in the hydrolysis of ABA glucose conjugate in leaves. J. Exp. Bot. 2000, 51, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Szostkiewicz, I.; Korte, A.; Moes, D.; Yang, Y.; Christmann, A.; Grill, E. Regulators of PP2C Phosphatase Activity Function as Abscisic Acid Sensors. Science 2009, 324, 1064–1068. [Google Scholar] [CrossRef]
- Park, S.Y.; Fung, P.; Nishimura, N.; Jensen, D.R.; Fujii, H.; Zhao, Y.; Lumba, S.; Santiago, J.; Rodrigues, A.; Chow, T.F.F.; et al. Abscisic Acid Inhibits Type 2C Protein Phosphatases via the PYR/PYL Family of START Proteins. Science 2009, 324, 1068–1071. [Google Scholar] [CrossRef] [Green Version]
- Melcher, K.; Ng, L.-M.; Zhou, X.E.; Soon, F.-F.; Xu, Y.; Suino-Powell, K.M.; Park, S.-Y.; Weiner, J.J.; Fujii, H.; Chinnusamy, V.; et al. A gate–latch–lock mechanism for hormone signalling by abscisic acid receptors. Nature 2009, 462, 602–608. [Google Scholar] [CrossRef] [Green Version]
- Miyazono, K.-I.; Miyakawa, T.; Sawano, Y.; Kubota, K.; Kang, H.-J.; Asano, A.; Miyauchi, Y.; Takahashi, M.; Zhi, Y.; Fujita, Y.; et al. Structural basis of abscisic acid signalling. Nature 2009, 462, 609–614. [Google Scholar] [CrossRef]
- Hubbard, K.; Nishimura, N.; Hitomi, K.; Getzoff, E.D.; Schroeder, J.I. Early abscisic acid signal transduction mechanisms: Newly discovered components and newly emerging questions. Genes Dev. 2010, 24, 1695–1708. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rinne, P.L.; Welling, A.; Van Der Schoot, C. Perennial Life Style of Populus: Dormancy Cycling and Overwintering. In Genetics and Genomics of Populus; Springer: New York, NY, USA, 2009; pp. 171–200. [Google Scholar] [CrossRef]
- Horvath, D.P.; Anderson, J.V.; Chao, W.S.; Foley, M.E. Knowing when to grow: Signals regulating bud dormancy. Trends Plant Sci. 2003, 8, 534–540. [Google Scholar] [CrossRef] [PubMed]
- Perry, T.O. Dormancy of Trees in Winter. Science 1971, 171, 29–36. [Google Scholar] [CrossRef]
- Savage, J.A.; Chuine, I. Coordination of spring vascular and organ phenology in deciduous angiosperms growing in seasonally cold climates. New Phytol. 2021, 230, 1700–1715. [Google Scholar] [CrossRef]
- Liu, J.; Sherif, S.M. Hormonal Orchestration of Bud Dormancy Cycle in Deciduous Woody Perennials. Front. Plant Sci. 2019, 10, 1136. [Google Scholar] [CrossRef]
- Falavigna, V.D.S.; Guitton, B.; Costes, E.; Andrés, F. I Want to (Bud) Break Free: The Potential Role of DAM and SVP-Like Genes in Regulating Dormancy Cycle in Temperate Fruit Trees. Front. Plant Sci. 2019, 9, 1990. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, R.-Y.; Niimi, Y.; Han, D.-S. Changes in endogenous abscisic acid and soluble sugars levels during dormancy-release in bulbs of Lilium rubellum. Sci. Hortic. 2006, 111, 68–72. [Google Scholar] [CrossRef]
- Wu, J.; Jin, Y.; Liu, C.; Vonapartis, E.; Liang, J.; Wu, W.; Gazzarrini, S.; He, J.; Yi, M. GhNAC83 inhibits corm dormancy release by regulating ABA signaling and cytokinin biosynthesis in Gladiolus hybridus. J. Exp. Bot. 2018, 70, 1221–1237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yasin, H.J.; Bufler, G. Dormancy and sprouting in onion (Allium cepa L.) bulbs. I. Changes in carbohydrate metabolism. J. Hortic. Sci. Biotechnol. 2007, 82, 89–96. [Google Scholar] [CrossRef]
- Wu, J.; Seng, S.S.; Sui, J.J.; Vonapartis, E.; Luo, X.; Gong, B.H.; Liu, C.; Wu, C.Y.; Liu, C.; Zhang, F.Q.; et al. Gladiolus hybridus ABSCISIC ACID INSENSITIVE 5 (GhABI5) is an important transcription factor in ABA signaling that can enhance Gladiolus corm dormancy and Arabidopsis seed dormancy. Front. Plant Sci. 2015, 6, 960. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lazare, S.; Bechar, D.; Fernie, A.R.; Brotman, Y.; Zaccai, M. The proof is in the bulb: Glycerol influences key stages of lily development. Plant J. 2019, 97, 321–340. [Google Scholar] [CrossRef] [Green Version]
- Benkeblia, N.; Selselet-Attou, G. Effects of low temperatures on changes in oligosaccharides, phenolics and peroxidase in inner bud of onion Allium cepa L. during break of dormancy. Acta Agric. Scand. B Plant Soil Sci. 1999, 49, 98–102. [Google Scholar] [CrossRef]
- Campoy, J.A.; Ruiz, D.; Egea, J. Dormancy in temperate fruit trees in a global warming context: A review. Sci. Hortic. 2011, 130, 357–372. [Google Scholar] [CrossRef]
- Tylewicz, S.; Petterle, A.; Marttila, S.; Miskolczi, P.; Azeez, A.; Singh, R.K.; Immanen, J.; Mähler, N.; Hvidsten, T.R.; Eklund, D.M.; et al. Photoperiodic control of seasonal growth is mediated by ABA acting on cell-cell communication. Science 2018, 360, 212–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rohde, A.; Bhalerao, R.P. Plant dormancy in the perennial context. Trends Plant Sci. 2007, 12, 217–223. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Deng, X.W. CBF-phyB-PIF Module Links Light and Low Temperature Signaling. Trends Plant Sci. 2020, 25, 952–954. [Google Scholar] [CrossRef] [PubMed]
- Legris, M.; Klose, C.; Burgie, E.S.; Rojas, C.C.; Neme, M.; Hiltbrunner, A.; Wigge, P.A.; Schafer, E.; Vierstra, R.D.; Casal, J.J. Phytochrome B integrates light and temperature signals in Arabidopsis. Science 2016, 354, 897–900. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Paik, I.; Huq, E. Plant photoreceptors: Multi-functional sensory proteins and their signaling networks. Semin. Cell Dev. Biol. 2019, 92, 114–121. [Google Scholar] [CrossRef]
- Bohlenius, H.; Huang, T.; Charbonnel-Campaa, L.; Brunner, A.M.; Jansson, S.; Strauss, S.H.; Nilsson, O. CO/FT regulatory module controls timing of flowering and seasonal growth cessation in trees. Science 2006, 312, 1040–1043. [Google Scholar] [CrossRef] [Green Version]
- Karlberg, A.; Bakó, L.; Bhalerao, R.P. Short Day–Mediated Cessation of Growth Requires the Downregulation of AINTEGUMENTALIKE1 Transcription Factor in Hybrid Aspen. PLoS Genet. 2011, 7, e1002361. [Google Scholar] [CrossRef] [Green Version]
- Kumar, G.; Arya, P.; Gupta, K.; Randhawa, V.; Acharya, V.; Singh, A.K. Comparative phylogenetic analysis and transcriptional profiling of MADS-box gene family identified DAM and FLC-like genes in apple (Malusx domestica). Sci. Rep. 2016, 6, 20695. [Google Scholar] [CrossRef] [Green Version]
- Singh, R.K.; Miskolczi, P.; Maurya, J.P.; Bhalerao, R.P. A Tree Ortholog of SHORT VEGETATIVE PHASE Floral Repressor Mediates Photoperiodic Control of Bud Dormancy. Curr. Biol. 2019, 29, 128–133.e2. [Google Scholar] [CrossRef] [Green Version]
- Gomez-Soto, D.; Ramos-Sanchez, J.M.; Alique, D.; Conde, D.; Triozzi, P.M.; Perales, M.; Allona, I. Overexpression of a SOC1-Related Gene Promotes Bud Break in Ecodormant Poplars. Front. Plant Sci. 2021, 12, 670497. [Google Scholar] [CrossRef]
- Singh, R.K.; Maurya, J.P.; Azeez, A.; Miskolczi, P.; Tylewicz, S.; Stojkovic, K.; Delhomme, N.; Busov, V.; Bhalerao, R.P. A genetic network mediating the control of bud break in hybrid aspen. Nat. Commun. 2018, 9. [Google Scholar] [CrossRef] [Green Version]
- Busov, V.B. Plant Development: Dual Roles of Poplar SVL in Vegetative Bud Dormancy. Curr. Biol. 2019, 29, R68–R70. [Google Scholar] [CrossRef] [Green Version]
- Maurya, J.P.; Singh, R.; Miskolczi, P.C.; Prasad, A.N.; Jonsson, K.; Wu, F.; Bhalerao, R.P. Branching Regulator BRC1 Mediates Photoperiodic Control of Seasonal Growth in Hybrid Aspen. Curr. Biol. 2020, 30, 122–126.e2. [Google Scholar] [CrossRef]
- Rohde, A.; Prinsen, E.; De Rycke, R.; Engler, G.; Van Montagu, M.; Boerjan, W. PtABI3 Impinges on the Growth and Differentiation of Embryonic Leaves during Bud Set in Poplar. Plant Cell 2002, 14, 1885–1901. [Google Scholar] [CrossRef] [Green Version]
- Maurya, J.P.; Bhalerao, R.P. Photoperiod- and temperature-mediated control of growth cessation and dormancy in trees: A molecular perspective. Ann. Bot. 2017, 120, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Kamenetsky, R.; Okubo, H. Ornamental Geophytes: From Basic Science to Sustainable Production; CRC Press: Boca Raton, FL, USA, 2012. [Google Scholar]
- Heide, O.M.; Prestrud, A.K. Low temperature, but not photoperiod, controls growth cessation and dormancy induction and release in apple and pear. Tree Physiol. 2005, 25, 109–114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Q.; Gao, Y.; Wu, X.; Moriguchi, T.; Bai, S.; Teng, Y. Bud endodormancy in deciduous fruit trees: Advances and prospects. Hortic. Res. 2021, 8, 139. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Yan, X.; Yang, Q.; Ma, Y.; Yang, B.; Tian, J.; Teng, Y.; Bai, S. PpCBFs selectively regulate PpDAMs and contribute to the pear bud endodormancy process. Plant Mol. Biol. 2019, 99, 575–586. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Xu, G.; Zhao, X.; Zhang, Z.; Wang, X.; Liu, X.; Xiao, W.; Fu, X.; Chen, X.; Gao, D.; et al. Transcription factor TCP20 regulates peach bud endodormancy by inhibiting DAM5/DAM6 and interacting with ABF2. J. Exp. Bot. 2020, 71, 1585–1597. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Yang, B.; Li, J.; Wang, Y.; Tao, R.; Yang, F.; Wu, X.; Yan, X.; Ahmad, M.; Shen, J.; et al. ABA-responsive ABRE-BINDING FACTOR3 activates DAM3 expression to promote bud dormancy in Asian pear. Plant Cell Environ. 2020, 43, 1360–1375. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Zhang, B.; Li, Y.; Andre, D.; Nilsson, O. Phytochrome B and PHYTOCHROME INTERACTING FACTOR8 modulate seasonal growth in trees. New Phytol. 2021. [Google Scholar] [CrossRef]
- Azeez, A.; Zhao, Y.C.; Singh, R.K.; Yordanov, Y.S.; Dash, M.; Miskolczi, P.; Stojkovic, K.; Strauss, S.H.; Bhalerao, R.P.; Busov, V.B. EARLY BUD-BREAK 1 and EARLY BUD-BREAK 3 control resumption of poplar growth after winter dormancy. Nat. Commun. 2021, 12, 1123. [Google Scholar] [CrossRef]
- Yordanov, Y.; Ma, C.; Strauss, S.H.; Busov, V.B. EARLY BUD-BREAK 1 (EBB1) is a regulator of release from seasonal dormancy in poplar trees. Proc. Natl. Acad. Sci. USA 2014, 111, 10001–10006. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.; Wu, W.; Liang, J.; Jin, Y.; Gazzarrini, S.; He, J.; Yi, M. GhTCP19 Transcription Factor Regulates Corm Dormancy Release by Repressing GHNCED Expression in Gladiolus. Plant Cell Physiol. 2018, 60, 52–62. [Google Scholar] [CrossRef]
- Lastdrager, J.; Hanson, J.; Smeekens, S. Sugar signals and the control of plant growth and development. J. Exp. Bot. 2014, 65, 799–807. [Google Scholar] [CrossRef]
- Wang, M.; Pérez-Garcia, M.-D.; Davière, J.-M.; Barbier, F.; Ogé, L.; Gentilhomme, J.; Voisine, L.; Péron, T.; Launay-Avon, A.; Clément, G.; et al. Outgrowth of the axillary bud in rose is controlled by sugar metabolism and signalling. J. Exp. Bot. 2021, 72, 3044–3060. [Google Scholar] [CrossRef]
- Tsai, A.Y.-L.; Gazzarrini, S. Trehalose-6-phosphate and SnRK1 kinases in plant development and signaling: The emerging picture. Front. Plant Sci. 2014, 5, 119. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gibson, S.I. Control of plant development and gene expression by sugar signaling. Curr. Opin. Plant Biol. 2005, 8, 93–102. [Google Scholar] [CrossRef] [PubMed]
- León, P. Sugar and hormone connections. Trends Plant Sci. 2003, 8, 110–116. [Google Scholar] [CrossRef]
- Gazzarrini, S.; McCourt, P. Genetic interactions between ABA, ethylene and sugar signaling pathways. Curr. Opin. Plant Biol. 2001, 4, 387–391. [Google Scholar] [CrossRef]
- Arora, R.; Rowland, L.J.; Tanino, K. Induction and release of bud dormancy in woody perennials: A science comes of age. Hortscience 2003, 38, 911–921. [Google Scholar] [CrossRef] [Green Version]
- Anderson, J.V.; Gesch, R.W.; Jia, Y.; Chao, W.S.; Horvath, D.P. Seasonal shifts in dormancy status, carbohydrate metabolism, and related gene expression in crown buds of leafy spurge. Plant Cell Environ. 2005, 28, 1567–1578. [Google Scholar] [CrossRef]
- Chao, W.S.; Serpe, M.D.; Anderson, J.V.; Gesch, R.W.; Horvath, D.P. Sugars, hormones, and environment affect the dormancy status in underground adventitious buds of leafy spurge (Euphorbia esula). Weed Sci. 2006, 54, 59–68. [Google Scholar] [CrossRef]
- Chao, W.S.; Foley, M.E.; Horvath, D.P.; Anderson, J.V. Signals regulating dormancy in vegetative buds. Int. J. Plant Dev. Biol. 2007, 1, 49–56. [Google Scholar]
- Zhang, Z.; Zhuo, X.; Zhao, K.; Zheng, T.; Han, Y.; Yuan, C.; Zhang, Q. Transcriptome Profiles Reveal the Crucial Roles of Hormone and Sugar in the Bud Dormancy of Prunus mume. Sci. Rep. 2018, 8, 5090. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, C.-Q.; Li, Q.-H.; Jiang, X.-Q.; Fan, Y.-W.; Gao, J.-P.; Zhang, C.-Q. Dynamic changes in α-and β-amylase activities and gene expression in bulbs of the Oriental hybrid lily ‘Siberia’during dormancy release. J. Hortic. Sci. Biotechnol. 2015, 90, 753–759. [Google Scholar] [CrossRef]
- Chmielewski, F.M.; Gotz, K.; Homann, T.; Huschek, G.; Rawel, H. Identification of endodormancy release for cherries (Prunus avium, L.) by abscisic acid and sugars. J. Hortic. 2017, 4, 210. [Google Scholar] [CrossRef]
- Zhang, Y.; Yuan, Y.; Liu, Z.; Zhang, T.; Li, F.; Liu, C.; Gai, S. GA3 is superior to GA4 in promoting bud endodormancy release in tree peony (Paeonia suffruticosa) and their potential working mechanism. BMC Plant Biol. 2021, 21, 323. [Google Scholar] [CrossRef]
- Sonnewald, S.; Sonnewald, U. Regulation of potato tuber sprouting. Planta 2014, 239, 27–38. [Google Scholar] [CrossRef] [Green Version]
- Baena-Gonzalez, E.; Lunn, J.E. SnRK1 and trehalose 6-phosphate—Two ancient pathways converge to regulate plant metabolism and growth. Curr. Opin. Plant Biol. 2020, 55, 52–59. [Google Scholar] [CrossRef]
- Debast, S.; Nunes-Nesi, A.; Hajirezaei, M.R.; Hofmann, J.; Sonnewald, U.; Fernie, A.R.; Bornke, F. Altering Trehalose-6-Phosphate Content in Transgenic Potato Tubers Affects Tuber Growth and Alters Responsiveness to Hormones during Sprouting. Plant Physiol. 2011, 156, 1754–1771. [Google Scholar] [CrossRef] [Green Version]
- Murcia, G.; Pontin, M.; Reinoso, H.; Baraldi, R.; Bertazza, G.; Gomez-Talquenca, S.; Bottini, R.; Piccoli, P.N. ABA and GA 3 increase carbon allocation in different organs of grapevine plants by inducing accumulation of non-structural carbohydrates in leaves, enhancement of phloem area and expression of sugar transporters. Physiol. Plant. 2016, 156, 323–337. [Google Scholar] [CrossRef]
- Rubio, S.; Noriega, X.; Pérez, F.J. ABA promotes starch synthesis and storage metabolism in dormant grapevine buds. J. Plant Physiol. 2019, 234–235, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Wu, Y.; Sheen, J. TOR signaling in plants: Conservation and innovation. Development 2018, 145, dev160887. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Crepin, N.; Rolland, F. SnRK1 activation, signaling, and networking for energy homeostasis. Curr. Opin. Plant Biol. 2019, 51, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.C.; Zhao, Y.; Li, Z.P.; Hsu, C.C.; Liu, X.; Fu, L.W.; Hou, Y.J.; Du, Y.; Xie, S.J.; Zhang, C.G.; et al. Reciprocal Regulation of the TOR Kinase and ABA Receptor Balances Plant Growth and Stress Response. Mol. Cell 2018, 69, 100–112. [Google Scholar] [CrossRef] [Green Version]
- Rodrigues, A.; Adamo, M.; Crozet, P.; Margalha, L.; Confraria, A.; Martinho, C.; Elias, A.; Rabissi, A.; Lumbreras, V.; Gonzalez-Guzman, M.; et al. ABI1 and PP2CA Phosphatases Are Negative Regulators of Snf1-Related Protein Kinase1 Signaling in Arabidopsis. Plant Cell 2013, 25, 3871–3884. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Parada, F.; Noriega, X.; Dantas, D.; Bressan-Smith, R.; Pérez, F.J. Differences in respiration between dormant and non-dormant buds suggest the involvement of ABA in the development of endodormancy in grapevines. J. Plant Physiol. 2016, 201, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Shi, Z.; Halaly-Basha, T.; Zheng, C.; Schwager, M.S.; Wang, C.; Galbraith, D.W.; Ophir, R.; Pang, X.; Or, E. Identification of potential post-ethylene events in the signaling cascade induced by stimuli of bud dormancy release in grapevine. Plant J. 2020, 104, 1251–1268. [Google Scholar] [CrossRef] [PubMed]
- Lv, L.; Huo, X.; Wen, L.; Gao, Z.; Khalil-Ur-Rehman, M. Isolation and Role of PmRGL2 in GA-mediated Floral Bud Dormancy Release in Japanese Apricot (Prunus mume Siebold et Zucc.). Front. Plant Sci. 2018, 9, 27. [Google Scholar] [CrossRef]
- Tosetti, R.; Waters, A.; Chope, G.; Cools, K.; Alamar, M.; McWilliam, S.; Thompson, A.; Terry, L. New insights into the effects of ethylene on ABA catabolism, sweetening and dormancy in stored potato tubers. Postharvest Biol. Technol. 2020, 173, 111420. [Google Scholar] [CrossRef]
- Bhattacharya, A. Chapter 6—Effect of High-Temperature Stress on the Metabolism of Plant Growth Regulators. In Effect of High Temperature on Crop Productivity and Metabolism of Macro Molecules; Bhattacharya, A., Ed.; Academic Press: Cambridge, MA, USA, 2019; pp. 485–591. [Google Scholar] [CrossRef]
- Sun, T.-P. Gibberellin metabolism, perception and signaling pathways in Arabidopsis. In The Arabidopsis Book/American Society of Plant Biologists; ASPB Press: Rockville, MD, USA, 2008; Volume 6. [Google Scholar] [CrossRef]
- Zheng, C.; Acheampong, A.K.; Shi, Z.; Halaly, T.; Kamiya, Y.; Ophir, R.; Galbraith, D.W.; Or, E. Distinct gibberellin functions during and after grapevine bud dormancy release. J. Exp. Bot. 2018, 69, 1635–1648. [Google Scholar] [CrossRef] [Green Version]
- Liu, X.; Hou, X. Antagonistic Regulation of ABA and GA in Metabolism and Signaling Pathways. Front. Plant Sci. 2018, 9, 251. [Google Scholar] [CrossRef] [Green Version]
- Fuentes, S.; Ljung, K.; Sorefan, K.; Alvey, E.; Harberd, N.P.; Østergaard, L. Fruit Growth in Arabidopsis Occurs via DELLA-Dependent and DELLA-Independent Gibberellin Responses. Plant Cell 2012, 24, 3982–3996. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yamazaki, H.; Nishijima, T.; Koshioka, M.; Miura, H. Gibberellins do not act against abscisic acid in the regulation of bulb dormancy of Allium wakegi Araki. Plant Growth Regul. 2002, 36, 223–229. [Google Scholar] [CrossRef]
- Yang, Q.; Niu, Q.; Tang, Y.; Ma, Y.; Yan, X.; Li, J.; Tian, J.; Bai, S.; Teng, Y. PpyGAST1 is potentially involved in bud dormancy release by integrating the GA biosynthesis and ABA signaling in ‘Suli’ pear (Pyrus pyrifolia White Pear Group). Environ. Exp. Bot. 2019, 162, 302–312. [Google Scholar] [CrossRef]
- Ram, R.; Mukherjee, D.; Manuja, S. Plant Growth Regulators Affect the Development of Both Corms and Cormels in Gladiolus. HortScience 2002, 37, 343–344. [Google Scholar] [CrossRef]
- Lionakis, S.M.; Schwabe, W.W. Bud Dormancy in the Kiwi Fruit, Actinidia chinensis Planch. Ann. Bot. 1984, 54, 467–484. [Google Scholar] [CrossRef]
- Mauriat, M.; Sandberg, L.G.; Moritz, T. Proper gibberellin localization in vascular tissue is required to control auxin-dependent leaf development and bud outgrowth in hybrid aspen. Plant J. 2011, 67, 805–816. [Google Scholar] [CrossRef]
- Ni, J.; Gao, C.; Chen, M.-S.; Pan, B.-Z.; Ye, K.; Xu, Z.-F. Gibberellin Promotes Shoot Branching in the Perennial Woody PlantJatropha curcas. Plant Cell Physiol. 2015, 56, 1655–1666. [Google Scholar] [CrossRef] [Green Version]
- Ogawa, M.; Hanada, A.; Yamauchi, Y.; Kuwahara, A.; Kamiya, Y.; Yamaguchi, S. Gibberellin Biosynthesis and Response during Arabidopsis Seed Germination[W]. Plant Cell 2003, 15, 1591–1604. [Google Scholar] [CrossRef] [Green Version]
- Ito, A.; Tuan, P.A.; Saito, T.; Bai, S.; Kita, M.; Moriguchi, T. Changes in phytohormone content and associated gene expression throughout the stages of pear (Pyrus pyrifolia Nakai) dormancy. Tree Physiol. 2021, 41, 529–543. [Google Scholar] [CrossRef]
- Footitt, S.; Soler, I.D.; Clay, H.; Finch-Savage, W.E. Dormancy cycling in Arabidopsis seeds is controlled by seasonally distinct hormone-signaling pathways. Proc. Natl. Acad. Sci. USA 2011, 108, 20236–20241. [Google Scholar] [CrossRef] [Green Version]
- Gazzarrini, S.; Tsai, A.Y.-L. Hormone cross-talk during seed germination. Essays Biochem. 2015, 58, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Nambara, E.; Okamoto, M.; Tatematsu, K.; Yano, R.; Seo, M.; Kamiya, Y. Abscisic acid and the control of seed dormancy and germination. Seed Sci. Res. 2010, 20, 55–67. [Google Scholar] [CrossRef]
- Shi, L.; Gast, R.T.; Gopalraj, M.; E Olszewski, N. Characterization of a shoot-specific, GA3- and ABA-Regulated gene from tomato. Plant J. 1992, 2, 153–159. [Google Scholar] [CrossRef]
- Dong, Z.; Xiao, Y.; Govindarajulu, R.; Feil, R.; Siddoway, M.L.; Nielsen, T.; Lunn, J.E.; Hawkins, J.; Whipple, C.; Chuck, G. The regulatory landscape of a core maize domestication module controlling bud dormancy and growth repression. Nat. Commun. 2019, 10, 3810. [Google Scholar] [CrossRef]
- Argueso, C.T.; Ferreira, F.J.; Kieber, J.J. Environmental perception avenues: The interaction of cytokinin and environmental response pathways. Plant Cell Environ. 2009, 32, 1147–1160. [Google Scholar] [CrossRef]
- Kieber, J.J.; Schaller, G.E. Cytokinins. In The Arabidopsis Book/American Society of Plant Biologists; ASPB Press: Rockville, MD, USA, 2014; Volume 12, p. 0168. [Google Scholar] [CrossRef] [Green Version]
- Werner, T.; Motyka, V.; Laucou, V.; Smets, R.; Van, O.H.; Schmülling, T. Cytokinin-Deficient Transgenic Arabidopsis Plants Show Multiple Developmental Alterations Indicating Opposite Functions of Cytokinins in the Regulation of Shoot and Root Meristem Activity. Plant Cell 2003, 15, 2532–2550. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Krall, L.; Raschke, M.; Zenk, M.H.; Baron, C. The Tzs protein from Agrobacterium tumefaciens C58 produces zeatin riboside 5′-phosphate from 4-hydroxy-3-methyl-2-(E)-butenyl diphosphate and AMP. Sci. Direct. 2002, 527, 315–318. [Google Scholar]
- Sakakibara, H. CYTOKININS: Activity, Biosynthesis, and Translocation. Annu. Rev. Plant Biol. 2006, 57, 431–449. [Google Scholar] [CrossRef] [Green Version]
- Li, S.-M.; Zheng, H.-X.; Zhang, X.-S.; Sui, N. Cytokinins as central regulators during plant growth and stress response. Plant Cell Rep. 2021, 40, 271–282. [Google Scholar] [CrossRef] [PubMed]
- Werner, T.; Schmülling, T. Cytokinin action in plant development. Curr. Opin. Plant Biol. 2009, 12, 527–538. [Google Scholar] [CrossRef]
- Hwang, I.; Sheen, J. Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 2001, 413, 383–389. [Google Scholar] [CrossRef]
- Kim, D.-J.; Forst, S. Genomic analysis of the histidine kinase family in bacteria and archaea. Microbiology 2001, 147, 1197–1212. [Google Scholar] [CrossRef] [Green Version]
- To, J.P.; Kieber, J.J. Cytokinin signaling: Two-components and more. Trends Plant Sci. 2008, 13, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Arkhipov, D.V.; Lomin, S.N.; Myakushina, Y.A.; Savelieva, E.M.; Osolodkin, D.I.; Romanov, G.A. Modeling of Protein–Protein Interactions in Cytokinin Signal Transduction. Int. J. Mol. Sci. 2019, 20, 2096. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Li, L.; Ye, T.; Zhao, S.; Liu, Z.; Feng, Y.-Q.; Wu, Y. Cytokinin antagonizes ABA suppression to seed germination of Arabidopsis by downregulating ABI5 expression. Plant J. 2011, 68, 249–261. [Google Scholar] [CrossRef]
- Thirugnanasambantham, K.; Prabu, G.; Mandal, A.K.A. Synergistic effect of cytokinin and gibberellins stimulates release of dormancy in tea (Camellia sinensis (L.) O. Kuntze) bud. Physiol. Mol. Biol. Plants 2020, 26, 1035–1045. [Google Scholar] [CrossRef]
- Qiu, Y.; Guan, S.C.; Wen, C.; Li, P.; Gao, Z.; Chen, X. Auxin and cytokinin coordinate the dormancy and outgrowth of axillary bud in strawberry runner. BMC Plant Biol. 2019, 19, 528. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, A.; Senning, M.; Hedden, P.; Sonnewald, U.; Sonnewald, S. Reactivation of Meristem Activity and Sprout Growth in Potato Tubers Require Both Cytokinin and Gibberellin. Plant Physiol. 2010, 155, 776–796. [Google Scholar] [CrossRef] [Green Version]
- Turnbull, C.G.N.; Hanke, D.E. The control of bud dormancy in potato tubers. Planta 1985, 165, 359–365. [Google Scholar] [CrossRef]
- Alvim, R.; Hewett, E.W.; Saunders, P.F. Seasonal variation in the hormone content of willow: I. Changes in abscisic Acid content and cytokinin activity in the xylem sap. Plant Physiol. 1976, 57, 474–476. [Google Scholar] [CrossRef] [Green Version]
- Cutting, J.; Strydom, D.K.; Jacobs, G.; Bellstedt, D.U.; Weiler, E.W. Changes in Xylem Constituents in Response to Rest-breaking Agents Applied to Apple before Budbreak. J. Am. Soc. Hortic. Sci. 1991, 116, 680–683. [Google Scholar] [CrossRef] [Green Version]
- Yamane, H.; Wada, M.; Honda, C.; Matsuura, T.; Ikeda, Y.; Hirayama, T.; Osako, Y.; Gao-Takai, M.; Kojima, M.; Sakakibara, H.; et al. Overexpression of Prunus DAM6 inhibits growth, represses bud break competency of dormant buds and delays bud outgrowth in apple plants. PLoS ONE 2019, 14, e0214788. [Google Scholar] [CrossRef] [Green Version]
- Cattani, A.M.; da Silveira Falavigna, V.; Silveira, C.P.; Buffon, V.; Dos Santos Maraschin, F.; Pasquali, G.; Revers, L.F. Type-B cytokinin response regulators link hormonal stimuli and molecular responses during the transition from endo- to ecodormancy in apple buds. Plant Cell Rep. 2020, 39, 1687–1703. [Google Scholar] [CrossRef]
- Tran, L.-S.P.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Role of cytokinin responsive two-component system in ABA and osmotic stress signalings. Plant Signal. Behav. 2010, 5, 148–150. [Google Scholar] [CrossRef] [Green Version]
- Dobisova, T.; Hrdinova, V.; Cuesta, C.; Michlickova, S.; Urbankova, I.; Hejatkova, R.; Zadnikova, P.; Pernisova, M.; Benkova, E.; Hejatko, J. Light Controls Cytokinin Signaling via Transcriptional Regulation of Constitutively Active Sensor Histidine Kinase CKI1. Plant Physiol. 2017, 174, 387–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roman, H.; Girault, T.; Barbier, F.; Péron, T.; Brouard, N.; Pěnčík, A.; Novák, O.; Vian, A.; Sakr, S.; Lothier, J.; et al. Cytokinins Are Initial Targets of Light in the Control of Bud Outgrowth. Plant Physiol. 2016, 172, 489–509. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Corot, A.; Roman, H.; Douillet, O.; Autret, H.; Perez-Garcia, M.D.; Citerne, S.; Bertheloot, J.; Sakr, S.; Leduc, N.; Demotes-Mainard, S. Cytokinins and Abscisic Acid Act Antagonistically in the Regulation of the Bud Outgrowth Pattern by Light Intensity. Front. Plant Sci. 2017, 8, 1724. [Google Scholar] [CrossRef]
- Martínez-Fernández, I.; de Moura, S.M.; Alves-Ferreira, M.; Ferrándiz, C.; Balanzà, V. Identification of Players Controlling Meristem Arrest Downstream of the FRUITFULL-APETALA2 Pathway. Plant Physiol. 2020, 184, 945–959. [Google Scholar] [CrossRef]
- Di, D.W.; Zhang, C.; Luo, P. An, C.W.; Guo, G.Q. The biosynthesis of auxin: How many paths truly lead to IAA? Plant Growth Regul. 2016, 78, 275–285. [Google Scholar] [CrossRef]
- Fendrych, M.; Leung, J.; Friml, J. TIR1/AFB-Aux/IAA auxin perception mediates rapid cell wall acidification and growth of Arabidopsis hypocotyls. Elife 2016, 5, 19048. [Google Scholar] [CrossRef]
- Zhao, Y. Auxin Biosynthesis: A Simple Two-Step Pathway Converts Tryptophan to Indole-3-Acetic Acid in Plants. Mol. Plant 2012, 5, 334–338. [Google Scholar] [CrossRef] [Green Version]
- Leyser, O. Auxin Signaling. Plant Physiol. 2017, 176, 465–479. [Google Scholar] [CrossRef] [Green Version]
- Aloni, R.; Peterson, C.A. Auxin promotes dormancy callose removal from the phloem of Magnolia kobus and callose accumulation and earlywood vessel differentiation in Quercus Robur. J. Plant Res. 1997, 110, 37. [Google Scholar] [CrossRef] [PubMed]
- Qiu, Z.; Wan, L.; Chen, T.; Wan, Y.; He, X.; Lu, S.; Wang, Y.; Lin, J. The regulation of cambial activity in Chinese fir (Cunninghamia lanceolata) involves extensive transcriptome remodeling. New Phytol. 2013, 199, 708–719. [Google Scholar] [CrossRef] [PubMed]
- Nagar, P.K.; Sood, S. Changes in endogenous auxins during winter dormancy in tea (Camellia sinensis L.) O. Kuntze. Acta Physiol. Plant. 2006, 28, 165–169. [Google Scholar] [CrossRef]
- Noriega, X.; Pérez, F.J. ABA Biosynthesis Genes are Down-regulated While Auxin and Cytokinin Biosynthesis Genes are Up-regulated During the Release of Grapevine Buds from Endodormancy. J. Plant Growth Regul. 2017, 36, 814–823. [Google Scholar] [CrossRef]
- Anderson, J.V.; Dogramaci, M.; Horvath, D.P.; Foley, M.E.; Chao, W.S.; Suttle, J.C.; Thimmapuram, J.; Hernandez, A.G.; Ali, S.; Mikel, M.A. Auxin and ABA act as central regulators of developmental networks associated with paradormancy in Canada thistle (Cirsium arvense). Funct. Integr. Genom. 2012, 12, 515–531. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Jiao, Y. Regulation of Axillary Meristem Initiation by Transcription Factors and Plant Hormones. Front. Plant Sci. 2016, 7, 183. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tantikanjana, T. Control of axillary bud initiation and shoot architecture in Arabidopsis through the SUPERSHOOT gene. Genes Dev. 2011, 15, 1577–1588. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Müller, D.; Leyser, O. Auxin, cytokinin and the control of shoot branching. Ann. Bot. 2011, 107, 1203–1212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Young, N.F.; Ferguson, B.; Antoniadi, I.; Bennett, M.H.; Beveridge, C.; Turnbull, C.G. Conditional Auxin Response and Differential Cytokinin Profiles in Shoot Branching Mutants. Plant Physiol. 2014, 165, 1723–1736. [Google Scholar] [CrossRef] [Green Version]
- Brewer, P.B.; Dun, E.A.; Ferguson, B.J.; Rameau, C.; Beveridge, C.A. Strigolactone acts downstream of auxin to regulate bud outgrowth in pea and Arabidopsis. Plant Physiol. 2009, 150, 482–493. [Google Scholar] [CrossRef] [Green Version]
- Chatfield, S.P.; Stirnberg, P.; Forde, B.G.; Leyser, O. The hormonal regulation of axillary bud growth in Arabidopsis. Plant J. 2000, 24, 159–169. [Google Scholar] [CrossRef]
- Rameau, C.; Bertheloot, J.; LeDuc, N.; Andrieu, B.; Foucher, F.; Sakr, S. Multiple pathways regulate shoot branching. Front. Plant Sci. 2015, 5, 741. [Google Scholar] [CrossRef] [Green Version]
- Le Bris, M.; Michaux-Ferrière, N.; Jacob, Y.; Poupet, A.; Barthe, P.; Guigonis, J.M.; Le Page-Degivry, M.T. Regulation of bud dormancy by manipulation of ABA in isolated buds of Rosa hybrida cultured in vitro. Funct. Plant Biol. 1999, 26, 273–281. [Google Scholar] [CrossRef]
- Arney, S.E.; Mitchell, D.L. The Effect of Abscisic Acid on Stem Elongation and Correlative Inhibition. New Phytol. 1969, 68, 1001–1015. [Google Scholar] [CrossRef]
- Arend, M.; Schnitzler, J.P.; Ehlting, B.; Hansch, R.; Lange, T.; Rennenberg, H.; Himmelbach, A.; Grill, E.; Fromm, J. Expression of the Arabidopsis mutant ABI1 gene alters abscisic acid sensitivity, stomatal development, and growth morphology in gray poplars. Plant Physiol. 2009, 151, 2110–2119. [Google Scholar] [CrossRef] [Green Version]
- Yao, C.; Finlayson, S.A. Abscisic Acid Is a General Negative Regulator of Arabidopsis Axillary Bud Growth. Plant Physiol. 2015, 169, 611–626. [Google Scholar] [CrossRef] [Green Version]
- Grandío, E.G.; Poza-Carrión, C.; Sorzano, C.O.S.; Cubas, P. BRANCHED1 Promotes Axillary Bud Dormancy in Response to Shade in Arabidopsis. Plant Cell 2013, 25, 834–850. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grandío, E.G.; Cubas, P. Identification of gene functions associated to active and dormant buds in Arabidopsis. Plant Signal. Behav. 2014, 9, e27994. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- González-Grandío, E.; Pajoro, A.; Franco-Zorrilla, J.M.; Tarancón, C.; Cubas, P. Abscisic acid signaling is controlled by a BRANCHED1/HD-ZIP I cascade in Arabidopsis axillary buds. Proc. Natl. Acad. Sci. USA 2016, 114, E245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aguilar-Martinez, J.A.; Poza-Carrion, C.; Cubas, P. Arabidopsis BRANCHED1 Acts as an Integrator of Branching Signals within Axillary Buds. J. Plant Cell 2007, 19, 458–472. [Google Scholar] [CrossRef]
- Johnson, P.R.; Ecker, J. The ethylene gas signal transduction pathway: A Molecular Perspective. Annu. Rev. Genet. 1998, 32, 227–254. [Google Scholar] [CrossRef]
- Bleecker, A.B.; Kende, H. Ethylene: A gaseous signal molecule in plants. Annu. Rev. Cell Dev. Biol. 2000, 16, 1–18. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, S.F. Ethylene biosynthesis and its regulation in higher plants. Annu. Rev. Plant Physiol. 1984, 35, 155–189. [Google Scholar] [CrossRef]
- Kende, H. Ethylene Biosynthesis. Annu. Rev. Plant Biol. 1993, 44, 283–307. [Google Scholar] [CrossRef]
- Wang, K.L.-C.; Li, H.; Ecker, J.R. Ethylene Biosynthesis and Signaling Networks. Plant Cell 2002, 14, S131–S151. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iqbal, N.; Trivellini, A.; Masood, A.; Ferrante, A.; Khan, N.A. Current understanding on ethylene signaling in plants: The influence of nutrient availability. Plant Physiol. Biochem. 2013, 73, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Kieber, J.J.; Rothenberg, M.; Roman, G.; Feldmann, K.A.; Ecker, J. CTR1, a negative regulator of the ethylene response pathway in arabidopsis, encodes a member of the Raf family of protein kinases. Cell 1993, 72, 427–441. [Google Scholar] [CrossRef]
- Huang, Y.; Hui, L.; Hutchison, C.E.; Laskey, J.; Kieber, J.J. Biochemical and functional analysis of CTR1, a protein kinase that negatively regulates ethylene signaling in Arabidopsis. Plant J. 2010, 33, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Ecker, J.R. The ethylene signaling pathway: New insights. Curr. Opin. Plant Biol. 2004, 7, 40–49. [Google Scholar] [CrossRef] [PubMed]
- Alonso, J.M.; Hirayama, T.; Roman, G.; Nourizadeh, S.; Ecker, J.R. EIN2, a bifunctional transducer of ethylene and stress responses in Arabidopsis. Science 1999, 284, 2148–2152. [Google Scholar] [CrossRef]
- An, F.; Zhao, Q.; Ji, Y.; Li, W.; Guo, H. Ethylene-induced stabilization of ETHYLENE INSENSITIVE3 and EIN3-LIKE1 is mediated by proteasomal degradation of EIN3 binding F-box 1 and 2 that requires EIN2 in arabidopsis. Plant Cell 2010, 22, 2384–2401. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.J.; Hong, S.H.; Kim, Y.W.; Lee, I.H.; Jun, J.H.; Phee, B.-K.; Rupak, T.; Jeong, H.; Lee, Y.; Hong, B.S.; et al. Gene regulatory cascade of senescence-associated NAC transcription factors activated by ETHYLENE-INSENSITIVE2-mediated leaf senescence signalling in Arabidopsis. J. Exp. Bot. 2014, 65, 4023–4036. [Google Scholar] [CrossRef] [Green Version]
- Solano, R.; Stepanova, A.; Chao, Q.; Ecker, J.R. Nuclear events in ethylene signaling: A transcriptional cascade mediated by ETHYLENE-INSENSITIVE3 and ETHYLENE-RESPONSE-FACTOR1. Genes Dev. 1998, 12, 3703–3714. [Google Scholar] [CrossRef] [Green Version]
- Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AP2/ERF family transcription factors in plant abiotic stress responses. Biochim. et Biophys. Acta (BBA) Bioenerg. 2012, 1819, 86–96. [Google Scholar] [CrossRef]
- Howe, G.T.; Horvath, D.P.; Dharmawardhana, P.; Priest, H.D.; Mockler, T.C.; Strauss, S.H. Extensive Transcriptome Changes During Natural Onset and Release of Vegetative Bud Dormancy in Populus. Front. Plant Sci. 2015, 6, 989. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suttle, J.C. Involvement of ethylene in potato microtuber dormancy. Plant Physiol. 1998, 118, 843–848. [Google Scholar] [CrossRef] [Green Version]
- Alamar, M.C.; Anastasiadi, M.; Lopez-Cobollo, R.; Bennett, M.H.; Thompson, A.J.; Turnbull, C.G.N.; Mohareb, F.; Terry, L.A. Transcriptome and phytohormone changes associated with ethylene-induced onion bulb dormancy. Postharvest Biol. Technol. 2020, 168, 111267. [Google Scholar] [CrossRef]
- Ruonala, R.; Rinne, P.L.; Baghour, M.; Moritz, T.; Tuominen, H.; Kangasjärvi, J. Transitions in the functioning of the shoot apical meristem in birch (Betula pendula) involve ethylene. Plant J. 2006, 46, 628–640. [Google Scholar] [CrossRef] [PubMed]
- Dubois, M.; Broeck, L.V.D.; Claeys, H.; Van Vlierberghe, K.; Matsui, M.; Inzé, D. The ETHYLENE RESPONSE FACTORs ERF6 and ERF11 Antagonistically Regulate Mannitol-Induced Growth Inhibition in Arabidopsis. Plant Physiol. 2015, 169, 166–179. [Google Scholar] [CrossRef] [PubMed]
- Pierik, R.; Cuppens, M.L.C.; Voesenek, L.A.C.J.; Visser, E.J.W. Interactions between Ethylene and Gibberellins in Phytochrome-Mediated Shade Avoidance Responses in Tobacco. J. Plant Physiol. 2004, 136, 2928–2936. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ophir, R.; Pang, X.; Halaly, T.; Venkateswari, J.; Lavee, S.; Galbraith, D.; Or, E. Gene-expression profiling of grape bud response to two alternative dormancy-release stimuli expose possible links between impaired mitochondrial activity, hypoxia, ethylene-ABA interplay and cell enlargement. Plant Mol. Biol. 2009, 71, 403–423. [Google Scholar] [CrossRef]
- Mathiason, K.; He, D.; Grimplet, J.; Venkateswari, J.; Galbraith, D.W.; Or, E.; Fennell, A. Transcript profiling in Vitis riparia during chilling requirement fulfillment reveals coordination of gene expression patterns with optimized bud break. Funct. Integr. Genom. 2009, 9, 81–96. [Google Scholar] [CrossRef]
- Sudawan, B.; Chang, C.S.; Chao, H.F.; Ku, M.; Yen, Y.F. Hydrogen cyanamide breaks grapevine bud dormancy in the summer through transient activation of gene expression and accumulation of reactive oxygen and nitrogen species. BMC Plant Biol. 2016, 16, 202. [Google Scholar] [CrossRef] [Green Version]
- Gao, Y.; Yang, Q.; Yan, X.; Wu, X.; Yang, F.; Li, J.; Wei, J.; Ni, J.; Ahmad, M.; Bai, S.; et al. High-quality genome assembly of ‘Cuiguan’ pear (Pyrus pyrifolia) as a reference genome for identifying regulatory genes and epigenetic modifications responsible for bud dormancy. Hortic. Res. 2021, 8, 197. [Google Scholar] [CrossRef]
- Browse, J. Jasmonate passes muster: A receptor and targets for the defense hormone. Annu. Rev. Plant Biol. 2009, 60, 183–205. [Google Scholar] [CrossRef] [PubMed]
- Sherif, S.; El-Sharkawy, I.; Mathur, J.; Ravindran, P.; Kumar, P.; Paliyath, G.; Jayasankar, S. A stable JAZ protein from peach mediates the transition from outcrossing to self-pollination. BMC Biol. 2015, 13, 11. [Google Scholar] [CrossRef] [Green Version]
- Goossens, J.; Fernández-Calvo, P.; Schweizer, F.; Goossens, A. Jasmonates: Signal transduction components and their roles in environmental stress responses. Plant Mol. Biol. 2016, 91, 673–689. [Google Scholar] [CrossRef]
- Hu, Y.; Jiang, Y.; Han, X.; Wang, H.; Pan, J.; Yu, D. Jasmonate regulates leaf senescence and tolerance to cold stress: Crosstalk with other phytohormones. J. Exp. Bot. 2017, 68, 1361–1369. [Google Scholar] [CrossRef] [PubMed]
- Swiatek, A.; Lenjou, M.; Van Bockstaele, D.; Inze, D.; Van Onckelen, H. Differential effect of jasmonic acid and abscisic acid on cell cycle progression in tobacco BY-2 cells. Plant Physiol. 2002, 128, 201–211. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Turner, J.G. Wound-Induced Endogenous Jasmonates Stunt Plant Growth by Inhibiting Mitosis. PLoS ONE 2008, 3, e3699. [Google Scholar] [CrossRef] [PubMed]
- Wilen, R.W.; Van Rooijen, G.J.H.; Pearce, D.W.; Pharis, R.P.; Holbrook, L.A.; Moloney, M.M. Effects of Jasmonic Acid on Embryo-Specific Processes in Brassica and Linum Oilseeds. Plant Physiol. 1991, 95, 399–405. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ellis, C.; Turner, J.G. A conditionally fertile coi1 allele indicates cross-talk between plant hormone signalling pathways in Arabidopsis thaliana seeds and young seedlings. Planta 2002, 215, 549–556. [Google Scholar] [CrossRef]
- Juvany, M.; Müller, M.; Munné-Bosch, S. Bud vigor, budburst lipid peroxidation, and hormonal changes during bud development in healthy and moribund beech (Fagus sylvatica L.) trees. Trees 2015, 29, 1781–1790. [Google Scholar] [CrossRef]
- Hao, X.; Yang, Y.; Yue, C.; Wang, L.; Horvath, D.P.; Wang, X.C. Comprehensive Transcriptome Analyses Reveal Differential Gene Expression Profiles of Camellia sinensis Axillary Buds at Para-, Endo-, Ecodormancy, and Bud Flush Stages. Front. Plant Sci. 2017, 8, 553. [Google Scholar] [CrossRef] [Green Version]
- Suttle, J.C.; Huckle, L.L.; Lulai, E.C. The effects of dormancy status on the endogenous contents and biological activities of jasmonic acid, n-(jasmonoyl)-isoleucine, and tuberonic acid in potato tubers. Am. J. Potato Res. 2011, 88, 283–293. [Google Scholar] [CrossRef]
- Fan, X.Y.; Yang, Y.; Li, M.; Fu, L.L.; Zang, Y.Q.; Wang, C.X.; Hao, T.Y.; Sun, H.M. Transcriptomics and targeted metabolomics reveal the regulatory network of Lilium davidii var. unicolor during bulb dormancy release. Planta 2021, 254, 59. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.L.; Wang, J.N.; Shan, W.; Fan, J.G.; Kuang, J.F.; Wu, K.Q.; Li, X.P.; Chen, W.X.; He, F.Y.; Chen, J.Y.; et al. Induction of jasmonate signalling regulators MaMYC2s and their physical interactions with MaICE1 in methyl jasmonate-induced chilling tolerance in banana fruit. Plant Cell Environ. 2012, 36, 30–51. [Google Scholar] [CrossRef]
- Hu, Y.; Jiang, L.; Wang, F.; Yu, D. Jasmonate Regulates the INDUCER OF CBF EXPRESSION–C-REPEAT BINDING FACTOR/DRE BINDING FACTOR1 Cascade and Freezing Tolerance in Arabidopsis. Plant Cell 2013, 25, 2907–2924. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cavalli, G.; Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 2019, 571, 489–499. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothkegel, K.; Sánchez, E.; Montes, C.; Greve, M.; Tapia, S.; Bravo, S.; Prieto, H.; Almeida, A.M. DNA methylation and small interference RNAs participate in the regulation of MADS-box genes involved in dormancy in sweet cherry (Prunus avium L.). Tree Physiol. 2017, 37, 1739–1751. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.A.; Kanno, T.; Matzke, A.J. RNA-Directed DNA Methylation: The Evolution of a Complex Epigenetic Pathway in Flowering Plants. Annu. Rev. Plant Biol. 2015, 66, 243–267. [Google Scholar] [CrossRef]
- Santamaría, M.; Hasbún, R.; Valera, M.; Meijón, M.; Valledor, L.; Rodríguez, J.L.; Toorop, P.E.; Canal-Villanueva, M.J.F.; Rodríguez, R. Acetylated H4 histone and genomic DNA methylation patterns during bud set and bud burst in Castanea sativa. J. Plant Physiol. 2009, 166, 1360–1369. [Google Scholar] [CrossRef]
- Rothkegel, K.; Sandoval, P.; Soto, E.; Ulloa, L.; Riveros, A.; Lillo-Carmona, V.; Caceres-Molina, J.; Almeida, A.M.; Meneses, C. Dormant but Active: Chilling Accumulation Modulates the Epigenome and Transcriptome of Prunus avium during Bud Dormancy. Front. Plant Sci. 2020, 11, 1115. [Google Scholar] [CrossRef]
- Chinnusamy, V.; Gong, Z.; Zhu, J.K. Abscisic acid-mediated epigenetic processes in plant development and stress responses. J. Integr. Plant Biol. 2008, 50, 1187–1195. [Google Scholar] [CrossRef]
- Albini, S.; Zakharova, V.; Ait-Si-Ali, S. Chapter 3—Histone Modifications. In Epigenetics and Regeneration; Palacios, D., Ed.; Academic Press: Cambridge, MA, USA, 2019; Volume 11, pp. 47–72. [Google Scholar] [CrossRef]
- David Law, R.; Suttle, J.C. Changes in histone H3 and H4 multi-acetylation during natural and forced dormancy break in potato tubers. Physiol. Plant 2004, 120, 642–649. [Google Scholar] [CrossRef] [PubMed]
- de la Fuente, L.; Conesa, A.; Lloret, A.; Badenes, M.L.; Ríos, G. Genome-wide changes in histone H3 lysine 27 trimethylation associated with bud dormancy release in peach. Tree Genet Genomes 2015, 11, 45. [Google Scholar] [CrossRef]
- Leida, C.; Conesa, A.; Llacer, G.; Badenes, M.L.; Rios, G. Histone modifications and expression of DAM6 gene in peach are modulated during bud dormancy release in a cultivar-dependent manner. New Phytol. 2012, 193, 67–80. [Google Scholar] [CrossRef] [PubMed]
- Horvath, D.P.; Sung, S.; Kim, D.; Chao, W.; Anderson, J. Characterization, expression and function of DORMANCY ASSOCIATED MADS-BOX genes from leafy spurge. Plant Mol. Biol. 2010, 73, 169–179. [Google Scholar] [CrossRef]
- Wu, R.; Wang, T.; Richardson, A.C.; Allan, A.C.; Macknight, R.C.; Varkonyi-Gasic, E. Histone modification and activation by SOC1-like and drought stress-related transcription factors may regulate AcSVP2 expression during kiwifruit winter dormancy. Plant Sci. 2019, 281, 242–250. [Google Scholar] [CrossRef]
- Vimont, N.; Quah, F.X.; Schöepfer, D.G.; Roudier, F.; Dirlewanger, E.; Wigge, P.A.; Wenden, B.; Cortijo, S. ChIP-seq and RNA-seq for complex and low-abundance tree buds reveal chromatin and expression co-dynamics during sweet cherry bud dormancy. Tree Genet Genomes 2019, 16, 9. [Google Scholar] [CrossRef] [Green Version]
- Bertrand-Lehouillier, V.; Legault, L.-M.; McGraw, S. Endocrine Epigenetics, Epigenetic Profiling and Biomarker Identification. In Encyclopedia of Endocrine Diseases, 2nd ed.; Huhtaniemi, I., Martini, L., Eds.; Academic Press: Cambridge, MA, USA, 2019; pp. 31–35. [Google Scholar] [CrossRef]
- Yu, J.; Bennett, D.; Dardick, C.; Zhebentyayeva, T.; Abbott, A.G.; Liu, Z.; Staton, M.E. Genome-Wide Changes of Regulatory Non-Coding RNAs Reveal Pollen Development Initiated at Ecodormancy in Peach. Front. Mol. Biosci. 2021, 8, 612881. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Liu, J.; Liang, Q.; Zhang, Y.; Kang, K.; Wang, W.; Feng, Y.; Wu, S.; Yang, C.; Li, Y. Genome-wide analysis of long noncoding RNAs affecting floral bud dormancy in pears in response to cold stress. Tree Physiol. 2021, 41, 771–790. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Xu, Q.; Yu, H.; Ma, H.; Li, X.; Yang, J.; Chu, J.; Xie, Q.; Wang, Y.; Smith, S.M.; et al. Strigolactone and Karrikin Signaling Pathways Elicit Ubiquitination and Proteolysis of SMXL2 to Regulate Hypocotyl Elongation in Arabidopsis. Plant Cell 2020, 32, 2251–2270. [Google Scholar] [CrossRef]
- Zhao, L.-H.; Zhou, X.E.; Wu, Z.-S.; Yi, W.; Xu, Y.; Li, S.; Xu, T.; Liu, Y.; Chen, R.-Z.; Kovach, A.; et al. Crystal structures of two phytohormone signal-transducing α/β hydrolases: Karrikin-signaling KAI2 and strigolactone-signaling DWARF14. Cell Res. 2013, 23, 436–439. [Google Scholar] [CrossRef]
- Imanishi, H.; Fortanier, E.J. Effects of exposing freesia corms to ethylene or to smoke on dormancy-breaking and flowering. Sci. Hortic. 1983, 18, 381–389. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pan, W.; Liang, J.; Sui, J.; Li, J.; Liu, C.; Xin, Y.; Zhang, Y.; Wang, S.; Zhao, Y.; Zhang, J.; et al. ABA and Bud Dormancy in Perennials: Current Knowledge and Future Perspective. Genes 2021, 12, 1635. https://doi.org/10.3390/genes12101635
Pan W, Liang J, Sui J, Li J, Liu C, Xin Y, Zhang Y, Wang S, Zhao Y, Zhang J, et al. ABA and Bud Dormancy in Perennials: Current Knowledge and Future Perspective. Genes. 2021; 12(10):1635. https://doi.org/10.3390/genes12101635
Chicago/Turabian StylePan, Wenqiang, Jiahui Liang, Juanjuan Sui, Jingru Li, Chang Liu, Yin Xin, Yanmin Zhang, Shaokun Wang, Yajie Zhao, Jie Zhang, and et al. 2021. "ABA and Bud Dormancy in Perennials: Current Knowledge and Future Perspective" Genes 12, no. 10: 1635. https://doi.org/10.3390/genes12101635
APA StylePan, W., Liang, J., Sui, J., Li, J., Liu, C., Xin, Y., Zhang, Y., Wang, S., Zhao, Y., Zhang, J., Yi, M., Gazzarrini, S., & Wu, J. (2021). ABA and Bud Dormancy in Perennials: Current Knowledge and Future Perspective. Genes, 12(10), 1635. https://doi.org/10.3390/genes12101635





