Satellite DNA in Neotropical Deer Species
Abstract
1. Introduction
2. Material and Methods
2.1. Species and Samples
2.2. Satellite DNA Isolation
2.3. Sequence Analysis
2.4. FISH
2.5. Phylogenetic Analysis
3. Results
3.1. Sequence Analysis
3.2. FISH
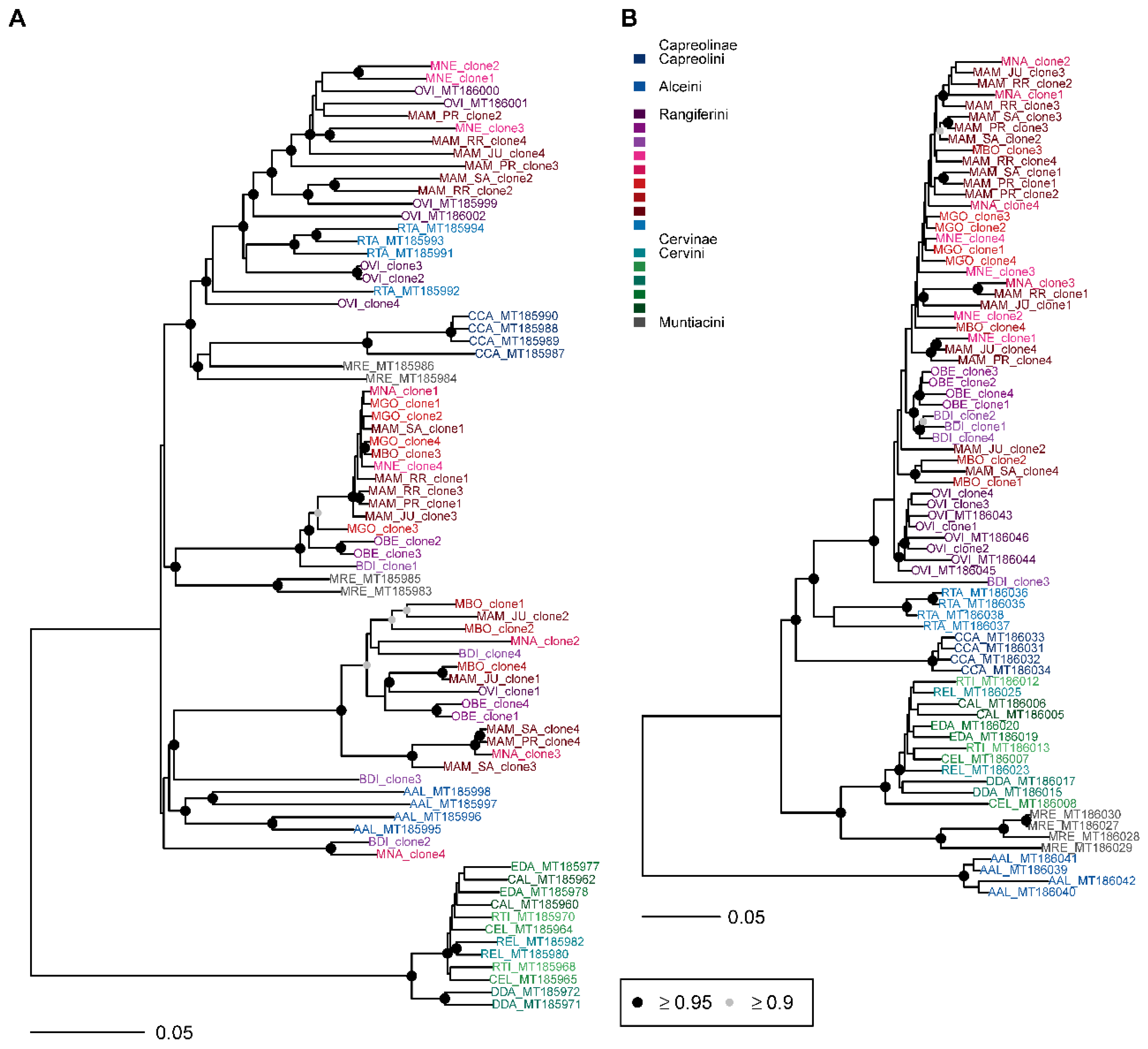
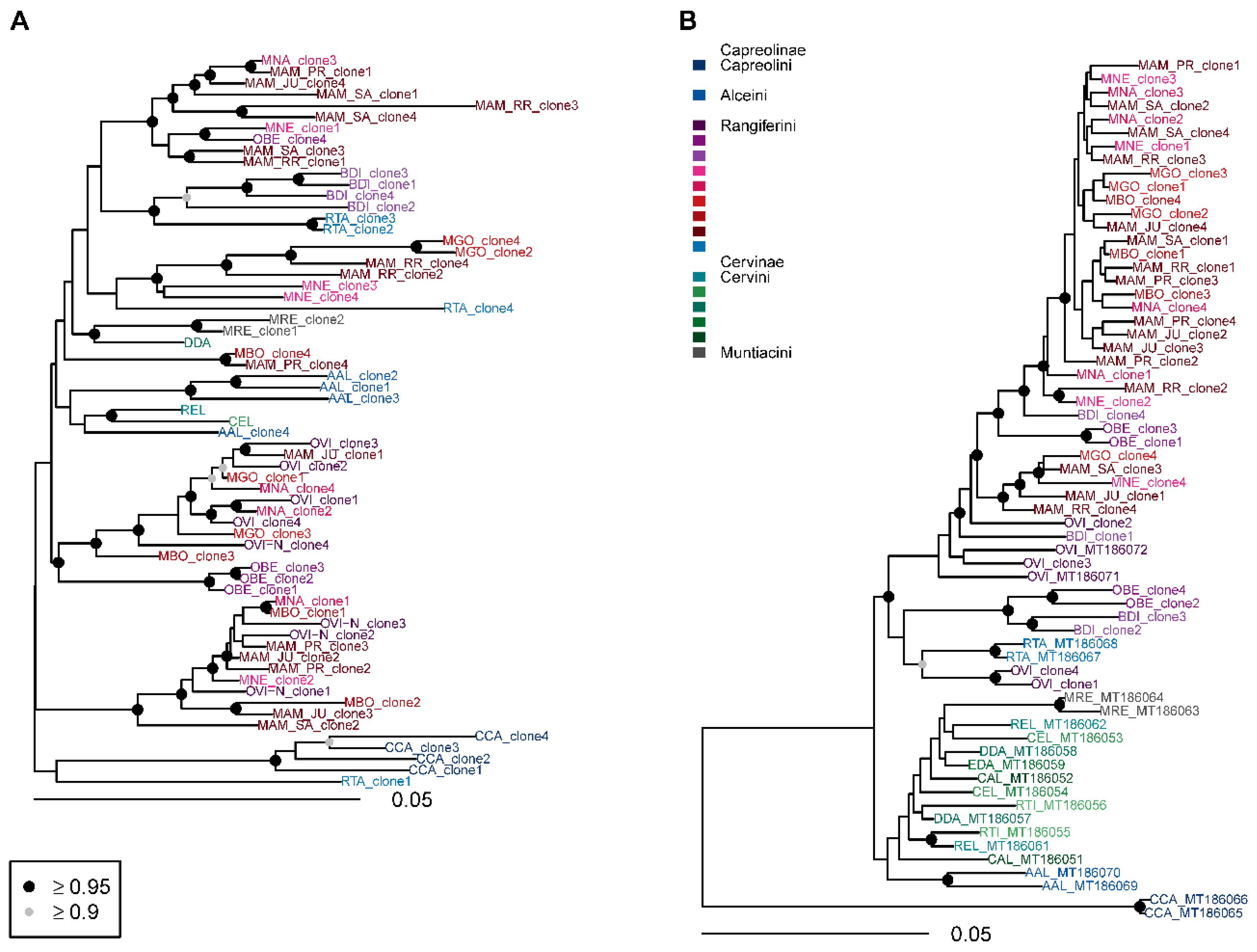
3.3. Phylogenetic Analysis
4. Discussion
Sequence Comparisons
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Ethical Statement
References
- Wilson, D.E.; Reeder, D.M. Mammal Species of the World: A Taxonomic and Geographic Reference; Johns Hopkins University Press: Baltimore, MD, USA, 2005; ISBN 978-0-8018-8221-0. [Google Scholar]
- Chen, L.; Qiu, Q.; Jiang, Y.; Wang, K.; Lin, Z.; Li, Z.; Bibi, F.; Yang, Y.; Wang, J.; Nie, W.; et al. Large-Scale Ruminant Genome Sequencing Provides Insights into Their Evolution and Distinct Traits. Science 2019, 364. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, E.E.; Helgen, K.M.; McDonough, M.M.; Bauer, F.; Hawkins, M.T.R.; Escobedo-Morales, L.A.; Patterson, B.D.; Maldonado, J.E. A Gene-Tree Test of the Traditional Taxonomy of American Deer: The Importance of Voucher Specimens, Geographic Data, and Dense Sampling. ZooKeys 2017, 697, 87–131. [Google Scholar] [CrossRef]
- Duarte, J.M.B.; González, S.; Maldonado, J.E. The Surprising Evolutionary History of South American Deer. Mol. Phylogenet. Evol. 2008, 49, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Groves, C.; Grubb, P. Ungulate Taxonomy, 1st ed.; Johns Hopkins University Press: Baltimore, MD, USA, 2011; ISBN 978-1-4214-0093-8. [Google Scholar]
- Fontana, F.; Rubini, M. Chromosomal Evolution in Cervidae. BioSystems 1990, 24, 157–174. [Google Scholar] [CrossRef]
- Duarte, J.M.B.; Jorge, W. Morphologic and Cytogenetic Description of the Small Red Brocket (Mazama Bororo Duarte,1996) in Brazil. Mammalia 2009, 67, 403–410. [Google Scholar] [CrossRef]
- Huang, L.; Chi, J.; Nie, W.; Wang, J.; Yang, F. Phylogenomics of Several Deer Species Revealed by Comparative Chromosome Painting with Chinese Muntjac Paints. Genetica 2006, 127, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Nietzel, H. Chromosome Evolution of Cervidae: Karyotypic and Molecular Aspects. In Cytogenetics: Basic and Applied Aspects; Obe, G., Basler, A., Eds.; Springer: Berlin/Heidelberg, Germany, 1987; ISBN 978-3-642-72804-4. [Google Scholar]
- Cifuentes-Rincón, A.; Morales-Donoso, J.A.; Sandoval, E.D.P.; Tomazella, I.M.; Mantellatto, A.M.B.; de Thoisy, B.; Duarte, J.M.B. Designation of a Neotype for Mazama Americana (Artiodactyla, Cervidae) Reveals a Cryptic New Complex of Brocket Deer Species. ZooKeys 2020, 958, 143–164. [Google Scholar] [CrossRef]
- Cursino, M.S.; Salviano, M.B.; Abril, V.V.; Zanetti, E.; dos, S.; Duarte, J.M. The Role of Chromosome Variation in the Speciation of the Red Brocket Deer Complex: The Study of Reproductive Isolation in Females. BMC Evol. Biol. 2014, 14, 40. [Google Scholar] [CrossRef]
- Abril, V.V.; Carnelossi, E.A.G.; González, S.; Duarte, J.M.B. Elucidating the Evolution of the Red Brocket Deer Mazama Americana Complex (Artiodactyla; Cervidae). Cytogenet. Genome Res. 2010, 128, 177–187. [Google Scholar] [CrossRef]
- Plohl, M.; Luchetti, A.; Mestrović, N.; Mantovani, B. Satellite DNAs between Selfishness and Functionality: Structure, Genomics and Evolution of Tandem Repeats in Centromeric (Hetero)Chromatin. Gene 2008, 409, 72–82. [Google Scholar] [CrossRef]
- Ugarković, D.; Plohl, M. Variation in Satellite DNA Profiles—Causes and Effects. EMBO J. 2002, 21, 5955–5959. [Google Scholar] [CrossRef] [PubMed]
- Schalch, T.; Steiner, F.A. Structure of Centromere Chromatin: From Nucleosome to Chromosomal Architecture. Chromosoma 2017, 126, 443–455. [Google Scholar] [CrossRef] [PubMed]
- Kunze, B.; Traut, W.; Garagna, S.; Weichenhan, D.; Redi, C.A.; Winking, H. Pericentric Satellite DNA and Molecular Phylogeny in Acomys (Rodentia). Chromosome Res. 1999, 7, 131. [Google Scholar] [CrossRef] [PubMed]
- Louzada, S.; Vieira-da-Silva, A.; Mendes-da-Silva, A.; Kubickova, S.; Rubes, J.; Adega, F.; Chaves, R. A Novel Satellite DNA Sequence in the Peromyscus Genome (PMSat): Evolution via Copy Number Fluctuation. Mol. Phylogenet. Evol. 2015, 92, 193–203. [Google Scholar] [CrossRef] [PubMed]
- Baicharoen, S.; Miyabe-Nishiwaki, T.; Arsaithamkul, V.; Hirai, Y.; Duangsa-ard, K.; Siriaroonrat, B.; Domae, H.; Srikulnath, K.; Koga, A.; Hirai, H. Locational Diversity of Alpha Satellite DNA and Intergeneric Hybridization Aspects in the Nomascus and Hylobates Genera of Small Apes. PLoS ONE 2014, 9, e109151. [Google Scholar] [CrossRef]
- Vozdova, M.; Kubickova, S.; Cernohorska, H.; Fröhlich, J.; Vodicka, R.; Rubes, J. Comparative Study of the Bush Dog (Speothos venaticus) Karyotype and Analysis of Satellite DNA Sequences and Their Chromosome Distribution in Six Species of Canidae. Cytogenet. Genome Res. 2019, 159, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Vozdova, M.; Kubickova, S.; Cernohorska, H.; Fröhlich, J.; Rubes, J. Satellite DNA Sequences in Canidae and Their Chromosome Distribution in Dog and Red Fox. Cytogenet. Genome Res. 2016, 150, 118–127. [Google Scholar] [CrossRef]
- Jobse, C.; Buntjer, J.B.; Haagsma, N.; Breukelman, H.J.; Beintema, J.J.; Lenstra, J.A. Evolution and Recombination of Bovine DNA Repeats. J. Mol. Evol. 1995, 41, 277–283. [Google Scholar] [CrossRef]
- Kopecna, O.; Kubickova, S.; Cernohorska, H.; Cabelova, K.; Vahala, J.; Martinkova, N.; Rubes, J. Tribe-Specific Satellite DNA in Non-Domestic Bovidae. Chromosome Res. 2014, 22, 277–291. [Google Scholar] [CrossRef]
- Chaves, R.; Guedes-Pinto, H.; Heslop-Harrison, J.S. Phylogenetic Relationships and the Primitive X Chromosome Inferred from Chromosomal and Satellite DNA Analysis in Bovidae. Proc. Biol. Sci. 2005, 272, 2009–2016. [Google Scholar] [CrossRef]
- Vozdova, M.; Kubickova, S.; Cernohorska, H.; Fröhlich, J.; Martínková, N.; Rubes, J. Sequence Analysis and FISH Mapping of Four Satellite DNA Families among Cervidae. Genes 2020, 11, 584. [Google Scholar] [CrossRef] [PubMed]
- Slamovits, H.C.; Cook, J.A.; Lessa, E.P.; Rossi, M.S. Recurrent Amplifications and Deletions of Satellite DNA Accompanied Chromosomal Diversification in South American Tuco-Tucos (Genus Ctenomys, Rodentia: Octodontidae): A Phylogenetic Approach. Mol. Biol. Evol. 2001, 18, 1708–1719. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, L.; Sperlich, D. Gradual Evolution of a Specific Satellite DNA Family in Drosophila Ambigua, D. Tristis, and D. Obscura. Mol. Biol. Evol. 1993, 10, 647–659. [Google Scholar] [PubMed]
- Garrido-Ramos, M.A.; de la Herrán, R.; Jamilena, M.; Lozano, R.; Ruiz Rejón, C.; Ruiz Rejón, M. Evolution of Centromeric Satellite DNA and Its Use in Phylogenetic Studies of the Sparidae Family (Pisces, Perciformes). Mol. Phylogenet. Evol. 1999, 12, 200–204. [Google Scholar] [CrossRef]
- Lee, C.; Court, D.R.; Cho, C.; Haslett, J.L.; Lin, C.C. Higher-Order Organization of Subrepeats and the Evolution of Cervid Satellite I DNA. J. Mol. Evol. 1997, 44, 327–335. [Google Scholar] [CrossRef]
- Li, Y.-C.; Lin, C.-C. Cervid Satellite DNA and Karyotypic Evolution of Indian Muntjac. Genes Genom. 2012, 34, 7–11. [Google Scholar] [CrossRef]
- Liu, Y.; Nie, W.; Huang, L.; Wang, J.; Su, W.; Lin, C.; Yang, F. Cloning, Characterization, and FISH Mapping of Four Satellite DNAs from Black Muntjac (Muntiacus crinifrons) and Fea’s Muntjac (M. feae). Zoolog. Res. 2008, 2008, 225–235. [Google Scholar] [CrossRef]
- Lin, C.C.; Li, Y.C. Chromosomal Distribution and Organization of Three Cervid Satellite DNAs in Chinese Water Deer (Hydropotes inermis). Cytogenet. Genome Res. 2006, 114, 147–154. [Google Scholar] [CrossRef]
- Li, Y.C.; Lee, C.; Chang, W.S.; Li, S.-Y.; Lin, C.C. Isolation and Identification of a Novel Satellite DNA Family Highly Conserved in Several Cervidae Species. Chromosoma 2002, 111, 176–183. [Google Scholar] [CrossRef]
- Blake, R.D.; Wang, J.Z.; Beauregard, L. Repetitive Sequence Families in Alces Alces Americana. J. Mol. Evol. 1997, 44, 509–520. [Google Scholar] [CrossRef]
- Li, Y.C.; Lee, C.; Hseu, T.H.; Li, S.Y.; Lin, C.C.; Hsu, T.H. Direct Visualization of the Genomic Distribution and Organization of Two Cervid Centromeric Satellite DNA Families. Cytogenet. Cell Genet. 2000, 89, 192–198. [Google Scholar] [CrossRef]
- Płucienniczak, A.; Skowroński, J.; Jaworski, J. Nucleotide Sequence of Bovine 1.715 Satellite DNA and Its Relation to Other Bovine Satellite Sequences. J. Mol. Biol. 1982, 158, 293–304. [Google Scholar] [CrossRef]
- Masumoto, H.; Masukata, H.; Muro, Y.; Nozaki, N.; Okazaki, T. A Human Centromere Antigen (CENP-B) Interacts with a Short Specific Sequence in Alphoid DNA, a Human Centromeric Satellite. J. Cell Biol. 1989, 109, 1963–1973. [Google Scholar] [CrossRef] [PubMed]
- Grant, C.E.; Bailey, T.L.; Noble, W.S. FIMO: Scanning for Occurrences of a given Motif. Bioinformatics 2011, 27, 1017–1018. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Rozewicki, J.; Yamada, D.K. MAFFT Online Service: Multiple Sequence Alignment, Interactive Sequence Choice and Visualization. Brief. Bioinform. 2019, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Katoh, K.; Kuma, K.; Toh, H.; Miyata, T. MAFFT Version 5: Improvement in Accuracy of Multiple Sequence Alignment. Nucleic Acids Res. 2005, 33, 511–518. [Google Scholar] [CrossRef]
- Machado, D.J.; Castroviejo-Fisher, S.; Grant, T. Evidence of Absence Treated as Absence of Evidence: The Effects of Variation in the Number and Distribution of Gaps Treated as Missing Data on the Results of Standard Maximum Likelihood Analysis. Mol. Phylogenet. Evol. 2021, 154, 106966. [Google Scholar] [CrossRef] [PubMed]
- Lefort, V.; Longueville, J.-E.; Gascuel, O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian Phylogenetic Inference and Model Choice across a Large Model Space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- Paradis, E.; Schliep, K. Ape 5.0: An Environment for Modern Phylogenetics and Evolutionary Analyses in R. Bioinformatics 2019, 35, 526–528. [Google Scholar] [CrossRef]
- Wang, L.-G.; Lam, T.T.-Y.; Xu, S.; Dai, Z.; Zhou, L.; Feng, T.; Guo, P.; Dunn, C.W.; Jones, B.R.; Bradley, T.; et al. Treeio: An R Package for Phylogenetic Tree Input and Output with Richly Annotated and Associated Data. Mol. Biol. Evol. 2020, 37, 599–603. [Google Scholar] [CrossRef]
- Revell, L.J. Phytools: An R Package for Phylogenetic Comparative Biology (and Other Things). Methods Ecol. Evol. 2012, 3, 217–223. [Google Scholar] [CrossRef]
- Neuwirth, E. RColorBrewer: ColorBrewer Palettes. 2014. Available online: https://CRAN.R-project.org/package=RColorBrewer (accessed on 7 December 2014).
- Barbanti Duarte, J.M.; González, S. Neotropical Cervidology, Biology and Medicine of Latin American Deer. Available online: https://www.iucn.org/content/neotropical-cervidology (accessed on 23 November 2020).
- Bogenberger, J.M.; Neumaier, P.S.; Fittler, F. The Muntjak Satellite IA Sequence Is Composed of 31-Base-Pair Internal Repeats That Are Highly Homologous to the 31-Base-Pair Subrepeats of the Bovine Satellite 1. Eur. J. Biochem. 1985, 148, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Vafa, O.; Shelby, R.D.; Sullivan, K.F. CENP-A Associated Complex Satellite DNA in the Kinetochore of the Indian Muntjac. Chromosoma 1999, 108, 367–374. [Google Scholar] [CrossRef] [PubMed]
- Carroll, C.W.; Silva, M.C.C.; Godek, K.M.; Jansen, L.E.T.; Straight, A.F. Centromere Assembly Requires the Direct Recognition of CENP-A Nucleosomes by CENP-N. Nat. Cell Biol. 2009, 11, 896–902. [Google Scholar] [CrossRef] [PubMed]
- Suntronpong, A.; Kugou, K.; Masumoto, H.; Srikulnath, K.; Ohshima, K.; Hirai, H.; Koga, A. CENP-B Box, a Nucleotide Motif Involved in Centromere Formation, Occurs in a New World Monkey. Biol. Lett. 2016, 12. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ruano, F.J.; López-León, M.D.; Cabrero, J.; Camacho, J.P.M. High-Throughput Analysis of the Satellitome Illuminates Satellite DNA Evolution. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Mestrović, N.; Plohl, M.; Mravinac, B.; Ugarković, D. Evolution of Satellite DNAs from the Genus Palorus--Experimental Evidence for the “Library” Hypothesis. Mol. Biol. Evol. 1998, 15, 1062–1068. [Google Scholar] [CrossRef] [PubMed]
- Palacios-Gimenez, O.M.; Milani, D.; Song, H.; Marti, D.A.; López-León, M.D.; Ruiz-Ruano, F.J.; Camacho, J.P.M.; Cabral-de-Mello, D.C. Eight Million Years of Satellite DNA Evolution in Grasshoppers of the Genus Schistocerca Illuminate the Ins and Outs of the Library Hypothesis. Genome Biol. Evol. 2020, 12, 88–102. [Google Scholar] [CrossRef]
- Fry, K.; Salser, W. Nucleotide Sequences of HS-α Satellite DNA from Kangaroo Rat Dipodomys Ordii and Characterization of Similar Sequences in Other Rodents. Cell 1977, 12, 1069–1084. [Google Scholar] [CrossRef]
- Walsh, J.B. Persistence of Tandem Arrays: Implications for Satellite and Simple-Sequence DNAs. Genetics 1987, 115, 553–567. [Google Scholar] [CrossRef] [PubMed]
- Okumura, K.; Kiyama, R.; Oishi, M. Sequence Analyses of Extrachromosomal Sau3A and Related Family DNA: Analysis of Recombination in the Excision Event. Nucleic Acids Res. 1987, 15, 7477–7489. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Alkan, C.; Eichler, E.E.; Bailey, J.A.; Sahinalp, S.C.; Tüzün, E. The Role of Unequal Crossover in Alpha-Satellite DNA Evolution: A Computational Analysis. J. Comput. Biol. 2004, 11, 933–944. [Google Scholar] [CrossRef] [PubMed]
- Stephan, W. Recombination and the Evolution of Satellite DNA. Genet. Res. 1986, 47, 167–174. [Google Scholar] [CrossRef]
- Dover, G.A. Molecular Drive in Multigene Families: How Biological Novelties Arise, Spread and Are Assimilated. Trends Genet. 1986, 2, 159–165. [Google Scholar] [CrossRef]
- Lorite, P.; Muñoz-López, M.; Carrillo, J.A.; Sanllorente, O.; Vela, J.; Mora, P.; Tinaut, A.; Torres, M.I.; Palomeque, T. Concerted Evolution, a Slow Process for Ant Satellite DNA: Study of the Satellite DNA in the Aphaenogaster Genus (Hymenoptera, Formicidae). Org. Divers. Evol. 2017, 17, 595–606. [Google Scholar] [CrossRef]
- Durfy, S.J.; Willard, H.F. Concerted Evolution of Primate Alpha Satellite DNA: Evidence for an Ancestral Sequence Shared by Gorilla and Human X Chromosome Alpha Satellite. J. Mol. Biol. 1990, 216, 555–566. [Google Scholar] [CrossRef]
- Mravinac, B.; Plohl, M. Parallelism in Evolution of Highly Repetitive DNAs in Sibling Species. Mol. Biol. Evol. 2010, 27, 1857–1867. [Google Scholar] [CrossRef]
- Mantellatto, A.M.B.; González, S.; Duarte, J.M.B. Molecular Identification of Mazama Species (Cervidae: Artiodactyla) from Natural History Collections. Genet. Mol. Biol. 2020, 43. [Google Scholar] [CrossRef]
- Escobedo-Morales, L.A.; Mandujano, S.; Eguiarte, L.E.; Rodríguez-Rodríguez, M.A.; Maldonado, J.E. First Phylogenetic Analysis of Mesoamerican Brocket Deer Mazama Pandora and Mazama Temama (Cetartiodactyla: Cervidae) Based on Mitochondrial Sequences: Implications for Neotropical Deer Evolution. Mammal. Biol. 2016, 81, 303–313. [Google Scholar] [CrossRef]
- Heckeberg, N.S. The Systematics of the Cervidae: A Total Evidence Approach. PeerJ 2020, 8, e8114. [Google Scholar] [CrossRef]
- Carrillo, J.D.; Faurby, S.; Silvestro, D.; Zizka, A.; Jaramillo, C.; Bacon, C.D.; Antonelli, A. Disproportionate Extinction of South American Mammals Drove the Asymmetry of the Great American Biotic Interchange. Proc. Natl. Acad. Sci. USA 2020, 117, 26281–26287. [Google Scholar] [CrossRef]
- Carrizo, L.V.; Tulli, M.J.; Santos, D.A.D.; Abdala, V. Interplay between Postcranial Morphology and Locomotor Types in Neotropical Sigmodontine Rodents. J. Anat. 2014, 224, 469–481. [Google Scholar] [CrossRef]
- Pečnerová, P.; Moravec, J.C.; Martínková, N. A Skull Might Lie: Modeling Ancestral Ranges and Diet from Genes and Shape of Tree Squirrels. Syst. Biol. 2015, 64, 1074–1088. [Google Scholar] [CrossRef]
- Barra, V.; Fachinetti, D. The Dark Side of Centromeres: Types, Causes and Consequences of Structural Abnormalities Implicating Centromeric DNA. Nat. Commun. 2018, 9, 4340. [Google Scholar] [CrossRef]
- Puppo, I.L.; Saifitdinova, A.F.; Tonyan, Z.N. The Role of Satellite DNA in Causing Structural Rearrangements in Human Karyotype. Russ. J. Genet. 2020, 56, 41–47. [Google Scholar] [CrossRef]
- Aquino, C.I.; Abril, V.V.; Duarte, J.M.B. Meiotic Pairing of B Chromosomes, Multiple Sexual System, and Robertsonian Fusion in the Red Brocket Deer Mazama Americana (Mammalia, Cervidae). Genet. Mol. Res. GMR 2013, 12, 3566–3574. [Google Scholar] [CrossRef]
- Abril, V.V.; Duarte, J.M.B. Chromosome Polymorphism in the Brazilian Dwarf Brocket Deer, Mazama Nana (Mammalia, Cervidae). Genet. Mol. Biol. 2008, 31, 53–57. [Google Scholar] [CrossRef]
- Valeri, P.M.; Tomazella, I.M.; Duarte, J.M.B. Intrapopulation Chromosomal Polymorphism in Mazama Gouazoubira (Cetartiodactyla; Cervidae): The Emergence of a New Species? Cytogenet. Genome Res. 2018, 154, 147–152. [Google Scholar] [CrossRef]
- Li, Y.C.; Lee, C.; Sanoudou, D.; Hseu, T.H.; Li, S.Y.; Lin, C.C.; Hsu, T.H. Interstitial Colocalization of Two Cervid Satellite DNAs Involved in the Genesis of the Indian Muntjac Karyotype. Chrom. Res. 2000, 8, 363–373. [Google Scholar] [CrossRef]
- Yang, F.; O’Brien, P.C.; Wienberg, J.; Neitzel, H.; Lin, C.C.; Ferguson-Smith, M.A. Chromosomal Evolution of the Chinese Muntjac (Muntiacus Reevesi). Chromosoma 1997, 106, 37–43. [Google Scholar] [CrossRef]
- Hartmann, N.; Scherthan, H. Characterization of Ancestral Chromosome Fusion Points in the Indian Muntjac Deer. Chromosoma 2004, 112, 213–220. [Google Scholar] [CrossRef]
- Fiorillo, B.F.; Sarria-Perea, J.A.; Abril, V.V.; Duarte, J.M.B. Cytogenetic Description of the Amazonian Brown Brocket Mazama Nemorivaga (Artiodactyla, Cervidae). Comp. cytogenet. 2013, 7, 25–31. [Google Scholar] [CrossRef]
- Yang, F.; Carter, N.P.; Shi, L.; Ferguson-Smith, M.A. A Comparative Study of Karyotypes of Muntjacs by Chromosome Painting. Chromosoma 1995, 103, 642–652. [Google Scholar] [CrossRef]
- Yang, F.; O’Brien, P.C.; Wienberg, J.; Ferguson-Smith, M.A. A Reappraisal of the Tandem Fusion Theory of Karyotype Evolution in Indian Muntjac Using Chromosome Painting. Chrom. Res. 1997, 5, 109–117. [Google Scholar] [CrossRef]
- Ashley, T. X-Autosome Translocations, Meiotic Synapsis, Chromosome Evolution and Speciation. Cytogenet. Genome Res. 2002, 96, 33–39. [Google Scholar] [CrossRef]
- Cotton, A.M.; Chen, C.-Y.; Lam, L.L.; Wasserman, W.W.; Kobor, M.S.; Brown, C.J. Spread of X-Chromosome Inactivation into Autosomal Sequences: Role for DNA Elements, Chromatin Features and Chromosomal Domains. Hum. Mol. Genet. 2014, 23, 1211–1223. [Google Scholar] [CrossRef][Green Version]
- Kalz-Füller, B.; Sleegers, E.; Schwanitz, G.; Schubert, R. Characterisation, Phenotypic Manifestations and X-Inactivation Pattern in 14 Patients with X-Autosome Translocations. Clin. Genet. 1999, 55, 362–366. [Google Scholar] [CrossRef]
- Dobigny, G.; Ozouf-Costaz, C.; Bonillo, C.; Volobouev, V. Viability of X-Autosome Translocations in Mammals: An Epigenomic Hypothesis from a Rodent Case-Study. Chromosoma 2004, 113, 34–41. [Google Scholar] [CrossRef]
- Ratomponirina, C.; Viegas-Péquignot, E.; Dutrillaux, B.; Petter, F.; Rumpler, Y. Synaptonemal Complexes in Gerbillidae: Probable Role of Intercalated Heterochromatin in Gonosome-Autosome Translocations. Cytogenet. Cell Genet. 1986, 43, 161–167. [Google Scholar] [CrossRef] [PubMed]
- Vozdova, M.; Ruiz-Herrera, A.; Fernandez, J.; Cernohorska, H.; Frohlich, J.; Sebestova, H.; Kubickova, S.; Rubes, J. Meiotic Behaviour of Evolutionary Sex-Autosome Translocations in Bovidae. Chrom. Res. 2016, 24, 325–338. [Google Scholar] [CrossRef] [PubMed]
- Camacho, J.P.; Sharbel, T.F.; Beukeboom, L.W. B-Chromosome Evolution. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2000, 355, 163–178. [Google Scholar] [CrossRef] [PubMed]
- Trifonov, V.A.; Dementyeva, P.V.; Larkin, D.M.; O’Brien, P.C.M.; Perelman, P.L.; Yang, F.; Ferguson-Smith, M.A.; Graphodatsky, A.S. Transcription of a Protein-Coding Gene on B Chromosomes of the Siberian Roe Deer (Capreolus pygargus). BMC Biol. 2013, 11, 90. [Google Scholar] [CrossRef] [PubMed]
- Duke Becker, S.E.; Thomas, R.; Trifonov, V.A.; Wayne, R.K.; Graphodatsky, A.S.; Breen, M. Anchoring the Dog to Its Relatives Reveals New Evolutionary Breakpoints across 11 Species of the Canidae and Provides New Clues for the Role of B Chromosomes. Chrom. Res. 2011, 19, 685–708. [Google Scholar] [CrossRef]
- Makunin, A.I.; Kichigin, I.G.; Larkin, D.M.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Yang, F.; Proskuryakova, A.A.; Vorobieva, N.V.; Chernyaeva, E.N.; O’Brien, S.J.; et al. Contrasting Origin of B Chromosomes in Two Cervids (Siberian Roe Deer and Grey Brocket Deer) Unravelled by Chromosome-Specific DNA Sequencing. BMC Genom. 2016, 17, 618. [Google Scholar] [CrossRef]
| Species | Latin Name | Abbr. a | 2n |
|---|---|---|---|
| Brown brocket deer | Mazama gouazoubira | MGO | 2n = 70 + B |
| Amazonian brown brocket deer | Mazama nemorivaga | MNE | 2n = 69 + B |
| Brazilian dwarf brocket deer | Mazama nana | MNA | 2n = 39 + Bs |
| Small red brocket deer | Mazama bororo | MBO | 2n = 33 + Bs |
| Red brocket deer-cytotype Paraná | Mazama americana | MAM-PR | 2n = 53 + Bs |
| Red brocket deer-cytotype Santarém | M. americana | MAM-SA | 2n = 51 + Bs |
| Red brocket deer-cytotype Juína | M. americana | MAM-JU | 2n = 45 + Bs |
| Red brocket deer-neotype from Roraima b | M. americana | MAM-RR | 2n = 46 + Bs |
| Marsh deer | Blastocerus dichotomus | BDI | 2n = 66 |
| Pampas deer | Ozotoceros bezoarticus | OBE | 2n = 68 |
| White-tailed deer of Brazilian origin | Odocoileus virginianus | OVI | 2n = 70 |
| Red deer | Cervus elaphus | CEL | 2n = 68 |
| Fallow deer | Dama dama | DDA | 2n = 68 |
| Eld’s deer | Rucervus eldii | REL | 2n = 58 |
| Chinese muntjac | Muntiacus reevesi | MRE | 2n = 46 |
| Roe deer | Capreolus capreolus | CCA | 2n = 70 |
| Reindeer | Rangifer tarandus | RTA | 2n = 70 |
| Moose | Alces alces | AAL | 2n = 68 |
| White-tailed deer of North American origin | Odocoileus virginianus | OVI-N | 2n = 70 |
| Species | SatI | SatII | SatIII | SatIV | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Length (bp) | GC Content (%) | Similarity (%) | No. of 31–bp Units | Length (bp) | GC Content (%) | Similarity (%) | Length (bp) | GC Content (%) | Similarity (%) | Length (bp) | GC Content (%) | Similarity (%) | |
| MGO | 910 | 51 | 96–99 | 19 | 579–581 | 67 | 97–99 | 583 | 53–56 | 89–98 | 727 | 45 | 97–99 |
| MNE | 910–913 | 51–53 | 76–92 | 19 | 578–580 | 65–67 | 92–96 | 580–583 | 55–58 | 90–93 | 726–727 | 44–45 | 96–99 |
| MNA | 904–919 | 46–51 | 75–84 | 20 | 575–579 | 64–67 | 90–93 | 579–583 | 56–59 | 89–97 | 727–728 | 45–46 | 98–99 |
| MBO | 910–917 | 49–51 | 76–92 | 18 | 578–579 | 64–66 | 92–94 | 579–583 | 55–58 | 90–94 | 727 | 45 | 99–100 |
| MAM_PR | 904–913 | 47–54 | 75–85 | 19 | 579 | 65–67 | 94–97 | 579–583 | 56–59 | 90–98 | 727 | 45 | 98–99 |
| MAM_SA | 904–910 | 47–51 | 72–94 | 18 | 578–579 | 64–67 | 92–98 | 581–583 | 56–59 | 90–95 | 727 | 44–46 | 97–99 |
| MAM_JU | 909–917 | 49–53 | 74–94 | 18 | 579–581 | 66–67 | 92–95 | 578–583 | 56–60 | 90–96 | 726–727 | 45–46 | 96–99 |
| MAM_RR | 908–910 | 51–52 | 77–98 | 20 | 575–579 | 64–67 | 93–96 | 580–583 | 54–59 | 86–93 | 727 | 45 | 97–99 |
| BDI | 911–919 | 48–50 | 77–80 | 21 | 579–580 | 63–67 | 88–98 | 579–584 | 56 | 92–97 | 726–737 | 45 | 93–97 |
| OBE | 911–917 | 48–51 | 76–98 | 18 | 580 | 67–68 | 96–99 | 582–583 | 55–58 | 91–99 | 727 | 45–46 | 92–99 |
| OVI | 907–915 | 48–51 | 74–99 | 19 | 578–579 | 66–67 | 97–98 | 581–584 | 55–56 | 94–97 | 727 | 44–46 | 95–99 |
| Species | SatI | SatII | SatIII–Partial | SatIV | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Neotropical Deer | Capreolinae | Cervinae | Neotropical Deer | Capreolinae | Cervinae | Neotropical Deer | Capreolinae | Cervinae | Neotropical Deer | Capreolinae | Cervinae | |
| MGO | 75–99 | 76–81 | 79–83 | 88–99 | 77–97 | 77–84 | 85–99 | 83–95 | 87–93 | 93–99 | 93–96 | 85–97 |
| MNE | 71–99 | 73–89 | 76–83 | 86–99 | 77–97 | 75–84 | 86–99 | 85–98 | 89–93 | 92–99 | 92–96 | 85–97 |
| MNA | 71–99 | 69–80 | 73–83 | 85–97 | 73–95 | 75–82 | 87–99 | 85–98 | 89–94 | 93–99 | 93–96 | 85–97 |
| MBO | 73–99 | 71–80 | 73–83 | 87–97 | 76–95 | 75–82 | 86–99 | 85–98 | 89–94 | 93–99 | 93–96 | 85–97 |
| MAM_PR | 72–99 | 73–89 | 74–83 | 87–99 | 77–96 | 76–84 | 87–99 | 85–98 | 89–94 | 93–99 | 93–96 | 85–96 |
| MAM_SA | 72–99 | 72–86 | 74–83 | 87–99 | 76–97 | 74–84 | 87–97 | 84–95 | 88–93 | 93–99 | 93–96 | 85–97 |
| MAM_JU | 72–99 | 70–86 | 73–83 | 86–97 | 76–96 | 74–83 | 88–99 | 85–98 | 89–94 | 93–99 | 93–96 | 85–97 |
| MAM_RR | 72–99 | 74–88 | 76–83 | 86–97 | 74–96 | 75–83 | 85–97 | 83–92 | 86–93 | 93–99 | 93–97 | 85–97 |
| BDI | 73–95 | 72–80 | 75–83 | 85–98 | 75–97 | 72–84 | 85–93 | 85–92 | 89–94 | 91–98 | 92–96 | 84–96 |
| OBE | 73–96 | 72–81 | 75–83 | 88–98 | 78–97 | 76–84 | 86–97 | 85–92 | 89–94 | 92–97 | 92–96 | 84–95 |
| OVI | 72–92 | 72–87 | 73–84 | 79–98 | 79–98 | 77–85 | 86–98 | 85–95 | 89–93 | 93–98 | 93–97 | 85–97 |
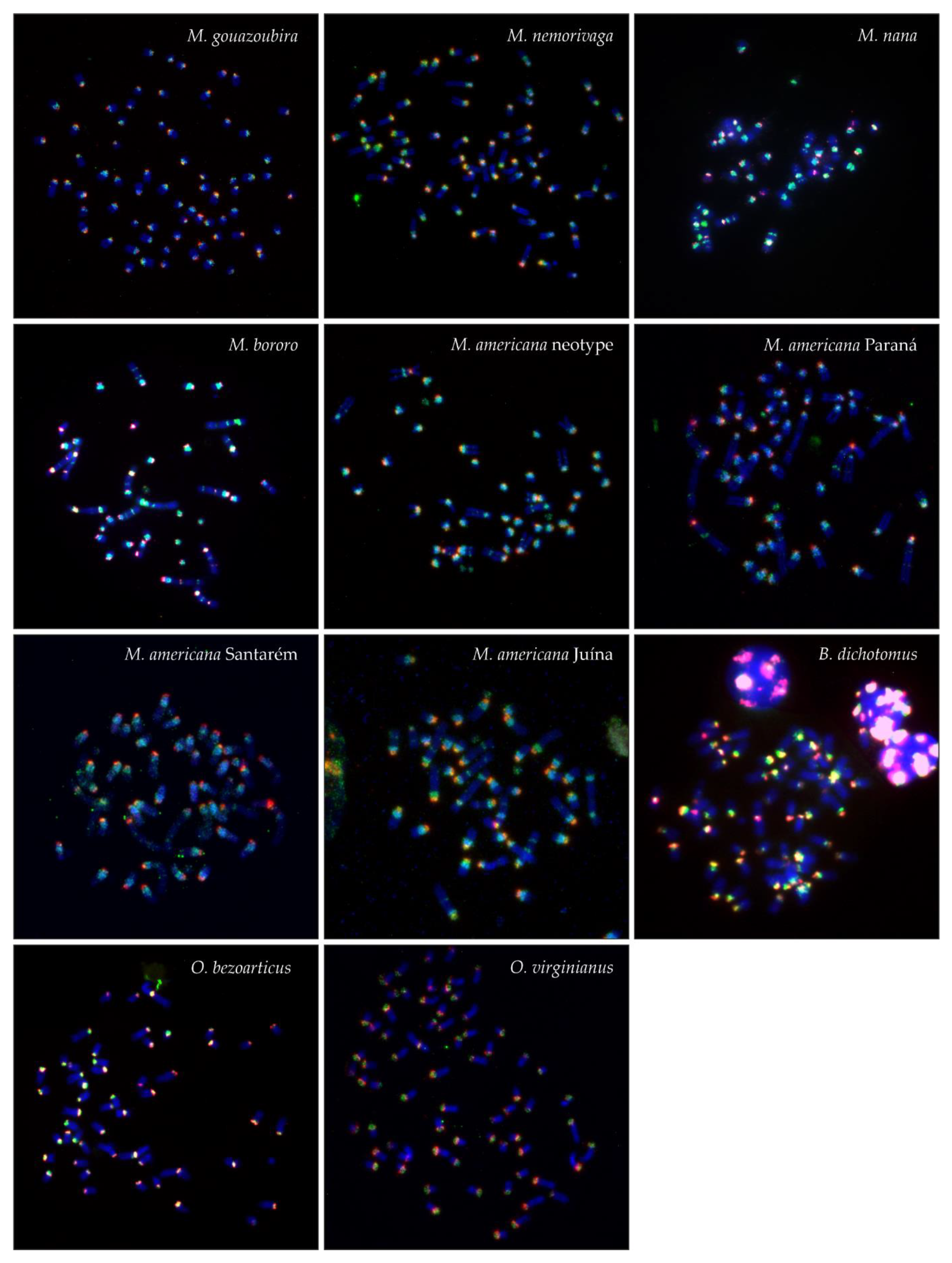
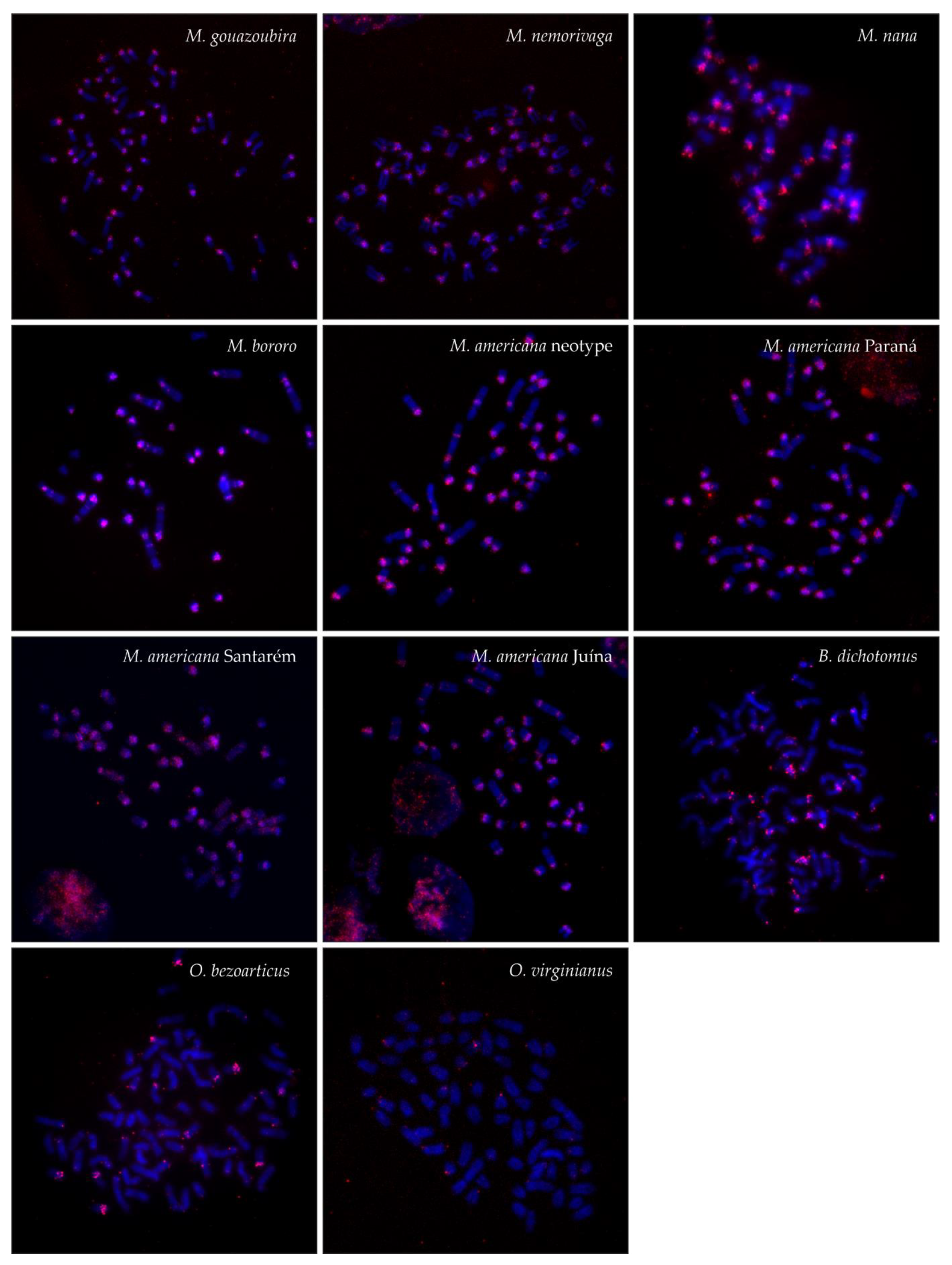
| Species | 2n | FN | B | SatI | SatII | SatIII | SatIV |
|---|---|---|---|---|---|---|---|
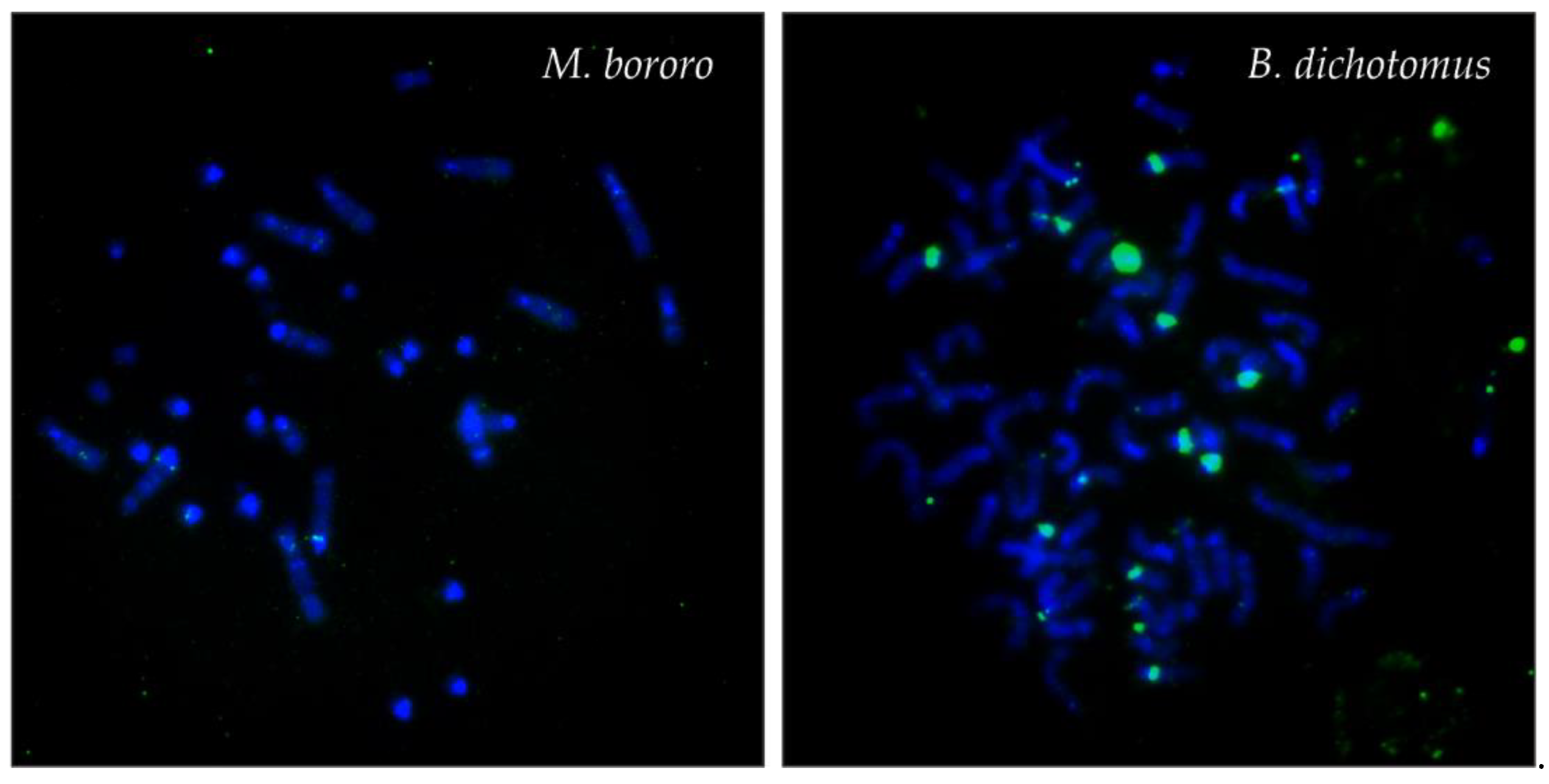
| MGO | 70 | 70 | + | all autosomes, X, Bs | all autosomes, X | a single autosome, weak | all autosomes, X |
| MNE | 69 | 72 | + | all autosomes, X, Bs | all autosomes, X | a few autosomes, weak | all autosomes |
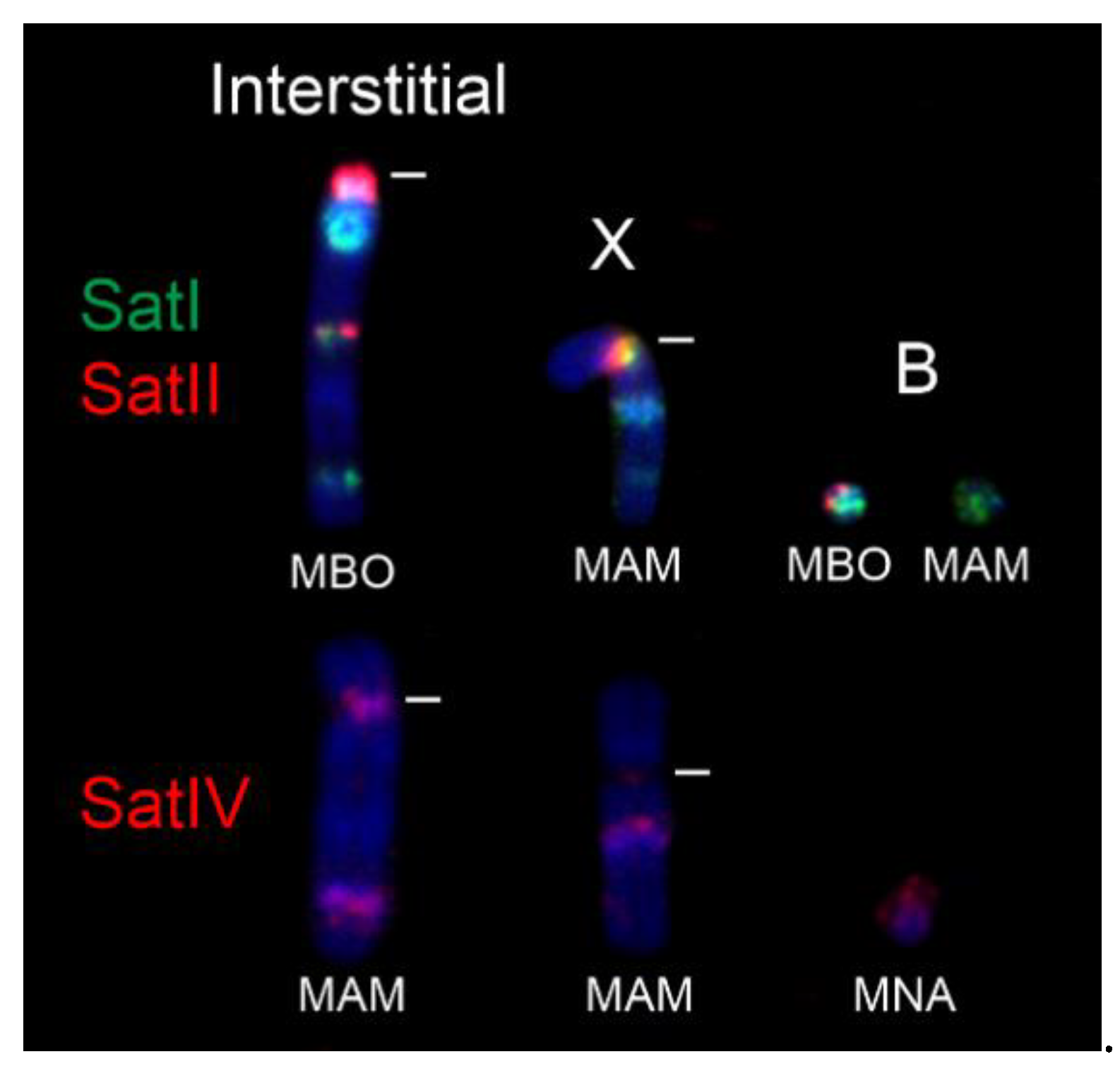
| MNA | 39 | 58 | + | all autosomes, X, Bs, interstitial | all autosomes, X, interstitial | a few autosomes, weak | all autosomes, X, Bs, interstitial |
| MBO | 33 | 46 | + | all autosomes, X, Bs, interstitial | all autosomes, X, Bs, interstitial | a few autosomes, weak | most autosomes, X, interstitial |
| MAM PR | 53 | 56 | + | all autosomes, X, Bs, interstitial | all autosomes, X | a few autosomes, weak | all autosomes, X, interstitial |
| MAM SA | 51 | 56 | + | all autosomes, X, Bs, interstitial | all autosomes, X, | a few autosomes, weak | all autosomes, interstitial |
| MAM JU | 45 | 48 | + | all autosomes, X, Bs, interstitial | all autosomes, X | a few autosomes, weak | all autosomes, X, interstitial |
| MAM RR | 46 | 51 | + | all autosomes, X, Bs, interstitial | all autosomes, X | a few autosomes, weak | all autosomes, interstitial |
| BDI | 66 | 74 | - | all autosomes, X | all autosomes, X | a few autosomes, big or weak | a few autosomes, big or weak |
| OBE | 68 | 74 | - | all autosomes, X | all autosomes, X | a few autosomes, weak | a few autosomes |
| OVI | 70 | 74 | - | all autosomes, X | all autosomes, X | a few autosomes, weak | a few autosomes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vozdova, M.; Kubickova, S.; Martínková, N.; Galindo, D.J.; Bernegossi, A.M.; Cernohorska, H.; Kadlcikova, D.; Musilová, P.; Duarte, J.M.; Rubes, J. Satellite DNA in Neotropical Deer Species. Genes 2021, 12, 123. https://doi.org/10.3390/genes12010123
Vozdova M, Kubickova S, Martínková N, Galindo DJ, Bernegossi AM, Cernohorska H, Kadlcikova D, Musilová P, Duarte JM, Rubes J. Satellite DNA in Neotropical Deer Species. Genes. 2021; 12(1):123. https://doi.org/10.3390/genes12010123
Chicago/Turabian StyleVozdova, Miluse, Svatava Kubickova, Natália Martínková, David Javier Galindo, Agda Maria Bernegossi, Halina Cernohorska, Dita Kadlcikova, Petra Musilová, Jose Mauricio Duarte, and Jiri Rubes. 2021. "Satellite DNA in Neotropical Deer Species" Genes 12, no. 1: 123. https://doi.org/10.3390/genes12010123
APA StyleVozdova, M., Kubickova, S., Martínková, N., Galindo, D. J., Bernegossi, A. M., Cernohorska, H., Kadlcikova, D., Musilová, P., Duarte, J. M., & Rubes, J. (2021). Satellite DNA in Neotropical Deer Species. Genes, 12(1), 123. https://doi.org/10.3390/genes12010123







