Large-Scale Cloning and Comparative Analysis of TaNAC Genes in Response to Stripe Rust and Powdery Mildew in Wheat (Triticum aestivum L.)
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials and Fungi
2.2. Reidentification and Sequence Analyses of NAC TFs in Wheat
2.3. RNA Extraction
2.4. Screening Fungus-Responsive NAC TFs and Evaluating Splice Variants
2.5. Real-Time Quantitative PCR Analysis
2.6. Analysis of TaNAC TF Protein-Sequence Characteristics
3. Results
3.1. Genome-Wide Reidentification of TaNAC TFs in IWGSC RefSeq v1.1
3.2. Cloning and Nomenclature of New TaNAC Transcripts in N9134
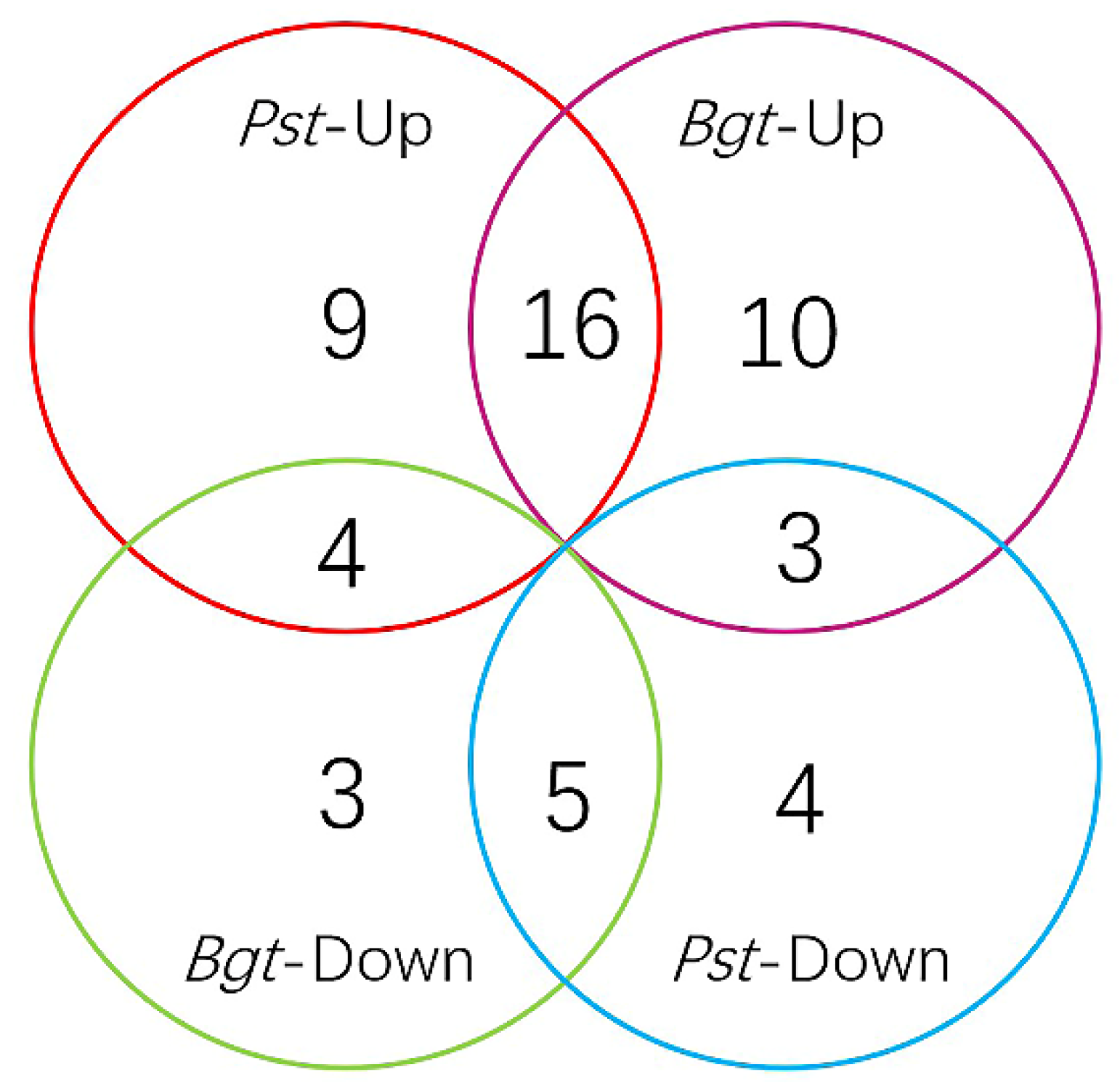
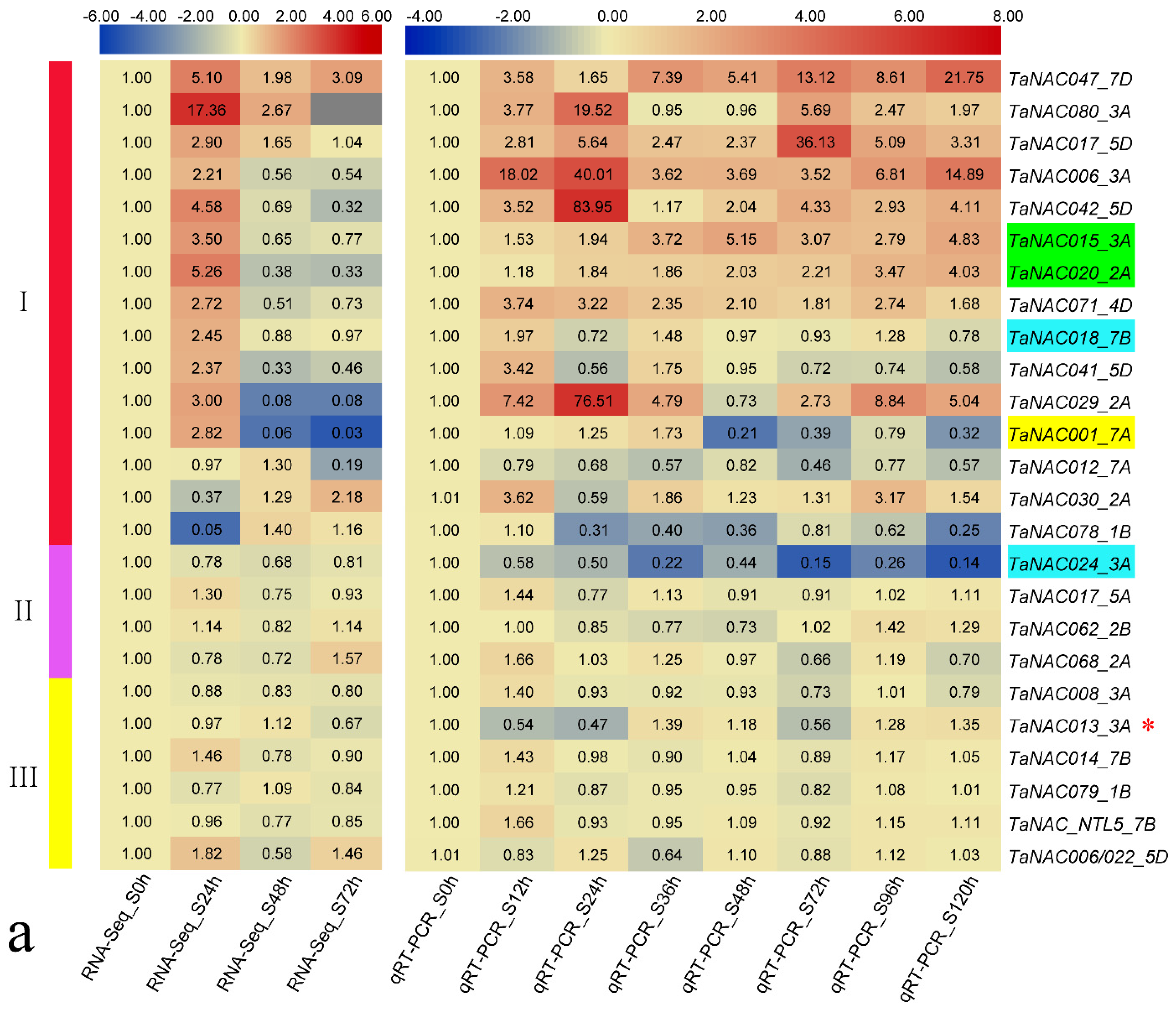
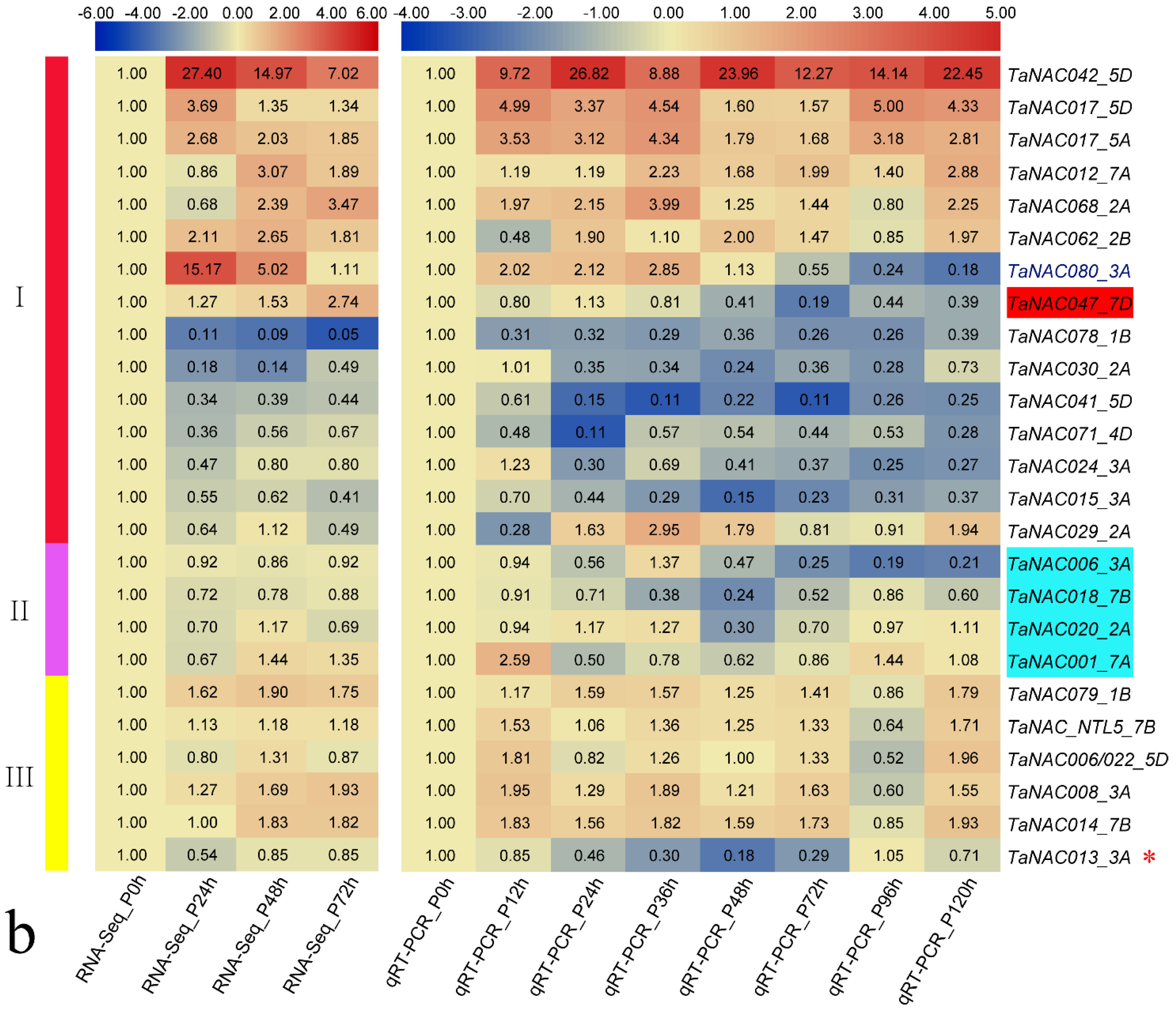
3.3. TaNAC Expression in Wheat under Fungus Infection
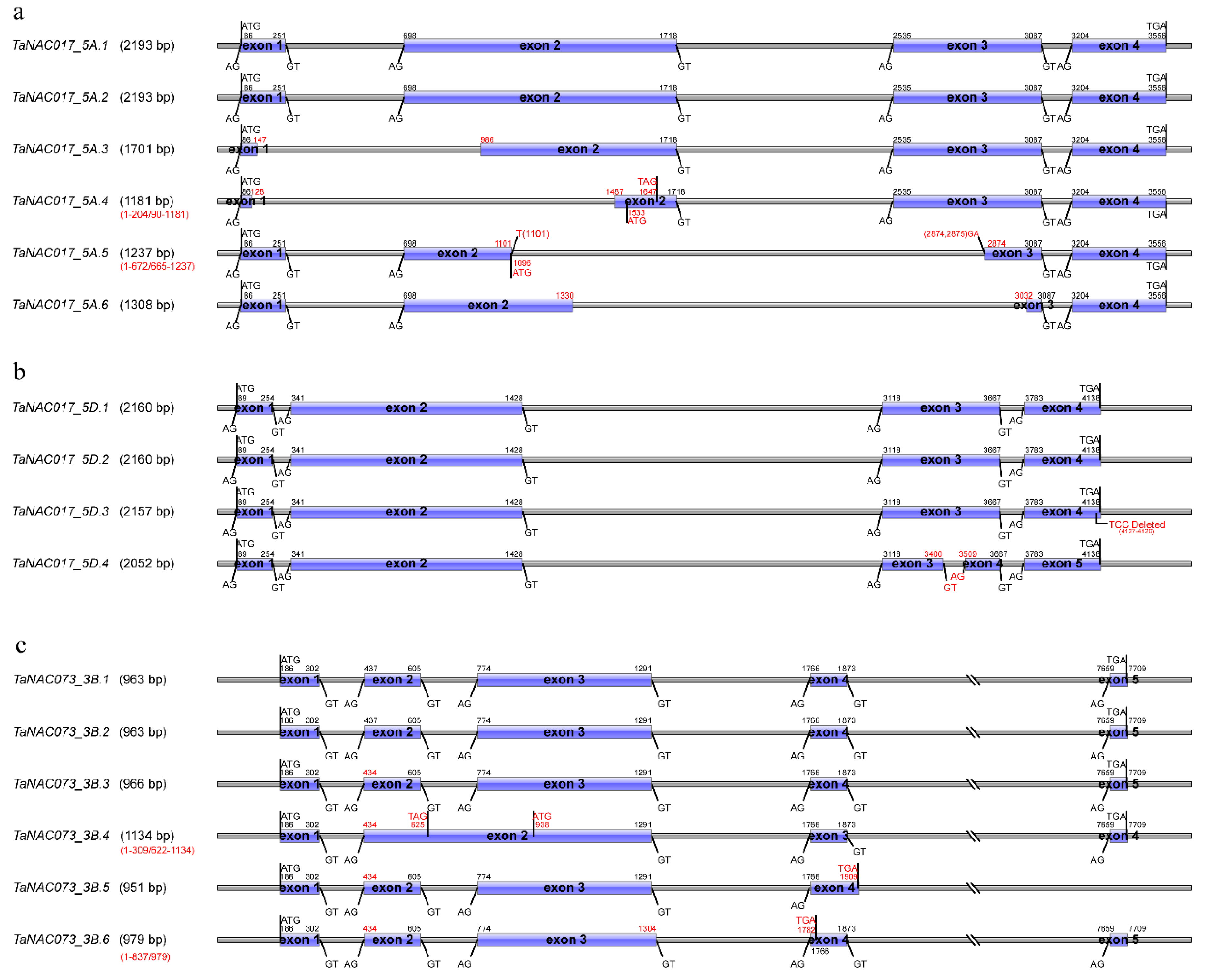
3.4. Structural Variations of TaNAC Transcripts in N9134 under Fungal Stress
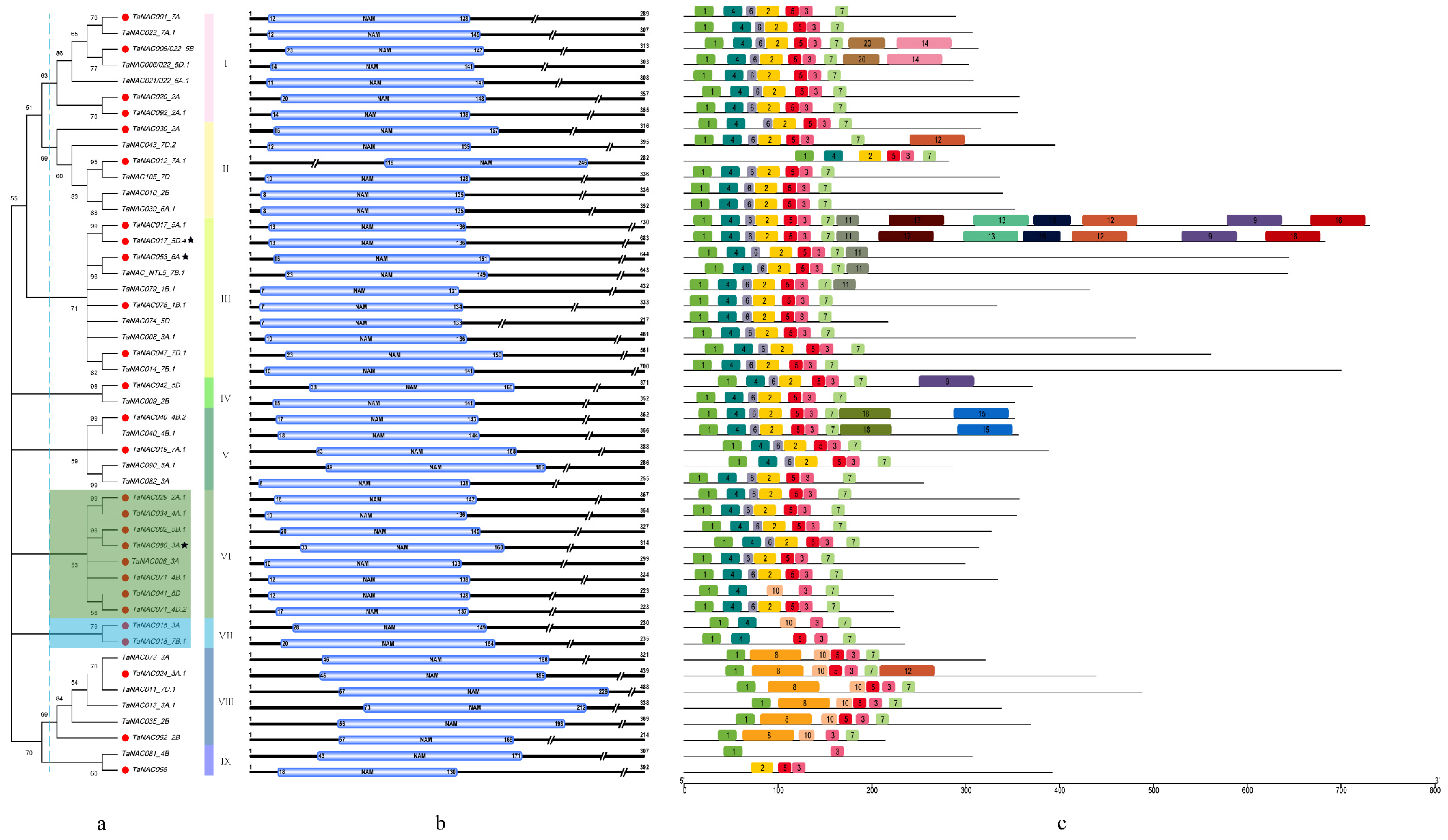
3.5. DE-TaNAC Characterization Based on Phylogenetic Analysis, Conserved Domain, and Motif.s
3.6. Comparative Analysis of TaNAC Membrane-Associated Transcription Factors (MTFs)
4. Discussion
4.1. Specific Spatiotemporal Expression and Alternative Splicing Suggested That TaNAC TF Family Transcripts Could Be Further Enriched and Improved
4.2. Small-Scale Duplications or Deletion Might Also Produce Structural Variations of Transcripts
4.3. Abnormalities in TaNAC Genes with Transmembrane Motifs Might Cause Different Functions by Varying Regulation
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Compliance with Ethical Standards
References
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef] [PubMed]
- Souer, E.; Van Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The no apical meristem gene of petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef]
- Aida, M.; Ishida, T.; Fukaki, H.; Fujisawa, H.; Tasaka, M. Genes involved in organ separation in Arabidopsis: An analysis of the cup-shaped cotyledon mutant. Plant Cell 1997, 9, 841–857. [Google Scholar] [CrossRef] [PubMed]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Olsen, A.N.; Ernst, H.A.; Leggio, L.L.; Skriver, K. NAC transcription factors: Structurally distinct, functionally diverse. Trends Plant Sci. 2005, 10, 79–87. [Google Scholar] [CrossRef]
- Puranik, S.; Sahu, P.P.; Srivastava, P.S.; Prasad, M. NAC proteins: Regulation and role in stress tolerance. Trends Plant Sci. 2012, 17, 369–381. [Google Scholar] [CrossRef]
- Fujita, M.; Fujita, Y.; Maruyama, K.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Tran, L.S.; Yamaguchi-Shinozaki, K.; Shinozaki, K. A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J. Cell Mol. Biol. 2004, 39, 863–876. [Google Scholar] [CrossRef]
- Tran, L.S.; Nakashima, K.; Sakuma, Y.; Simpson, S.D.; Fujita, Y.; Maruyama, K.; Fujita, M.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 2004, 16, 2481–2498. [Google Scholar] [CrossRef]
- Guo, Y.; Gan, S. AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J. Cell Mol. Biol. 2006, 46, 601–612. [Google Scholar] [CrossRef]
- Hu, H.; Dai, M.; Yao, J.; Xiao, B.; Li, X.; Zhang, Q.; Xiong, L. Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc. Natl. Acad. Sci. USA 2006, 103, 12987–12992. [Google Scholar] [CrossRef]
- Xu, B.; Ohtani, M.; Yamaguchi, M.; Toyooka, K.; Wakazaki, M.; Sato, M.; Kubo, M.; Nakano, Y.; Sano, R.; Hiwatashi, Y.; et al. Contribution of NAC transcription factors to plant adaptation to land. Science 2014, 343, 1505–1508. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Montesino, R.; Bouza-Morcillo, L.; Marquez, J.; Ghita, M.; Duran-Nebreda, S.; Gomez, L.; Holdsworth, M.J.; Bassel, G.; Onate-Sanchez, L. A regulatory module controlling ga-mediated endosperm cell expansion is critical for seed germination in Arabidopsis. Mol. Plant 2019, 12, 71–85. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.Y.; Kim, S.Y.; do Hyeon, Y.; Kim, D.H.; Dong, T.; Park, Y.; Jin, J.B.; Joo, S.H.; Kim, S.K.; Hong, J.C.; et al. The Arabidopsis NAC transcription factor ANAC096 cooperates with bZIP-type transcription factors in dehydration and osmotic stress responses. Plant Cell 2013, 25, 4708–4724. [Google Scholar] [CrossRef] [PubMed]
- Liang, C.; Wang, Y.; Zhu, Y.; Tang, J.; Hu, B.; Liu, L.; Ou, S.; Wu, H.; Sun, X.; Chu, J.; et al. OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 10013–10018. [Google Scholar] [CrossRef] [PubMed]
- Pascual, M.B.; Canovas, F.M.; Avila, C. The NAC transcription factor family in maritime pine (Pinus Pinaster): Molecular regulation of two genes involved in stress responses. BMC Plant Biol. 2015, 15, 254. [Google Scholar] [CrossRef]
- Shan, W.; Chen, J.Y.; Kuang, J.F.; Lu, W.J. Banana fruit NAC transcription factor MaNAC5 cooperates with MaWRKYs to enhance the expression of pathogenesis-related genes against Colletotrichum musae. Mol. Plant Pathol. 2016, 17, 330–338. [Google Scholar] [CrossRef]
- Ohtani, M.; Demura, T. The quest for transcriptional hubs of lignin biosynthesis: Beyond the NAC–MYB-gene regulatory network model. Curr. Opin. Biotechnol. 2019, 56, 82–87. [Google Scholar] [CrossRef]
- Murozuka, E.; Massange-Sanchez, J.A.; Nielsen, K.; Gregersen, P.L.; Braumann, I. Genome wide characterization of barley NAC transcription factors enables the identification of grain-specific transcription factors exclusive for the Poaceae family of monocotyledonous plants. PLoS ONE 2018, 13, e0209769. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef]
- Guerin, C.; Roche, J.; Allard, V.; Ravel, C.; Mouzeyar, S.; Bouzidi, M.F. Genome-wide analysis, expansion and expression of the NAC family under drought and heat stresses in bread wheat (T. aestivum L.). PLoS ONE 2019, 14, e0213390. [Google Scholar] [CrossRef]
- Borrill, P.; Harrington, S.A.; Uauy, C. Genome-wide sequence and expression analysis of the NAC transcription factor family in polyploid wheat. G3 (Bethesda) 2017, 7, 3019–3029. [Google Scholar] [CrossRef] [PubMed]
- Cowger, C.; Miranda, L.; Griffey, C.; Hall, M.; Murphy, J.P.; Maxwell, J.; Sharma, I. Wheat Powdery Mildew; Disease Resistance in Wheat, CBA International: Oxfordshire, UK, 2012; pp. 84–119. [Google Scholar]
- Collinge, M.; Boller, T. Differential induction of two potato genes, Stprx2 and StNAC, in response to infection by Phytophthora infestans and to wounding. Plant Mol. Biol. 2001, 46, 521–529. [Google Scholar] [CrossRef] [PubMed]
- Nuruzzaman, M.; Sharoni, A.M.; Kikuchi, S. Roles of NAC transcription factors in the regulation of biotic and abiotic stress responses in plants. Front. Microbiol. 2013, 4, 248. [Google Scholar] [CrossRef]
- Jensen, M.K.; Rung, J.H.; Gregersen, P.L.; Gjetting, T.; Fuglsang, A.T.; Hansen, M.; Joehnk, N.; Lyngkjaer, M.F.; Collinge, D.B. The HvNAC6 transcription factor: A positive regulator of penetration resistance in barley and Arabidopsis. Plant Mol. Biol. 2007, 65, 137–150. [Google Scholar] [CrossRef] [PubMed]
- Kaneda, T.; Taga, Y.; Takai, R.; Iwano, M.; Matsui, H.; Takayama, S.; Isogai, A.; Che, F.S. The transcription factor OsNAC4 is a key positive regulator of plant hypersensitive cell death. EMBO J. 2009, 28, 926–936. [Google Scholar] [CrossRef]
- Zhang, X.M.; Zhang, Q.; Pei, C.L.; Li, X.; Huang, X.L.; Chang, C.Y.; Wang, X.J.; Huang, L.L.; Kang, Z.S. TaNAC2 is a negative regulator in the wheat-stripe rust fungus interaction at the early stage. Physiol. Mol. Plant Pathol. 2018, 102, 144–153. [Google Scholar] [CrossRef]
- Perochon, A.; Kahla, A.; Vranic, M.; Jia, J.; Malla, K.B.; Craze, M.; Wallington, E.; Doohan, F.M. A wheat NAC interacts with an orphan protein and enhances resistance to Fusarium head blight disease. Plant Biotechnol. J. 2019, 17, 1892–1904. [Google Scholar] [CrossRef]
- Feng, H.; Duan, X.; Zhang, Q.; Li, X.; Wang, B.; Huang, L.; Wang, X.; Kang, Z. The target gene of tae-miR164, a novel NAC transcription factor from the NAM subfamily, negatively regulates resistance of wheat to stripe rust. Mol. Plant Pathol. 2014, 15, 284–296. [Google Scholar] [CrossRef]
- Zhang, H.; Yang, Y.; Wang, C.; Liu, M.; Li, H.; Fu, Y.; Wang, Y.; Nie, Y.; Liu, X.; Ji, W. Large-scale transcriptome comparison reveals distinct gene activations in wheat responding to stripe rust and powdery mildew. BMC Genom. 2014, 15, 898. [Google Scholar] [CrossRef]
- International Wheat Genome Sequencing Consortium; Appels, R.; Eversole, K.; Feuillet, C.; Keller, B.; Rogers, J.; Stein, N.; Pozniak, C.J.; Stein, N.; Choulet, F.; et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 2018, 361, eaar7191. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C (T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 1992, 8, 275–282. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evol. Int. J. Org. Evol. 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Distelfeld, A.; Pearce, S.P.; Avni, R.; Scherer, B.; Uauy, C.; Piston, F.; Slade, A.; Zhao, R.; Dubcovsky, J. Divergent functions of orthologous NAC transcription factors in wheat and rice. Plant Mol. Biol. 2012, 78, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Park, J.H.; Kim, J.; Kim, J.J.; Hong, S.; Kim, J.; Kim, J.H.; Woo, H.R.; Hyeon, C.; Lim, P.O.; et al. Time-evolving genetic networks reveal a NAC troika that negatively regulates leaf senescence in Arabidopsis. Proc. Natl. Acad. Sci. USA 2018, 115, E4930–E4939. [Google Scholar] [CrossRef]
- Kim, Y.S.; Kim, S.G.; Park, J.E.; Park, H.Y.; Lim, M.H.; Chua, N.H.; Park, C.M. A membrane-bound NAC transcription factor regulates cell division in Arabidopsis. Plant Cell 2006, 18, 3132–3144. [Google Scholar] [CrossRef]
- Ostria-Gallardo, E.; Ranjan, A.; Chitwood, D.H.; Kumar, R.; Townsley, B.T.; Ichihashi, Y.; Corcuera, L.J.; Sinha, N.R. Transcriptomic analysis suggests a key role for squamosa promoter binding protein like, NAC and YUCCA genes in the heteroblastic development of the temperate rainforest tree Gevuina avellana (Proteaceae). New Phytol. 2016, 210, 694–708. [Google Scholar] [CrossRef]
- Xia, N.; Zhang, G.; Liu, X.Y.; Deng, L.; Cai, G.L.; Zhang, Y.; Wang, X.J.; Zhao, J.; Huang, L.L.; Kang, Z.S. Characterization of a novel wheat NAC transcription factor gene involved in defense response against stripe rust pathogen infection and abiotic stresses. Mol. Biol. Rep. 2010, 37, 3703–3712. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, L.; Xia, C.; Zhao, G.; Jia, J.; Kong, X. The novel wheat transcription factor TaNAC47 enhances multiple abiotic stress tolerances in transgenic plants. Front. Plant Sci. 2015, 6, 1174. [Google Scholar] [CrossRef] [PubMed]
- Jensen, M.K.; Kjaersgaard, T.; Nielsen, M.M.; Galberg, P.; Petersen, K.; O’Shea, C.; Skriver, K. The Arabidopsis thaliana NAC transcription factor family: Structure-function relationships and determinants of ANAC019 stress signalling. Biochem. J. 2010, 426, 183–196. [Google Scholar] [CrossRef] [PubMed]
- Shahnejat-Bushehri, S.; Tarkowska, D.; Sakuraba, Y.; Balazadeh, S. Arabidopsis NAC transcription factor JUB1 regulates GA/BR metabolism and signalling. Nat. Plants 2016, 2, 16013. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, A.J.; Gelbart, W.M.; Lewontin, R.C.; Miller, J.H. Modern Genetic Analysis: Integrating Genes and Genomes; Macmillan: London, UK, 2002; Volume 1. [Google Scholar]
- Huang, Q.; Wang, Y. Overexpression of TaNAC2D displays opposite responses to abiotic stresses between seedling and mature stage of transgenic Arabidopsis. Front. Plant Sci. 2016, 7, 1754. [Google Scholar] [CrossRef]
- Xia, N.; Zhang, G.; Sun, Y.F.; Zhu, L.; Xu, L.S.; Chen, X.M.; Liu, B.; Yu, Y.T.; Wang, X.J.; Huang, L.L.; et al. TaNAC8, a novel NAC transcription factor gene in wheat, responds to stripe rust pathogen infection and abiotic stresses. Physiol. Mol. Plant Pathol. 2010, 74, 394–402. [Google Scholar] [CrossRef]
- Wang, F.; Lin, R.; Feng, J.; Chen, W.; Qiu, D.; Xu, S. TaNAC1 acts as a negative regulator of stripe rust resistance in wheat, enhances susceptibility to Pseudomonas syringae, and promotes lateral root development in transgenic Arabidopsis thaliana. Front. Plant Sci. 2015, 6, 108. [Google Scholar] [CrossRef]
- Uauy, C.; Distelfeld, A.; Fahima, T.; Blechl, A.; Dubcovsky, J. A NAC Gene regulating senescence improves grain protein, zinc, and iron content in wheat. Science 2006, 314, 1298–1301. [Google Scholar] [CrossRef]
- Zhang, H.; Mao, R.; Wang, Y.; Zhang, L.; Wang, C.; Lv, S.; Liu, X.; Wang, Y.; Ji, W. Transcriptome-wide alternative splicing modulation during plant-pathogen interactions in wheat. Plant Sci. 2019, 288, 110160. [Google Scholar] [CrossRef] [PubMed]
- Mastrangelo, A.M.; Marone, D.; Laido, G.; De Leonardis, A.M.; De Vita, P. Alternative splicing: Enhancing ability to cope with stress via transcriptome plasticity. Plant Sci. 2012, 185–186, 40–49. [Google Scholar] [CrossRef]
- Fan, K.; Wang, M.; Miao, Y.; Ni, M.; Bibi, N.; Yuan, S.; Li, F.; Wang, X. Molecular evolution and expansion analysis of the NAC transcription factor in Zea mays. PLoS ONE 2014, 9, e111837. [Google Scholar] [CrossRef]
- Fan, K.; Li, F.; Chen, J.; Li, Z.; Lin, W.; Cai, S.; Liu, J.; Lin, W. Asymmetric evolution and expansion of the NAC transcription factor in polyploidized cotton. Front. Plant Sci. 2018, 9, 47. [Google Scholar] [CrossRef] [PubMed]
- Shang, H.; Li, W.; Zou, C.; Yuan, Y. Analyses of the NAC transcription factor gene family in Gossypium raimondii Ulbr: Chromosomal location, structure, phylogeny, and expression patterns. J. Integr. Plant Biol. 2013, 55, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Glover, N.M.; Daron, J.; Pingault, L.; Vandepoele, K.; Paux, E.; Feuillet, C.; Choulet, F. Small-scale gene duplications played a major role in the recent evolution of wheat chromosome 3B. Genome Biol. 2015, 16, 188. [Google Scholar] [CrossRef] [PubMed]
- Stein, G.S.; van Wijnen, A.J.; Stein, J.L.; Lian, J.B.; Montecino, M.; Choi, J.; Zaidi, K.; Javed, A. Intranuclear trafficking of transcription factors: Implications for biological control. J. Cell Sci. 2000, 113 Pt 14, 2527–2533. [Google Scholar]
- Kim, S.G.; Kim, S.Y.; Park, C.M. A membrane-associated NAC transcription factor regulates salt-responsive flowering via FLOWERING LOCUS T in Arabidopsis. Planta 2007, 226, 647–654. [Google Scholar] [CrossRef]
- Kim, S.Y.; Kim, S.G.; Kim, Y.S.; Seo, P.J.; Bae, M.; Yoon, H.K.; Park, C.M. Exploring membrane-associated NAC transcription factors in Arabidopsis: Implications for membrane biology in genome regulation. Nucleic Acids Res. 2007, 35, 203–213. [Google Scholar] [CrossRef]
- Seo, P.J.; Park, C.M. A membrane-bound NAC transcription factor as an integrator of biotic and abiotic stress signals. Plant Signal. Behav. 2010, 5, 481–483. [Google Scholar] [CrossRef]
- Lee, S.; Seo, P.J.; Lee, H.J.; Park, C.M. A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J. Cell Mol. Biol. 2012, 70, 831–844. [Google Scholar] [CrossRef]

| Number | Gene Names | Types |
|---|---|---|
| 3 | TraesCS5A02G271500.1, TraesCS5B02G271800.1, TraesCS5D02G279100.1 | No domain |
| 8 | TraesCS2A02G326600.3, TraesCS2D02G101300.1, TraesCS2D02G336300.2, TraesCS4A02G213100.1, TraesCS5B02G480900.1, TraesCS5D02G481200.2, TraesCS6D02G266000.1, TraesCS7A02G152400.1 | One short NAM domain |
| 1 | TraesCS1D02G004800.1 | One normal NAM, one DnaJ, and one ZnF_BED domains |
| 1 | TraesCS7B02G461700.1 | One normal NAM domain and two AA_Kinase domains |
| 8 | TraesCS2A02G328100.1, TraesCS2A02G363700.1, TraesCS2B02G381700.1, TraesCS2D02G334800.1, TraesCS2D02G361500.1, TraesCS3A02G269900.1, TraesCS3B02G303800.1, TraesCS3D02G269600.1 | Two normal NAM domains |
| 1 | TraesCS2A02G326600.2 | One normal NAM domain and one signal peptide in N-terminal |
| 537 | All other transcripts | One normal NAM Domain |
| No. | Types | DEG | Non-DEG | p Value |
|---|---|---|---|---|
| 1 | Selected unigenes | 28 | 22 | 1.000 |
| 2 | Total transcripts | 117 | 69 | 0.058 |
| 3 | Cloned transcripts | 101 | 66 | 0.244 |
| 4 | Assembly transcripts | 9 | 2 | 0.085 |
| 5 | Transcripts from assembly and IWGSC | 7 | 1 | 0.073 |
| 6 | Normal encoded transcripts | 93 | 61 | 0.272 |
| 7 | Cloned and normal encoded transcripts | 78 | 59 | 0.826 |
| 8 | Abnormal encoded transcripts | 24 | 8 | 0.030 * |
| 9 | Cloned and abnormal encoded transcripts | 23 | 7 | 0.023 * |
| Sequence Types | DEG | Non-DEG | p-Value |
|---|---|---|---|
| Selected unigenes from N9134 | 28 | 22 | 1.000 |
| Transcripts with transmembrane motifs obtained from N9134 | 12 | 7 | 0.530 |
| Transcripts with transmembrane motifs cloned from N9134 | 9 | 7 | 0.984 |
| Transcript Types | With Transmembrane Motifs | Without Transmembrane Motifs | p-Value |
|---|---|---|---|
| Transcripts from IWGSC RefSeq v1.1 | 36 | 523 | 1.000 |
| Total normal encoded transcripts from N9134 | 19 | 135 | 0.003 ** |
| Cloned and normal encoded transcripts from N9134 | 16 | 121 | 0.012 * |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lv, S.; Guo, H.; Zhang, M.; Wang, Q.; Zhang, H.; Ji, W. Large-Scale Cloning and Comparative Analysis of TaNAC Genes in Response to Stripe Rust and Powdery Mildew in Wheat (Triticum aestivum L.). Genes 2020, 11, 1073. https://doi.org/10.3390/genes11091073
Lv S, Guo H, Zhang M, Wang Q, Zhang H, Ji W. Large-Scale Cloning and Comparative Analysis of TaNAC Genes in Response to Stripe Rust and Powdery Mildew in Wheat (Triticum aestivum L.). Genes. 2020; 11(9):1073. https://doi.org/10.3390/genes11091073
Chicago/Turabian StyleLv, Shikai, Huan Guo, Min Zhang, Qiaohui Wang, Hong Zhang, and Wanquan Ji. 2020. "Large-Scale Cloning and Comparative Analysis of TaNAC Genes in Response to Stripe Rust and Powdery Mildew in Wheat (Triticum aestivum L.)" Genes 11, no. 9: 1073. https://doi.org/10.3390/genes11091073
APA StyleLv, S., Guo, H., Zhang, M., Wang, Q., Zhang, H., & Ji, W. (2020). Large-Scale Cloning and Comparative Analysis of TaNAC Genes in Response to Stripe Rust and Powdery Mildew in Wheat (Triticum aestivum L.). Genes, 11(9), 1073. https://doi.org/10.3390/genes11091073





