Distribution of the Warmblood Fragile Foal Syndrome Type 1 Mutation (PLOD1 c.2032G>A) in Different Horse Breeds from Europe and the United States
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Description Population Screening
2.2. DNA Isolation and Genotyping of WFFS
2.3. Historic DNA—Bairactar Or. Ar.
3. Results
3.1. Distribution of the WFFS Allele
| Breed | Total | Carriers | Carrier Frequency % | Allele Frequency % | 95% CI of WFFS Allele Frequency | Breed Information |
|---|---|---|---|---|---|---|
| Akhal Teke | 35 | 0 | 0 | 0 | ||
| American Sport Pony | 12 | 1 | 8.33 | 4.17 | Warmblood-derived, studbook not closed | |
| American Warmblood | 57 | 8 | 14.04 | 7.02 | 3.41 to 13.43 | Warmblood |
| Appaloosa | 43 | 0 | 0 | 0 | ||
| Arabian | 302 | 0 | 0 | 0 | ||
| Baden-Württemberger | 3 | 1 | 33.33 | 16.67 | Warmblood | |
| Belgian Sport Horse | 10 | 1 | 10.00 | 5.00 | Warmblood | |
| Belgian Warmblood | 44 | 5 | 11.36 | 5.68 | 2.14 to 12.93 | Warmblood |
| Canadian Warmblood | 29 | 3 | 10.34 | 5.17 | Warmblood | |
| Danish Warmblood | 127 | 22 | 17.32 | 8.66 | 5.74 to 12.82 | Warmblood |
| Dutch Warmblood | 249 | 19 | 7.63 | 3.82 | 2.42 to 5.92 | Warmblood |
| Friesian | 197 | 0 | 0 | 0 | ||
| Friesian Cross | 72 | 0 | 0 | 0 | ||
| Haflinger | 48 | 2 | 4.17 | 2.08 | 0.12 to 7.74 | Origin in Europe with influence from several breeds including Arabians, studbook not closed until 1946 |
| Hanoverian | 283 | 49 | 17.31 | 9.01 | 6.90 to 11.67 | Warmblood |
| Polish Heavy Draft | 209 | 0 | 0 | 0 | ||
| Hessen | 2 | 1 | 50.00 | 25.00 | Warmblood | |
| Holsteiner | 132 | 11 | 8.33 | 4.17 | 2.26 to 7.39 | Warmblood |
| Hucul | 146 | 0 | 0 | 0 | ||
| Knabstrupper | 46 | 3 | 6.52 | 3.26 | 0.72 to 9.55 | Baroque type; Warmblood crosses allowed, studbook not closed |
| Lesser Poland Warmblood | 157 | 3 | 1.91 | 0.96 | 0.19 to 2.91 | Warmblood |
| Lippizan | 42 | 0 | 0 | 0 | ||
| Norwegian Fjord | 42 | 0 | 0 | 0 | ||
| Oldenburg | 219 | 34 | 15.53 | 7.76 | 5.58 to 10.68 | Warmblood |
| Polish Konik | 96 | 0 | 0 | 0 | ||
| Quarter Horse | 112 | 0 | 0 | 0 | ||
| Rheinland | 12 | 2 | 16.67 | 8.33 | Warmblood | |
| Rocky Mountain Horse | 89 | 0 | 0 | 0 | ||
| Selle Français | 52 | 3 | 5.77 | 2.88 | 0.62 to 8.50 | Warmblood |
| Shetland Pony | 40 | 0 | 0 | 0 | ||
| Shire | 39 | 0 | 0 | 0 | ||
| Silesian Horse | 96 | 12 | 12.50 | 6.25 | 3.51 to 10.71 | Warmblood |
| Swedish Warmblood | 16 | 0 | 0 | 0 | ||
| Tennessee Walker | 39 | 0 | 0 | 0 | ||
| Thoroughbred * | 146 | 0 | 0 | 0 | ||
| Thoroughbred ** | 716 | 17 | 2.37 | 1.19 | 0.73 to 1.91 | Arabian, Barb, and Turkoman ancestry |
| Trakehner | 64 | 1 | 1.56 | 0.78 | 0.01 to 4.73 | Warmblood |
| Westfalen | 47 | 2 | 4.26 | 2.13 | 0.12 to 7.89 | Warmblood |
| Zangersheide | 11 | 0 | 0 | 0 | ||
| total | 4081 | 200 | 4.90 | 2.47 | 2.16 to 2.84 |
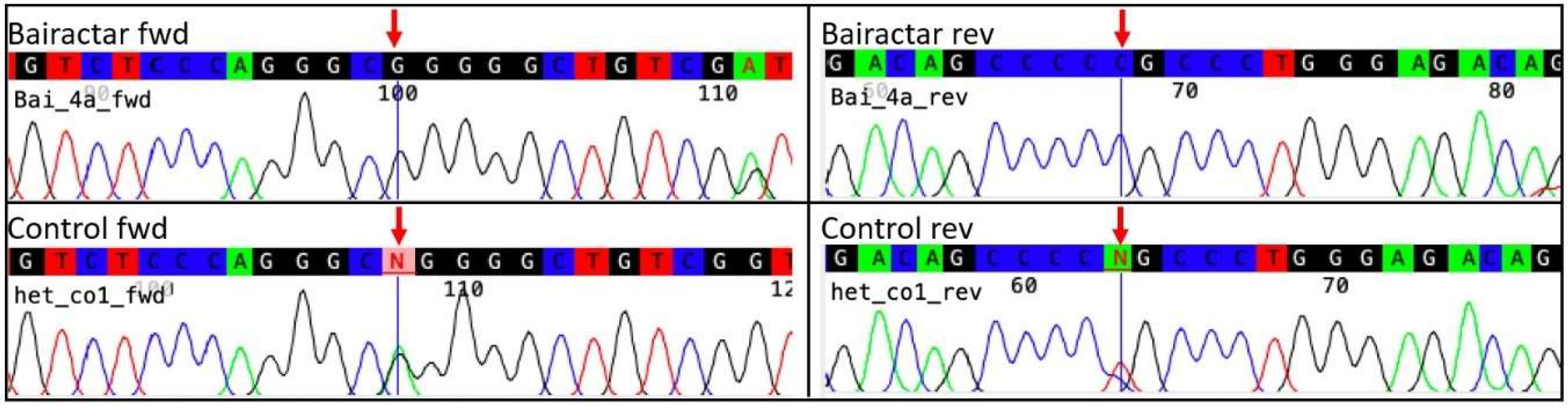
3.2. Historic DNA–Bairactar Or. Ar.
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ehlers-Danlos Syndrome. Available online: https://ghr.nlm.nih.gov/condition/ehlers-danlos-syndrome (accessed on 17 September 2020).
- Malfait, F.; De Paepe, A. The Ehlers-Danlos syndrome. Adv. Exp. Med. Biol. 2014, 802, 129–143. [Google Scholar] [CrossRef]
- Halper, J. Connective tissue disorders in domestic animals. Adv. Exp. Med. Biol. 2014, 802, 231–240. [Google Scholar] [CrossRef]
- Eßer, M.; Niederacher, V.; Pfeffer, K.; Scheuer, H. Über die selten auftretende Dermatosparaxie (Ehlers-Danlos-Syndrom) bei einem Fohlen—Ein Fallbericht. Pferdeheilkunde 1999, 15, 434–436. [Google Scholar] [CrossRef]
- Witzig, P.; Suter, M.; Wild, P.; Rao, V.H.; Steinmann, B.; von Rotz, A. Dermatosparaxis in a foal and a cow—A rare disease? Schweiz. Arch. Tierheilkd. 1984, 126, 589–596. [Google Scholar]
- Marshall, V.L.; Secombe, C.; Nicholls, P.K. Cutaneous asthenia in a Warmblood foal. Aust. Vet. J. 2011, 89, 77–81. [Google Scholar] [CrossRef]
- Rüfenacht, S.; Straub, R.; Steinmann, B.; Winand, N.; Bidaut, A.; Stoffel, M.H.; Gerber, V.; Wyder, M.; Müller, E.; Roosje, P.J. Swiss warmblood horse with symptoms of hereditary equine regional dermal asthenia without mutation in the cyclophylin B gene (PPIB). Schweiz. Arch. Tierheilkd. 2010, 152, 188–192. [Google Scholar] [CrossRef]
- Winter, T.; Börgel, C.; Aupperle, H.; Schoon, H. Eine dem Ehlers-Danlos Syndrom des Menschen ähnliche Bindegewebserkrankung der Haut bei einem Fohlen. Pferdeheilkunde 2004, 20, 19–22. [Google Scholar] [CrossRef]
- Rueff, A.G. Eigenthümliche Krankheit neugeborener Fohlen (a peculiar disease of newborn foals). In Jahrbuch für Pferdezucht; Verlag Voigt: Weimar, Germany, 1855; pp. 267–277. [Google Scholar]
- Gunson, D.E.; Halliwell, R.E.; Minor, R.R. Dermal collagen degradation and phagocytosis. Occurrence in a horse with hyperextensible fragile skin. Arch. Dermatol. 1984, 120, 599–604. [Google Scholar] [CrossRef]
- Rashmir-Raven, A. Heritable equine regional dermal asthenia. Vet. Clin. N. Am. Equine Pract. 2013, 29, 689–702. [Google Scholar] [CrossRef]
- White, S.D.; Affolter, V.K.; Bannasch, D.L.; Schultheiss, P.C.; Hamar, D.W.; Chapman, P.L.; Naydan, D.; Spier, S.J.; Rosychuk, R.A.W.; Rees, C.; et al. Hereditary equine regional dermal asthenia (“hyperelastosis cutis”) in 50 horses: Clinical, histological, immunohistological and ultrastructural findings. Vet. Dermatol. 2004, 15, 207–217. [Google Scholar] [CrossRef]
- Steelman, S.M.; Jackson, N.D.; Conant, E.; Juras, R.; Cothran, E.G.; Edwards, J.F.; Chowdhary, B.P. Ehlers-danlos syndrome in a quarter horse gelding: A case report of PPIB-independent hereditary equine regional dermal asthenia. J. Equine Vet. Sci. 2014, 34, 565–568. [Google Scholar] [CrossRef]
- Borges, A.S.; Conceição, L.G.; Alves, A.L.G.; Fabris, V.E.; Pessoa, M.A. Hereditary equine regional dermal asthenia in three related Quarter horses in Brazil. Vet. Dermatol. 2005, 16, 125–130. [Google Scholar] [CrossRef] [PubMed]
- Lerner, D.; McCracken, M. Hyperelastosis cutis in 2 Horses. J. Equine Med. Surg. 1978, 2, 350–352. [Google Scholar]
- White, S.D.; Affolter, V.K.; Schultheiss, P.C.; Ball, B.A.; Wessel, M.T.; Kass, P.; Molinaro, A.M.; Bannasch, D.L.; Ihrke, P.J. Clinical and pathological findings in a HERDA-affected foal for 1.5 years of life. Vet. Dermatol. 2007, 18, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Hardy, M.H.; Fisher, K.R.; Vrablic, O.E.; Yager, J.A.; Nimmo-Wilkie, J.S.; Parker, W.; Keeley, F.W. An inherited connective tissue disease in the horse. Lab. Investig. 1988, 59, 253–262. [Google Scholar] [PubMed]
- Brounts, S.H.; Rashmir-Raven, A.M.; Black, S.S. Zonal dermal separation: A distinctive histopathological lesion associated with hyperelastosis cutis in a Quarter Horse. Vet. Dermatol. 2001, 12, 219–224. [Google Scholar] [CrossRef] [PubMed]
- Bowser, J.E.; Elder, S.H.; Pasquali, M.; Grady, J.G.; Rashmir-Raven, A.M.; Wills, R.; Swiderski, C.E. Tensile properties in collagen-rich tissues of Quarter Horses with hereditary equine regional dermal asthenia (HERDA). Equine Vet. J. 2014, 46, 216–222. [Google Scholar] [CrossRef]
- Grady, J.G.; Elder, S.H.; Ryan, P.L.; Swiderski, C.E.; Rashmir-Raven, A.M. Biomechanical and molecular characteristics of hereditary equine regional dermal asthenia in Quarter Horses. Vet. Dermatol. 2009, 20, 591–599. [Google Scholar] [CrossRef]
- Mochal, C.A.; Miller, W.W.; Cooley, A.J.; Linford, R.L.; Ryan, P.L.; Rashmir-Raven, A.M. Ocular findings in Quarter Horses with hereditary equine regional dermal asthenia. J. Am. Vet. Med. Assoc. 2010, 237, 304–310. [Google Scholar] [CrossRef]
- Tryon, R.C.; Penedo, M.C.T.; McCue, M.E.; Valberg, S.J.; Mickelson, J.R.; Famula, T.R.; Wagner, M.L.; Jackson, M.; Hamilton, M.J.; Nooteboom, S.; et al. Evaluation of allele frequencies of inherited disease genes in subgroups of American Quarter Horses. J. Am. Vet. Med. Assoc. 2009, 234, 120–125. [Google Scholar] [CrossRef]
- Finno, C.J.; Spier, S.J.; Valberg, S.J. Equine diseases caused by known genetic mutations. Vet. J. 2009, 179, 336–347. [Google Scholar] [CrossRef] [PubMed]
- Solomons, B. Equine cutis hyperelastica. Equine Vet. J. 1984, 16, 541–542. [Google Scholar] [CrossRef] [PubMed]
- Tryon, R.C.; White, S.D.; Bannasch, D.L. Homozygosity mapping approach identifies a missense mutation in equine cyclophilin B (PPIB) associated with HERDA in the American Quarter Horse. Genomics 2007, 90, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Monthoux, C.; de Brot, S.; Jackson, M.; Bleul, U.; Walter, J. Skin malformations in a neonatal foal tested homozygous positive for Warmblood Fragile Foal Syndrome. BMC Vet. Res. 2015, 11, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Winand, N.J. Identification of the Causative Mutation for Inherited Connective Tissue Disorders in Equines, United States Department of Commerce Application Number: 61/486,464; (Filing Date: May 16th, 2011). Available online: https://patentscope.wipo.int/search/en/detail.jsf?docId=WO2012158711&recNum=1&maxRec=1&office=&prevFilter=&sortOption=Pub+Date+Desc&queryString=FP%3A%28WO2012158711%29&tab=PCT+Biblio (accessed on 20 September 2020).
- Aurich, C.; Müller-Herbst, S.; Reineking, W.; Müller, E.; Wohlsein, P.; Gunreben, B.; Aurich, J. Characterization of abortion, stillbirth and non-viable foals homozygous for the Warmblood Fragile Foal Syndrome. Anim. Reprod. Sci. 2019, 211, 106202. [Google Scholar] [CrossRef] [PubMed]
- Metzger, J.; Kreft, O.; Sieme, H.; Martinsson, G.; Reineking, W.; Hewicker-Trautwein, M.; Distl, O. Hanoverian F/W-line contributes to segregation of Warmblood fragile foal syndrome type 1 variant PLOD1:c.2032G>A in Warmblood horses. Equine Vet. J. 2020, 22, 1–9. [Google Scholar] [CrossRef]
- Bellone, R.R.; Ocampo, N.R.; Hughes, S.S.; Le, V.; Arthur, R.; Finno, C.J.; Penedo, M.C.T. Warmblood fragile foal syndrome type 1 mutation (PLOD1 c.2032G>A) is not associated with catastrophic breakdown and has a low allele frequency in the Thoroughbred breed. Equine Vet. J. 2020, 52, 411–414. [Google Scholar] [CrossRef]
- Wege zum Angemessenen Umgang mit WFFS und Anderen Genetischen Eigenschaften. Available online: https://www.vit.de/aktuelles/20191/wege-zum-angemessenen-umgang-mit-wffs-und-anderen-genetischen-eigenschaften/ (accessed on 17 December 2020).
- Zhang, X.; Hirschfeld, M.; Schafberg, R.; Swalve, H.; Brenig, B. Skin exhibits of Dark Ronald XX are homozygous wild type at the Warmblood fragile foal syndrome causative missense variant position in lysyl hydroxylase gene PLOD1. Anim. Genet. 2020, 1–2. [Google Scholar] [CrossRef]
- Locke, M.M.; Penedo, M.C.T.; Bricker, S.J.; Millon, L.V.; Murray, J.D. Linkage of the grey coat colour locus to microsatellites on horse chromosome 25. Anim. Genet. 2002, 33, 329–337. [Google Scholar] [CrossRef]
- Agrest, A.; Coull, B.S. Approximate is Better than “Exact” for Interval Estimation of Binomial Proportions. Am. Stat. 1998, 52, 119–126. [Google Scholar] [CrossRef]
- Dias, N.M.; de Andrade, D.G.A.; Teixeira-Neto, A.R.; Trinque, C.M.; de Oliveira-Filho, J.P.; Winand, N.J.; Araújo, J.P.; Borges, A.S. Warmblood Fragile Foal Syndrome causative single nucleotide polymorphism frequency in Warmblood horses in Brazil. Vet. J. 2019, 248, 101–102. [Google Scholar] [CrossRef]
- Sandoval-Castellanos, E.; Wutke, S.; Gonzalez-Salazar, C.; Ludwig, A. Coat colour adaptation of post-glacial horses to increasing forest vegetation. Nat. Ecol. Evol. 2017, 1, 1816–1819. [Google Scholar] [CrossRef]
- Georgescu, S.; Manea, M.; Dudu, A.; Costache, M.; Mol, G. Phylogenetic relationships of the Hucul horse from Romania inferred from mitochondrial D-loop variation. Genet. Mol. Res. 2011, 10, 4104–4113. [Google Scholar] [CrossRef]
- Polak, G. Genetic variability of cold-blooded horses participating in genetic resources conservation programs, using pedigree analysis. Ann. Anim. Sci. 2019, 19, 49–60. [Google Scholar] [CrossRef]
- Fung, T.; Kennan, K. General case of sampling form a finite diploid population of any size. PLoS ONE 2014, 9, e85925. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reiter, S.; Wallner, B.; Brem, G.; Haring, E.; Hoelzle, L.; Stefaniuk-Szmukier, M.; Długosz, B.; Piórkowska, K.; Ropka-Molik, K.; Malvick, J.; et al. Distribution of the Warmblood Fragile Foal Syndrome Type 1 Mutation (PLOD1 c.2032G>A) in Different Horse Breeds from Europe and the United States. Genes 2020, 11, 1518. https://doi.org/10.3390/genes11121518
Reiter S, Wallner B, Brem G, Haring E, Hoelzle L, Stefaniuk-Szmukier M, Długosz B, Piórkowska K, Ropka-Molik K, Malvick J, et al. Distribution of the Warmblood Fragile Foal Syndrome Type 1 Mutation (PLOD1 c.2032G>A) in Different Horse Breeds from Europe and the United States. Genes. 2020; 11(12):1518. https://doi.org/10.3390/genes11121518
Chicago/Turabian StyleReiter, Simone, Barbara Wallner, Gottfried Brem, Elisabeth Haring, Ludwig Hoelzle, Monika Stefaniuk-Szmukier, Bogusława Długosz, Katarzyna Piórkowska, Katarzyna Ropka-Molik, Julia Malvick, and et al. 2020. "Distribution of the Warmblood Fragile Foal Syndrome Type 1 Mutation (PLOD1 c.2032G>A) in Different Horse Breeds from Europe and the United States" Genes 11, no. 12: 1518. https://doi.org/10.3390/genes11121518
APA StyleReiter, S., Wallner, B., Brem, G., Haring, E., Hoelzle, L., Stefaniuk-Szmukier, M., Długosz, B., Piórkowska, K., Ropka-Molik, K., Malvick, J., Penedo, M. C. T., & Bellone, R. R. (2020). Distribution of the Warmblood Fragile Foal Syndrome Type 1 Mutation (PLOD1 c.2032G>A) in Different Horse Breeds from Europe and the United States. Genes, 11(12), 1518. https://doi.org/10.3390/genes11121518






