Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium
Abstract
1. Introduction
2. Materials and Methods
2.1. TargetScan (v7.2)
2.2. RNA22 (v2)
2.3. TCGA-LUAD Dataset
2.4. Viral Genome Analysis
2.5. miRNA Differential Expression Analysis
3. Results
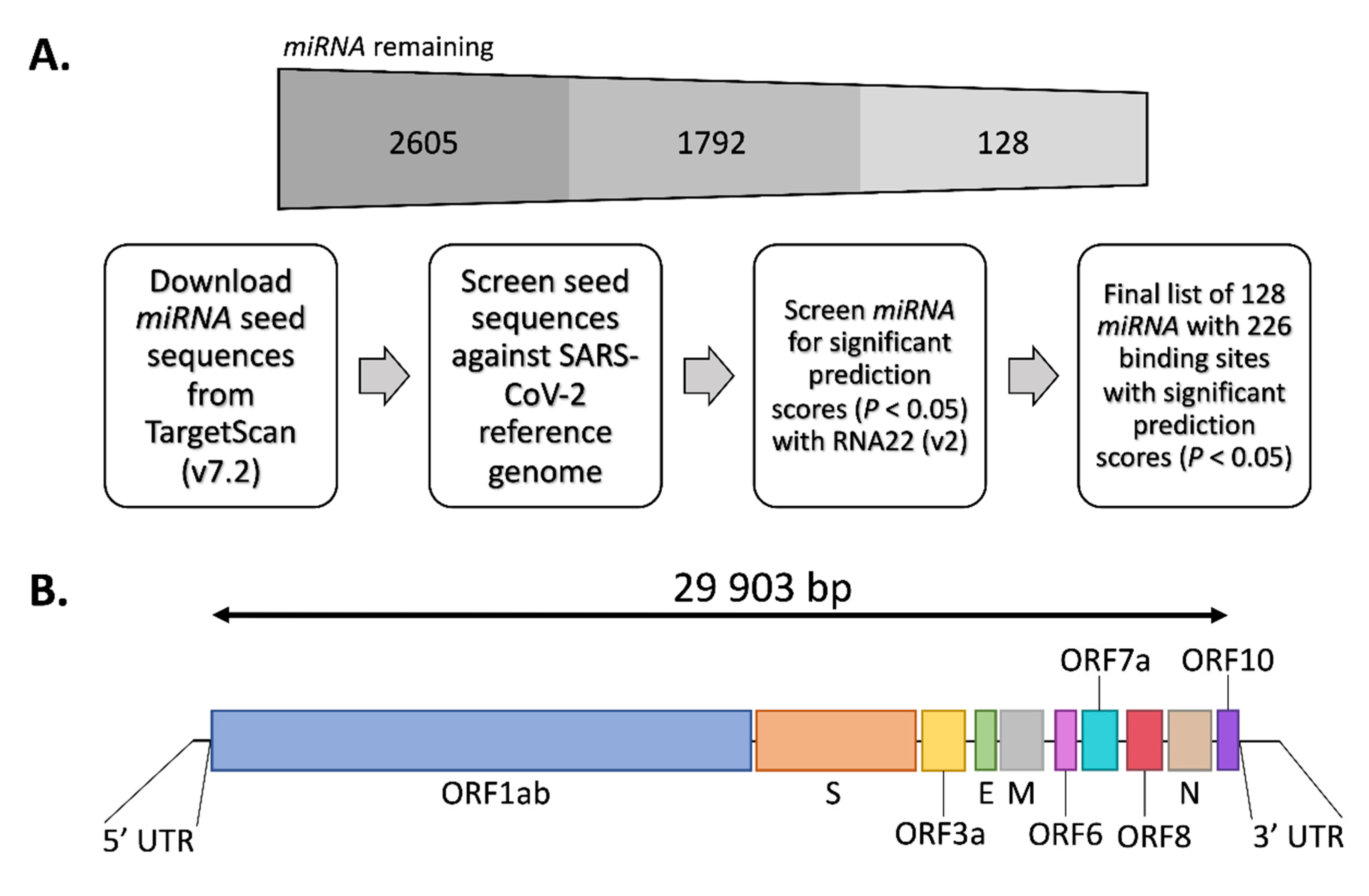
3.1. Target Prediction in SARS-CoV-2
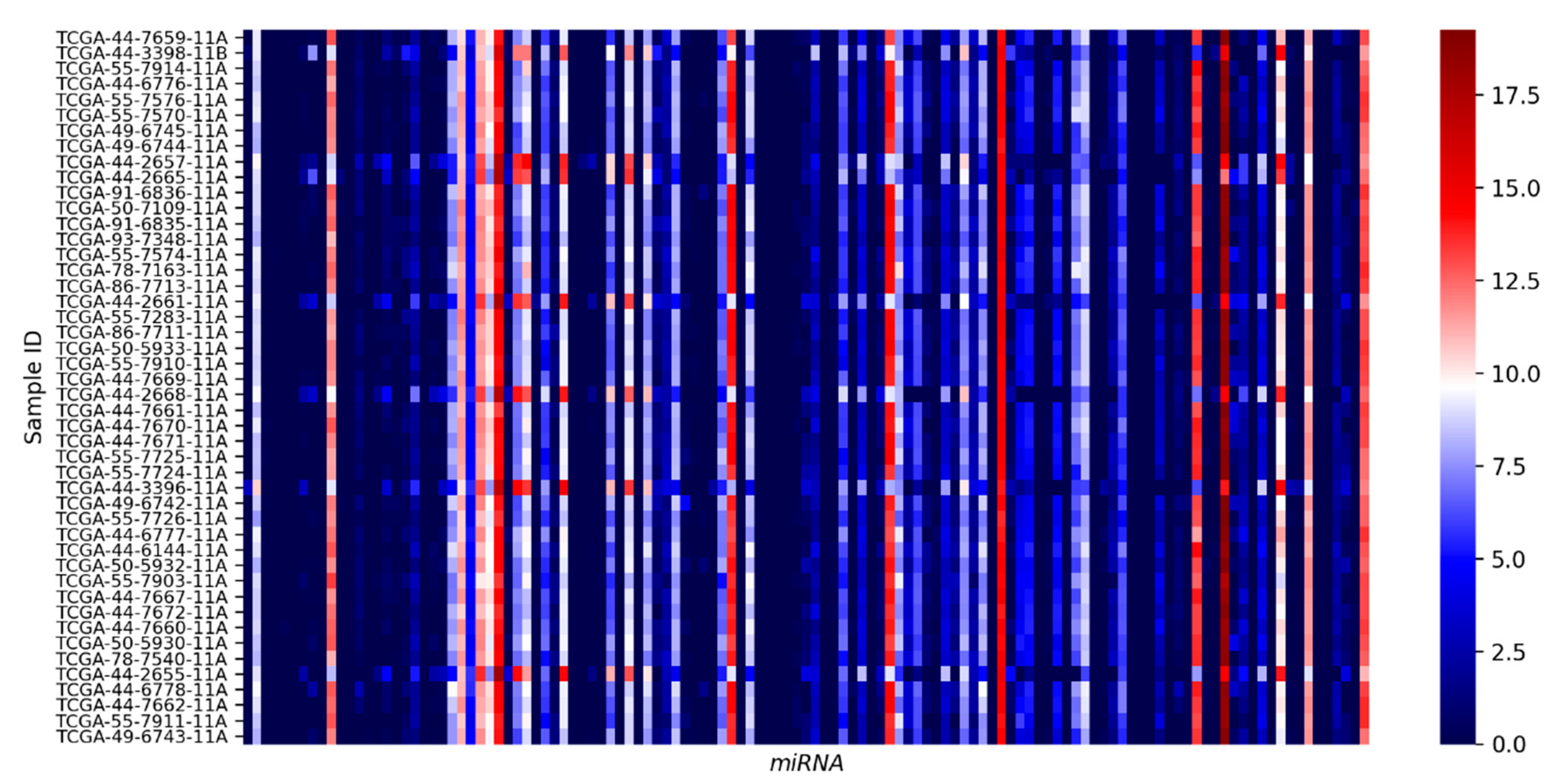
3.2. Candidate miRNA Expression in Normal Lung Epithelia
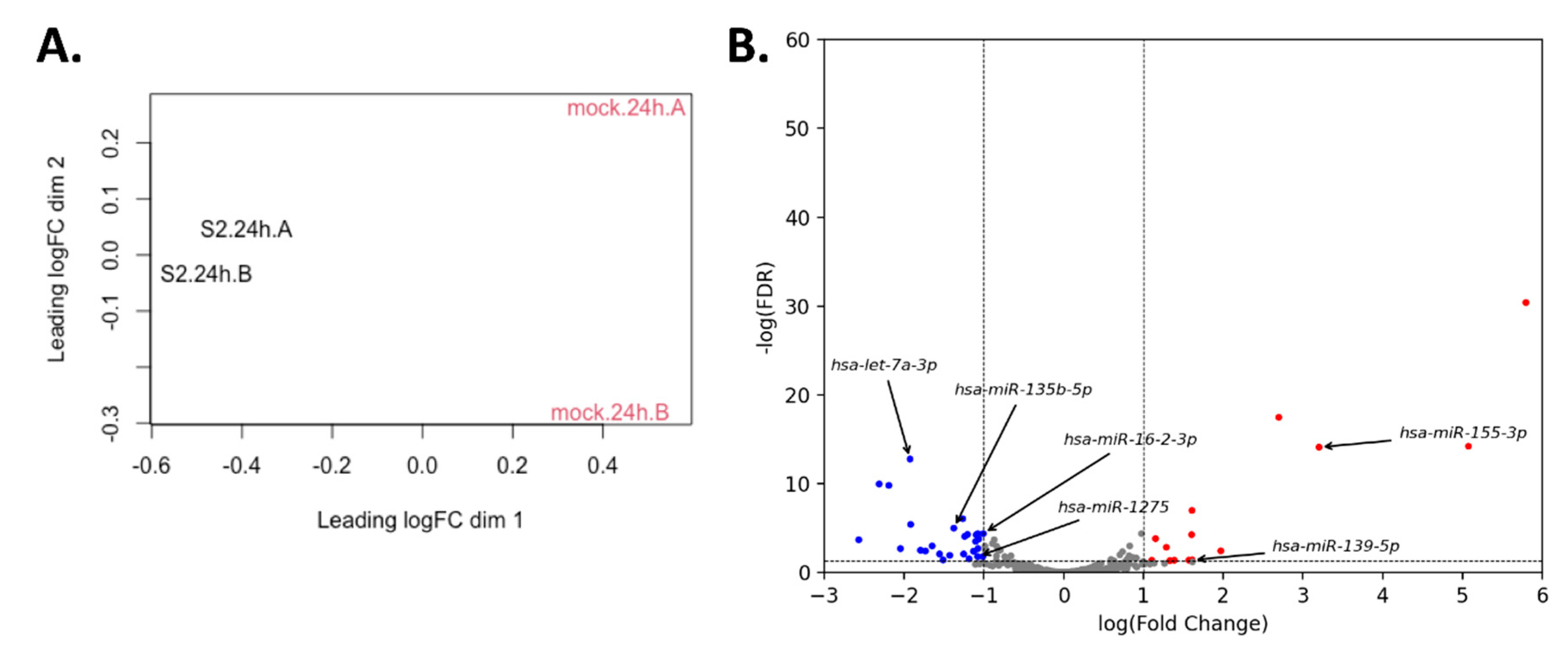
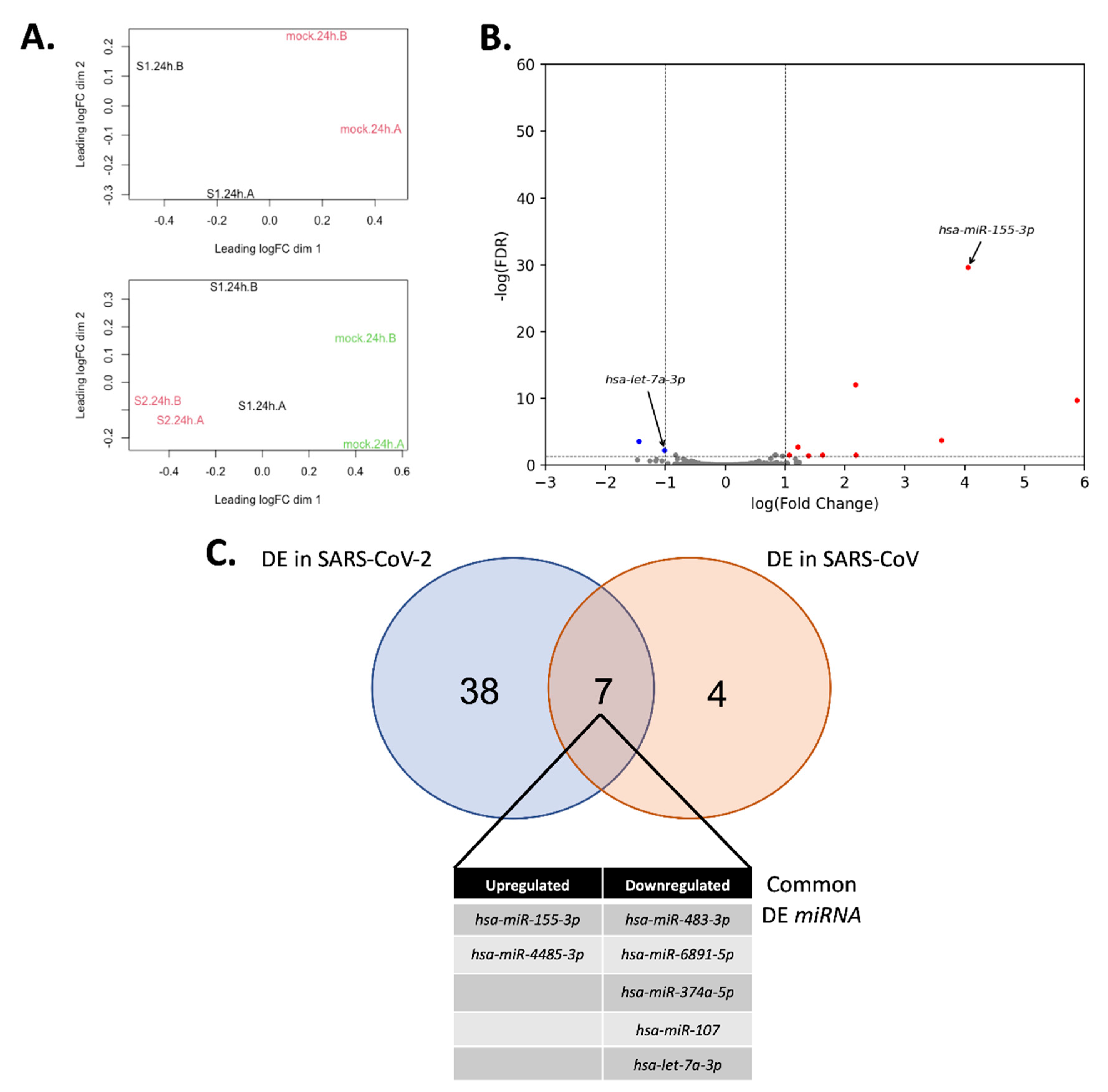
3.3. miRNA Expression Changes upon SARS-CoV-2 Infection
3.4. Conservation of miRNA Binding in Coronaviruses
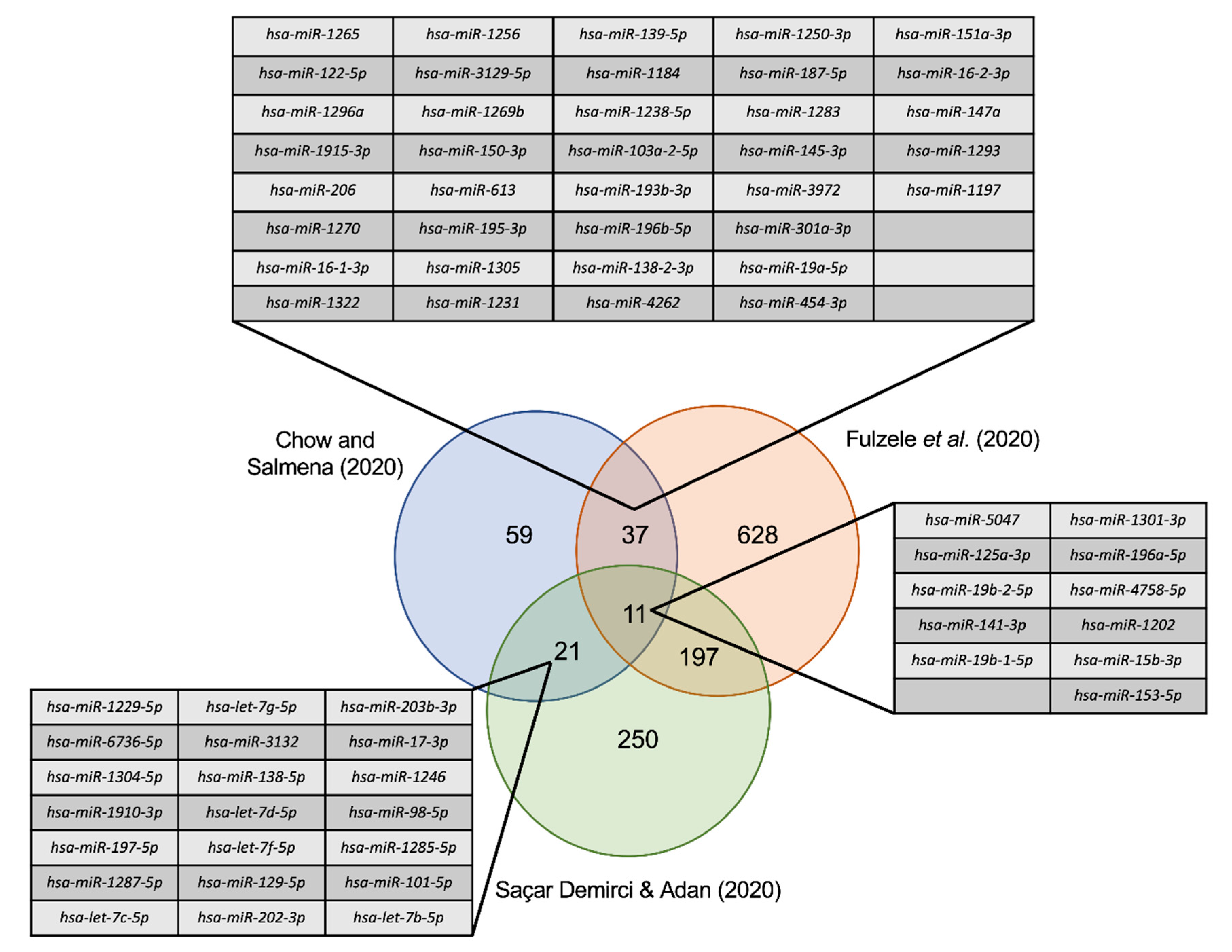
3.5. Comparison to Other miRNA SARS-CoV-2 Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Madjid, M.; Safavi-Naeini, P.; Solomon, S.D.; Vardeny, O. Potential Effects of Coronaviruses on the Cardiovascular System: A Review. JAMA Cardiol. 2020. [Google Scholar] [CrossRef]
- Bhatti, J.S.; Bhatti, G.K.; Khullar, N.; Reddy, A.P.; Reddy, P.H. Therapeutic Strategies in the Development of Anti-viral Drugs and Vaccines against SARS-CoV-2 Infection. Mol. Neurobiol. 2020, 1–22. [Google Scholar] [CrossRef]
- Wiersinga, W.J.; Rhodes, A.; Cheng, A.C.; Peacock, S.J.; Prescott, H.C. Pathophysiology, Transmission, Diagnosis, and Treatment of Coronavirus Disease 2019 (COVID-19). JAMA 2020. [Google Scholar] [CrossRef] [PubMed]
- Al-Kassmy, J.; Pedersen, J.; Kobinger, G. Vaccine Candidates Against Coronavirus Infections. Where Does COVID-19 Stand? Viruses 2020, 12, 861. [Google Scholar] [CrossRef]
- Bartel, B. Metazoan MicroRNAs. Cell 2018, 173, 20–51. [Google Scholar] [CrossRef] [PubMed]
- Kawamata, T.; Tomari, Y. Making RISC. Trends Biochem. Sci. 2010, 35, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, C.-D.; Joshua-Tor, L. Eukaryotic Argonautes come into focus. Trends Biochem. Sci. 2013, 38, 263–271. [Google Scholar] [CrossRef] [PubMed]
- Macfarlane, L.-A.; Murphy, P.R. MicroRNA: Biogenesis, Function and Role in Cancer. Curr. Genom. 2010, 11, 537–561. [Google Scholar] [CrossRef]
- Sontheimer, E.J. Assembly and function of RNA silencing complexes. Nat. Rev. Mol. Cell Biol. 2005, 6, 127–138. [Google Scholar] [CrossRef]
- Lewis, B.P.; Burge, C.B.; Bartel, B. Conserved Seed Pairing, Often Flanked by Adenosines, Indicates that Thousands of Human Genes are MicroRNA Targets. Cell 2005, 120, 15–20. [Google Scholar] [CrossRef] [PubMed]
- Brennecke, J.; Stark, A.; Russell, R.B.; Cohen, S.M. Principles of microRNA-target recognition. PLoS Biol. 2005, 3, e85. [Google Scholar] [CrossRef] [PubMed]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef] [PubMed]
- Trobaugh, D.; Klimstra, W.B. MicroRNA Regulation of RNA Virus Replication and Pathogenesis. Trends Mol. Med. 2016, 23, 80–93. [Google Scholar] [CrossRef]
- Pedersen, I.M.; Cheng, G.; Wieland, S.; Volinia, S.; Croce, C.M.; Chisari, F.V.; David, M. Interferon modulation of cellular microRNAs as an antiviral mechanism. Nature 2007, 449, 919–922. [Google Scholar] [CrossRef]
- Peng, S.; Wang, J.; Wei, S.; Li, C.; Zhou, K.; Hu, J.; Ye, X.; Yan, J.; Liu, W.; Gao, G.F.; et al. Endogenous Cellular MicroRNAs Mediate Antiviral Defense against Influenza A Virus. Mol. Ther. Nucleic Acids 2018, 10, 361–375. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Kumar, A.; Ingle, H.; Kumar, S.; Mishra, R.; Verma, M.K.; Biswas, D.; Kumar, N.S.; Mishra, A.; Raut, A.A.; et al. MicroRNA hsa-miR-324-5p Suppresses H5N1 Virus Replication by Targeting the Viral PB1 and Host CUEDC2. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Gao, J.; Gao, L.; Li, R.; Lai, Z.; Zhang, Z.; Fan, X.-H. Integrated analysis of microRNA-mRNA expression in A549 cells infected with influenza A viruses (IAVs) from different host species. Virus Res. 2019, 263, 34–46. [Google Scholar] [CrossRef]
- Wyler, E.; Mösbauer, K.; Franke, V.; Diag, A.; Gottula, L.T.; Arsie, R.; Klironomos, F.; Koppstein, D.; Ayoub, S.; Buccitelli, C.; et al. Bulk and single-cell gene expression profiling of SARS-CoV-2 infected human cell lines identifies molecular targets for therapeutic intervention. BioRxiv 2020. [Google Scholar] [CrossRef]
- Fulzele, S.; Sahay, B.; Yusufu, I.; Lee, T.J.; Sharma, A.; Kolhe, R.; Isales, C.M. COVID-19 Virulence in Aged Patients Might Be Impacted by the Host Cellular MicroRNAs Abundance/Profile. Aging Dis. 2020, 11, 509–522. [Google Scholar] [CrossRef]
- Demirci, M.D.S.; Adan, A. Computational analysis of microRNA-mediated interactions in SARS-CoV-2 infection. PeerJ 2020, 8, e9369. [Google Scholar] [CrossRef]
- Miranda, K.C.; Huynh, T.; Tay, Y.; Ang, Y.-S.; Tam, W.-L.; Thomson, A.M.; Lim, B.; Rigoutsos, I. A Pattern-Based Method for the Identification of MicroRNA Binding Sites and Their Corresponding Heteroduplexes. Cell 2006, 126, 1203–1217. [Google Scholar] [CrossRef] [PubMed]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshal, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets--update. Nucleic Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Collisson, E.A.; Berger, A.H.; Beer, D.G.; Hammerman, P.S.; Herman, J.G.; Jurisica, I.; Kwiatkowski, D.; Robertson, G.; The Cancer Genome Atlas Research Network; Cancer Genome Atlas Research Network; et al. Comprehensive molecular profiling of lung adenocarcinoma. Nature 2014, 511, 543–550. [Google Scholar] [CrossRef]
- De Wit, E.; Van Doremalen, N.; Falzarano, D.; Munster, V. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Genet. 2016, 14, 523–534. [Google Scholar] [CrossRef]
- Peiris, J.S.M.; Guan, Y.; Yuen, K.-Y. Severe acute respiratory syndrome. Nat. Med. 2004, 10, S88–S97. [Google Scholar] [CrossRef]
- Chafekar, A.; Fielding, B.C. MERS-CoV: Understanding the Latest Human Coronavirus Threat. Viruses 2018, 10, 93. [Google Scholar] [CrossRef]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Girardi, E.; López, P.; Pfeffer, S. On the Importance of Host MicroRNAs During Viral Infection. Front. Genet. 2018, 9, 439. [Google Scholar] [CrossRef]
- Ameres, S.L.; Martinez, J.; Schroeder, R. Molecular Basis for Target RNA Recognition and Cleavage by Human RISC. Cell 2007, 130, 101–112. [Google Scholar] [CrossRef]
- Kertesz, M.; Iovino, N.; Unnerstall, U.; Gaul, U.; Segal, E. The role of site accessibility in microRNA target recognition. Nat. Genet. 2007, 39, 1278–1284. [Google Scholar] [CrossRef] [PubMed]
- Long, D.; Lee, R.; Williams, P.; Chan, C.Y.; Ambros, V.; Ding, Y. Potent effect of target structure on microRNA function. Nat. Struct. Mol. Biol. 2007, 14, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Yang, N.; Leibowitz, J.L. The structure and functions of coronavirus genomic 3′ and 5′ ends. Virus Res. 2015, 206, 120–133. [Google Scholar] [CrossRef] [PubMed]
- Sola, I.; Almazán, F.; Zuñiga, S.; Enjuanes, L. Continuous and Discontinuous RNA Synthesis in Coronaviruses. Annu. Rev. Virol. 2015, 2, 265–288. [Google Scholar] [CrossRef] [PubMed]
- Hamada-Tsutsumi, S.; Naito, Y.; Sato, S.; Takaoka, A.; Kawashima, K.; Isogawa, M.; Ochiya, T.; Tanaka, Y. The antiviral effects of human microRNA miR-302c-3p against hepatitis B virus infection. Aliment. Pharmacol. Ther. 2019, 49, 1060–1070. [Google Scholar] [CrossRef]
- Skalsky, R.L.; Cullen, B.R. Viruses, microRNAs, and host interactions. Annu. Rev. Microbiol. 2010, 64, 123–141. [Google Scholar] [CrossRef]
- Tycowski, K.T.; Guo, Y.E.; Lee, N.; Moss, W.N.; Vallery, T.K.; Xie, M.; Steitz, J.A. Viral noncoding RNAs: More surprises. Genes Dev. 2015, 29, 567–584. [Google Scholar] [CrossRef]
- Perez, J.T.; Varble, A.; Sachidanandam, R.; Zlatev, I.; Manoharan, M.; García-Sastre, A.; Tenoever, B.R. Influenza A virus-generated small RNAs regulate the switch from transcription to replication. Proc. Natl. Acad. Sci. USA 2010, 107, 11525–11530. [Google Scholar] [CrossRef]
- Morales, L.; Oliveros, J.C.; Fernández-Delgado, R.; Tenoever, B.R.; Enjuanes, L.; Sola, I. SARS-CoV-Encoded Small RNAs Contribute to Infection-Associated Lung Pathology. Cell Host Microbe 2017, 21, 344–355. [Google Scholar] [CrossRef]
- Bogerd, H.P.; Skalsky, R.L.; Kennedy, E.M.; Furuse, Y.; Whisnant, A.W.; Flores, O.; Schultz, K.L.W.; Putnam, N.; Barrows, N.J.; Sherry, B.; et al. Replication of Many Human Viruses Is Refractory to Inhibition by Endogenous Cellular MicroRNAs. J. Virol. 2014, 88, 8065–8076. [Google Scholar] [CrossRef] [PubMed]
- Backes, S.; Langlois, R.A.; Schmid, S.; Varble, A.; Shim, J.V.; Sachs, D.; Tenoever, B.R. The Mammalian response to virus infection is independent of small RNA silencing. Cell Rep. 2014, 8, 114–125. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, M.; Jing, Q.; Georgel, P.; New, L.; Chen, J.; Mols, J.; Kang, Y.J.; Jiang, Z.; Du, X.; Cook, R.; et al. Hypersusceptibility to Vesicular Stomatitis Virus Infection in Dicer1-Deficient Mice Is Due to Impaired miR24 and miR93 Expression. Immunity 2007, 27, 123–134. [Google Scholar] [CrossRef] [PubMed]
| miRNA with a Significant Predicted Binding Site | ||||
|---|---|---|---|---|
| hsa-let-7i-5p | hsa-miR-182-3p | hsa-let-7d-5p | hsa-miR-19b-2-5p | hsa-miR-142-3p.1 |
| hsa-miR-4701-3p | hsa-miR-1270 | hsa-miR-1184 | hsa-miR-138-1-3p | hsa-miR-1284 |
| hsa-miR-1273g-5p | hsa-miR-4298 | hsa-miR-150-3p | hsa-miR-202-3p | hsa-miR-138-2-3p |
| hsa-miR-1265 | hsa-miR-16-1-3p | hsa-miR-15b-3p | hsa-miR-103a-2-5p | hsa-miR-1208 |
| hsa-miR-6736-5p | hsa-let-7b-5p | hsa-let-7f-5p | hsa-miR-1185-2-3p | hsa-miR-1273g-3p |
| hsa-miR-122-5p | hsa-miR-197-5p | hsa-miR-129-5p | hsa-miR-1976 | hsa-miR-4262 |
| hsa-miR-1229-5p | hsa-miR-1322 | hsa-miR-1202 | hsa-miR-301a-3p | hsa-miR-1273e |
| hsa-miR-6511a-5p | hsa-miR-1275 | hsa-miR-4665-5p | hsa-miR-206 | hsa-miR-17-3p |
| hsa-miR-1269a | hsa-miR-196a-5p | hsa-miR-1267 | hsa-miR-98-5p | hsa-miR-1304-3p |
| hsa-miR-4420 | hsa-miR-1305 | hsa-miR-1231 | hsa-miR-4500 | hsa-miR-19a-5p |
| hsa-miR-1301-3p | hsa-miR-152-5p | hsa-miR-1238-5p | hsa-miR-147a | hsa-miR-1302 |
| hsa-miR-1256 | hsa-miR-3132 | hsa-miR-19b-1-5p | hsa-miR-3116 | hsa-miR-1237-3p |
| hsa-miR-1910-3p | hsa-miR-138-5p | hsa-miR-3118 | hsa-miR-141-3p | hsa-miR-191-3p |
| hsa-miR-1915-5p | hsa-miR-134-5p | hsa-miR-6874-3p | hsa-miR-185-3p | hsa-miR-1299 |
| hsa-miR-1292-5p | hsa-miR-1298-3p | hsa-miR-1199-5p | hsa-miR-134-3p | hsa-miR-146a-3p |
| hsa-let-7g-5p | hsa-miR-195-3p | hsa-miR-1254 | hsa-miR-1238-3p | hsa-miR-613 |
| hsa-miR-3129-5p | hsa-miR-1250-5p | hsa-miR-1224-5p | hsa-miR-193b-5p | hsa-let-7f-2-3p |
| hsa-miR-1287-5p | hsa-miR-1269b | hsa-miR-550a-3p | hsa-miR-1250-3p | hsa-miR-196b-5p |
| hsa-let-7c-5p | hsa-miR-153-5p | hsa-miR-202-5p | hsa-miR-142-3p.2 | hsa-miR-3972 |
| hsa-let-7e-5p | hsa-miR-125a-3p | hsa-miR-187-5p | hsa-miR-135a-5p | hsa-miR-143-5p |
| hsa-miR-1304-5p | hsa-miR-139-5p | hsa-miR-101-5p | hsa-miR-1251-3p | hsa-miR-151a-3p |
| hsa-miR-4758-5p | hsa-miR-5047 | hsa-miR-1972 | hsa-miR-1185-1-3p | hsa-miR-203a-3p.2 |
| hsa-miR-135b-5p | hsa-miR-155-3p | hsa-miR-145-3p | hsa-miR-1233-3p | hsa-miR-1293 |
| hsa-miR-203b-3p | hsa-miR-4458 | hsa-miR-454-3p | hsa-miR-4518 | hsa-miR-1291 |
| hsa-let-7a-3p | hsa-miR-16-2-3p | hsa-miR-193b-3p | hsa-miR-1283 | hsa-miR-1285-5p |
| hsa-miR-1246 | hsa-miR-1197 | hsa-miR-124-5p | ||
| Differentially Expressed miRNA | |||
|---|---|---|---|
| Upregulated miRNA | Downregulated miRNA | ||
| hsa-miR-4485-3p | hsa-miR-501-5p | hsa-let-7a-3p | hsa-miR-26b-3p |
| hsa-miR-483-3p | hsa-miR-181-5p | hsa-miR-374a-5p | hsa-miR-23c |
| hsa-miR-6891-5p | hsa-miR-4745-3p | hsa-miR-374a-3p | hsa-miR-374c-5p |
| hsa-miR-4284 | hsa-miR-194-5p | hsa-miR-374b-3p | |
| hsa-miR-4463 | hsa-miR-4454 | hsa-miR-26a-1-3p | |
| hsa-miR-155-3p | hsa-miR-135b-5p | hsa-miR-365a-3p | |
| hsa-miR-483-5p | hsa-miR-16-2-3p | hsa-miR-365b-3p | |
| hsa-miR-12136 | hsa-miR-23b-3p | hsa-miR-181-3p | |
| hsa-miR-155-5p | hsa-miR-21-5p | hsa-miR-940 | |
| hsa-miR-107 | hsa-let-7f-1-3p | hsa-miR-362-3p | |
| hsa-miR-125b-5p | hsa-miR-429 | hsa-miR-1275 | |
| hsa-miR-29b-2-5p | hsa-miR-5701 | hsa-miR-1296-5p | |
| hsa-miR-139-5p | hsa-miR-450b-5p | hsa-miR-126-5p | |
| hsa-miR-299-5p | hsa-miR-7-1-3p | hsa-miR-548d-3p | |
| SARS-CoV | MERS-CoV | ||
|---|---|---|---|
| hsa-let-7i-5p | hsa-miR-1208 | hsa-let-7i-5p | hsa-let-7a-2-3p |
| hsa-let-7b-5p | hsa-miR-4500 | hsa-let-7c-5p | hsa-miR-103a-2-5p |
| hsa-let-7c-5p | hsa-miR-101-3p.2 | hsa-let-7e-5p | hsa-miR-4500 |
| hsa-let-7e-5p | hsa-let-7a-2-3p | hsa-let-7g-5p | hsa-miR-10a-5p |
| hsa-let-7g-5p | hsa-miR-98-5p | hsa-let-7b-5p | hsa-miR-101-5p |
| hsa-miR-1202 | hsa-let-7i-3p | hsa-miR-1202 | hsa-miR-10b-5p |
| hsa-miR-105-3p | hsa-let-7g-3p | hsa-let-7d-5p | hsa-miR-1185-2-3p |
| hsa-miR-1224-5p | hsa-miR-1197 | hsa-let-7f-5p | hsa-miR-103b |
| hsa-miR-4458 | hsa-miR-101-3p.1 | hsa-miR-98-5p | hsa-miR-1185-1-3p |
| hsa-let-7f-5p | hsa-miR-1199-5p | hsa-miR-1224-5p | |
| hsa-miR-1205 | hsa-miR-103b | hsa-miR-1184 | |
| hsa-miR-1184 | hsa-miR-10b-5p | hsa-miR-105-3p | |
| hsa-miR-1183 | hsa-miR-1178-3p | hsa-miR-4458 | |
| hsa-miR-103a-2-5p | hsa-let-7d-5p | hsa-miR-1183 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chow, J.T.-S.; Salmena, L. Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium. Genes 2020, 11, 1002. https://doi.org/10.3390/genes11091002
Chow JT-S, Salmena L. Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium. Genes. 2020; 11(9):1002. https://doi.org/10.3390/genes11091002
Chicago/Turabian StyleChow, Jonathan Tak-Sum, and Leonardo Salmena. 2020. "Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium" Genes 11, no. 9: 1002. https://doi.org/10.3390/genes11091002
APA StyleChow, J. T.-S., & Salmena, L. (2020). Prediction and Analysis of SARS-CoV-2-Targeting MicroRNA in Human Lung Epithelium. Genes, 11(9), 1002. https://doi.org/10.3390/genes11091002





