HSP play crucial functions, including stress tolerance, cell survival, protein refolding, and prevention of protein denaturation. Moreover, HSP also facilitate the regulation of specific cellular processes such as signal transduction, protein synthesis, protein trafficking, and DNA replication [
4,
25,
26]. It is well established that
HSP90 plays pivotal role in controlling cell cycle, transport, folding, and degradation of proteins as well as thermotolerance and induction of signal transduction. Studies have revealed that
HSP90 is present in various organisms and this HSP family is highly conserved [
27]. HSPs are the molecular chaperones that help to maintain activity of cells in different stressful states. Some of these proteins are produced due to different stressful stimuli (inducible), while others are expressed constitutively. Cellular metabolism, ultraviolet radiations, alteration in protein structures, cytotoxic drugs, and heat are some of the stress stimuli that usually induce HSP expression. These proteins are highly involved in facilitating the degradation of misfolded proteins and regulating proper protein folding [
28]. In this study, we analyzed a total of 64 HSP genes in buffalo. Members of
HSP90 are found to be present on four chromosomes in buffalo similar to
Bos taurus [
29], suggesting that these protein molecules are highly conserved and share a common ancestor (
Figure S3).
4.1. Physiochemical Properties of HSP Gene Families
The characterization of the physiochemical properties of proteins of various gene families is essential to identify the functions and properties of proteins encoded by these genes. Isoelectric point indicates that members of
HSP90 in buffalo are acidic in nature (pI < 7.0), except for the
TRAP1 gene, which was basic in nature similar to human
HSP90 [
30].
AI of the globular proteins is associated with their thermostability [
31]. The AI, the respective volume of a protein occupied by aliphatic side chains (alanine, valine, isoleucine, and leucine), may be interpreted as a positive factor for increasing thermostability of globular proteins [
31]. As the AI is associated with the thermostability of globular proteins, so a higher value of the AI reveals the high thermostability of protein. Hence, a higher AI of HSP90 and HSP70 showed that members of these proteins are more thermostable compared to HSP40 members. Similar results have also been observed in cattle genome [
29]. In the present study, the
HSP90 gene family of buffalo had AI greater than 65, indicating the stability of proteins at high temperatures. Similarly, the higher expression of
HSP90 has been found in goat during the peak summer season, revealing the extent of thermostability of HSPs [
32].
The GRAVY value is used to predict water and protein interaction. The GRAVY value for a protein or a peptide is defined as the sum of hydropathy values of all amino acids divided by the protein length. GRAVY qualifies and quantifies the hydrophobic or hydrophilic property of a protein. A positive GRAVY score indicates a globally hydrophobic protein, whereas a negative GRAVY score is related to hydrophilic protein [
33]. Hence, more the negative GRAVY score, more soluble the protein is. All members of HSP40, HSP70, and HSP90 had a negative GRAVY score in the present study, suggesting these all members are soluble proteins. However, highly negative GRAVY values of
HSP90 observed in the present study indicate the greater hydrophilic nature of its members. It can help to enhance the functional properties of protein binding and oligomerization [
34,
35]. Among heat-shock proteins,
HSP90 is considered an essential and highly conserved molecular chaperone widely present in eukaryotic cells [
36].
HSP90 in buffalo share high sequence similarities with other species (
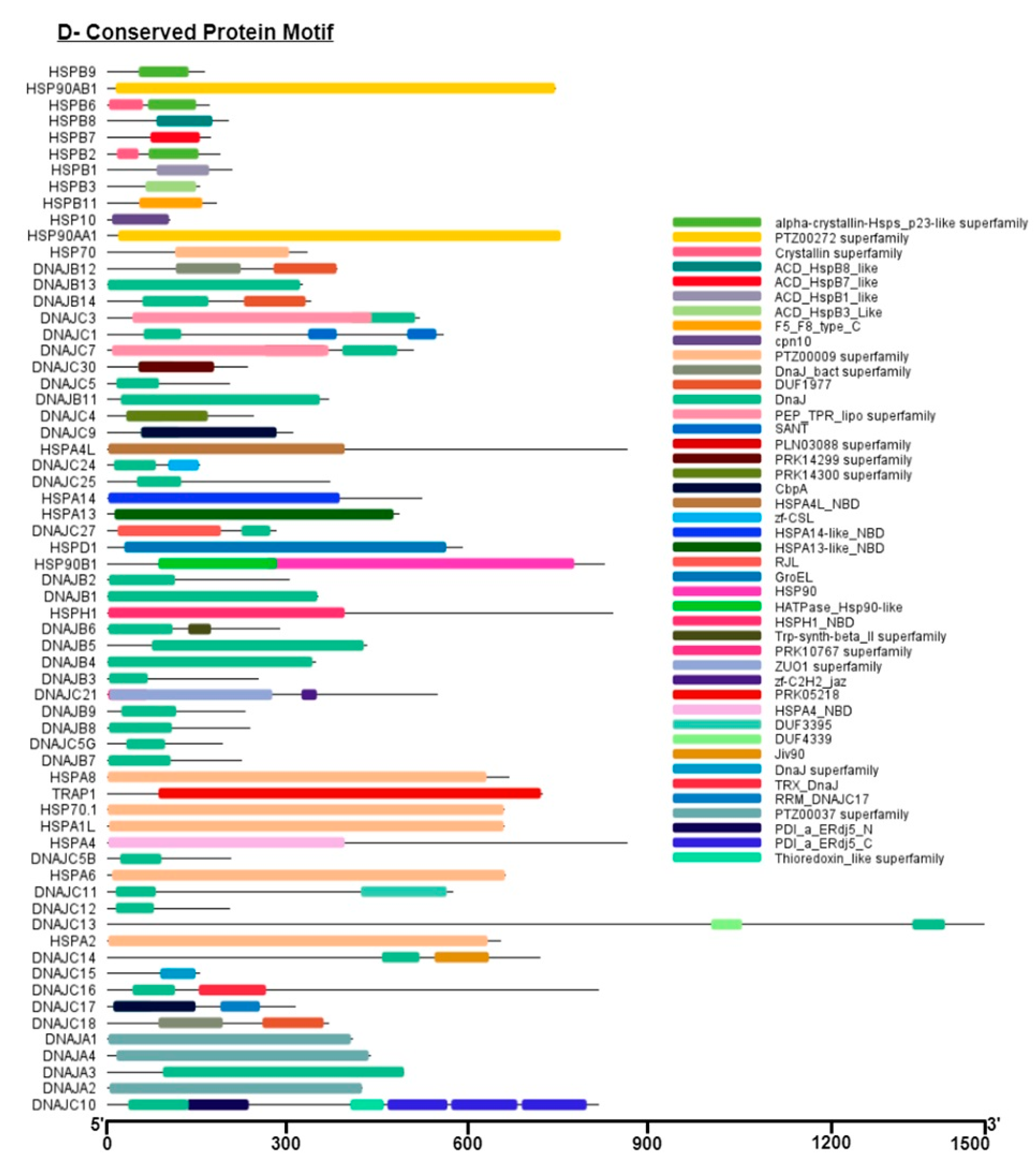
Figure S3). The presence of conserved motifs helps interact with other proteins and increases their chaperone function [
37,
38].
A total of 10
HSP70 members were observed in buffalo. Interestingly, all members of
HSP70 showed acidic nature and almost similar isoelectric points indicating the possibility of functional similarity among buffalo
HSP70 members. It has been suggested earlier that
HSP70 has conserved biological functions [
39]. GRAVY values of
HSP70 protein of buffalo indicates their hydrophilic nature and high values of AI revealed their thermal stability in a wide range of temperatures. These findings confirm the consistency of its chaperoning role in protection against specific cellular and systemic stresses that can lead to protein denaturation [
40]. In evolution,
HSP70 is the most conserved protein, and its conserved domains and functions are considered responsible for across species conservation [
11,
41]. In the present study, a tree with an alignment view of
HSP70 showed the presence of conserved domains. Buffalo is closely related to
Bos taurus, followed by goat and sheep (
Figure S2). The comparative analysis of the
HSP70 protein in water buffalo revealed that most of the sequences were conserved across species. Comparative sequence analysis for bubaline
HSP70.1 indicated the high similarity of genes among mammalian species, such as cattle, goat, human, mouse, and pig, respectively. Furthermore, phylogenetic analysis of bovine and bubaline
HSP70.1 gene with other species revealed Indian zebu cattle grouped close to
Bos taurus followed by buffalo and goat, while porcine, ruminants, and murine formed distinct cluster [
42,
43].
Heat-shock protein 40 (
HSP40/DnaJ) is evolutionary conserved and thought to play an essential role in translocation, degradation, translation, folding, and unfolding of proteins by stimulating the ATPase activity of
HSP70 protein chaperones [
44,
45]. All members of
HSP40 have domain J, which helps in binding with
HSP70. Also, 41
DnaJ domains have been reported in humans [
46]. In the present study, 39 putative members of
HSP40 were found. GRAVY value of
HSP40 members represents the hydrophilic nature of these proteins. Furthermore, pI values of
HSP40 protein members suggest that some of these proteins possess acidic nature while other proteins appear to be basic. It may explain the different functional properties of these proteins and provides insights for designing further experiments to elucidate their functional diversity.
HSP40 represents a large and diverse protein family and is usually divided into three classes [
47]. Type I
HSP40s is known to have a glycine and phenylalanine (G/F)-rich region and a cysteine-rich, zinc finger-like region (ZFLR). Type II
HSP40s only contain a G/F-rich region, while type III
HSP40s have neither a G/F-rich region nor a ZFLR [
48]. Comparative sequence and phylogenetic analyses revealed that the
HSP40 family of buffalo is closely related to
Bos taurus (
Figure S1). Another study conducted on the bovine genome also indicated the presence of evolutionary conserved residues in
HSP40 [
29].
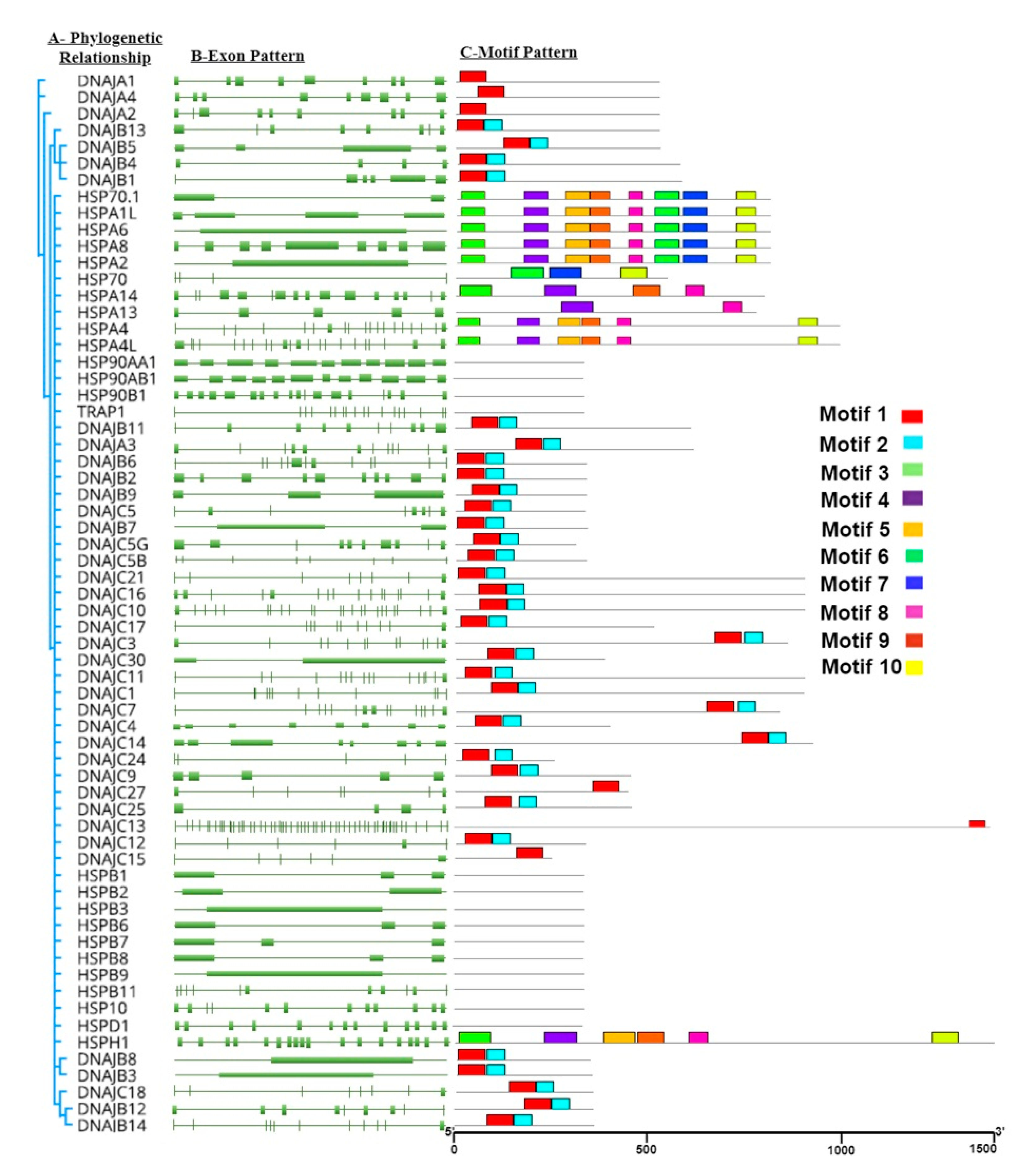
4.3. Phylogenetic Analysis
Molecular phylogenetic tree of heat-shock proteins (
HSP40,
HSP70,
HSP90,
HSPB,
HSPD,
HSP10, and
HSPH1) was constructed by using the Maximum Likelihood method based on the JTT matrix-based model [
22]. The bootstrap consensus tree inferred from 1000 replicates was taken to represent the evolutionary history of the taxa analyzed (
Figure 1). Overall phylogenetic relationships revealed that buffalo is more closely related to goat than
Bos taurus, and sheep along with horse and camel (
Figure 2). Moreover, buffalo is distantly related to other mammals, including pig, dog, rat, mouse and zebra fish. A study conducted on cattle also revealed similar results as phylogenetic analysis of
HSPA8 and
HSPA1A indicted the high similarity between buffalo and sheep [
51]. Previously, molecular characterization of
HSP70.1 of goat have shown that buffalo, cattle, sheep, and goat share a common ancestor whereas human, horse and pig showed dissimilarities thus having different ancestors [
52]. Moreover, Karan Fries cattle was evolutionary related to
Bos taurus,
Bos indicus, and buffalo, whereas distinctly related to rat and mouse [
53].
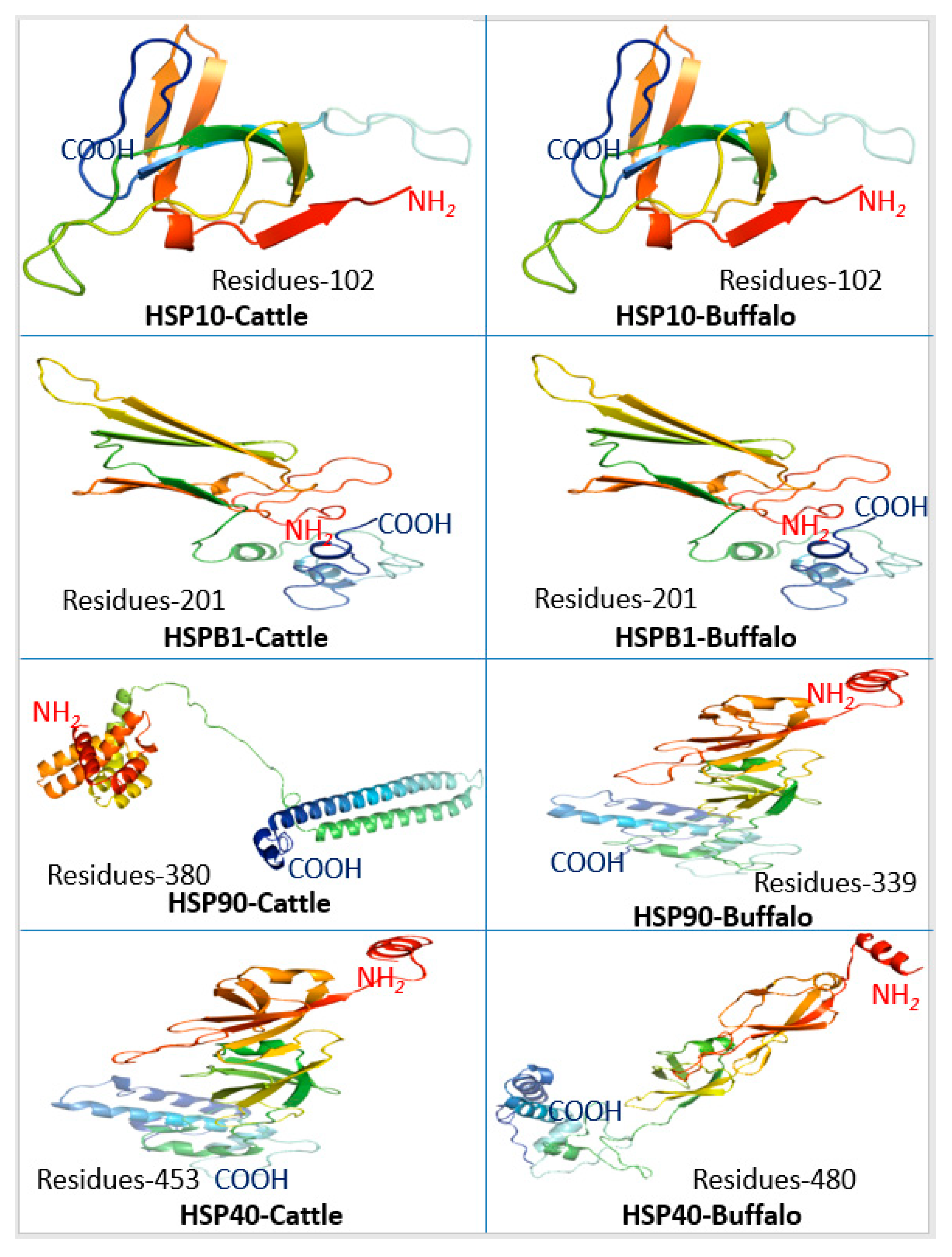
4.4. Protein Structural Configuration
Three-dimensional (3-D) protein configurations of HSP in
Bos taurus and buffalo was predicted. The analysis of 3-D secondary protein structures revealed the structural variations only in
HSP90 and
HSP40 of
Bos taurus and buffalo, while other HSPs were similar or structurally related in both species. These findings confirm the results obtained in gene structure and phylogenetic analysis that HSPs are conserved proteins. Eukaryotic
HSP90 proteins had a modular structure with an N-terminal ATP binding domain connected to a middle domain by a ‘linker’ of variable length. The N-terminal and middle domains together form a “split” ATPase site that is also the binding pocket for GA [
54].
HSP40 can function as a molecular chaperone to bind non-native polypeptides.
HSP40 pairs with
HSP70 to achieve its function of protein folding, transport, and degradation functions. The J-domain of
HSP40 may stimulate the ATPase activity of
HSP70 and the C-terminal peptide-binding fragment of
HSP40 can load the bound non-native polypeptide to
HSP70 [
55].
HSP70 and
HSP40 are essential chaperons affecting the protein folding in the cell. Three-dimensional structural configurations of
HSP40 in cattle and buffalo varied in number of amino acid residues and overall secondary structural attributes that may be associated with differential chaperon functions and stress responses in both species. Further investigations at the transcriptome level are required to corroborate these findings.
4.5. SNP Analysis
The present study revealed that
HSP90AA1,
HSP90AB1 and
HSP90B1 are more conserved genes than TRP1 due to a higher number of non-synonymous SNPs in
TRP1. Similarly, our findings indicate that
HSP70.1, HSPA14, and
HSPA1L are more conserved than other genes of
HSP70 family. We observed a relatively higher ratio of non-synonymous to synonymous SNPs in the
HSP70 gene (0.59) as compared to other genes of the
HSP70 family. It indicates that
HSP70 gene is under selection owing to its strong association with cellular thermotolerance [
1]. Moreover, the expression dynamics of the
HSP70 gene are used as a potential biomarker to reveal the level of heat stress and the ability of animals to withstand climate stress [
56]. It is mainly attributed to the fact that HSP expression indicates cellular response and intensity of stress in animals [
57]. At the transcription level, persuaded expression of
HSP70 genes is coordinated as upstream elements at promoter region directly control the expression profiles of
HSP70. Studies have shown a higher expression of HSP70 in Murrah and Tarai buffaloes [
58] during the summer compared to other seasons (or thermoneutral zone). Moreover, increased expression of
HSP70 in bovine lymphocytes has also been observed after heat stress [
59].
Due to subsequent induction of
HSP70 after heat shock and its association with physiological parameters (rectal temperature, pulse, and respiratory rates) of animals [
59], serum
HSP70 is considered a sensitive biomarker for heat stress management in livestock [
58]. In addition to thermotolerance, studies have also reported other roles of the HSP family in animal physiology. In particular, SNPs identified in the 5′-UTR regions of HSP70 gene have been linked with different performance traits of livestock such as milk production, thermal stress and disease vulnerability [
1,
43]. Moreover, SNPs have also been identified in the
HSP70 gene of beef cattle breeds (Angus, Brahman, and their crossbreds) with little effect on milk protein and milk fat [
60]. Polymorphism (SNP) at the promoter region has been shown to alter the expression level of bovine
HSP70.1 gene, and exhibited its significant association with heat tolerance and higher milk production in Frieswal cattle [
43]. These findings indicate the putative effects of selective pressure (natural or artificial selection) on the
HSP70 gene to ultimately favor animals with better thermotolerance, performance, and stress resilience. Our findings are in line with recent whole genome-sequencing studies on buffalo that reported evidence for the selection of genes related to social behavior, digestive physiology, fitness, and milk production [
61].
HSPs are highly conserved in nature and play crucial role in stress stimulus and in several normal cellular functions. The genomic characterization of the HSP gene family and elucidation of their physiochemical properties performed in the present study would help in better understanding HSPs in buffalo. HSPs are highly conserved proteins and important from evolutionary point of view, so further investigations are warranted to study expression dynamics under various physiological conditions to elucidate their putative roles in thermotolerance and stress physiology in buffalo. In particular, non-synonymous SNPs and indels observed in the present study need to be further explored to detect their effect on protein function and subsequent physiological manifestations in buffalo. These findings might explain the better thermotolerance and fitness of buffalo compared to Bos taurus, particularly under tropical climate.