Literature Review of BARD1 as a Cancer Predisposing Gene with a Focus on Breast and Ovarian Cancers
Abstract
1. Introduction
2. BARD1 as a Candidate Cancer Predisposing Gene
Subsequent Studies with Inconsistent Results
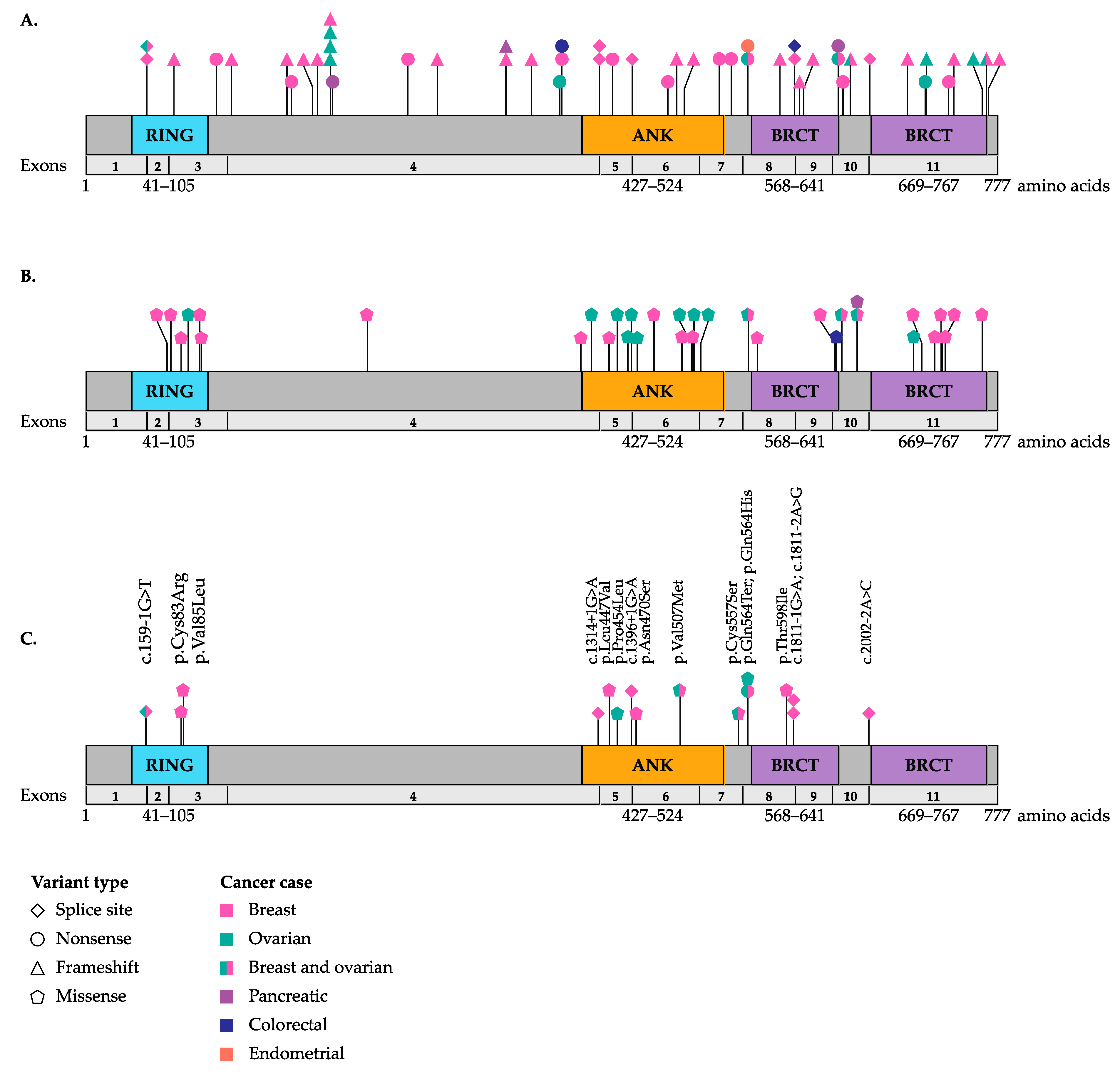
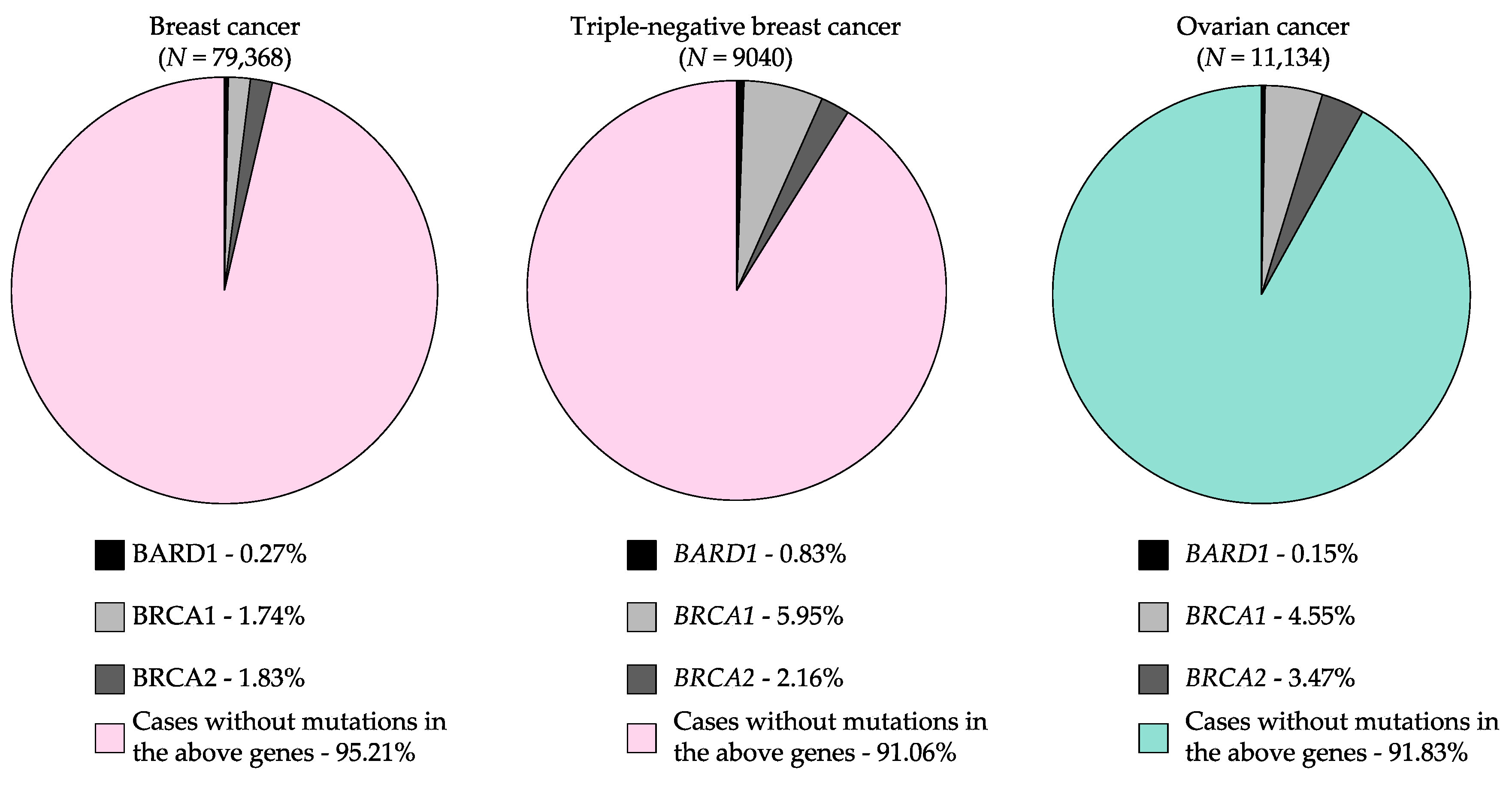
3. Potentially Pathogenic Germline BARD1 Variants Identified in Cancer Cases
3.1. Targeted BARD1 Studies of Breast and Ovarian Cancer
3.2. Multi-Gene Panel and Next-Generation Sequencing Studies Including BARD1 of Breast and Ovarian Cancers
3.3. BARD1 in Other Cancers
3.4. Carriers of a BARD1 Variant and a Pathogenic Variant in a Known High-Risk Cancer Predisposing Gene
4. The Biological Effect of Potentially Pathogenic BARD1 Variants
4.1. BARD1 Expression in Mammalian Cells
4.2. BARD1 in Animal and Cell Line Models
4.3. The Biological Impact of Potentially Pathogenic Variants in Different BARD1 Domains
5. Risk Assessments for Cancer in Carriers of BARD1 Variants
| Reference | Cancer Type | Selection of Cancer Cases | Number of Cases | Number of Controls | Population | Type of Variants Included | OR (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|---|
| [11] | Breast | Unselected | 28,536 | 26,078 b | Mixed e | LoF or missense f | 2.16 (1.31–3.63) | 2.26 × 10−3 |
| [52] | Breast | Familial | 2134 | 26,375 b | Mixed e | LoF or missense g | 3.18 (1.34–7.36) | 1.22 × 10−2 |
| [13] | TNBC | High risk | 4090 | 26,079 b | White | LoF or missense f | 5.92 (3.36–10.27) | 2.20 × 10−9 |
| TNBC | High risk | 2003 | 26,079 b | White | LoF or missense f | 4.35 (2.02–9.30) | 7.6 × 10−4 | |
| [23] | Breast | Familial | 3667 | – b | French | LoF or missense g | 2.00 (0.74–4.10) | – |
| TNBC | Familial | 3667 | – b | French | LoF or missense g | 11.27 (3.37–25.01) | – | |
| [33] | Breast | Familial | 4469 | 37,265 c | Unknown | LoF or missense f | 5.35 (3.17–9.04) | <10−5 |
| [22] | TNBC | Unselected | 4824 | 123,136 d | Mixed e | LoF or missense f | 9.76 (6.77–13.87) | – |
| [60] | Ovarian a | Unselected | 1915 | 36,276 b | Mixed e | LoF or missense g | 4.2 (1.4–12.5) | 0.02 |
| [63] | Ovarian a | High risk | 4122 | 4688 | White | LoF | 1.59 (0.31–9.56) h | 0.57 |
| Reference | Coding DNA Reference Sequence a | Predicted Amino Acid Change | Cancer Type | Selection of Cancer Cases | Number of Cases | Number of Controls | Population | OR (95% CI) | p-Value |
|---|---|---|---|---|---|---|---|---|---|
| [28] | c.70C>T | p.Pro24Ser | Breast | Unselected | 507 | 539 | China | 0.68 (0.52–0.9) | – |
| [34] | c.722C>G | p.Ser241Cys | Breast | Unselected | 143 | 155 | Japan | 1.57 (0.61–4.05) b | – |
| [34] | c.1139_1159del | p.Asn380_Phe387delinsIle | Breast | Unselected | 143 | 155 | Japan | 0.96 (0.49–1.86) c | – |
| [34] | c.1134G>C | p.Arg378Ser | Breast | Unselected | 143 | 155 | Japan | 1.35 (0.7–2.61) d | – |
| [28] | c.1134G>C | p.Arg378Ser | Breast | Unselected | 507 | 539 | China | 0.94 (0.72–1.24) | – |
| [26] | c.1972C>T | p.Arg658Cys | Breast | Unselected | 12,476 | 4707 | Poland | 1.16 (0.75–1.81) | 0.51 |
| [26] | c.1972C>T | p.Arg658Cys | TNBC | Unselected | 1120 | 4707 | Poland | 0.97 (0.40–2.36) | 0.95 |
| [26] | c.1977A>G | p.Arg659= | Breast | Unselected | 12,476 | 4707 | Poland | 1.32 (0.73–2.40) | 0.36 |
| [26] | c.1977A>G | p.Arg659= | TNBC | Unselected | 1120 | 4707 | Poland | 0.60 (0.14–2.64) | 0.5 |
| [34] | c.1518C>T | p.His506= | Breast | Unselected | 143 | 155 | Japan | 0.71 (0.36–1.42) e | – |
| [34] | c.1519G>A | p.Val507Met | Breast | Unselected | 143 | 155 | Japan | 1.28 (0.8–2.04) f | – |
| [8] | c.1519G>A | p.Val507Met | Breast | Familial | 663 | 718 | Finland | 1.04 (0.8–1.34) | 0.79 |
| [8] | c.1519G>A | p.Val507Met | Breast | Unselected | 867 | 718 | Finland | 1.27 (0.99–1.63) | 0.06 |
| [28] | c.1519G>A | p.Val507Met | Breast | Unselected | 507 | 539 | China | 0.98 (0.75–1.29) | – |
| [7] | c.1670G>C | p.Cys557Ser | Breast or ovarian | Familial | 126 | 1018 | Finland | 4.2 (1.7–10.7) | 5 × 10−3 |
| [8] | c.1670G>C | p.Cys557Ser | Breast | Familial | 926 | 725 | Finland | 0.49 (0.24–1.01) | 0.05 |
| [8] | c.1670G>C | p.Cys557Ser | Breast | Familial | 255 | 358 | Finland | 0.70 (0.21–2.34) | 0.56 |
| [8] | c.1670G>C | p.Cys557Ser | Breast | Unselected | 868 | 725 | Finland | 0.74 (0.38–1.44) | 0.38 |
| [8] | c.1670G>C | p.Cys557Ser | Breast | Unselected | 697 | 358 | Finland | 1.09 (0.47–2.56) | 0.84 |
| [14] | c.1670G>C | p.Cys557Ser | Breast or ovarian | Familial | 757 | 3591 | Nordic | 1.9 (1.32–2.83) | 10−3 |
| [14] | c.1670G>C | p.Cys557Ser | Breast | Unselected | 1984 | 3591 | Nordic | 1.7 (1.23–2.22) | 10−3 |
| [15] | c.1670G>C | p.Cys557Ser | Breast | Unselected | 992 | 703 | Iceland | 1.82 (1.11–3.01) | 0.014 |
| [17] | c.1670G>C | p.Cys557Ser | Breast | High risk | 3188 | 1038 | Poland | 1.2 (0.9–1.7) | 0.3 |
| [16] | c.1670G>C | p.Cys557Ser | Breast | Unselected | 1136 | 623 | Australia | 0.80 (0.50–1.27) g | 0.3 |
| [35] | c.1670G>C | p.Cys557Ser | Breast | High risk | 322 | 570 | Chile | 1.4 (0.6–3.7) | 0.47 |
| [64] | c.1690C>T | p.Gln564Ter | Breast | Familial | 1018 | 2036 | Poland | 1.0 (0.1–8.6) | 1 |
| [26] | c.1690C>T | p.Gln564Ter | Breast | Unselected | 13,935 | 5896 | Poland, Belarus | 2.30 (1.03–5.15) h | 0.04 |
| [26] | c.1690C>T | p.Gln564Ter | TNBC | Unselected | 1120 | 4707 | Poland | 3.62 (1.21–10.78) | 0.02 |
6. BARD1 in Medical Genetics Settings
7. Our Perspective of BARD1 as a Cancer Predisposing Gene
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Miki, Y.; Swensen, J.; Shattuck-Eidens, D.; Futreal, P.A.; Harshman, K.; Tavtigian, S.; Liu, Q.; Cochran, C.; Bennett, L.M.; Ding, W.; et al. Strong Candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 1994, 266, 66–71. [Google Scholar] [CrossRef]
- Rahman, N. Realizing the promise of cancer predisposition genes. Nature 2014, 505, 302–308. [Google Scholar] [CrossRef]
- Fearon, E.R. Human cancer syndromes: Clues to the origin and nature of cancer. Science 1997, 278, 1043–1050. [Google Scholar] [CrossRef]
- Nielsen, F.C.; van Overeem Hansen, T.; Sørensen, C.S. Hereditary breast and ovarian cancer: New genes in confined pathways. Nat. Rev. Cancer 2016, 16, 599–612. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.C.; Wang, Z.W.; Tsan, J.T.; Spillman, M.A.; Phung, A.; Xu, X.L.; Yang, M.C.; Hwang, L.Y.; Bowcock, A.M.; Baer, R. Identification of a RING protein that can interact in vivo with the BRCA1 gene product. Nat. Genet. 1996, 14, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Thai, T.H.; Du, F.; Tsan, J.T.; Jin, Y.; Phung, A.; Spillman, M.A.; Massa, H.F.; Muller, C.Y.; Ashfaq, R.; Mathis, J.M.; et al. Mutations in the BRCA1-associated RING domain (BARD1) gene in primary breast, ovarian and uterine cancers. Hum. Mol. Genet. 1998, 7, 195–202. [Google Scholar] [CrossRef] [PubMed]
- Karppinen, S.-M.; Heikkinen, K.; Rapakko, K.; Winqvist, R. Mutation screening of the BARD1 gene: Evidence for involvement of the Cys557Ser allele in hereditary susceptibility to breast cancer. J. Med. Genet. 2004, 41, e114. [Google Scholar] [CrossRef]
- Karppinen, S.-M.; Barkardottir, R.B.; Backenhorn, K.; Sydenham, T.; Syrjäkoski, K.; Schleutker, J.; Ikonen, T.; Pylkäs, K.; Rapakko, K.; Erkko, H.; et al. Nordic collaborative study of the BARD1 Cys557Ser allele in 3956 patients with cancer: Enrichment in familial BRCA1/BRCA2 mutation-negative breast cancer but not in other malignancies. J. Med. Genet. 2006, 43, 856–862. [Google Scholar] [CrossRef][Green Version]
- Stacey, S.N.; Sulem, P.; Johannsson, O.T.; Helgason, A.; Gudmundsson, J.; Kostic, J.P.; Kristjansson, K.; Jonsdottir, T.; Sigurdsson, H.; Hrafnkelsson, J.; et al. The BARD1 Cys557Ser variant and breast cancer risk in Iceland. PLoS Med. 2006, 3, e217. [Google Scholar] [CrossRef]
- Vahteristo, P.; Syrjäkoski, K.; Heikkinen, T.; Eerola, H.; Aittomäki, K.; von Smitten, K.; Holli, K.; Blomqvist, C.; Kallioniemi, O.-P.; Nevanlinna, H. BARD1 variants Cys557Ser and Val507Met in breast cancer predisposition. Eur. J. Hum. Genet. 2006, 14, 167–172. [Google Scholar] [CrossRef]
- Johnatty, S.E.; Beesley, J.; Chen, X.; Hopper, J.L.; Southey, M.C.; Giles, G.G.; Goldgar, D.E.; Chenevix-Trench, G.; Spurdle, A.B. The BARD1 Cys557Ser polymorphism and breast cancer risk: An Australian case-control and family analysis. Breast Cancer Res. Treat. 2009, 115, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Jakubowska, A.; Cybulski, C.; Szymańska, A.; Huzarski, T.; Byrski, T.; Gronwald, J.; Debniak, T.; Górski, B.; Kowalska, E.; Narod, S.A.; et al. BARD1 and breast cancer in Poland. Breast Cancer Res. Treat. 2008, 107, 119–122. [Google Scholar] [CrossRef] [PubMed]
- Karczewski, K.J.; Francioli, L.C.; Tiao, G.; Cummings, B.B.; Alföldi, J.; Wang, Q.; Collins, R.L.; Laricchia, K.M.; Ganna, A.; Birnbaum, D.P.; et al. The mutational constraint spectrum quantified from variation in 141, 456 humans. Nature 2020, 581, 434–443. [Google Scholar] [CrossRef]
- Guénard, F.; Labrie, Y.; Ouellette, G.; Beauparlant, C.J.; Durocher, F. Genetic sequence variations of BRCA1-interacting genes AURKA, BAP1, BARD1 and DHX9 in French Canadian families with high risk of breast cancer. J. Hum. Genet. 2009, 54, 152–161. [Google Scholar] [CrossRef]
- Ramus, S.J.; Song, H.; Dicks, E.; Tyrer, J.P.; Rosenthal, A.N.; Intermaggio, M.P.; Fraser, L.; Gentry-Maharaj, A.; Hayward, J.; Philpott, S.; et al. Germline mutations in the BRIP1, BARD1, PALB2, and NBN genes in women with ovarian cancer. J. Natl. Cancer Inst. 2015, 107. [Google Scholar] [CrossRef] [PubMed]
- Couch, F.J.; Hart, S.N.; Sharma, P.; Toland, A.E.; Wang, X.; Miron, P.; Olson, J.E.; Godwin, A.K.; Pankratz, V.S.; Olswold, C.; et al. Inherited mutations in 17 breast cancer susceptibility genes among a large triple-negative breast cancer cohort unselected for family history of breast cancer. J. Clin. Oncol. 2015, 33, 304–311. [Google Scholar] [CrossRef] [PubMed]
- Gorringe, K.L.; Choong, D.Y.H.; Visvader, J.E.; Lindeman, G.J.; Campbell, I.G. BARD1 variants are not associated with breast cancer risk in Australian familial breast cancer. Breast Cancer Res. Treat. 2008, 111, 505–509. [Google Scholar] [CrossRef]
- Couch, F.J.; Shimelis, H.; Hu, C.; Hart, S.N.; Polley, E.C.; Na, J.; Hallberg, E.; Moore, R.; Thomas, A.; Lilyquist, J.; et al. Associations between cancer predisposition testing panel genes and breast cancer. JAMA Oncol. 2017, 3, 1190–1196. [Google Scholar] [CrossRef]
- Hu, C.; Polley, E.C.; Yadav, S.; Lilyquist, J.; Shimelis, H.; Na, J.; Hart, S.N.; Goldgar, D.E.; Shah, S.; Pesaran, T.; et al. The contribution of germline predisposition gene mutations to clinical subtypes of invasive breast cancer from a clinical genetic testing cohort. J. Natl. Cancer Inst. 2020. [Google Scholar] [CrossRef]
- Shimelis, H.; LaDuca, H.; Hu, C.; Hart, S.N.; Na, J.; Thomas, A.; Akinhanmi, M.; Moore, R.M.; Brauch, H.; Cox, A.; et al. Triple-Negative breast cancer risk genes identified by multigene hereditary cancer panel testing. J. Natl. Cancer Inst. 2018, 110, 855–862. [Google Scholar] [CrossRef]
- Castéra, L.; Harter, V.; Muller, E.; Krieger, S.; Goardon, N.; Ricou, A.; Rousselin, A.; Paimparay, G.; Legros, A.; Bruet, O.; et al. Landscape of pathogenic variations in a panel of 34 genes and cancer risk estimation from 5131 HBOC families. Genet. Med. 2018, 20, 1677–1686. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Aggarwal, R. An overview of triple-negative breast cancer. Arch. Gynecol. Obstet. 2016, 293, 247–269. [Google Scholar] [CrossRef] [PubMed]
- Ryu, D.W.; Jung, M.J.; Choi, W.S.; Lee, C.H. Clinical significance of morphologic characteristics in triple negative breast cancer. J. Korean Surg. Soc. 2011, 80, 301. [Google Scholar] [CrossRef] [PubMed]
- Suszynska, M.; Kluzniak, W.; Wokolorczyk, D.; Jakubowska, A.; Huzarski, T.; Gronwald, J.; Debniak, T.; Szwiec, M.; Ratajska, M.; Klonowska, K.; et al. BARD1 is a low/moderate breast cancer risk gene: Evidence Based on an association study of the central European p.Q564X Recurrent mutation. Cancers (Basel) 2019, 11, 740. [Google Scholar] [CrossRef]
- Ratajska, M.; Antoszewska, E.; Piskorz, A.; Brozek, I.; Borg, Å.; Kusmierek, H.; Biernat, W.; Limon, J. Cancer predisposing BARD1 mutations in breast-ovarian cancer families. Breast Cancer Res. Treat. 2012, 131, 89–97. [Google Scholar] [CrossRef]
- Huo, X.; Hu, Z.; Zhai, X.; Wang, Y.; Wang, S.; Wang, X.; Qin, J.; Chen, W.; Jin, G.; Liu, J.; et al. Common non-synonymous polymorphisms in the BRCA1 Associated RING Domain (BARD1) gene are associated with breast cancer susceptibility: A case-control analysis. Breast Cancer Res. Treat. 2007, 102, 329–337. [Google Scholar] [CrossRef]
- Sauer, M.K.; Andrulis, I.L. Identification and characterization of missense alterations in the BRCA1 associated RING domain (BARD1) gene in breast and ovarian cancer. J. Med. Genet. 2005, 42, 633–638. [Google Scholar] [CrossRef]
- De Brakeleer, S.; De Grève, J.; Loris, R.; Janin, N.; Lissens, W.; Sermijn, E.; Teugels, E. Cancer predisposing missense and protein truncating BARD1 mutations in non-BRCA1 or BRCA2 breast cancer families. Hum. Mutat. 2010, 31, E1175-85. [Google Scholar] [CrossRef]
- De Brakeleer, S.; De Grève, J.; Desmedt, C.; Joris, S.; Sotiriou, C.; Piccart, M.; Pauwels, I.; Teugels, E. Frequent incidence of BARD1-truncating mutations in germline DNA from triple-negative breast cancer patients. Clin. Genet. 2016, 89, 336–340. [Google Scholar] [CrossRef]
- Toh, M.R.; Chong, S.T.; Chan, S.H.; Low, C.E.; Ishak, N.D.B.; Lim, J.Q.; Courtney, E.; Ngeow, J. Functional analysis of clinical BARD1 germline variants. Cold Spring Harb. Mol. Case Stud. 2019, 5. [Google Scholar] [CrossRef]
- Weber-Lassalle, N.; Borde, J.; Weber-Lassalle, K.; Horváth, J.; Niederacher, D.; Arnold, N.; Kaulfuß, S.; Ernst, C.; Paul, V.G.; Honisch, E.; et al. Germline loss-of-function variants in the BARD1 gene are associated with early-onset familial breast cancer but not ovarian cancer. Breast Cancer Res. 2019, 21, 55. [Google Scholar] [CrossRef] [PubMed]
- Ishitobi, M.; Miyoshi, Y.; Hasegawa, S.; Egawa, C.; Tamaki, Y.; Monden, M.; Noguchi, S. Mutational analysis of BARD1 in familial breast cancer patients in Japan. Cancer Lett. 2003, 200, 1–7. [Google Scholar] [CrossRef]
- Gonzalez-Hormazabal, P.; Reyes, J.M.; Blanco, R.; Bravo, T.; Carrera, I.; Peralta, O.; Gomez, F.; Waugh, E.; Margarit, S.; Ibañez, G.; et al. The BARD1 Cys557Ser variant and risk of familial breast cancer in a South-American population. Mol. Biol. Rep. 2012, 39, 8091–8098. [Google Scholar] [CrossRef] [PubMed]
- Gass, J.; Tatro, M.; Blackburn, P.; Hines, S.; Atwal, P.S. BARD1 nonsense variant c.1921C>T in a patient with recurrent breast cancer. Clin. Case Rep. 2017, 5, 104–107. [Google Scholar] [CrossRef]
- Rana, H.Q.; Sacca, R.; Drogan, C.; Gutierrez, S.; Schlosnagle, E.; Regan, M.M.; Speare, V.; LaDuca, H.; Dolinsky, J.; Garber, J.E.; et al. Prevalence of germline variants in inflammatory breast cancer. Cancer 2019, 125, 2194–2202. [Google Scholar] [CrossRef]
- Churpek, J.E.; Walsh, T.; Zheng, Y.; Moton, Z.; Thornton, A.M.; Lee, M.K.; Casadei, S.; Watts, A.; Neistadt, B.; Churpek, M.M.; et al. Inherited predisposition to breast cancer among African American women. Breast Cancer Res. Treat. 2015, 149, 31–39. [Google Scholar] [CrossRef]
- Norquist, B.M.; Harrell, M.I.; Brady, M.F.; Walsh, T.; Lee, M.K.; Gulsuner, S.; Bernards, S.S.; Casadei, S.; Yi, Q.; Burger, R.A.; et al. Inherited mutations in women with ovarian carcinoma. JAMA Oncol. 2016, 2, 482–490. [Google Scholar] [CrossRef]
- Walsh, T.; Casadei, S.; Lee, M.K.; Pennil, C.C.; Nord, A.S.; Thornton, A.M.; Roeb, W.; Agnew, K.J.; Stray, S.M.; Wickramanayake, A.; et al. Mutations in 12 genes for inherited ovarian, fallopian tube, and peritoneal carcinoma identified by massively parallel sequencing. Proc. Natl. Acad. Sci. USA 2011, 108, 18032–18037. [Google Scholar] [CrossRef]
- Caminsky, N.G.; Mucaki, E.J.; Perri, A.M.; Lu, R.; Knoll, J.H.M.; Rogan, P.K. Prioritizing variants in complete hereditary breast and ovarian cancer genes in patients lacking known BRCA mutations. Hum. Mutat. 2016, 37, 640–652. [Google Scholar] [CrossRef]
- Lhotova, K.; Stolarova, L.; Zemankova, P.; Vocka, M.; Janatova, M.; Borecka, M.; Cerna, M.; Jelinkova, S.; Kral, J.; Volkova, Z.; et al. Multigene panel germline testing of 1333 czech patients with ovarian cancer. Cancers (Basel) 2020, 12, 956. [Google Scholar] [CrossRef]
- Buys, S.S.; Sandbach, J.F.; Gammon, A.; Patel, G.; Kidd, J.; Brown, K.L.; Sharma, L.; Saam, J.; Lancaster, J.; Daly, M.B. A study of over 35,000 women with breast cancer tested with a 25-gene panel of hereditary cancer genes. Cancer 2017, 123, 1721–1730. [Google Scholar] [CrossRef] [PubMed]
- Koczkowska, M.; Krawczynska, N.; Stukan, M.; Kuzniacka, A.; Brozek, I.; Sniadecki, M.; Debniak, J.; Wydra, D.; Biernat, W.; Kozlowski, P.; et al. Spectrum and prevalence of pathogenic variants in ovarian cancer susceptibility genes in a group of 333 patients. Cancers (Basel) 2018, 10, 442. [Google Scholar] [CrossRef] [PubMed]
- Jian, W.; Shao, K.; Qin, Q.; Wang, X.; Song, S.; Wang, X. Clinical and genetic characterization of hereditary breast cancer in a Chinese population. Hered. Cancer Clin. Pract. 2017, 15, 19. [Google Scholar] [CrossRef] [PubMed]
- Pennington, K.P.; Walsh, T.; Harrell, M.I.; Lee, M.K.; Pennil, C.C.; Rendi, M.H.; Thornton, A.; Norquist, B.M.; Casadei, S.; Nord, A.S.; et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 2014, 20, 764–775. [Google Scholar] [CrossRef]
- Park, J.S.; Lee, S.-T.; Nam, E.J.; Han, J.W.; Lee, J.-Y.; Kim, J.; Kim, T.I.; Park, H.S. Variants of cancer susceptibility genes in Korean BRCA1/2 mutation-negative patients with high risk for hereditary breast cancer. BMC Cancer 2018, 18, 83. [Google Scholar] [CrossRef]
- González-Rivera, M.; Lobo, M.; López-Tarruella, S.; Jerez, Y.; Del Monte-Millán, M.; Massarrah, T.; Ramos-Medina, R.; Ocaña, I.; Picornell, A.; Santillán Garzón, S.; et al. Frequency of germline DNA genetic findings in an unselected prospective cohort of triple-negative breast cancer patients participating in a platinum-based neoadjuvant chemotherapy trial. Breast Cancer Res. Treat. 2016, 156, 507–515. [Google Scholar] [CrossRef]
- Corredor, J.; Woodson, A.H.; Gutierrez Barrera, A.; Arun, B. Multigene panel testing results in patients with multiple breast cancer primaries. Breast J. 2020, 26, 1337–1342. [Google Scholar] [CrossRef]
- Ng, P.S.; Wen, W.X.; Fadlullah, M.Z.H.; Yoon, S.Y.; Lee, S.Y.; Thong, M.K.; Yip, C.H.; Mohd Taib, N.A.; Teo, S.H. Identification of germline alterations in breast cancer predisposition genes among Malaysian breast cancer patients using panel testing. Clin. Genet. 2016, 90, 315–323. [Google Scholar] [CrossRef]
- Lolas Hamameh, S.; Renbaum, P.; Kamal, L.; Dweik, D.; Salahat, M.; Jaraysa, T.; Abu Rayyan, A.; Casadei, S.; Mandell, J.B.; Gulsuner, S.; et al. Genomic analysis of inherited breast cancer among Palestinian women: Genetic heterogeneity and a founder mutation in TP53. Int. J. Cancer 2017, 141, 750–756. [Google Scholar] [CrossRef]
- Li, J.; Meeks, H.; Feng, B.-J.; Healey, S.; Thorne, H.; Makunin, I.; Ellis, J.; Campbell, I.; Southey, M.; Mitchell, G.; et al. Targeted massively parallel sequencing of a panel of putative breast cancer susceptibility genes in a large cohort of multiple-case breast and ovarian cancer families. J. Med. Genet. 2016, 53, 34–42. [Google Scholar] [CrossRef]
- Crawford, B.; Adams, S.B.; Sittler, T.; van den Akker, J.; Chan, S.; Leitner, O.; Ryan, L.; Gil, E.; van ’t Veer, L. Multi-Gene panel testing for hereditary cancer predisposition in unsolved high-risk breast and ovarian cancer patients. Breast Cancer Res. Treat. 2017, 163, 383–390. [Google Scholar] [CrossRef] [PubMed]
- Silva Carvalho, S.D.C.E.; Cury, N.M.; Brotto, D.B.; de Araujo, L.F.; Rosa, R.C.A.; Texeira, L.A.; Plaça, J.R.; Marques, A.A.; Peronni, K.C.; de Cássia Ruy, P.; et al. Germline variants in DNA repair genes associated with hereditary breast and ovarian cancer syndrome: Analysis of a 21 gene panel in the Brazilian population. BMC Med. Genom. 2020, 13, 21. [Google Scholar] [CrossRef]
- Amemiya, Y.; Bacopulos, S.; Al-Shawarby, M.; Al-Tamimi, D.; Naser, W.; Ahmed, A.; Khalifa, M.; Slodkowska, E.; Seth, A. A comparative analysis of breast and ovarian cancer-related gene mutations in canadian and saudi arabian patients with breast cancer. Anticancer Res. 2015, 35, 2601–2610. [Google Scholar] [PubMed]
- Rodríguez-Balada, M.; Roig, B.; Melé, M.; Albacar, C.; Serrano, S.; Salvat, M.; Querol, M.; Borràs, J.; Martorell, L.; Gumà, J. Identification of germline pathogenic variants in DNA damage repair genes by a next-generation sequencing multigene panel in BRCAX patients. Clin. Biochem. 2020, 76, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Slavin, T.P.; Maxwell, K.N.; Lilyquist, J.; Vijai, J.; Neuhausen, S.L.; Hart, S.N.; Ravichandran, V.; Thomas, T.; Maria, A.; Villano, D.; et al. The contribution of pathogenic variants in breast cancer susceptibility genes to familial breast cancer risk. NPJ Breast Cancer 2017, 3, 22. [Google Scholar] [CrossRef] [PubMed]
- Castéra, L.; Krieger, S.; Rousselin, A.; Legros, A.; Baumann, J.-J.; Bruet, O.; Brault, B.; Fouillet, R.; Goardon, N.; Letac, O.; et al. Next-Generation sequencing for the diagnosis of hereditary breast and ovarian cancer using genomic capture targeting multiple candidate genes. Eur. J. Hum. Genet. 2014, 22, 1305–1313. [Google Scholar] [CrossRef]
- Adedokun, B.; Zheng, Y.; Ndom, P.; Gakwaya, A.; Makumbi, T.; Zhou, A.Y.; Yoshimatsu, T.F.; Rodriguez, A.; Madduri, R.K.; Foster, I.T.; et al. Prevalence of inherited mutations in breast cancer predisposition genes among women in uganda and cameroon. Cancer Epidemiol. Biomark. Prev. 2020, 29, 359–367. [Google Scholar] [CrossRef] [PubMed]
- Li, J.-Y.; Jing, R.; Wei, H.; Wang, M.; Xiaowei, Q.; Liu, H.; Jian, L.; Ou, J.-H.; Jiang, W.-H.; Tian, F.-G.; et al. Germline mutations in 40 cancer susceptibility genes among Chinese patients with high hereditary risk breast cancer. Int. J. Cancer 2019, 144, 281–289. [Google Scholar] [CrossRef] [PubMed]
- Young, E.L.; Feng, B.J.; Stark, A.W.; Damiola, F.; Durand, G.; Forey, N.; Francy, T.C.; Gammon, A.; Kohlmann, W.K.; Kaphingst, K.A.; et al. Multigene testing of moderate-risk genes: Be mindful of the missense. J. Med. Genet. 2016, 53, 366–376. [Google Scholar] [CrossRef] [PubMed]
- Bonache, S.; Esteban, I.; Moles-Fernández, A.; Tenés, A.; Duran-Lozano, L.; Montalban, G.; Bach, V.; Carrasco, E.; Gadea, N.; López-Fernández, A.; et al. Multigene panel testing beyond BRCA1/2 in breast/ovarian cancer Spanish families and clinical actionability of findings. J. Cancer Res. Clin. Oncol. 2018, 144, 2495–2513. [Google Scholar] [CrossRef] [PubMed]
- Scarpitta, R.; Zanna, I.; Aretini, P.; Gambino, G.; Scatena, C.; Mei, B.; Ghilli, M.; Rossetti, E.; Roncella, M.; Congregati, C.; et al. Germline investigation in male breast cancer of DNA repair genes by next-generation sequencing. Breast Cancer Res. Treat. 2019, 178, 557–564. [Google Scholar] [CrossRef] [PubMed]
- Kaur, R.P.; Kumar, V.; Shafi, G.; Vashistha, R.; Kulharia, M.; Munshi, A. A study of mechanistic mapping of novel SNPs to male breast cancer. Med. Oncol. 2019, 36, 70. [Google Scholar] [CrossRef] [PubMed]
- Arvai, K.J.; Roberts, M.E.; Torene, R.I.; Susswein, L.R.; Marshall, M.L.; Zhang, Z.; Carter, N.J.; Yackowski, L.; Rinella, E.S.; Klein, R.T.; et al. Age-Adjusted association of homologous recombination genes with ovarian cancer using clinical exomes as controls. Hered. Cancer Clin. Pract. 2019, 17, 19. [Google Scholar] [CrossRef] [PubMed]
- Cybulski, C.; Kluźniak, W.; Huzarski, T.; Wokołorczyk, D.; Kashyap, A.; Rusak, B.; Stempa, K.; Gronwald, J.; Szymiczek, A.; Bagherzadeh, M.; et al. The spectrum of mutations predisposing to familial breast cancer in Poland. Int. J. Cancer 2019, 145, 3311–3320. [Google Scholar] [CrossRef] [PubMed]
- Zeng, C.; Guo, X.; Wen, W.; Shi, J.; Long, J.; Cai, Q.; Shu, X.-O.; Xiang, Y.; Zheng, W. Evaluation of pathogenetic mutations in breast cancer predisposition genes in population-based studies conducted among Chinese women. Breast Cancer Res. Treat. 2020, 181, 465–473. [Google Scholar] [CrossRef] [PubMed]
- Shahi, R.B.; De Brakeleer, S.; Caljon, B.; Pauwels, I.; Bonduelle, M.; Joris, S.; Fontaine, C.; Vanhoeij, M.; Van Dooren, S.; Teugels, E.; et al. Identification of candidate cancer predisposing variants by performing whole-exome sequencing on index patients from BRCA1 and BRCA2-negative breast cancer families. BMC Cancer 2019, 19, 313. [Google Scholar] [CrossRef]
- Torrezan, G.T.; de Almeida, F.G.D.S.R.; Figueiredo, M.C.P.; de Figueiredo Barros, B.D.; de Paula, C.A.A.; Valieris, R.; de Souza, J.E.S.; Ramalho, R.F.; da Silva, F.C.C.; Ferreira, E.N.; et al. Complex landscape of germline variants in brazilian patients with hereditary and early onset breast cancer. Front. Genet. 2018, 9, 161. [Google Scholar] [CrossRef]
- Rouleau, E.; Jesson, B.; Briaux, A.; Nogues, C.; Chabaud, V.; Demange, L.; Sokolowska, J.; Coulet, F.; Barouk-Simonet, E.; Bignon, Y.J.; et al. Rare germline large rearrangements in the BRCA1/2 genes and eight candidate genes in 472 patients with breast cancer predisposition. Breast Cancer Res. Treat. 2012, 133, 1179–1190. [Google Scholar] [CrossRef]
- Esteban-Jurado, C.; Vila-Casadesús, M.; Garre, P.; Lozano, J.J.; Pristoupilova, A.; Beltran, S.; Muñoz, J.; Ocaña, T.; Balaguer, F.; López-Cerón, M.; et al. Whole-Exome sequencing identifies rare pathogenic variants in new predisposition genes for familial colorectal cancer. Genet. Med. 2015, 17, 131–142. [Google Scholar] [CrossRef]
- Ring, K.L.; Bruegl, A.S.; Allen, B.A.; Elkin, E.P.; Singh, N.; Hartman, A.-R.; Daniels, M.S.; Broaddus, R.R. Germline multi-gene hereditary cancer panel testing in an unselected endometrial cancer cohort. Mod. Pathol. 2016, 29, 1381–1389. [Google Scholar] [CrossRef]
- Chaffee, K.G.; Oberg, A.L.; McWilliams, R.R.; Majithia, N.; Allen, B.A.; Kidd, J.; Singh, N.; Hartman, A.-R.; Wenstrup, R.J.; Petersen, G.M. Prevalence of germ-line mutations in cancer genes among pancreatic cancer patients with a positive family history. Genet. Med. 2018, 20, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Hart, S.N.; Bamlet, W.R.; Moore, R.M.; Nandakumar, K.; Eckloff, B.W.; Lee, Y.K.; Petersen, G.M.; McWilliams, R.R.; Couch, F.J. Prevalence of pathogenic mutations in cancer predisposition genes among pancreatic cancer patients. Cancer Epidemiol. Biomark. Prev. 2016, 25, 207–211. [Google Scholar] [CrossRef] [PubMed]
- Meynard, G.; Mansi, L.; Lebahar, P.; Villanueva, C.; Klajer, E.; Calcagno, F.; Vivalta, A.; Chaix, M.; Collonge-Rame, M.-A.; Populaire, C.; et al. First description of a double heterozygosity for BRCA1 and BRCA2 pathogenic variants in a French metastatic breast cancer patient: A case report. Oncol. Rep. 2017, 37, 1573–1578. [Google Scholar] [CrossRef] [PubMed]
- Palmirotta, R.; Lovero, D.; Stucci, L.S.; Silvestris, E.; Quaresmini, D.; Cardascia, A.; Silvestris, F. Double heterozygosity for BRCA1 pathogenic variant and BRCA2 polymorphic stop codon K3326X: A case report in a Southern Italian family. Int. J. Mol. Sci. 2018, 19, 285. [Google Scholar] [CrossRef] [PubMed]
- Ramus, S.J.; Friedman, L.S.; Gayther, S.A.; Ponder, B.A.; Bobrow, L.; van der Looji, M.; Papp, J.; Olah, E. A breast/ovarian cancer patient with germline mutations in both BRCA1 and BRCA2. Nat. Genet. 1997, 15, 14–15. [Google Scholar] [CrossRef] [PubMed]
- Lavie, O.; Narod, S.; Lejbkowicz, F.; Dishon, S.; Goldberg, Y.; Gemer, O.; Rennert, G. Double heterozygosity in the BRCA1 and BRCA2 genes in the Jewish population. Ann. Oncol. 2011, 22, 964–966. [Google Scholar] [CrossRef]
- Friedman, E.; Bar-Sade Bruchim, R.; Kruglikova, A.; Risel, S.; Levy-Lahad, E.; Halle, D.; Bar-On, E.; Gershoni-Baruch, R.; Dagan, E.; Kepten, I.; et al. Double heterozygotes for the Ashkenazi founder mutations in BRCA1 and BRCA2 genes. Am. J. Hum. Genet. 1998, 63, 1224–1227. [Google Scholar] [CrossRef]
- Le Page, C.; Rahimi, K.; Rodrigues, M.; Heinzelmann-Schwarz, V.; Recio, N.; Tommasi, S.; Bataillon, G.; Portelance, L.; Golmard, L.; Meunier, L.; et al. Clinicopathological features of women with epithelial ovarian cancer and double heterozygosity for BRCA1 and BRCA2: A systematic review and case report analysis. Gynecol. Oncol. 2020, 156, 377–386. [Google Scholar] [CrossRef]
- Rebbeck, T.R.; Mitra, N.; Domchek, S.M.; Wan, F.; Chuai, S.; Friebel, T.M.; Panossian, S.; Spurdle, A.; Chenevix-Trench, G.; Singer, C.F.; et al. Modification of ovarian cancer risk by BRCA1/2-interacting genes in a multicenter cohort of BRCA1/2 mutation carriers. Cancer Res. 2009, 69, 5801–5810. [Google Scholar] [CrossRef]
- Spurdle, A.B.; Marquart, L.; McGuffog, L.; Healey, S.; Sinilnikova, O.; Wan, F.; Chen, X.; Beesley, J.; Singer, C.F.; Dressler, A.-C.; et al. Common genetic variation at BARD1 is not associated with breast cancer risk in BRCA1 or BRCA2 mutation carriers. Cancer Epidemiol. Biomark. Prev. 2011, 20, 1032–1038. [Google Scholar] [CrossRef]
- Moore, J.H.; Williams, S.M. Epistasis and its implications for personal genetics. Am. J. Hum. Genet. 2009, 85, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Gibson, G. Rare and common variants: Twenty arguments. Nat. Rev. Genet. 2012, 13, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Slatkin, M. Exchangeable models of complex inherited diseases. Genetics 2008, 179, 2253–2261. [Google Scholar] [CrossRef] [PubMed]
- Belanger, M.H.; Dolman, L.; Arcand, S.L.; Shen, Z.; Chong, G.; Mes-Masson, A.-M.; Provencher, D.; Tonin, P.N. A targeted analysis identifies a high frequency of BRCA1 and BRCA2 mutation carriers in women with ovarian cancer from a founder population. J. Ovarian Res. 2015, 8, 1. [Google Scholar] [CrossRef] [PubMed]
- Behl, S.; Hamel, N.; de Ladurantaye, M.; Lepage, S.; Lapointe, R.; Mes-Masson, A.-M.; Foulkes, W.D. Founder BRCA1/BRCA2/PALB2 pathogenic variants in French-Canadian breast cancer cases and controls. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef]
- Tonin, P.N.; Mes-Masson, A.M.; Futreal, P.A.; Morgan, K.; Mahon, M.; Foulkes, W.D.; Cole, D.E.; Provencher, D.; Ghadirian, P.; Narod, S.A. Founder BRCA1 and BRCA2 mutations in French Canadian breast and ovarian cancer families. Am. J. Hum. Genet. 1998, 63, 1341–1351. [Google Scholar] [CrossRef]
- Tonin, P.N.; Mes-Masson, A.-M.; Narod, S.A.; Ghadirian, P.; Provencher, D. Founder BRCA1 and BRCA2 mutations in French Canadian ovarian cancer cases unselected for family history. Clin. Genet. 1999, 55, 318–324. [Google Scholar] [CrossRef]
- Tonin, P.N.; Perret, C.; Lambert, J.A.; Paradis, A.J.; Kantemiroff, T.; Benoît, M.H.; Martin, G.; Foulkes, W.D.; Ghadirian, P. Founder BRCA1 and BRCA2 mutations in early-onset french Canadian breast cancer cases unselected for family history. Int. J. Cancer 2001, 95, 189–193. [Google Scholar] [CrossRef]
- Oros, K.K.; Ghadirian, P.; Greenwood, C.M.T.; Perret, C.; Shen, Z.; Paredes, Y.; Arcand, S.L.; Mes-Masson, A.M.; Narod, S.A.; Foulkes, W.D.; et al. Significant proportion of breast and/or ovarian cancer families of French Canadian descent harbor 1 of 5 BRCA1 and BRCA2 mutations. Int. J. Cancer 2004, 112, 411–419. [Google Scholar] [CrossRef]
- Tischkowitz, M.; Sabbaghian, N.; Hamel, N.; Pouchet, C.; Foulkes, W.D.; Mes-Masson, A.-M.; Provencher, D.M.; Tonin, P.N. Contribution of the PALB2 c.2323C>T [p. Q775X] Founder mutation in well-defined breast and/or ovarian cancer families and unselected ovarian cancer cases of French Canadian descent. BMC Med. Genet. 2013, 14, 5. [Google Scholar] [CrossRef]
- Rivera, B.; Di Iorio, M.; Frankum, J.; Nadaf, J.; Fahiminiya, S.; Arcand, S.L.; Burk, D.L.; Grapton, D.; Tomiak, E.; Hastings, V.; et al. Functionally null RAD51D missense mutation associates strongly with ovarian carcinoma. Cancer Res. 2017, 77, 4517–4529. [Google Scholar] [CrossRef] [PubMed]
- Castellsagué, E.; Liu, J.; Volenik, A.; Giroux, S.; Gagné, R.; Maranda, B.; Roussel-Jobin, A.; Latrielle, J.; Laframboise, R.; Palma, L.; et al. Characterization of a novel founder MSH6 mutation causing Lynch syndrome in the French Canadian population. Clin. Genet. 2015, 87, 536–542. [Google Scholar] [CrossRef] [PubMed]
- Fierheller, C.T.; Guitton-Sert, L.; Alenezi, W.M.; Revil, T.; Oros, K.K.; Bedard, K.; Arcand, S.L.; Serruya, C.; Behl, S.; Meunier, L.; et al. The genetic analysis of a founder Northern American population of European descent identifies FANCI as a candidate familial ovarian cancer risk gene. medRxiv 2020. [Google Scholar] [CrossRef]
- Jin, Y.; Xu, X.L.; Yang, M.C.; Wei, F.; Ayi, T.C.; Bowcock, A.M.; Baer, R. Cell cycle-dependent colocalization of BARD1 and BRCA1 proteins in discrete nuclear domains. Proc. Natl. Acad. Sci. USA 1997, 94, 12075–12080. [Google Scholar] [CrossRef] [PubMed]
- Irminger-Finger, I.; Soriano, J.V.; Vaudan, G.; Montesano, R.; Sappino, A.P. In vitro repression of Brca1-associated RING domain gene, Bard1, induces phenotypic changes in mammary epithelial cells. J. Cell Biol. 1998, 143, 1329–1339. [Google Scholar] [CrossRef]
- Cimmino, F.; Formicola, D.; Capasso, M. Dualistic role of BARD1 in cancer. Genes (Basel) 2017, 8, 375. [Google Scholar] [CrossRef]
- Bell, D.; Berchuck, A.; Birrer, M.; Chien, J.; Cramer, D.W.; Dao, F.; Dhir, R.; Disaia, P.; Gabra, H.; Glenn, P.; et al. Integrated genomic analyses of ovarian carcinoma. Nature 2011, 474, 609–615. [Google Scholar] [CrossRef]
- Koboldt, D.C.; Fulton, R.S.; McLellan, M.D.; Schmidt, H.; Kalicki-Veizer, J.; McMichael, J.F.; Fulton, L.L.; Dooling, D.J.; Ding, L.; Mardis, E.R.; et al. Comprehensive molecular portraits of human breast tumours. Nature 2012, 490, 61–70. [Google Scholar] [CrossRef]
- Zack, T.I.; Schumacher, S.E.; Carter, S.L.; Cherniack, A.D.; Saksena, G.; Tabak, B.; Lawrence, M.S.; Zhang, C.Z.; Wala, J.; Mermel, C.H.; et al. Pan-Cancer patterns of somatic copy number alteration. Nat. Genet. 2013, 45, 1134–1140. [Google Scholar] [CrossRef]
- Campbell, P.J.; Getz, G.; Korbel, J.O.; Stuart, J.M.; Jennings, J.L.; Stein, L.D.; Perry, M.D.; Nahal-Bose, H.K.; Ouellette, B.F.F.; Li, C.H.; et al. Pan-Cancer analysis of whole genomes. Nature 2020, 578, 82–93. [Google Scholar] [CrossRef]
- McCarthy, E.E.; Celebi, J.T.; Baer, R.; Ludwig, T. Loss of Bard1, the heterodimeric partner of the brca1 tumor suppressor, results in early embryonic lethality and chromosomal instability. Mol. Cell. Biol. 2003, 23, 5056–5063. [Google Scholar] [CrossRef]
- Shakya, R.; Szabolcs, M.; McCarthy, E.; Ospina, E.; Basso, K.; Nandula, S.; Murty, V.; Baer, R.; Ludwig, T. The basal-like mammary carcinomas induced by Brca1 or Bard1 inactivation implicate the BRCA1/BARD1 heterodimer in tumor suppression. Proc. Natl. Acad. Sci. USA 2008, 105, 7040–7045. [Google Scholar] [CrossRef] [PubMed]
- Ayi, T.C.; Tsan, J.T.; Hwang, L.Y.; Bowcock, A.M.; Baer, R. Conservation of function and primary structure in the BRCA1-associated RING domain (BARD1) protein. Oncogene 1998, 17, 2143–2148. [Google Scholar] [CrossRef] [PubMed]
- Fox, D., III; Le Trong, I.; Rajagopal, P.; Brzovic, P.S.; Stenkamp, R.E.; Klevit, R.E. Crystal structure of the BARD1 ankyrin repeat domain and its functional consequences. J. Biol. Chem. 2008, 283, 21179–21186. [Google Scholar] [CrossRef] [PubMed]
- Tarsounas, M.; Sung, P. The antitumorigenic roles of BRCA1–BARD1 in DNA repair and replication. Nat. Rev. Mol. Cell Biol. 2020, 21, 284–299. [Google Scholar] [CrossRef]
- Meza, J.E.; Brzovic, P.S.; King, M.C.; Klevit, R.E. Mapping the functional domains of BRCA1. Interaction of the ring finger domains of BRCA1 and BARD1. J. Biol. Chem. 1999, 274, 5659–5665. [Google Scholar] [CrossRef]
- Brzovic, P.S.; Rajagopal, P.; Hoyt, D.W.; King, M.C.; Klevit, R.E. Structure of a BRCA1-BARD1 heterodimeric RING-RING complex. Nat. Struct. Biol. 2001, 8, 833–837. [Google Scholar] [CrossRef]
- Joukov, V.; Chen, J.; Fox, E.A.; Green, J.B.; Livingston, D.M. Functional communication between endogenous BRCA1 and its partner, BARD1, during Xenopus laevis development. Proc. Natl. Acad. Sci. USA 2001, 98, 12078–12083. [Google Scholar] [CrossRef]
- Choudhury, A.D.; Xu, H.; Baer, R. Ubiquitination and proteasomal degradation of the BRCA1 tumor suppressor is regulated during cell cycle progression. J. Biol. Chem. 2004, 279, 33909–33918. [Google Scholar] [CrossRef]
- Lorick, K.L.; Jensen, J.P.; Fang, S.; Ong, A.M.; Hatakeyama, S.; Weissman, A.M. RING fingers mediate ubiquitin-conjugating enzyme (E2)-dependent ubiquitination. Proc. Natl. Acad. Sci. USA 1999, 96, 11364–11369. [Google Scholar] [CrossRef]
- Buetow, L.; Huang, D.T. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat. Rev. Mol. Cell Biol. 2016, 17, 626–642. [Google Scholar] [CrossRef]
- Senft, D.; Qi, J.; Ronai, Z.A. Ubiquitin ligases in oncogenic transformation and cancer therapy. Nat. Rev. Cancer 2018, 18, 69–88. [Google Scholar] [CrossRef] [PubMed]
- Hashizume, R.; Fukuda, M.; Maeda, I.; Nishikawa, H.; Oyake, D.; Yabuki, Y.; Ogata, H.; Ohta, T. The RING heterodimer BRCA1-BARD1 is a ubiquitin ligase inactivated by a breast cancer-derived mutation. J. Biol. Chem. 2001, 276, 14537–14540. [Google Scholar] [CrossRef] [PubMed]
- Ruffner, H.; Joazeiro, C.A.P.; Hemmati, D.; Hunter, T.; Verma, I.M. Cancer-Predisposing mutations within the RING domain of BRCA1: Loss of ubiquitin protein ligase activity and protection from radiation hypersensitivity. Proc. Natl. Acad. Sci. USA. 2001, 98, 5134–5139. [Google Scholar] [CrossRef] [PubMed]
- Brzovic, P.S.; Keeffe, J.R.; Nishikawa, H.; Miyamoto, K.; Fox, D., III; Fukuda, M.; Ohta, T.; Klevit, R. Binding and recognition in the assembly of an active BRCA1/BARD1 ubiquitin-ligase complex. Proc. Natl. Acad. Sci. USA 2003, 100, 5646–5651. [Google Scholar] [CrossRef] [PubMed]
- Brzovic, P.S.; Meza, J.E.; King, M.C.; Klevit, R.E. BRCA1 RING domain cancer-predisposing mutations. Structural consequences and effects on protein-protein interactions. J. Biol. Chem. 2001, 276, 41399–41406. [Google Scholar] [CrossRef]
- Stewart, M.D.; Zelin, E.; Dhall, A.; Walsh, T.; Upadhyay, E.; Corn, J.E.; Chatterjee, C.; King, M.-C.; Klevit, R.E. BARD1 is necessary for ubiquitylation of nucleosomal histone H2A and for transcriptional regulation of estrogen metabolism genes. Proc. Natl. Acad. Sci. USA 2018, 115, 1316–1321. [Google Scholar] [CrossRef]
- Adamovich, A.I.; Banerjee, T.; Wingo, M.; Duncan, K.; Ning, J.; Martins Rodrigues, F.; Huang, K.-L.; Lee, C.; Chen, F.; Ding, L.; et al. Functional analysis of BARD1 missense variants in homology-directed repair and damage sensitivity. PLoS Genet. 2019, 15, e1008049. [Google Scholar] [CrossRef]
- Irminger-Finger, I.; Jefford, C.E. Is there more to BARD1 than BRCA1? Nat. Rev. Cancer 2006, 6, 382–391. [Google Scholar] [CrossRef]
- Nakamura, K.; Saredi, G.; Becker, J.R.; Foster, B.M.; Nguyen, N.V.; Beyer, T.E.; Cesa, L.C.; Faull, P.A.; Lukauskas, S.; Frimurer, T.; et al. H4K20me0 recognition by BRCA1–BARD1 directs homologous recombination to sister chromatids. Nat. Cell Biol. 2019, 21, 311–318. [Google Scholar] [CrossRef]
- Tsuzuki, M.; Wu, W.; Nishikawa, H.; Hayami, R.; Oyake, D.; Yabuki, Y.; Fukuda, M.; Ohta, T. A truncated splice variant of human BARD1 that lacks the RING finger and ankyrin repeats. Cancer Lett. 2006, 233, 108–116. [Google Scholar] [CrossRef]
- Ratajska, M.; Matusiak, M.; Kuzniacka, A.; Wasag, B.; Brozek, I.; Biernat, W.; Koczkowska, M.; Debniak, J.; Sniadecki, M.; Kozlowski, P.; et al. Cancer predisposing BARD1 mutations affect exon skipping and are associated with overexpression of specific BARD1 isoforms. Oncol. Rep. 2015, 34, 2609–2617. [Google Scholar] [CrossRef] [PubMed]
- Billing, D.; Horiguchi, M.; Wu-Baer, F.; Taglialatela, A.; Leuzzi, G.; Nanez, S.A.; Jiang, W.; Zha, S.; Szabolcs, M.; Lin, C.-S.; et al. The BRCT domains of the BRCA1 and BARD1 tumor suppressors differentially regulate homology-directed repair and stalled fork protection. Mol. Cell 2018, 72, 127–139.e8. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Yu, X. Function of BRCA1 in the DNA damage response is mediated by ADP-ribosylation. Cancer Cell 2013, 23, 693–704. [Google Scholar] [CrossRef] [PubMed]
- Choudhary, R.K.; Siddiqui, M.Q.; Thapa, P.S.; Gadewal, N.; Nachimuthu, S.K.; Varma, A.K. Structural basis to stabilize the domain motion of BARD1-ARD BRCT by CstF50. Sci. Rep. 2017, 7, 3849. [Google Scholar] [CrossRef] [PubMed]
- Laufer, M.; Nandula, S.V.; Modi, A.P.; Wang, S.; Jasin, M.; Murty, V.V.V.S.; Ludwig, T.; Baer, R. Structural requirements for the BARD1 tumor suppressor in chromosomal stability and homology-directed DNA repair. J. Biol. Chem. 2007, 282, 34325–34333. [Google Scholar] [CrossRef]
- Irminger-Finger, I.; Leung, W.-C.; Li, J.; Dubois-Dauphin, M.; Harb, J.; Feki, A.; Jefford, C.E.; Soriano, J.V.; Jaconi, M.; Montesano, R.; et al. Identification of BARD1 as Mediator between proapoptotic stress and p53-Dependent apoptosis. Mol. Cell 2001, 8, 1255–1266. [Google Scholar] [CrossRef]
- Kleiman, F.E.; Manley, J.L. The BARD1-CstF-50 interaction links mRNA 3′ end formation to DNA damage and tumor suppression. Cell 2001, 104, 743–753. [Google Scholar] [CrossRef]
- Lee, C.; Banerjee, T.; Gillespie, J.; Ceravolo, A.; Parvinsmith, M.R.; Starita, L.M.; Fields, S.; Toland, A.E.; Parvin, J.D. Functional analysis of BARD1 missense variants in homology-directed repair of DNA double strand breaks. Hum. Mutat. 2015, 36, 1205–1214. [Google Scholar] [CrossRef]
- Casadei, S.; Gulsuner, S.; Shirts, B.H.; Mandell, J.B.; Kortbawi, H.M.; Norquist, B.S.; Swisher, E.M.; Lee, M.K.; Goldberg, Y.; O’Connor, R.; et al. Characterization of splice-altering mutations in inherited predisposition to cancer. Proc. Natl. Acad. Sci. USA 2019, 116, 26798–26807. [Google Scholar] [CrossRef]
- Yoshino, Y.; Qi, H.; Fujita, H.; Shirota, M.; Abe, S.; Komiyama, Y.; Shindo, K.; Nakayama, M.; Matsuzawa, A.; Kobayashi, A.; et al. BRCA1-Interacting Protein OLA1 requires interaction with BARD1 to regulate centrosome number. Mol. Cancer Res. 2018, 16, 1499–1511. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Cohen, P.; Chen, S. How big is a big odds ratio? Interpreting the magnitudes of odds ratios in epidemiological studies. Commun. Stat. Simul. Comput. 2010, 39, 860–864. [Google Scholar] [CrossRef]
- Liu, J.; Cristea, M.C.; Frankel, P.; Neuhausen, S.L.; Steele, L.; Engelstaedter, V.; Matulonis, U.; Sand, S.; Tung, N.; Garber, J.E.; et al. Clinical characteristics and outcomes of BRCA-associated ovarian cancer: Genotype and survival. Cancer Genet. 2012, 205, 34–41. [Google Scholar] [CrossRef] [PubMed]
- Suszynska, M.; Klonowska, K.; Jasinska, A.J.; Kozlowski, P. Large-Scale meta-analysis of mutations identified in panels of breast/ovarian cancer-related genes—Providing evidence of cancer predisposition genes. Gynecol. Oncol. 2019, 153, 452–462. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.-P.; Zhang, Y.; Ma, W.-L.; He, X.-F.; Wang, W.; Yu, H.-L.; Guo, Y.-B.; Zheng, W.-L. Lack of association between BARD1 Cys557Ser variant and breast cancer risk: A meta-analysis of 11,870 cases and 7,687 controls. J. Cancer Res. Clin. Oncol. 2011, 137, 1463–1468. [Google Scholar] [CrossRef]
- Tung, N.M.; Boughey, J.C.; Pierce, L.J.; Robson, M.E.; Bedrosian, I.; Dietz, J.R.; Dragun, A.; Gelpi, J.B.; Hofstatter, E.W.; Isaacs, C.J.; et al. Management of hereditary breast cancer: American society of clinical oncology, American society for radiation oncology, and society of surgical oncology guideline. J. Clin. Oncol. 2020, 38, 2080–2106. [Google Scholar] [CrossRef]
- Pal, T.; Agnese, D.; Daly, M.; La Spada, A.; Litton, J.; Wick, M.; Klugman, S.; Esplin, E.D.; Jarvik, G.P. Points to consider: Is there evidence to support BRCA1/2 and other inherited breast cancer genetic testing for all breast cancer patients? A statement of the American College of Medical Genetics and Genomics (ACMG). Genet. Med. 2020, 22, 681–685. [Google Scholar] [CrossRef]
- Plon, S.E.; Eccles, D.M.; Easton, D.; Foulkes, W.D.; Genuardi, M.; Greenblatt, M.S.; Hogervorst, F.B.L.; Hoogerbrugge, N.; Spurdle, A.B.; Tavtigian, S.V. Sequence variant classification and reporting: Recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum. Mutat. 2008, 29, 1282–1291. [Google Scholar] [CrossRef]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American college of medical genetics and genomics and the association for molecular pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Konstantinopoulos, P.A.; Norquist, B.; Lacchetti, C.; Armstrong, D.; Grisham, R.N.; Goodfellow, P.J.; Kohn, E.C.; Levine, D.A.; Liu, J.F.; Lu, K.H.; et al. Germline and somatic tumor testing in epithelial ovarian cancer: ASCO guideline. J. Clin. Oncol. 2020, 38, 1222–1245. [Google Scholar] [CrossRef]
- Landrum, M.J.; Lee, J.M.; Benson, M.; Brown, G.R.; Chao, C.; Chitipiralla, S.; Gu, B.; Hart, J.; Hoffman, D.; Jang, W.; et al. ClinVar: Improving access to variant interpretations and supporting evidence. Nucleic Acids Res. 2018, 46, 1062–1067. [Google Scholar] [CrossRef] [PubMed]
- Cline, M.S.; Liao, R.G.; Parsons, M.T.; Paten, B.; Alquaddoomi, F.; Antoniou, A.; Baxter, S.; Brody, L.; Cook-Deegan, R.; Coffin, A.; et al. BRCA Challenge: BRCA Exchange as a global resource for variants in BRCA1 and BRCA2. PLoS Genet. 2018, 14, e1007752. [Google Scholar] [CrossRef]
- Hart, S.N.; Polley, E.C.; Yussuf, A.; Yadav, S.; Goldgar, D.E.; Hu, C.; LaDuca, H.; Smith, L.P.; Fujimoto, J.; Li, S.; et al. Mutation prevalence tables for hereditary cancer derived from multi-gene panel testing. Hum. Mutat. 2020, 92, 220–225. [Google Scholar] [CrossRef]
- PDQ Cancer Genetics Editorial Board. Genetics of Breast and Gynecologic Cancers (PDQ®): Health Professional Version; National Cancer Institute (US): Bethesda, MD, USA, 2020.
- Yang, X.; Song, H.; Leslie, G.; Engel, C.; Hahnen, E.; Auber, B.; Horváth, J.; Kast, K.; Niederacher, D.; Turnbull, C.; et al. Ovarian and breast cancer risks associated with pathogenic variants in RAD51C and RAD51D. J. Natl. Cancer Inst. 2020, djaa030. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Leslie, G.; Doroszuk, A.; Schneider, S.; Allen, J.; Decker, B.; Dunning, A.M.; Redman, J.; Scarth, J.; Plaskocinska, I.; et al. Cancer risks associated with germline PALB2 pathogenic variants: An international study of 524 families. J. Clin. Oncol. 2020, 38, 674–685. [Google Scholar] [CrossRef] [PubMed]
- Meeks, H.D.; Song, H.; Michailidou, K.; Bolla, M.K.; Dennis, J.; Wang, Q.; Barrowdale, D.; Frost, D.; McGuffog, L.; Ellis, S.; et al. BRCA2 Polymorphic Stop Codon K3326X and the risk of breast, prostate, and ovarian cancers. J. Natl. Cancer Inst. 2016, 108. [Google Scholar] [CrossRef]
- Peterlongo, P.; Catucci, I.; Colombo, M.; Caleca, L.; Mucaki, E.; Bogliolo, M.; Marin, M.; Damiola, F.; Bernard, L.; Pensotti, V.; et al. FANCM c.5791C>T nonsense mutation (rs144567652) induces exon skipping, affects DNA repair activity and is a familial breast cancer risk factor. Hum. Mol. Genet. 2015, 24, 5345–5355. [Google Scholar] [CrossRef]
- Arem, H.; Loftfield, E. Cancer epidemiology: A survey of modifiable risk factors for prevention and survivorship. Am. J. Lifestyle Med. 2018, 12, 200–210. [Google Scholar] [CrossRef]
- Moorman, P.G.; Havrilesky, L.J.; Gierisch, J.M.; Coeytaux, R.R.; Lowery, W.J.; Urrutia, R.P.; Dinan, M.; Mcbroom, A.J.; Hasselblad, V.; Sanders, G.D.; et al. Oral contraceptives and risk of ovarian cancer and breast cancer among high-risk women: A systematic review and meta-analysis. J. Clin. Oncol. 2013, 31, 4188–4198. [Google Scholar] [CrossRef]
- Milne, R.L.; Osorio, A.; Ramón y Cajal, T.; Baiget, M.; Lasa, A.; Diaz-Rubio, E.; de la Hoya, M.; Caldés, T.; Teulé, A.; Lázaro, C.; et al. Parity and the risk of breast and ovarian cancer in BRCA1 and BRCA2 mutation carriers. Breast Cancer Res. Treat. 2010, 119, 221–232. [Google Scholar] [CrossRef]
- Hartmann, L.C.; Sellers, T.A.; Schaid, D.J.; Frank, T.S.; Soderberg, C.L.; Sitta, D.L.; Frost, M.H.; Grant, C.S.; Donohue, J.H.; Woods, J.E.; et al. Efficacy of bilateral prophylactic mastectomy in BRCA1 and BRCA2 gene mutation carriers. J. Natl. Cancer Inst. 2001, 93, 1633–1637. [Google Scholar] [CrossRef] [PubMed]
- Rebbeck, T.R.; Lynch, H.T.; Neuhausen, S.L.; Narod, S.A.; van’t Veer, L.; Garber, J.E.; Evans, G.; Isaacs, C.; Daly, M.B.; Matloff, E.; et al. Prophylactic oophorectomy in carriers of BRCA1 or BRCA2 mutations. N. Engl. J. Med. 2002, 346, 1616–1622. [Google Scholar] [CrossRef] [PubMed]
- Meijers-Heijboer, H.; van Geel, B.; van Putten, W.L.; Henzen-Logmans, S.C.; Seynaeve, C.; Menke-Pluymers, M.B.; Bartels, C.C.; Verhoog, L.C.; van den Ouweland, A.M.; Niermeijer, M.F.; et al. Breast cancer after prophylactic bilateral mastectomy in women with a BRCA1 or BRCA2 mutation. N. Engl. J. Med. 2001, 345, 159–164. [Google Scholar] [CrossRef] [PubMed]
- Bolton, K.L.; Chenevix-Trench, G.; Goh, C.; Sadetzki, S.; Ramus, S.J.; Karlan, B.Y.; Lambrechts, D.; Despierre, E.; Barrowdale, D.; McGuffog, L.; et al. Association between BRCA1 and BRCA2 mutations and survival in women with invasive epithelial ovarian cancer. JAMA 2012, 307, 382–390. [Google Scholar] [CrossRef]
- Mylavarapu, S.; Das, A.; Roy, M. Role of BRCA mutations in the modulation of response to platinum therapy. Front. Oncol. 2018, 8, 16. [Google Scholar] [CrossRef]
- Fong, P.C.; Yap, T.A.; Boss, D.S.; Carden, C.P.; Mergui-Roelvink, M.; Gourley, C.; De Greve, J.; Lubinski, J.; Shanley, S.; Messiou, C.; et al. Poly (ADP)-Ribose polymerase inhibition: Frequent durable responses in BRCA carrier ovarian cancer correlating with platinum-free interval. J. Clin. Oncol. 2010, 28, 2512–2519. [Google Scholar] [CrossRef]
- Audeh, M.W.; Carmichael, J.; Penson, R.T.; Friedlander, M.; Powell, B.; Bell-Mcguinn, K.M.; Scott, C.; Weitzel, J.N.; Oaknin, A.; Loman, N.; et al. Oral poly (ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: A proof-of-concept trial. Lancet Oncol. 2010, 376, 245–251. [Google Scholar] [CrossRef]
- Gelmon, K.A.; Tischkowitz, M.; Mackay, H.; Swenerton, K.; Robidoux, A.; Tonkin, K.; Hirte, H.; Huntsman, D.; Clemons, M.; Gilks, B.; et al. Olaparib in patients with recurrent high-grade serous or poorly differentiated ovarian carcinoma or triple-negative breast cancer: A phase 2, multicentre, open-label, non-randomised study. Lancet Oncol. 2011, 12, 852–861. [Google Scholar] [CrossRef]
- Kaufman, B.; Shapira-Frommer, R.; Schmutzler, R.K.; Audeh, M.W.; Friedlander, M.; Balmana, J.; Mitchell, G.; Fried, G.; Stemmer, S.M.; Hubert, A.; et al. Olaparib monotherapy in patients with advanced cancer and a germline BRCA1/2 mutation. J. Clin. Oncol. 2015, 33, 244–250. [Google Scholar] [CrossRef]
- Pujade-Lauraine, E.; Ledermann, J.A.; Selle, F.; Gebski, V.; Penson, R.T.; Oza, A.M.; Korach, J.; Huzarski, T.; Poveda, A.; Pignata, S.; et al. Olaparib tablets as maintenance therapy in patients with platinum-sensitive, relapsed ovarian cancer and a BRCA1/2 randomised, placebo-controlled, phase 3 trial. Lancet Oncol. 2017, 18, 1274–1284. [Google Scholar] [CrossRef]
- Ledermann, J.; Harter, P.; Gourley, C.; Friedlander, M.; Vergote, I.; Rustin, G.; Scott, C.L.; Meier, W.; Shapira-Frommer, R.; Safra, T.; et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: A preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 2014, 15, 852–861. [Google Scholar] [CrossRef]

| Reference | Resource Name | Acronym | Resource Type | BARD1 Included in Resource |
|---|---|---|---|---|
| www.nccn.org | National Comprehensive Cancer Network | NCCN | Guideline | Yes |
| [138,140] | American Society of Clinical Oncology | ASCO | Guideline | No |
| [139] | American College of Medical Genetics and Genomics | ACMG | Statement | No |
| www.inesss.qc.ca | Institut national d’excellence en santé et en services sociaux | INESSS | Statement | No |
| Context | Number of Cases Tested | Number of Carriers | % |
|---|---|---|---|
| Personal history of breast cancer | 59,375 | 158 | 0.27 |
| ER+ | 30,950 | 51 | 0.16 |
| PR+ | NA | NA | NA |
| HER2+ | 1670 | 4 | 0.24 |
| TNBC | 6745 | 56 | 0.83 |
| Family history of cancer | |||
| Breast | 37,372 | 107 | 0.29 |
| Ovarian | 7990 | 20 | 0.25 |
| Breast and ovarian | 5138 | 18 | 0.35 |
| No cancer | 3491 | 8 | 0.23 |
| Personal history of ovarian cancer | 9149 | 14 | 0.15 |
| Family history of cancer | |||
| Breast | 3867 | 9 | 0.23 |
| Ovarian | NA | NA | NA |
| Breast and ovarian | NA | NA | NA |
| No cancer | NA | NA | NA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alenezi, W.M.; Fierheller, C.T.; Recio, N.; Tonin, P.N. Literature Review of BARD1 as a Cancer Predisposing Gene with a Focus on Breast and Ovarian Cancers. Genes 2020, 11, 856. https://doi.org/10.3390/genes11080856
Alenezi WM, Fierheller CT, Recio N, Tonin PN. Literature Review of BARD1 as a Cancer Predisposing Gene with a Focus on Breast and Ovarian Cancers. Genes. 2020; 11(8):856. https://doi.org/10.3390/genes11080856
Chicago/Turabian StyleAlenezi, Wejdan M., Caitlin T. Fierheller, Neil Recio, and Patricia N. Tonin. 2020. "Literature Review of BARD1 as a Cancer Predisposing Gene with a Focus on Breast and Ovarian Cancers" Genes 11, no. 8: 856. https://doi.org/10.3390/genes11080856
APA StyleAlenezi, W. M., Fierheller, C. T., Recio, N., & Tonin, P. N. (2020). Literature Review of BARD1 as a Cancer Predisposing Gene with a Focus on Breast and Ovarian Cancers. Genes, 11(8), 856. https://doi.org/10.3390/genes11080856





