The Roles of Peptide Hormones and Their Receptors during Plant Root Development
Abstract
1. Introduction
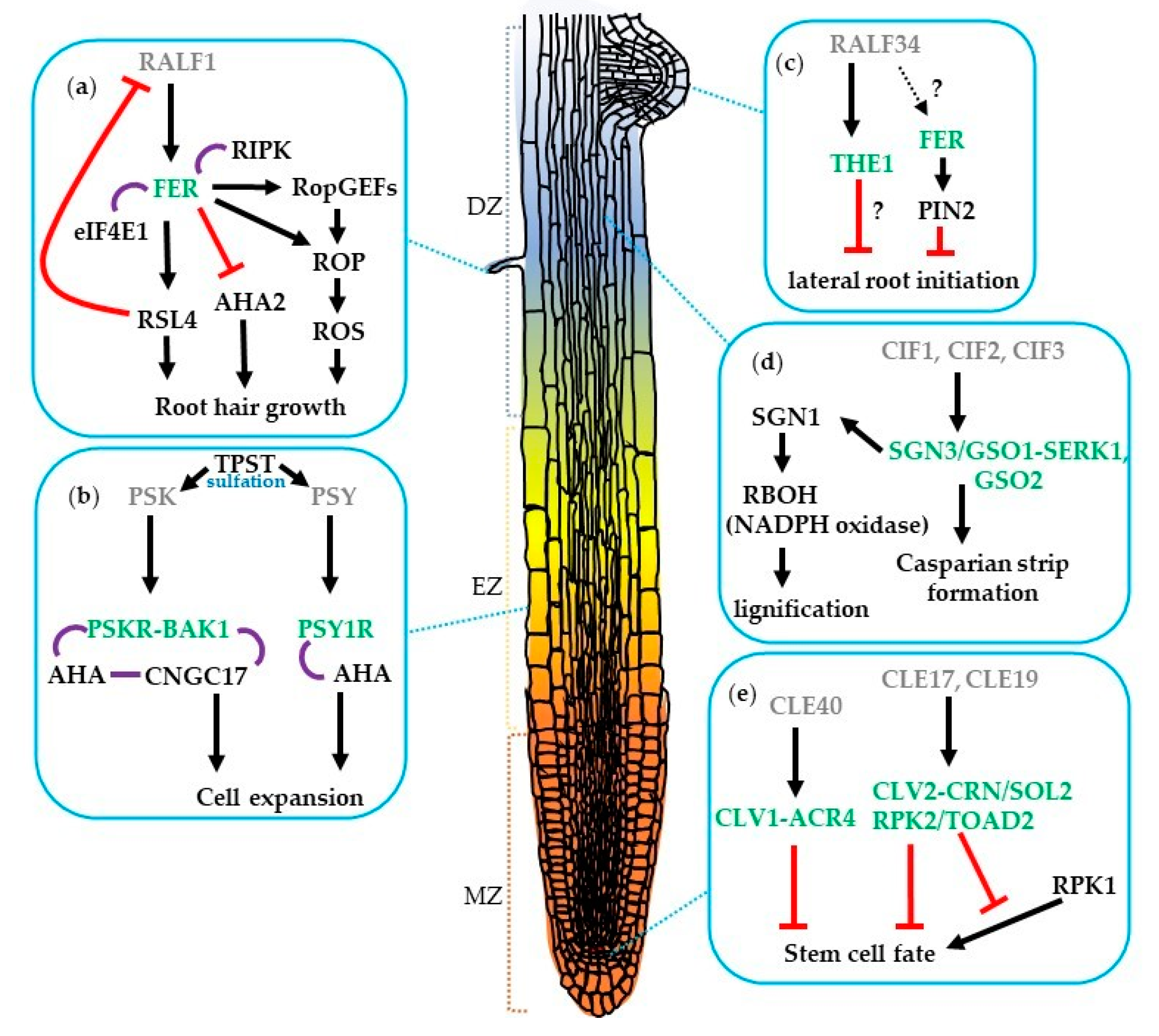
2. The Role of CLE Peptides and Their Receptors during Root Development
3. The Roles of Tyrosine-Sulfated Peptides and Their Receptors in Root Development
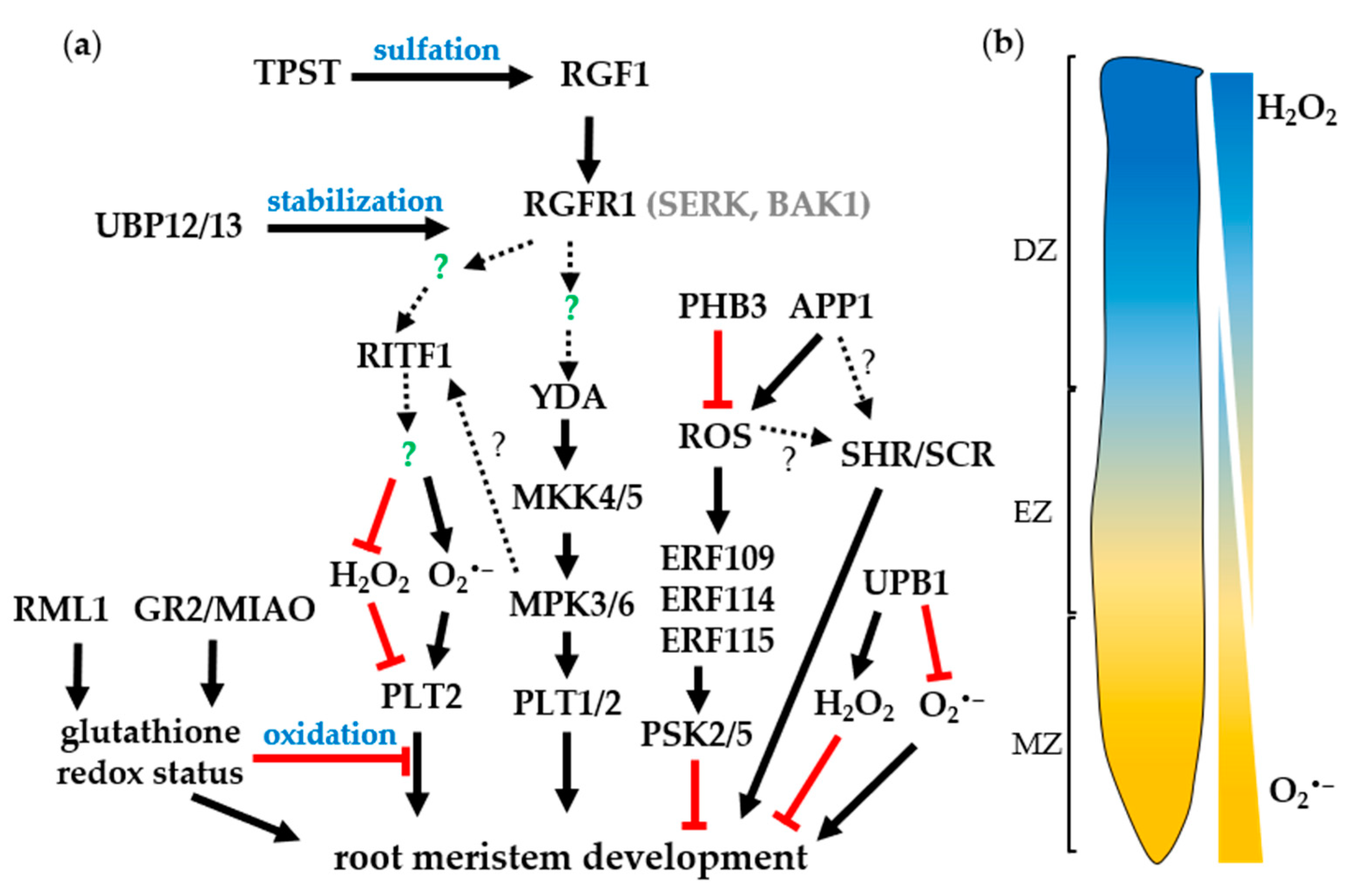
4. The Roles of ROS in Root Development
5. The Roles of RALF Peptides and Their Receptors in Root Development
6. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Pearce, G.; Strydom, D.; Johnson, S.; Ryan, C.A. A Polypeptide from Tomato Leaves Induces Wound-Inducible Proteinase Inhibitor Proteins. Science 1991, 253, 895–897. [Google Scholar] [CrossRef] [PubMed]
- Whitford, R.; Fernandez, A.; Tejos, R.; Pérez, A.C.; Kleine-Vehn, J.; Vanneste, S.; Drozdzecki, A.; Leitner, J.; Abas, L.; Aerts, M.; et al. GOLVEN Secretory Peptides Regulate Auxin Carrier Turnover during Plant Gravitropic Responses. Dev. Cell 2012, 22, 678–685. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Buchanan, B.B.; Feldman, L.J.; Luan, S. CLE-like (CLEL) peptides control the pattern of root growth and lateral root development in Arabidopsis. Proc. Natl. Acad. Sci. USA 2012, 109, 1760–1765. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Nakanomyo, I.; Motose, H.; Iwamoto, K.; Sawa, S.; Dohmae, N.; Fukuda, H. Dodeca-CLE Peptides as Suppressors of Plant Stem Cell Differentiation. Science 2006, 313, 842–845. [Google Scholar] [CrossRef]
- Haruta, M.; Sabat, G.; Stecker, K.; Minkoff, B.B.; Sussman, M.R. A Peptide Hormone and Its Receptor Protein Kinase Regulate Plant Cell Expansion. Science 2014, 343, 408–411. [Google Scholar] [CrossRef]
- Matsubayashi, Y.; Ogawa, M.; Morita, A.; Sakagami, Y. An LRR Receptor Kinase Involved in Perception of a Peptide Plant Hormone, Phytosulfokine. Science 2002, 296, 1470–1472. [Google Scholar] [CrossRef]
- Matsuzaki, Y.; Ogawa-Ohnishi, M.; Mori, A.; Matsubayashi, Y. Secreted Peptide Signals Required for Maintenance of Root Stem Cell Niche in Arabidopsis. Science 2010, 329, 1065–1067. [Google Scholar] [CrossRef]
- Doblas, V.G.; Smakowska-Luzan, E.; Fujita, S.; Alassimone, J.; Barberon, M.; Madalinski, M.; Belkhadir, Y.; Geldner, N. Root diffusion barrier control by a vasculature-derived peptide binding to the SGN3 receptor. Science 2017, 355, 280–284. [Google Scholar] [CrossRef]
- Amano, Y.; Tsubouchi, H.; Shinohara, H.; Ogawa, M.; Matsubayashi, Y. Tyrosine-sulfated glycopeptide involved in cellular proliferation and expansion in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 18333–18338. [Google Scholar] [CrossRef]
- Gou, X.; He, K.; Yang, H.; Yuan, T.; Lin, H.; Clouse, S.D.; Li, J. Genome-wide cloning and sequence analysis of leucine-rich repeat receptor-like protein kinase genes in Arabidopsis thaliana. BMC Genom. 2010, 11, 19. [Google Scholar] [CrossRef]
- Trotochaud, A.E.; Jeong, S.; Clark, S.E. CLAVATA3, a Multimeric Ligand for the CLAVATA1 Receptor-Kinase. Science 2000, 289, 613–617. [Google Scholar] [CrossRef] [PubMed]
- Brand, U. Dependence of Stem Cell Fate in Arabidopsis on a Feedback Loop Regulated by CLV3 Activity. Science 2000, 289, 617–619. [Google Scholar] [CrossRef] [PubMed]
- Schoof, H.; Lenhard, M.; Haecker, A.; Mayer, K.F.; Jürgens, G.; Laux, T. The Stem Cell Population of Arabidopsis Shoot Meristems Is Maintained by a Regulatory Loop between the CLAVATA and WUSCHEL Genes. Cell 2000, 100, 635–644. [Google Scholar] [CrossRef]
- Sánchez, W.C.; García-Ponce, B.; Sánchez, M.D.L.P.; Álvarez-Buylla, E.R.; Garay-Arroyo, A. Identifying the transition to the maturation zone in three ecotypes of Arabidopsis thaliana roots. Commun. Integr. Biol. 2018, 11, e1395993. [Google Scholar] [CrossRef] [PubMed]
- Qin, H.; He, L.; Huang, R. The Coordination of Ethylene and Other Hormones in Primary Root Development. Front. Plant Sci. 2019, 10, 874. [Google Scholar] [CrossRef] [PubMed]
- Pacifici, E.; Polverari, L.; Sabatini, S. Plant hormone cross-talk: The pivot of root growth. J. Exp. Bot. 2015, 66, 1113–1121. [Google Scholar] [CrossRef]
- De Tullio, M.C.; Jiang, K.; Feldman, L.J. Redox regulation of root apical meristem organization: Connecting root development to its environment. Plant Physiol. Biochem. 2010, 48, 328–336. [Google Scholar] [CrossRef]
- Dunand, C.; Crèvecoeur, M.; Penel, C. Distribution of superoxide and hydrogen peroxide in Arabidopsis root and their influence on root development: Possible interaction with peroxidases. New Phytol. 2007, 174, 332–341. [Google Scholar] [CrossRef]
- Tsukagoshi, H.; Busch, W.; Benfey, P.N. Transcriptional Regulation of ROS Controls Transition from Proliferation to Differentiation in the Root. Cell 2010, 143, 606–616. [Google Scholar] [CrossRef]
- Mhamdi, A.; Van Breusegem, F. Reactive oxygen species in plant development. Development 2018, 145, dev164376. [Google Scholar] [CrossRef]
- Mittler, R. ROS Are Good. Trends Plant Sci. 2017, 22, 11–19. [Google Scholar] [CrossRef]
- Gapper, C.; Dolan, L. Control of Plant Development by Reactive Oxygen Species. Plant Physiol. 2006, 141, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Ullah, F.; Zhou, D.-X.; Yi, M.; Zhao, Y. Mechanisms of ROS Regulation of Plant Development and Stress Responses. Front. Plant Sci. 2019, 10, 800. [Google Scholar] [CrossRef] [PubMed]
- Mangano, S.; Juárez, S.P.D.; Estevez, J.M. ROS Regulation of Polar Growth in Plant Cells. Plant Physiol. 2016, 171, 1593–1605. [Google Scholar] [CrossRef]
- Yamaguchi, Y.L.; Ishida, T.; Sawa, S. CLE peptides and their signaling pathways in plant development. J. Exp. Bot. 2016, 67, 4813–4826. [Google Scholar] [CrossRef] [PubMed]
- Whitford, R.; Fernandez, A.; De Groodt, R.; Ortega, E.; Hilson, P. Plant CLE peptides from two distinct functional classes synergistically induce division of vascular cells. Proc. Natl. Acad. Sci. USA 2008, 105, 18625–18630. [Google Scholar] [CrossRef]
- Jun, J.; Fiume, E.; Roeder, A.H.; Meng, L.; Sharma, V.K.; Osmont, K.S.; Baker, C.; Ha, C.M.; Meyerowitz, E.M.; Feldman, L.J.; et al. Comprehensive Analysis of CLE Polypeptide Signaling Gene Expression and Overexpression Activity in Arabidopsis. Plant Physiol. 2010, 154, 1721–1736. [Google Scholar] [CrossRef] [PubMed]
- Stahl, Y.; Grabowski, S.; Bleckmann, A.; Kühnemuth, R.; Weidtkamp-Peters, S.; Pinto, K.G.; Kirschner, G.K.; Schmid, J.B.; Wink, R.H.; Hülsewede, A.; et al. Moderation of Arabidopsis Root Stemness by CLAVATA1 and ARABIDOPSIS CRINKLY4 Receptor Kinase Complexes. Curr. Biol. 2013, 23, 362–371. [Google Scholar] [CrossRef]
- Casamitjana-Martínez, E.; Hofhuis, H.F.; Xu, J.; Liu, C.-M.; Heidstra, R.; Scheres, B. Root-Specific CLE19 Overexpression and the sol1/2 Suppressors Implicate a CLV-like Pathway in the Control of Arabidopsis Root Meristem Maintenance. Curr. Biol. 2003, 13, 1435–1441. [Google Scholar] [CrossRef]
- Bleckmann, A.; Weidtkamp-Peters, S.; Seidel, C.A.M.; Simon, R. Stem Cell Signaling in Arabidopsis Requires CRN to Localize CLV2 to the Plasma Membrane. Plant Physiol. 2009, 152, 166–176. [Google Scholar] [CrossRef]
- Zhu, Y.; Wang, Y.; Li, R.; Song, X.; Wang, Q.; Huang, S.; Jin, J.B.; Liu, C.; Lin, J. Analysis of interactions among the CLAVATA3 receptors reveals a direct interaction between CLAVATA2 and CORYNE in Arabidopsis. Plant J. 2009, 61, 223–233. [Google Scholar] [CrossRef]
- Miwa, H.; Betsuyaku, S.; Iwamoto, K.; Kinoshita, A.; Fukuda, H.; Sawa, S. The Receptor-Like Kinase SOL2 Mediates CLE Signaling in Arabidopsis. Plant Cell Physiol. 2008, 49, 1752–1757. [Google Scholar] [CrossRef]
- Racolta, A.; Nodine, M.D.; Davies, K.G.; Lee, C.; Rowe, S.C.; Velazco, Y.; Wellington, R.; Tax, F.E. A Common Pathway of Root Growth Control and Response to CLE Peptides Through Two Receptor Kinases inArabidopsis. Genetics 2017, 208, 687–704. [Google Scholar] [CrossRef] [PubMed]
- Komori, R.; Amano, Y.; Ogawa-Ohnishi, M.; Matsubayashi, Y. Identification of tyrosylprotein sulfotransferase in Arabidopsis. Proc. Natl. Acad. Sci. USA 2009, 106, 15067–15072. [Google Scholar] [CrossRef] [PubMed]
- Heyman, J.; Cools, T.; Vandenbussche, F.; Heyndrickx, K.S.; Van Leene, J.; Vercauteren, I.; Vanderauwera, S.; Vandepoele, K.; De Jaeger, G.; Van Der Straeten, D.; et al. ERF115 Controls Root Quiescent Center Cell Division and Stem Cell Replenishment. Science 2013, 342, 860–863. [Google Scholar] [CrossRef] [PubMed]
- Matsubayashi, Y.; Shinohara, H.; Ogawa, M. Identification and functional characterization of phytosulfokine receptor using a ligand-based approach. Chem. Rec. 2006, 6, 356–364. [Google Scholar] [CrossRef]
- Ladwig, F.; Dahlke, R.I.; Stührwohldt, N.; Hartmann, J.; Harter, K.; Sauter, M. Phytosulfokine Regulates Growth in Arabidopsis through a Response Module at the Plasma Membrane That Includes CYCLIC NUCLEOTIDE-GATED CHANNEL17, H+-ATPase, and BAK1. Plant Cell 2015, 27, 1718–1729. [Google Scholar] [CrossRef]
- Fuglsang, A.T.; Kristensen, A.; Cuin, T.; Schulze, W.X.; Persson, J.; Thuesen, K.H.; Ytting, C.K.; Oehlenschlaeger, C.B.; Mahmood, K.; Sondergaard, T.E.; et al. Receptor kinase-mediated control of primary active proton pumping at the plasma membrane. Plant J. 2014, 80, 951–964. [Google Scholar] [CrossRef]
- Ou, Y.; Lu, X.; Zi, Q.; Xun, Q.; Zhang, J.; Wu, Y.; Shi, H.; Wei, Z.; Zhao, B.; Zhang, X.; et al. RGF1 INSENSITIVE 1 to 5, a group of LRR receptor-like kinases, are essential for the perception of root meristem growth factor 1 in Arabidopsis thaliana. Cell Res. 2016, 26, 686–698. [Google Scholar] [CrossRef]
- Song, W.; Liu, L.; Wang, J.; Wu, Z.; Zhang, H.; Tang, J.; Lin, G.; Wang, Y.; Wenyang, L.; Li, W.; et al. Signature motif-guided identification of receptors for peptide hormones essential for root meristem growth. Cell Res. 2016, 26, 674–685. [Google Scholar] [CrossRef]
- Shinohara, H.; Mori, A.; Yasue, N.; Sumida, K.; Matsubayashi, Y. Identification of three LRR-RKs involved in perception of root meristem growth factor in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, 3897–3902. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Shi, H.; Ou, Y.; Cui, Y.; Chang, J.; Peng, L.; Gou, X.; He, K.; Li, J. RGF1-RGI1, a Peptide-Receptor Complex, Regulates Arabidopsis Root Meristem Development via a MAPK Signaling Cascade. Mol. Plant 2020, 13, 1594–1607. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Yu, X.; Xu, X.; Li, Y.; Yuan, W.; Xu, Y.; Mao, C.; Zhang, S.; Xu, J. The YDA-MKK4/MKK5-MPK3/MPK6 cascade functions downstream of the RGF1-RGI ligand-receptor pair in regulating mitotic activity in the root apical meristem. Mol. Plant 2020. [Google Scholar] [CrossRef] [PubMed]
- An, Z.; Liu, Y.; Ou, Y.; Li, J.; Zhang, B.; Sun, D.; Sun, Y.; Tang, W. Regulation of the stability of RGF1 receptor by the ubiquitin-specific proteases UBP12/UBP13 is critical for root meristem maintenance. Proc. Natl. Acad. Sci. USA 2018, 115, 1123–1128. [Google Scholar] [CrossRef]
- Yamada, M.; Han, X.; Benfey, P.N. RGF1 controls root meristem size through ROS signalling. Nat. Cell Biol. 2020, 577, 85–88. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yamada, M.; Han, X.; Ohler, U.; Benfey, P.N. High-Resolution Expression Map of the Arabidopsis Root Reveals Alternative Splicing and lincRNA Regulation. Dev. Cell 2016, 39, 508–522. [Google Scholar] [CrossRef]
- Nakayama, T.; Shinohara, H.; Tanaka, M.; Baba, K.; Ogawa-Ohnishi, M.; Matsubayashi, Y. A peptide hormone required for Casparian strip diffusion barrier formation in Arabidopsis roots. Science 2017, 355, 284–286. [Google Scholar] [CrossRef]
- Drapek, C.; Sparks, E.E.; Marhavý, P.; Taylor, I.; Andersen, T.G.; Hennacy, J.H.; Geldner, N.; Benfey, P.N. Minimum requirements for changing and maintaining endodermis cell identity in the Arabidopsis root. Nat. Plants 2018, 4, 586–595. [Google Scholar] [CrossRef]
- Okuda, S.; Fujita, S.; Moretti, A.; Hohmann, U.; Doblas, V.G.; Ma, Y.; Pfister, A.; Brandt, B.; Geldner, N.; Hothorn, M. Molecular mechanism for the recognition of sequence-divergent CIF peptides by the plant receptor kinases GSO1/SGN3 and GSO2. Proc. Natl. Acad. Sci. USA 2020, 117, 2693–2703. [Google Scholar] [CrossRef]
- Alassimone, J.; Fujita, S.; Doblas, V.G.; Van Dop, M.; Barberon, M.; Kalmbach, L.; Vermeer, J.E.M.; Rojas-Murcia, N.; Santuari, L.; Hardtke, C.S.; et al. Polarly localized kinase SGN1 is required for Casparian strip integrity and positioning. Nat. Plants 2016, 2, 16113. [Google Scholar] [CrossRef]
- Fujita, S.; De Bellis, D.; Edel, K.H.; Köster, P.; Andersen, T.G.; Schmid-Siegert, E.; Tendon, V.D.; Pfister, A.; Marhavý, P.; Ursache, R.; et al. SCHENGEN receptor module drives localized ROS production and lignification in plant roots. EMBO J. 2020, 39, e103894. [Google Scholar] [CrossRef] [PubMed]
- Cobbett, C.S.; May, M.J.; Howden, R.; Rolls, B.D. The glutathione-deficient, cadmium-sensitive mutant, cad2-1, of Arabidopsis thaliana is deficient in γ-glutamylcysteine synthetase. Plant J. 1998, 16, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Vernoux, T.; Wilson, R.C.; Seeley, K.A.; Reichheld, J.-P.; Muroy, S.; Brown, S.; Maughan, S.C.; Cobbett, C.S.; Van Montagu, M.; Inzé, D.; et al. The ROOT MERISTEMLESS1/CADMIUM SENSITIVE2 Gene Defines a Glutathione-Dependent Pathway Involved in Initiation and Maintenance of Cell Division during Postembryonic Root Development. Plant Cell 2000, 12, 97–109. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Pasternak, T.; Eiblmeier, M.; Ditengou, F.A.; Kochersperger, P.; Sun, J.; Wang, H.; Rennenberg, H.; Teale, W.; Paponov, I.; et al. Plastid-Localized Glutathione Reductase2-Regulated Glutathione Redox Status Is Essential for Arabidopsis Root Apical Meristem Maintenance. Plant Cell 2013, 25, 4451–4468. [Google Scholar] [CrossRef] [PubMed]
- Yu, Q.; Tian, H.; Yue, K.; Liu, J.; Zhang, B.; Li, X.; Ding, Z. A P-Loop NTPase Regulates Quiescent Center Cell Division and Distal Stem Cell Identity through the Regulation of ROS Homeostasis in Arabidopsis Root. PLoS Genet. 2016, 12, e1006175. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Tian, H.; Yu, Q.; Zhang, F.; Wang, R.; Gao, S.; Xu, W.; Liu, J.; Shani, E.; Fu, C.; et al. PHB3 Maintains Root Stem Cell Niche Identity through ROS-Responsive AP2/ERF Transcription Factors in Arabidopsis. Cell Rep. 2018, 22, 1350–1363. [Google Scholar] [CrossRef]
- Pearce, G.; Moura, D.S.; Stratmann, J.; Ryan, C.A. RALF, a 5-kDa ubiquitous polypeptide in plants, arrests root growth and development. Proc. Natl. Acad. Sci. USA 2001, 98, 12843–12847. [Google Scholar] [CrossRef]
- Haruta, M.; Monshausen, G.; Gilroy, S.; Sussman, M.R. A Cytoplasmic Ca2+ Functional Assay for Identifying and Purifying Endogenous Cell Signaling Peptides inArabidopsisSeedlings: Identification of AtRALF1 Peptide. Biochemistry 2008, 47, 6311–6321. [Google Scholar] [CrossRef]
- Cao, J.; Shi, F. Evolution of the RALF Gene Family in Plants: Gene Duplication and Selection Patterns. Evol. Bioinform. 2012, 8, EBO.S9652–92. [Google Scholar] [CrossRef]
- Canto, A.M.; Ceciliato, P.H.; Ribeiro, B.; Morea, F.A.O.; Garcia, A.A.F.; Silva-Filho, M.C.; Moura, D.S. Biological activity of nine recombinant AtRALF peptides: Implications for their perception and function in Arabidopsis. Plant Physiol. Biochem. 2014, 75, 45–54. [Google Scholar] [CrossRef]
- Matos, J.L.; Fiori, C.S.; Silva-Filho, M.C.; Moura, D.S. A conserved dibasic site is essential for correct processing of the peptide hormone AtRALF1 in Arabidopsis thaliana. FEBS Lett. 2008, 582, 3343–3347. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, R.; Liu, J.-X.; Guo, H.; Yin, Y.; Howell, S.H. Regulation and processing of a plant peptide hormone, AtRALF23, in Arabidopsis. Plant J. 2009, 59, 930–939. [Google Scholar] [CrossRef]
- Bergonci, T.; Ribeiro, B.; Ceciliato, P.H.; Guerrero-Abad, J.C.; Silva-Filho, M.C.; Moura, D.S. Arabidopsis thaliana RALF1 opposes brassinosteroid effects on root cell elongation and lateral root formation. J. Exp. Bot. 2014, 65, 2219–2230. [Google Scholar] [CrossRef] [PubMed]
- Ghorbani, S.; Lin, Y.-C.; Parizot, B.; Fernandez, A.; Njo, M.F.; Van De Peer, Y.; Beeckman, T.; Hilson, P. Expanding the repertoire of secretory peptides controlling root development with comparative genome analysis and functional assays. J. Exp. Bot. 2015, 66, 5257–5269. [Google Scholar] [CrossRef] [PubMed]
- Escobar-Restrepo, J.-M.; Huck, N.; Kessler, S.; Gagliardini, V.; Gheyselinck, J.; Yang, W.-C.; Grossniklaus, U. The FERONIA Receptor-like Kinase Mediates Male-Female Interactions During Pollen Tube Reception. Science 2007, 317, 656–660. [Google Scholar] [CrossRef]
- Du, C.; Li, X.; Chen, J.; Chen, W.; Li, B.; Li, C.; Wang, L.; Li, J.; Zhao, X.; Lin, J.; et al. Receptor kinase complex transmits RALF peptide signal to inhibit root growth in Arabidopsis. Proc. Natl. Acad. Sci. USA 2016, 113, E8326–E8334. [Google Scholar] [CrossRef]
- Abarca, A.; Franck, C.M.; Zipfel, C. Family-wide evaluation of RALF peptides in Arabidopsis thaliana. bioRxiv 2020. [Google Scholar] [CrossRef]
- Huang, G.-Q.; Li, E.; Ge, F.-R.; Li, S.; Wang, Q.; Zhang, C.-Q.; Zhang, Y. Arabidopsis RopGEF4 and RopGEF10 are important for FERONIA-mediated developmental but not environmental regulation of root hair growth. New Phytol. 2013, 200, 1089–1101. [Google Scholar] [CrossRef]
- Zhu, S.; Estévez, J.M.; Liao, H.; Zhu, Y.; Yang, T.; Li, C.; Wang, Y.; Li, L.; Liu, X.; Pacheco, J.M.; et al. The RALF1–FERONIA Complex Phosphorylates eIF4E1 to Promote Protein Synthesis and Polar Root Hair Growth. Mol. Plant 2020, 13, 698–716. [Google Scholar] [CrossRef]
- Zhu, S.; Pacheco, J.M.; Estevez, J.M.; Yu, F. Autocrine regulation of root hair size by the RALF-FERONIA-RSL4 signaling pathway. New Phytol. 2020, 227, 45–49. [Google Scholar] [CrossRef]
- Gonneau, M.; Desprez, T.; Martin, M.; Doblas, V.G.; Bacete, L.; Miart, F.; Sormani, R.; Hématy, K.; Renou, J.; Landrein, B.; et al. Receptor Kinase THESEUS1 Is a Rapid Alkalinization Factor 34 Receptor in Arabidopsis. Curr. Biol. 2018, 28, 2452–2458.e4. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.; Vu, L.D.; Broeck, L.V.D.; Lin, Z.; Ramakrishna, P.; Van De Cotte, B.; Gaudinier, A.; Goh, T.; Slane, D.; Beeckman, T.; et al. RALFL34 regulates formative cell divisions in Arabidopsis pericycle during lateral root initiation. J. Exp. Bot. 2016, 67, 4863–4875. [Google Scholar] [CrossRef] [PubMed]
- Dong, Q.; Zhang, Z.; Liu, Y.; Tao, L.; Liu, H. FERONIA regulates auxin-mediated lateral root development and primary root gravitropism. FEBS Lett. 2019, 593, 97–106. [Google Scholar] [CrossRef] [PubMed]
- Ohyama, K.; Shinohara, H.; Ogawa-Ohnishi, M.; Matsubayashi, Y. A glycopeptide regulating stem cell fate in Arabidopsis thaliana. Nat. Chem. Biol. 2009, 5, 578–580. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hsiao, Y.-C.; Yamada, M. The Roles of Peptide Hormones and Their Receptors during Plant Root Development. Genes 2021, 12, 22. https://doi.org/10.3390/genes12010022
Hsiao Y-C, Yamada M. The Roles of Peptide Hormones and Their Receptors during Plant Root Development. Genes. 2021; 12(1):22. https://doi.org/10.3390/genes12010022
Chicago/Turabian StyleHsiao, Yu-Chun, and Masashi Yamada. 2021. "The Roles of Peptide Hormones and Their Receptors during Plant Root Development" Genes 12, no. 1: 22. https://doi.org/10.3390/genes12010022
APA StyleHsiao, Y.-C., & Yamada, M. (2021). The Roles of Peptide Hormones and Their Receptors during Plant Root Development. Genes, 12(1), 22. https://doi.org/10.3390/genes12010022




