GWAS Meta-Analysis Reveals Shared Genes and Biological Pathways between Major Depressive Disorder and Insomnia
Abstract
:1. Introduction
2. Materials and Methods
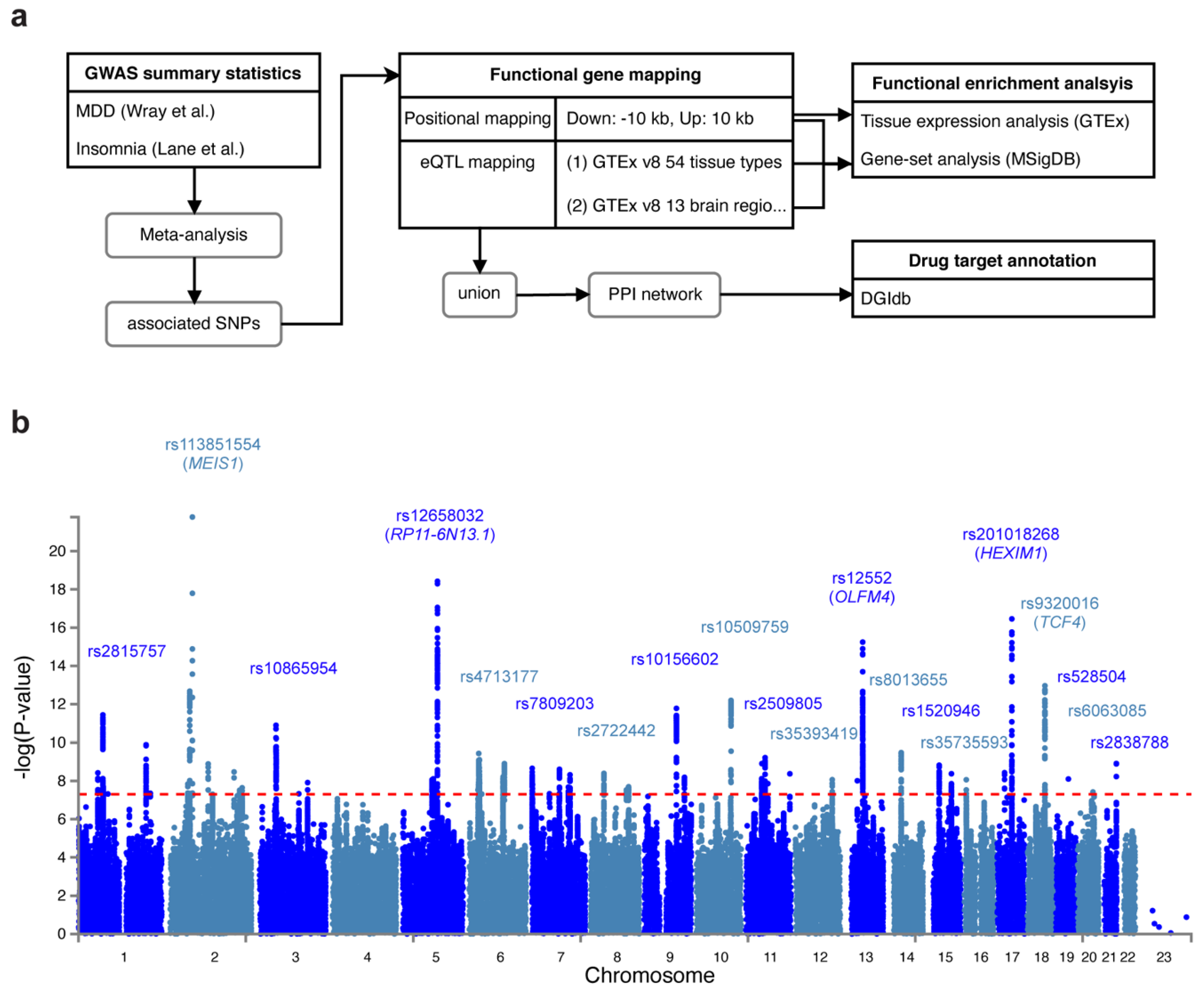
2.1. GWAS Data and Meta-Analysis
2.2. Identification of Candidate SNPs, Gene Mapping and Functional Annotation
2.3. MAGMA Gene-Based Tests
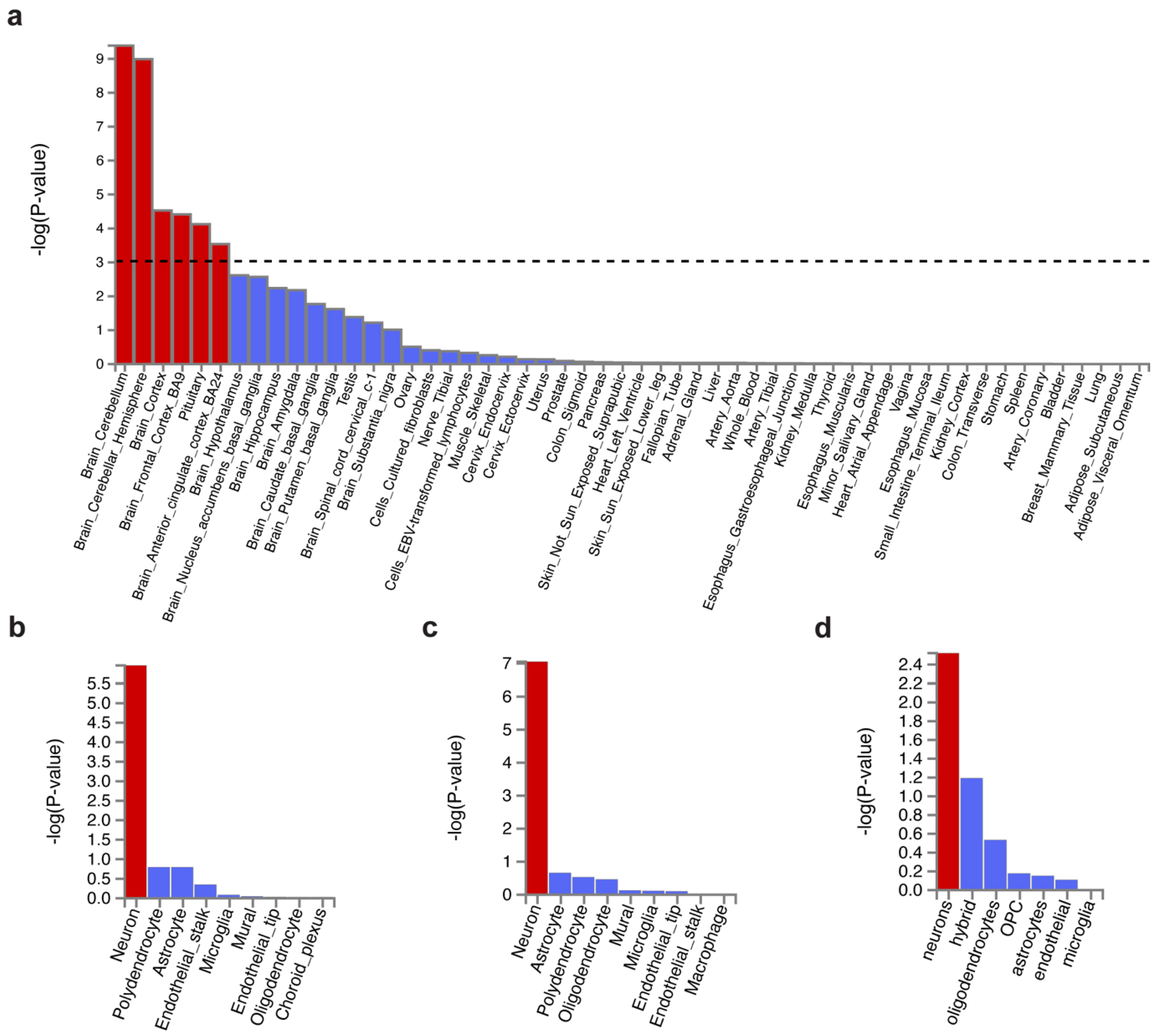
2.4. Cell Type Specificity Analysis
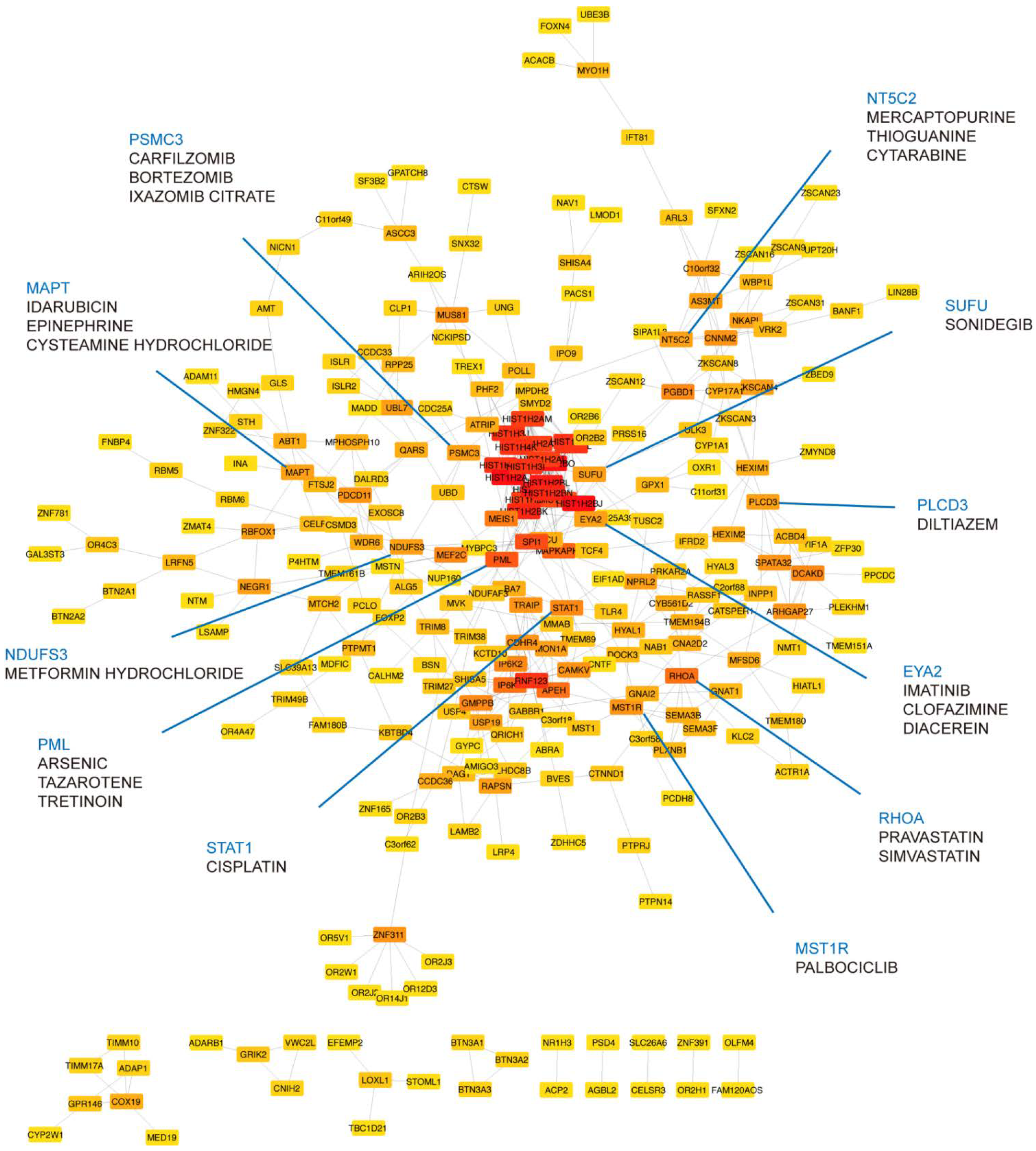
2.5. Identification of Druggable Targets
3. Results
3.1. Shared Genetic Variants
3.2. Tissue Expression and Cell Type Specificity
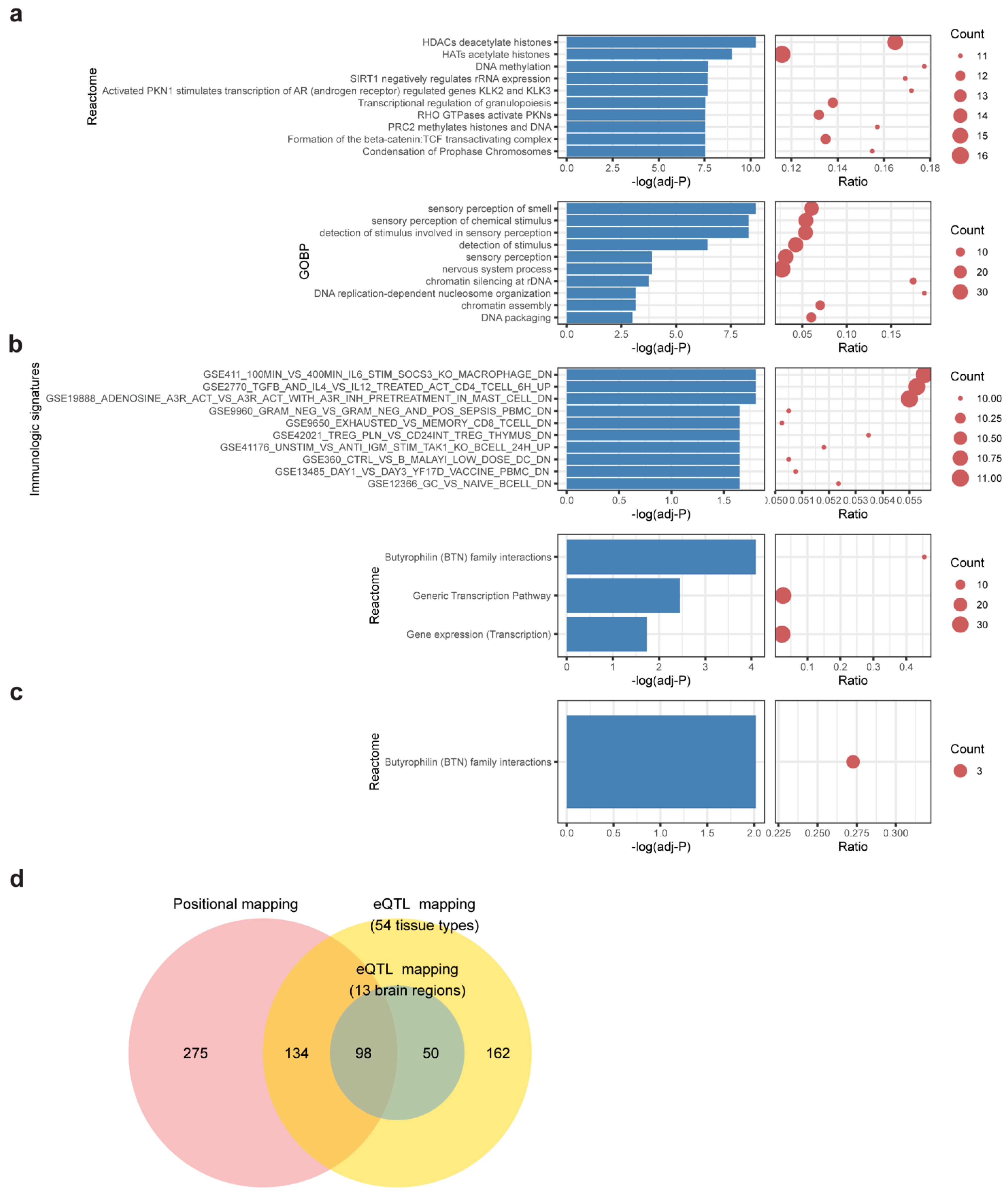
3.3. Gene Mapping and Functional Enrichment
3.4. Druggable Targets Identified by Network Approach
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gutiérrez-Rojas, L.; Porras-Segovia, A.; Dunne, H.; Andrade-González, N.; Cervilla, J.A. Prevalence and correlates of major depressive disorder: A systematic review. Rev. Bras. de Psiquiatr. 2020, 42, 657–672. [Google Scholar] [CrossRef]
- Tsuno, N.; Besset, A.; Ritchie, K. Sleep and depression. J. Clin. Psychiatry 2005, 66, 1254–1269. [Google Scholar] [CrossRef]
- Nutt, D.; Wilson, S.; Paterson, L. Sleep disorders as core symptoms of depression. Dialogues Clin. Neurosci. 2008, 10, 329–336. [Google Scholar]
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders, 5th ed.; American Psychiatric Association: Washington, DC, USA, 2013. [Google Scholar]
- Roth, T. Insomnia: Definition, Prevalence, Etiology, and Consequences. J. Clin. Sleep. Med. 2007, 3, S7–S10. [Google Scholar] [CrossRef] [Green Version]
- Taylor, D.J.; Lichstein, K.L.; Durrence, H.H.; Reidel, B.W.; Bush, A.J. Epidemiology of Insomnia, Depression, and Anxiety. Sleep 2005, 28, 1457–1464. [Google Scholar] [CrossRef]
- Richardson, J.D.; Thompson, A.; King, L.; Corbett, B.; Shnaider, P.; Cyr, K.S.; Nelson, J.; Sareen, J.; Elhai, J.; Zamorski, M. Insomnia, psychiatric disorders and suicidal ideation in a National Representative Sample of active Canadian Forces members. BMC Psychiatry 2017, 17, 211. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCall, W.V.; Blocker, J.N.; D’Agostino, R.; Kimball, J.; Boggs, N.; Lasater, B.; Haskett, R.; Krystal, A.; McDonald, W.M.; Rosenquist, R.B. Treatment of insomnia in depressed insomniacs: Effects on health-related quality of life, objective and self-reported sleep, and depression. J. Clin. Sleep Med. 2010, 6, 322–329. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Manber, R.; Edinger, J.D.; Gress, J.L.; San Pedro-Salcedo, M.G.; Kuo, T.F.; Kalista, T. Cognitive Behavioral Therapy for Insomnia Enhances Depression Outcome in Patients with Comorbid Major Depressive Disorder and Insomnia. Sleep 2008, 31, 489–495. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franzen, P.L.; Buysse, D.J. Sleep disturbances and depression: Risk relationships for subsequent depression and therapeutic implications. Dialogues Clin. Neurosci. 2008, 10, 473–481. [Google Scholar]
- Gulec, M.; Ozcan, H.; Oral, E.; Selvi, Y.; Aydin, A. The relationship between insomnia and major depressive disorder: A chicken and egg situation? Psychiatry Behav. Sci. 2012, 2, 28–33. [Google Scholar] [CrossRef] [Green Version]
- Okbay, A.; LifeLines Cohort Study; Baselmans, B.M.L.; De Neve, J.-E.; Turley, P.; Nivard, M.G.; Fontana, M.A.; Meddens, S.F.W.; Linnér, R.K.; Rietveld, A.C.; et al. Genetic variants associated with subjective well-being, depressive symptoms, and neuroticism identified through genome-wide analyses. Nat. Genet. 2016, 48, 624–633. [Google Scholar] [CrossRef] [Green Version]
- Hyde, C.L.; Nagle, M.; Tian, C.; Chen, C.L.H.X.; Paciga, S.A.; Wendland, J.R.; Tung, J.Y.; A Hinds, C.T.J.Y.T.D.; Perlis, R.H.; Winslow, M.W.N.S.A.P.J.R.W.A.R. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat. Genet. 2016, 48, 1031–1036. [Google Scholar] [CrossRef]
- Direk, N.; Williams, S.; Smith, J.A.; Ripke, S.; Air, T.; Amare, A.T.; Amin, N.; Baune, B.T.; Bennett, D.A.; Blackwood, D.H.R.; et al. An Analysis of Two Genome-wide Association Meta-analyses Identifies a New Locus for Broad Depression Phenotype. Biol. Psychiatry 2017, 82, 322–329. [Google Scholar] [CrossRef] [Green Version]
- Howard, D.M.; Adams, M.J.; Shirali, M.; Clarke, T.-K.; Marioni, R.E.; Davies, G.; Coleman, J.R.I.; Alloza, C.; Shen, X.; Barbu, M.C.; et al. Genome-wide association study of depression phenotypes in UK Biobank identifies variants in excitatory synaptic pathways. Nat. Commun. 2018, 9, 1470. [Google Scholar] [CrossRef] [Green Version]
- Wray, N.R.; Ripke, S.; Mattheisen, M.; Trzaskowski, M.; Byrne, E.M.; Abdellaoui, A.; Adams, M.J.; Esben Agerbo, E.; Air, T.M.; Andlauer, T.M.F.; et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 2018, 50, 668–681. [Google Scholar] [CrossRef] [Green Version]
- Howard, D.M.; Adams, M.J.; Clarke, T.-K.; Hafferty, J.D.; Gibson, J.; Shirali, M.; Coleman, J.R.I.; Hagenaars, S.R.; Ward, J.; Wigmore, E.M.; et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 2019, 22, 343–352. [Google Scholar] [CrossRef] [Green Version]
- Lane, J.M.; Liang, J.; Vlasac, I.; Anderson, S.G.; Bechtold, D.A.; Bowden, J.; Emsley, R.; Gill, S.; Little, M.A.; I Luik, A.; et al. Genome-wide association analyses of sleep disturbance traits identify new loci and highlight shared genetics with neuropsychiatric and metabolic traits. Nat. Genet. 2016, 49, 274–281. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hammerschlag, A.R.; Stringer, S.; de Leeuw, C.; Sniekers, S.; Taskesen, E.; Watanabe, K.; Blanken, T.F.; Dekker, K.; Lindert, B.H.W.T.; Wassing, R.; et al. Genome-wide association analysis of insomnia complaints identifies risk genes and genetic overlap with psychiatric and metabolic traits. Nat. Genet. 2017, 49, 1584–1592. [Google Scholar] [CrossRef] [PubMed]
- Stein, M.B.; McCarthy, M.J.; Chen, C.-Y.; Jain, S.; Gelernter, J.; He, F.; Heeringa, S.G.; Kessler, R.C.; Nock, M.K.; Ripke, S.; et al. Genome-wide analysis of insomnia disorder. Mol. Psychiatry 2018, 23, 2238–2250. [Google Scholar] [CrossRef]
- Lane, J.M.; Sleep, H.A.I.; Jones, S.E.; Dashti, H.S.; Wood, A.R.; Aragam, K.G.; van Hees, V.T.; Strand, L.B.; Winsvold, B.S.; Wang, H.; et al. Biological and clinical insights from genetics of insomnia symptoms. Nat. Genet. 2019, 51, 387–393. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jansen, P.R.; Watanabe, K.; Stringer, S.; Skene, N.; Bryois, J.; Hammerschlag, A.R.; De Leeuw, C.A.; Benjamins, J.S.; Manchado, A.M.; Nagel, M.; et al. Genome-wide analysis of insomnia in 1,331,010 individuals identifies new risk loci and functional pathways. Nat. Genet. 2019, 51, 394–403. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Song, W.; Torous, J.; Kossowsky, J.; Chen, C.-Y.; Huang, H.; Wright, A. Genome-wide association analysis of insomnia using data from Partners Biobank. Sci. Rep. 2020, 10, 6928. [Google Scholar] [CrossRef]
- Tam, V.; Patel, N.; Turcotte, M.; Bossé, Y.; Paré, G.; Meyre, D. Benefits and limitations of genome-wide association studies. Nat. Rev. Genet. 2019, 20, 467–484. [Google Scholar] [CrossRef] [PubMed]
- Byrne, E.M. The relationship between insomnia and complex diseases—insights from genetic data. Genome Med. 2019, 11, 57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Willer, C.J.; Li, Y.; Abecasis, G.R. METAL: Fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010, 26, 2190–2191. [Google Scholar] [CrossRef]
- Watanabe, K.; Taskesen, E.; van Bochoven, A.; Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 2017, 8, 1826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- The 1000 Genomes Project Consortium; Auton, A.; Abecasis, G.R.; Altshuler, D.M.; Durbin, R.M.; Bentley, D.R.; Chakravarti, A.; Clark, A.G.; Donnelly, P.; Eichler, E.E.; et al. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [CrossRef] [Green Version]
- Wang, K.; Li, M.; Hakonarson, H. Annovar: Functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010, 38, e164. [Google Scholar] [CrossRef]
- Consortium TGte. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 2020, 369, 1318–1330. [Google Scholar]
- Liberzon, A.; Subramanian, A.; Pinchback, R.; Thorvaldsdóttir, H.; Tamayo, P.; Mesirov, J.P. Molecular signatures database (MSigDB) 3.0. Bioinformatics 2011, 27, 1739–1740. [Google Scholar] [CrossRef]
- De Leeuw, C.A.; Mooij, J.M.; Heskes, T.; Posthuma, D. MAGMA: Generalized Gene-Set Analysis of GWAS Data. PLoS Comput. Biol. 2015, 11, e1004219. [Google Scholar] [CrossRef] [PubMed]
- Saunders, A.; Macosko, E.Z.; Wysoker, A.; Goldman, M.; Krienen, F.M.; de Rivera, H.; Bien, E.; Baum, M.; Bortolin, L.; Wang, S.; et al. Molecular Diversity and Specializations among the Cells of the Adult Mouse Brain. Cell 2018, 174, 1015–1030.e16. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Darmanis, S.; Sloan, S.A.; Zhang, Y.; Enge, M.; Caneda, C.; Shuer, L.M.; Gephart, M.G.H.; Barres, B.A.; Quake, S.R. A survey of human brain transcriptome diversity at the single cell level. Proc. Natl. Acad. Sci. USA 2015, 112, 7285–7290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [Green Version]
- 36. Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, D.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Chin, C.-H.; Chen, S.-H.; Wu, H.-H.; Ho, C.-W.; Ko, M.-T.; Lin, C.-Y. cytoHubba: Identifying hub objects and sub-networks from complex interactome. BMC Syst. Biol. 2014, 8, S11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Freshour, S.L.; Kiwala, S.; Cotto, K.C.; Coffman, A.C.; McMichael, J.F.; Song, J.J.; Griffith, M.; Griffith, O.L.; Wagner, A.H. Integration of the Drug–Gene Interaction Database (DGIdb 4.0) with open crowdsource efforts. Nucleic Acids Res. 2020, 49, D1144–D1151. [Google Scholar] [CrossRef]
- Ochoa, D.; Hercules, A.; Carmona, M.; Suveges, D.; Gonzalez-Uriarte, A.; Malangone, C.; Miranda, A.; Fumis, L.; Carvalho-Silva, D.; Spitzer, M.; et al. Open Targets Platform: Supporting systematic drug–target identification and prioritisation. Nucleic Acids Res. 2020, 49, D1302–D1310. [Google Scholar] [CrossRef]
- Sarayloo, F.; Dion, P.A.; Rouleau, G.A. MEIS1 and Restless Legs Syndrome: A Comprehensive Review. Front Neurol. 2019, 10, 935. [Google Scholar] [CrossRef]
- Powell, V.; Martin, J.; Thapar, A.; Rice, F.; Anney, R.J.L. Investigating regions of shared genetic variation in attention deficit/hyperactivity disorder and major depressive disorder: A GWAS meta-analysis. BioRxiv 2021, 11, 1–10. [Google Scholar]
- Davidson, S.; Miller, K.A.; Dowell, A.; Gildea, A.; Mackenzie, A. A remote and highly conserved enhancer supports amygdala specific expression of the gene encoding the anxiogenic neuropeptide substance-P. Mol. Psychiatry 2006, 11, 410–421. [Google Scholar] [CrossRef] [Green Version]
- Dashti, H.S.; Daghlas, I.; Lane, J.; Huang, Y.; Udler, M.; Wang, H.; Ollila, H.M.; Jones, S.E.; Kim, J.; Wood, A.R.; et al. Genetic determinants of daytime napping and effects on cardiometabolic health. MedRxiv 2021, 12, 1–5. [Google Scholar]
- Gerring, Z.F.; Gamazon, E.R.; Derks, E.M. Consortium for the MDDWG of the PG. A gene co-expression network-based analysis of multiple brain tissues reveals novel genes and molecular pathways underlying major depression. PLoS Genet. 2019, 15, e1008245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, R.; Yik, J.H.N.; Lew, Q.J.; Chao, S.-H. Brd4 and HEXIM1: Multiple roles in P-TEFb regulation and cancer. Biomed. Res. Int. 2014, 2014, 232870. [Google Scholar] [CrossRef]
- Shao, H.; Zhu, Q.; Lu, H.; Chang, A.; Gao, C.; Zhou, Q.; Luo, K. HEXIM1 controls P-TEFb processing and regulates drug sensitivity in triple-negative breast cancer. Mol. Biol. Cell 2020, 31, 1867–1878. [Google Scholar] [CrossRef] [PubMed]
- Stefansson, H.; Genetic Risk and Outcome in Psychosis (GROUP); Ophoff, R.A.; Steinberg, S.; Andreassen, O.A.; Cichon, S.; Rujescu, D.; Werge, T.; Pietiläinen, O.P.H.; Mors, O.; et al. Common variants conferring risk of schizophrenia. Nature 2009, 460, 744–747. [Google Scholar] [CrossRef] [Green Version]
- Schmedt, T.; Silva, M.M.; Ziaei, A.; Jurkunas, U. Molecular Bases of Corneal Endothelial Dystrophies. Exp. Eye Res. 2012, 95, 24–34. [Google Scholar] [CrossRef] [Green Version]
- Pontual, L.; Mathieu, Y.; Golzio, C.; Rio, M.; Malan, V.; Boddaert, N.; Soufflet, C.; Picard, C.; Durandy, A.; Dobbie, A.; et al. Mutational, Functional, and Expression Studies of the TCF4 Gene in Pitt-Hopkins Syndrome. Hum. Mutat. 2009, 30, 669–676. [Google Scholar] [CrossRef]
- Forrest, M.; Chapman, R.M.; Doyle, A.M.; Tinsley, C.L.; Waite, A.; Blake, D.J. Functional analysis of TCF4 missense mutations that cause Pitt–Hopkins syndrome. Hum. Mutat. 2012, 33, 1676–1686. [Google Scholar] [CrossRef]
- Cai, L.; Bao, Y.; Fu, X.; Cao, H.; Baranova, A.; Zhang, X.; Sun, J.; Zhang, F. Causal links between major depressive disorder and insomnia: A Mendelian randomisation study. Gene 2020, 768, 145271. [Google Scholar] [CrossRef]
- Rigucci, S.; Serafini, G.; Pompili, M.; Kotzalidis, G.D.; Tatarelli, R. Anatomical and functional correlates in major depressive disorder: The contribution of neuroimaging studies. World J. Biol. Psychiatry 2010, 11, 165–180. [Google Scholar] [CrossRef]
- Pandya, M.; Altinay, M.; Malone, D.A.; Anand, A. Where in the Brain Is Depression? Curr. Psychiatry Rep. 2012, 14, 634–642. [Google Scholar] [CrossRef] [Green Version]
- Drevets, W.C.; Savitz, J.; Trimble, M. The Subgenual Anterior Cingulate Cortex in Mood Disorders. CNS Spectr. 2008, 13, 663–681. [Google Scholar] [CrossRef]
- Kang, L.; Zhang, A.; Sun, N.; Liu, P.; Yang, C.; Li, G.; Liu, Z.; Wang, Y.; Zhang, K. Functional connectivity between the thalamus and the primary somatosensory cortex in major depressive disorder: A resting-state fMRI study. BMC Psychiatry 2018, 18, 339. [Google Scholar] [CrossRef] [Green Version]
- Lacerda, A.L.; Nicoletti, M.A.; Brambilla, P.; Sassi, R.B.; Mallinger, A.G.; Frank, E.; Kupfer, D.J.; Keshavan, M.S.; Soares, J.C. Anatomical MRI study of basal ganglia in major depressive disorder. Psychiatry Res. Neuroimaging 2003, 124, 129–140. [Google Scholar] [CrossRef]
- Liu, C.-H.; Guo, J.; Lu, S.-L.; Tang, L.-R.; Fan, J.; Wang, C.-Y.; Wang, L.; Liu, Q.-Q.; Liu, C.-Z. Increased Salience Network Activity in Patients With Insomnia Complaints in Major Depressive Disorder. Front. Psychiatry 2018, 9. [Google Scholar] [CrossRef] [Green Version]
- Yu, S.; Shen, Z.; Lai, R.; Feng, F.; Guo, B.; Wang, Z.; Yang, J.; Hu, Y.; Gong, L. The Orbitofrontal Cortex Gray Matter Is Associated With the Interaction Between Insomnia and Depression. Front. Psychiatry 2018, 9, 651. [Google Scholar] [CrossRef] [Green Version]
- Tao, S.; Chattun, M.R.; Yan, R.; Geng, J.T.; Zhu, R.X.; Shao, J.N.; Lu, Q.; Yao, Z.J. TPH-2 Gene Polymorphism in Major Depressive Disorder Patients With Early-Wakening Symptom. Front. Neurosci. 2018, 12, 827. [Google Scholar] [CrossRef] [Green Version]
- Schmahmann, J.D.; Caplan, D. Cognition, emotion and the cerebellum. Brain 2006, 129, 290–292. [Google Scholar] [CrossRef]
- Depping, M.S.; Schmitgen, M.M.; Kubera, K.M.; Wolf, R.C. Cerebellar Contributions to Major Depression. Front. Psychiatry 2018, 9, 634. [Google Scholar] [CrossRef] [Green Version]
- Bogoian, H.R.; King, T.Z.; Turner, J.A.; Semmel, E.S.; Dotson, V.M. Linking depressive symptom dimensions to cerebellar subregion volumes in later life. Transl. Psychiatry 2020, 10, 1–8. [Google Scholar] [CrossRef]
- DelRosso, L.M.; Hoque, R. The cerebellum and sleep. Neurol. Clin. 2014, 32, 893–900. [Google Scholar] [CrossRef]
- Canto, C.B.; Onuki, Y.; Bruinsma, B.; Werf, Y.D.; van der Zeeuw, C.I.D. The Sleeping Cerebellum. Trends Neurosci. 2017, 40, 309–323. [Google Scholar] [CrossRef] [Green Version]
- Vreeburg, S.; Hoogendijk, W.; Pelt, J.; Derijk, R.; Verhagen, J.; van Dyck, R.; Smit, J.H.; Zitman, F.G.; Penninx, B.W.J.H. Major Depressive Disorder and Hypothalamic-Pituitary-Adrenal Axis Activity: Results From a Large Cohort Study. Arch. Gen. Psychiatry 2009, 66, 617–626. [Google Scholar] [CrossRef] [Green Version]
- Keller, J.; Gomez, R.; Williams, G.; Lembke, A.; Lazzeroni, L.; Murphy, G.M.; Schatzberg, A.F. HPA axis in major depression: Cortisol, clinical symptomatology and genetic variation predict cognition. Mol. Psychiatry 2016, 22, 527–536. [Google Scholar] [CrossRef]
- Vgontzas, A.N.; Bixler, E.O.; Lin, H.-M.; Prolo, P.; Mastorakos, G.; Vela-Bueno, A.; Kales, A.; Chrousos, G.P. Chronic Insomnia Is Associated with Nyctohemeral Activation of the Hypothalamic-Pituitary-Adrenal Axis: Clinical Implications. J. Clin. Endocrinol. Metab. 2001, 86, 3787–3794. [Google Scholar] [CrossRef]
- Duman, R.S.; Sanacora, G.; Krystal, J.H. Altered Connectivity in Depression: GABA and Glutamate Neurotransmitter Deficits and Reversal by Novel Treatments. Neuron 2019, 102, 75–90. [Google Scholar] [CrossRef]
- Benson, K.L.; Bottary, R.; Schoerning, L.; Baer, L.; Gonenc, A.; Jensen, J.E.; Winkelman, J.W. 1H MRS Measurement of Cortical GABA and Glutamate in Primary Insomnia and Major Depressive Disorder: Relationship to Sleep Quality and Depression Severity. J. Affect. Disord. 2020, 274, 624–631. [Google Scholar] [CrossRef]
- Moriguchi, S.; Takamiya, A.; Noda, Y.; Horita, N.; Wada, M.; Tsugawa, S.; Plitman, E.; Sano, Y.; Tarumi, R.; ElSalhy, M.; et al. Glutamatergic neurometabolite levels in major depressive disorder: A systematic review and meta-analysis of proton magnetic resonance spectroscopy studies. Mol. Psychiatry 2018, 24, 952–964. [Google Scholar] [CrossRef] [Green Version]
- Plante, D.T.; Jensen, J.E.; Schoerning, L.; Winkelman, J.W. Reduced γ-aminobutyric acid in occipital and anterior cingulate cortices in primary insomnia: A link to major depressive disorder? Neuropsychopharmacology 2012, 37, 1548–1557. [Google Scholar] [CrossRef] [Green Version]
- Gottesmann, C. GABA mechanisms and sleep. Neuroscience 2002, 111, 231–239. [Google Scholar] [CrossRef]
- Luscher, B.; Shen, Q.; Sahir, N. The GABAergic deficit hypothesis of major depressive disorder. Mol. Psychiatry 2011, 16, 383–406. [Google Scholar] [CrossRef] [Green Version]
- Wu, J.; Liu, C.; Zhang, L.; Qu, C.-H.; Sui, X.-L.; Zhu, H.; Huang, L.; Xu, Y.-F.; Han, Y.-L.; Qin, C. Histone deacetylase-2 is involved in stress-induced cognitive impairment via histone deacetylation and PI3K/AKT signaling pathway modification. Mol. Med. Rep. 2017, 16, 1846–1854. [Google Scholar] [CrossRef] [Green Version]
- Renthal, W.; Maze, I.; Krishnan, V.; Covington, H.E.; Xiao, G.; Kumar, A.; Russo, S.J.; Graham, A.; Tsankova, N.; Kippin, T.E.; et al. Histone Deacetylase 5 Epigenetically Controls Behavioral Adaptations to Chronic Emotional Stimuli. Neuron 2007, 56, 517–529. [Google Scholar] [CrossRef] [Green Version]
- Schmauss, C. An HDAC-dependent epigenetic mechanism that enhances the efficacy of the antidepressant drug fluoxetine. Sci. Rep. 2015, 5, 8171. [Google Scholar] [CrossRef] [Green Version]
- Misztak, P.; Pańczyszyn-Trzewik, P.; Sowa-Kućma, M. Histone deacetylases (HDACs) as therapeutic target for depressive disorders. Pharmacol. Rep. 2018, 70, 398–408. [Google Scholar] [CrossRef]
- Ruifeng, D.; Liu, X.; Wang, T.; Wu, L.; Gao, X.; Zhang, Z. Histone Acetylation Regulation in Sleep Deprivation-Induced Spatial Memory Impairment. Neurochem. Res. 2016, 41, 2223–2232. [Google Scholar]
- Bahna, S.G.; Niles, L.P. Epigenetic regulation of melatonin receptors in neuropsychiatric disorders. Br. J. Pharmacol. 2018, 175, 3209–3219. [Google Scholar] [CrossRef] [Green Version]
- Wang, F.; Jin, J.; Wang, J.; He, R.; Li, K.; Hu, X.; Li, Y.; Zhu, Y. Association between olfactory function and inhibition of emotional competing distractors in major depressive disorder. Sci. Rep. 2020, 10, 1–8. [Google Scholar] [CrossRef]
- Smith, I.A.; Knezevic, B.R.; Ammann, J.U.; Rhodes, D.A.; Aw, D.; Palmer, D.B.; Mather, I.H.; Trowsdale, J. BTN1A1, the Mammary Gland Butyrophilin, and BTN2A2 Are Both Inhibitors of T Cell Activation. J. Immunol. 2010, 184, 3514–3525. [Google Scholar] [CrossRef]
- Ammann, J.U.; Cooke, A.; Trowsdale, J. Butyrophilin Btn2a2 Inhibits TCR Activation and Phosphatidylinositol 3-Kinase/Akt Pathway Signaling and Induces Foxp3 Expression in T Lymphocytes. J. Immunol. 2013, 190, 5030–5036. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gober, H.-J.; Kistowska, M.; Angman, L.; Jenö, P.; Mori, L.; De Libero, G. Human T cell receptor gammadelta cells recognize endogenous mevalonate metabolites in tumor cells. J. Exp. Med. 2003, 197, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.A.; Chen, H.-C.; Price, A.J.; Keeble, A.H.; Davey, M.S.; James, L.C.; Eberl, M.; Trowsdale, J. Activation of human γδ T cells by cytosolic interactions of BTN3A1 with soluble phosphoantigens and the cytoskeletal adaptor periplakin. J. Immunol. 2015, 194, 2390–2398. [Google Scholar] [CrossRef] [Green Version]
- Maes, M.; Bosmans, E.; De Jongh, R.; Kenis, G.; Vandoolaeghe, E.; Neels, H. Increased Serum Il-6 And Il-1 Receptor Antagonist Concentrations in Major Depression and Treatment Resistant Depression. Cytokine 1997, 9, 853–858. [Google Scholar] [CrossRef]
- Tuglu, C.; Kara, S.H.; Caliyurt, O.; Vardar, E.; Abay, E. Increased serum tumor necrosis factor-alpha levels and treatment response in major depressive disorder. Psychopharmacology 2003, 170, 429–433. [Google Scholar] [CrossRef]
- Das, R.; Emon, P.Z.; Shahriar, M.; Nahar, Z.; Islam, S.M.A.; Bhuiyan, M.A.; Islam, S.N.; Islam, R. Higher levels of serum IL-1β and TNF-α are associated with an increased probability of major depressive disorder. Psychiatry Res. 2020, 295, 113568. [Google Scholar] [CrossRef]
- Milenkovic, V.M.; Stanton, E.H.; Nothdurfter, C.; Rupprecht, R.; Wetzel, C.H. The Role of Chemokines in the Pathophysiology of Major Depressive Disorder. Int. J. Mol. Sci. 2019, 20, 2283. [Google Scholar] [CrossRef] [Green Version]
- Vgontzas, A.; Zoumakis, M.; Papanicolaou, D.; Bixler, E.; Prolo, P.; Lin, H.-M.; Vela-Bueno, A.; Kales, A.; Chrousos, G. Chronic insomnia is associated with a shift of interleukin-6 and tumor necrosis factor secretion from nighttime to daytime. Metabolism 2002, 51, 887–892. [Google Scholar] [CrossRef]
- Thase, M.E. Antidepressant treatment of the depressed patient with insomnia. J. Clin. Psychiatry 1999, 60 (Suppl. S17), 28–31, discussion 46–48. [Google Scholar]
- Barchas, J.D.; Altemus, M. Monoamine Hypotheses of Mood Disorders. In Basic Neurochemistry: Molecular, Cellular and Medical Aspects, 6th ed.; American Society for Neurochemistry, Sage Publications: Thousand Oaks, CA, USA, 1999. [Google Scholar]
- Moraes, W.A.; Dos, S.; Burke, P.R.; Coutinho, P.L.; Guilleminault, C.; Bittencourt, A.G.; Tufik, S.; Poyares, D. Sedative antidepressants and insomnia. Braz. J. Psychiatry 2011, 33, 91–95. [Google Scholar] [CrossRef] [Green Version]
- Clark, M.S.; Smith, P.O.; Jamieson, B. Antidepressants for the Treatment of Insomnia in Patients with Depression. AFP 2011, 84, 1. [Google Scholar]
- Vellante, F.; Cornelio, M.; Acciavatti, T.; Cinosi, E.; Marini, S.; Dezi, S.; Iorio, G.D.; Martinotti, G.; Giannantonio, M.D.; Di Giannantonio, M. Treatment of resistant insomnia and major depression. Clin. Ter. 2013, 164, 429–435. [Google Scholar] [PubMed]
- Brett, J.; Murnion, B. Management of benzodiazepine misuse and dependence. Aust. Prescr. 2015, 38, 152–155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferracioli-Oda, E.; Qawasmi, A.; Bloch, M.H. Meta-analysis: Melatonin for the treatment of primary sleep disorders. PLoS ONE 2013, 8, e63773. [Google Scholar]
| Name of Drug | ||
|---|---|---|
| Abiraterone Acitretin Alcohol Alendronic Acid Alprostadil Aminohippuric Acid Amodiaquine Apomorphine Atenolol Baclofen Bepridil Hydrochloride Bortezomib Butabarbital Butalbital Butethal Capecitabine Carbamazepine Carbidopa Carboplatin Carfilzomib Cefaclor Cholecalciferol Cisplatin Cyclosporine Cysteamine Hydrochloride Cytarabine Dacarbazine Dantrolene Dantrolene Sodium Daunorubicin Daunorubicin Hydrochloride Deferasirox Dexamethasone Dexketoprofen Diacerein Didanosine Dihydroergotamine Dihydroergotamine Mesylate Diltiazem Docetaxel Doxorubicin | Doxorubicin Hydrochloride Enzalutamide Epinephrine Epinephrine Bitartrate Ethopropazine Hydrochloride Felodipine Gabapentin Gabapentin Enacarbil Gefitinib Gemcitabine Gentian Violet Granisetron Hexachlorophene Hydroxyzine Pamoate Idarubicin Imatinib Inamrinone Infliximab Itraconazole Lansoprazole Menadione Mephobarbital Mercaptopurine Mesalamine Mesna Metformin Metformin Hydrochloride Metharbital Methotrexate Methylene Blue Mitoxantrone Hydrochloride Mycophenolate Mofetil Mycophenolic Acid Nelfinavir Niclosamide Nifedipine Nifuroxazide Nitazoxanide Norepinephrine Olanzapine Omeprazole | Oxitriptan Oxytetracycline Oxytetracycline - Hydrochloride Palbociclib Pantoprazole Phenazopyridine - Hydrochloride Phenobarbital Phenytoin Pravastatin Pregabalin Primidone Progesterone Promethazine Pyrantel Pamoate Rabeprazole Raloxifene Raloxifene -Hydrochloride Ribavirin Risperidone Ritonavir Safinamide Saquinavir Simvastatin Sodium Oxybate Sonidegib Spironolactone Sulfasalazine Tacrolimus Talbutal Tazarotene Thioguanine Thiopental Topiramate Trastuzumab Tretinoin Triclabendazole Trimetrexate Verapamil Vigabatrin Warfarin |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, Y.-S.; Wang, C.-C.; Chen, C.-Y. GWAS Meta-Analysis Reveals Shared Genes and Biological Pathways between Major Depressive Disorder and Insomnia. Genes 2021, 12, 1506. https://doi.org/10.3390/genes12101506
Lin Y-S, Wang C-C, Chen C-Y. GWAS Meta-Analysis Reveals Shared Genes and Biological Pathways between Major Depressive Disorder and Insomnia. Genes. 2021; 12(10):1506. https://doi.org/10.3390/genes12101506
Chicago/Turabian StyleLin, Yi-Sian, Chia-Chun Wang, and Cho-Yi Chen. 2021. "GWAS Meta-Analysis Reveals Shared Genes and Biological Pathways between Major Depressive Disorder and Insomnia" Genes 12, no. 10: 1506. https://doi.org/10.3390/genes12101506
APA StyleLin, Y.-S., Wang, C.-C., & Chen, C.-Y. (2021). GWAS Meta-Analysis Reveals Shared Genes and Biological Pathways between Major Depressive Disorder and Insomnia. Genes, 12(10), 1506. https://doi.org/10.3390/genes12101506






