Genome Mining of the Genus Streptacidiphilus for Biosynthetic and Biodegradation Potential
Abstract
1. Introduction
2. Materials and Methods
2.1. Streptacidiphilus Strains for Genome Analysis
2.2. Genome Sequencing, Assembly and Annotation
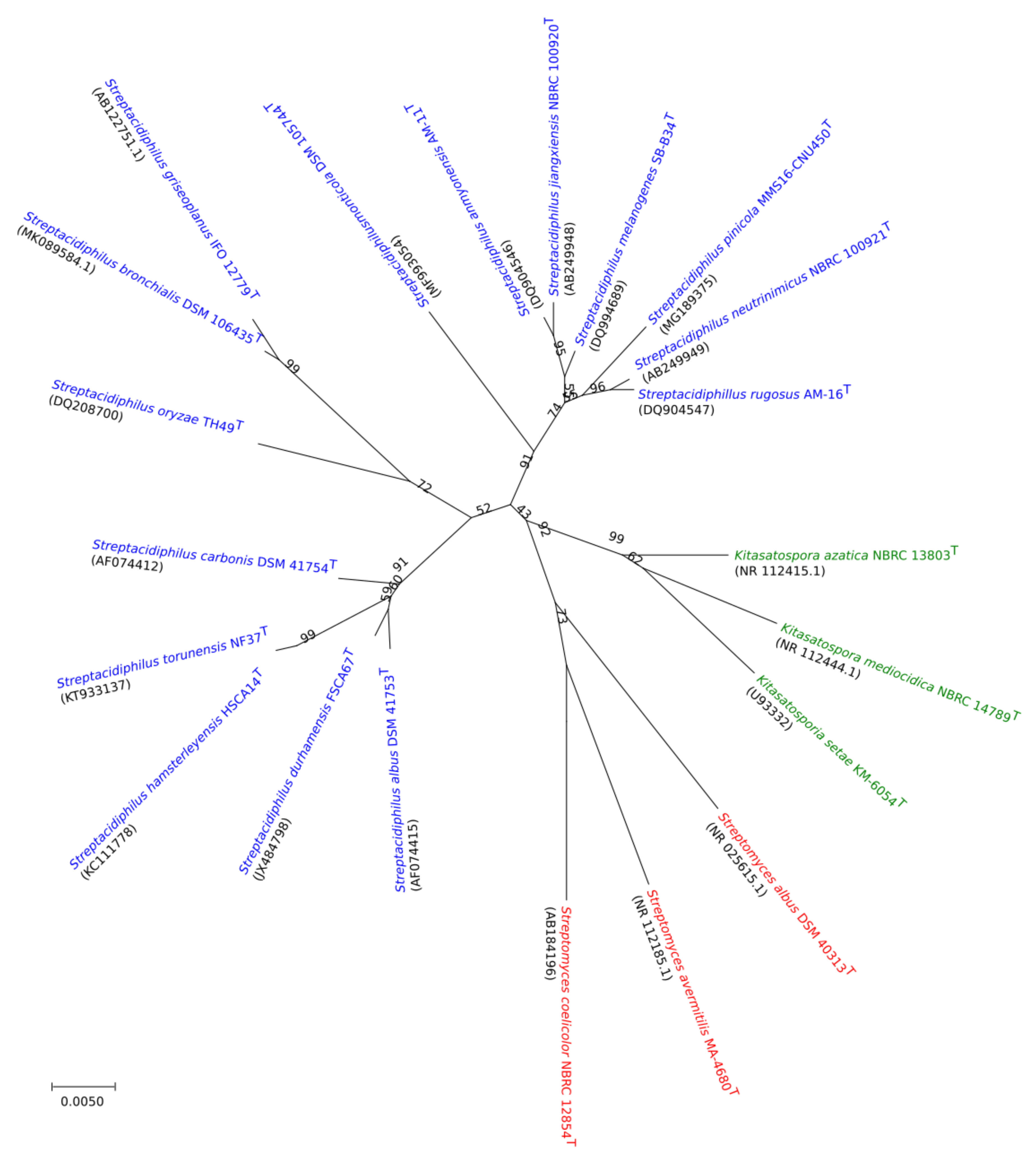
2.3. Comparative Genomic and Phylogenetic Analysis
3. Results and Discussion
3.1. General Features of Streptacidiphilus Genomes
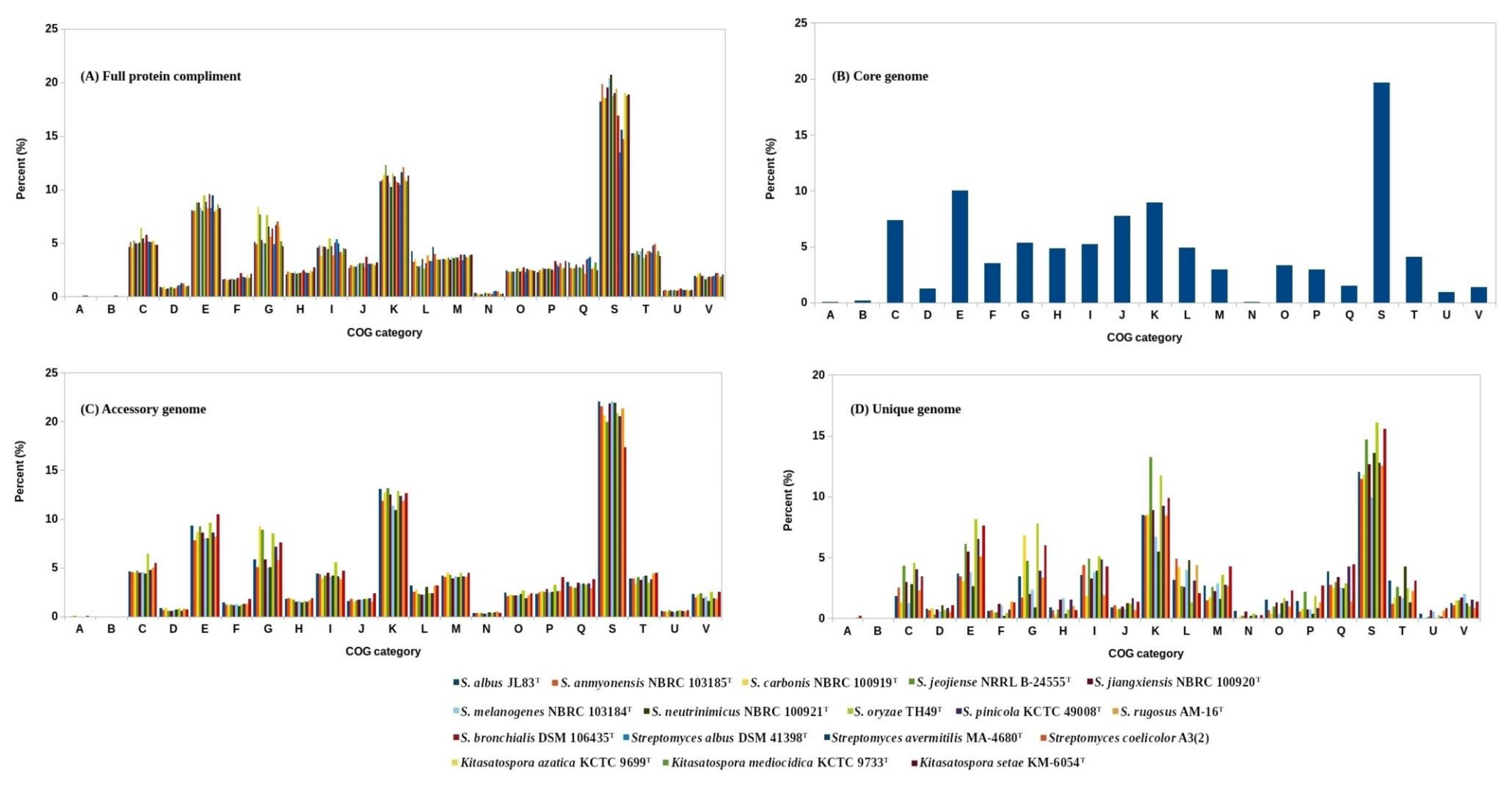
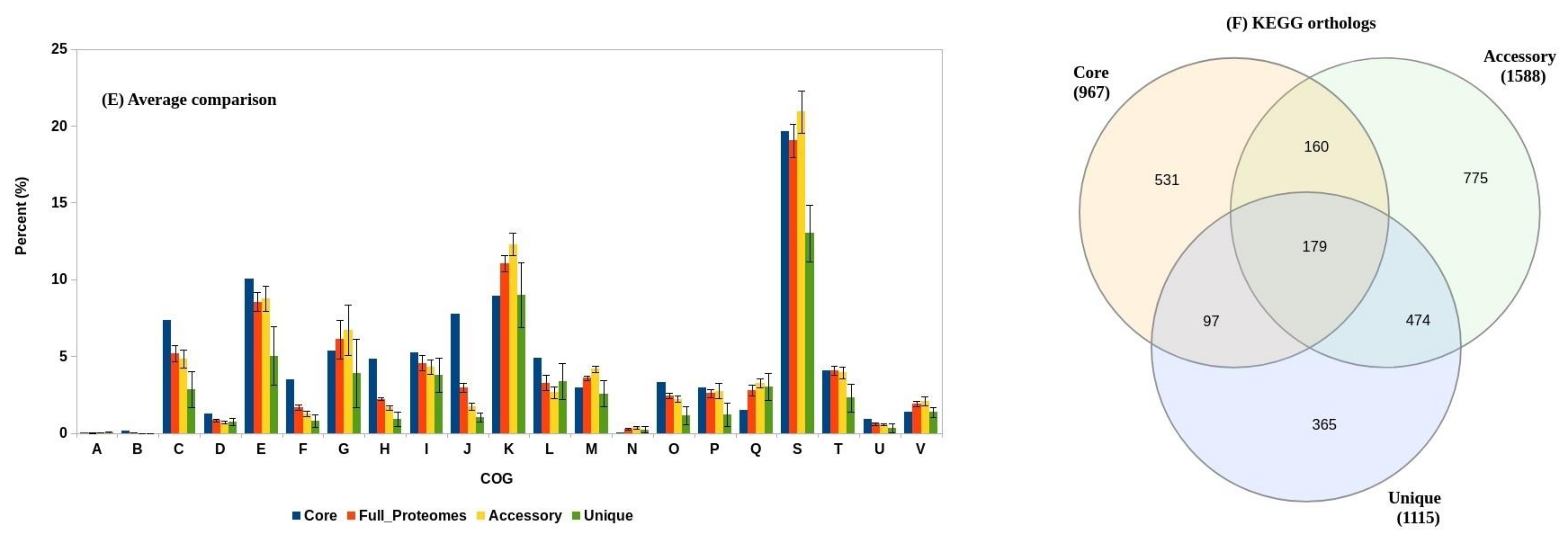
3.2. Functional Annotation
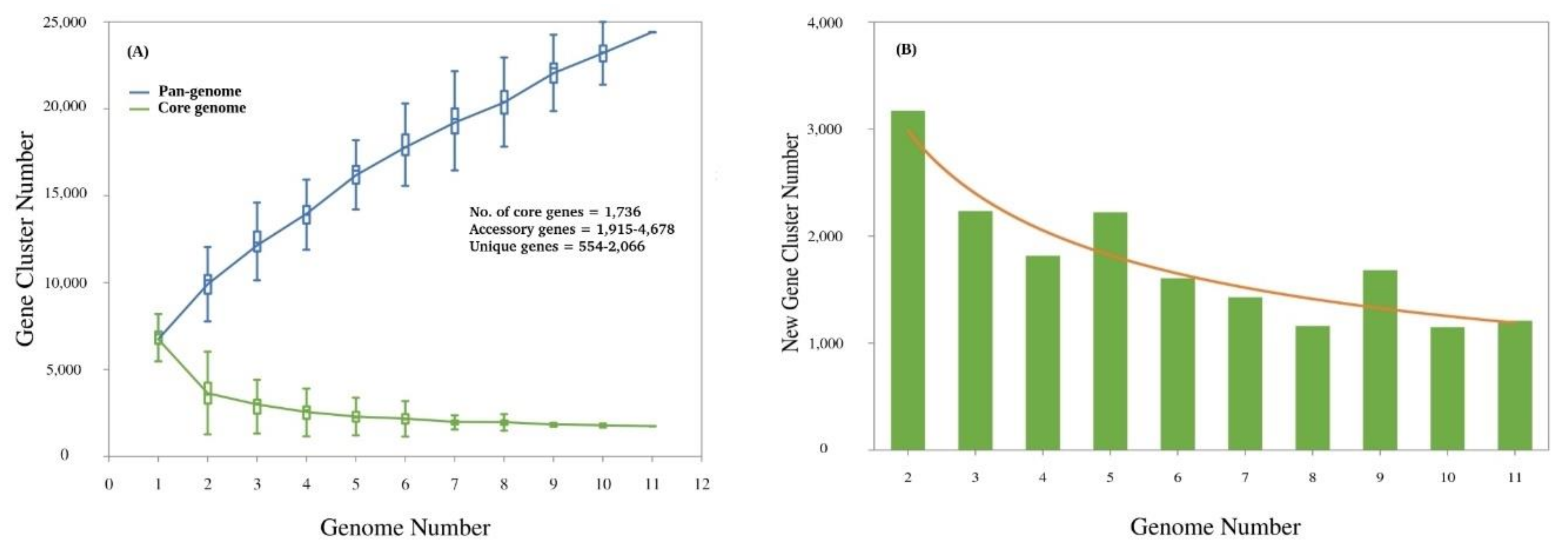
3.3. Pan-Genome Analysis
3.3.1. Core Genome
3.3.2. Accessory Genome
3.3.3. Unique Genome
3.4. Genes Related to Morphological Properties
3.5. Predicted BGCs of Streptacidiphilus
3.5.1. Type 1 Polyketide Synthase (T1PKS) BGCs
3.5.2. Non-Ribosomal Peptide Synthetase (NRPS) BGCs
3.5.3. T1PKS-NRPS Hybrid Gene Clusters
3.5.4. Other Hybrid Gene Clusters
3.5.5. Lanthipeptide Gene Clusters
3.5.6. Terpene Gene Clusters
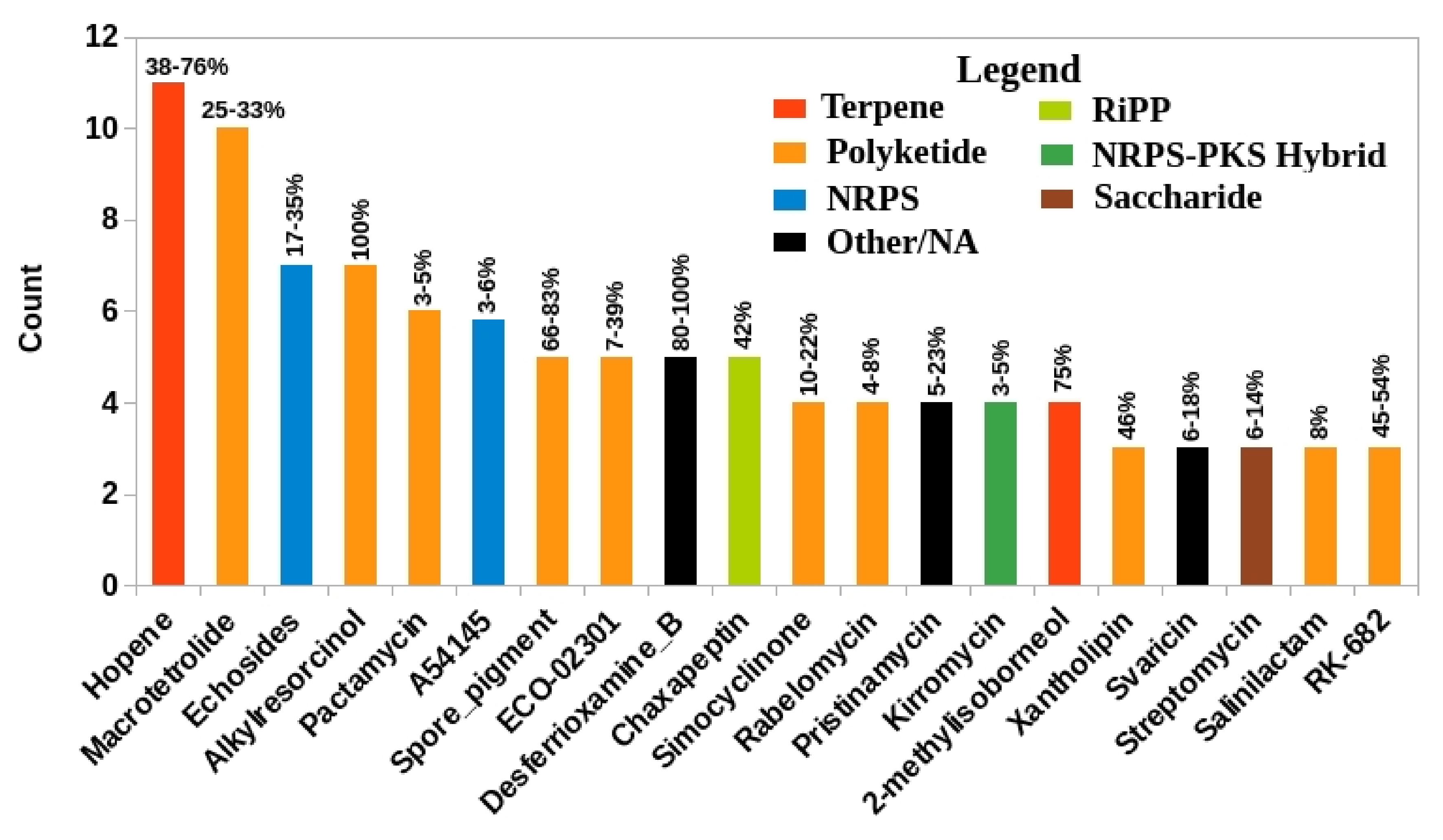
3.5.7. Distribution of Known BGCs in Streptomycetaceae
3.6. Diversity of Carbohydrate-Active Enzymes
3.6.1. Cellulose Degrading CAZymes or Cellulases
3.6.2. Hemi-Cellulose Degrading CAZymes
3.6.3. Chitin Degrading CAZymes
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hodgson, D.A. Primary metabolism and its control in streptomycetes: A most unusual group of bacteria. Adv. Microb. Physiol. 2000, 42, 47–238. [Google Scholar] [PubMed]
- Girard, G.; Willemse, J.; Zhu, H.; Claessen, D.; Bukarasam, K.; Goodfellow, M.; van Wezel, G.P. Analysis of novel kitasatosporae reveals significant evolutionary changes in conserved developmental genes between Kitasatospora and Streptomyces. Antonie Van Leeuwenhoek 2014, 106, 365–380. [Google Scholar] [CrossRef] [PubMed]
- Bentley, S.D.; Chater, K.F.; Cerdeno-Tarraga, A.M.; Challis, G.L.; Thomson, N.R.; James, K.D.; Harris, D.E.; Quail, M.A.; Kieser, H.; Harper, D.; et al. Complete genome sequence of the model actinomycete Streptomyces coelicolor A3(2). Nature 2002, 417, 141–147. [Google Scholar] [CrossRef] [PubMed]
- Roh, S.G.; Kim, M.K.; Park, S.; Yun, B.R.; Park, J.; Kim, M.J.; Kim, Y.S.; Kim, S.B. Streptacidiphilus pinicola sp. nov., isolated from pine grove soil. Int. J. Syst. Evol. Microbiol. 2018, 68, 3149–3155. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.B.; Lonsdale, J.; Seong, C.N.; Goodfellow, M. Streptacidiphilus gen. nov., acidophilic actinomycetes with wall chemotype I and emendation of the family Streptomycetaceae (Waksman and Henrici (1943) AL) emend. Rainey et al. 1997. Antonie Van Leeuwenhoek 2003, 83, 107–116. [Google Scholar] [CrossRef]
- Takahashi, Y. Genus Kitasatospora, taxonomic features and diversity of secondary metabolites. J. Antibiot. 2017, 70, 506–513. [Google Scholar] [CrossRef]
- Lyons, A.J.; Pridham, T.G. Streptomyces griseus (Krainsky) Waksman and Henrici: A Taxonomic Study of Some Strains; United States Department of Agriculture Economic Research Service: Washington, DC, USA, 1966; Available online: http://ageconsearch.umn.edu/record/171447/files/tb1360.pdf (accessed on 17 July 2020).
- Burg, R.W.; Miller, B.M.; Baker, E.E.; Birnbaum, J.; Currie, S.A.; Hartman, R.; Kong, Y.L.; Monaghan, R.L.; Olson, G.; Putter, I.; et al. Avermectins, new family of potent anthelmintic agents: Producing organism and fermentation. Antimicrob. Agents Chemother. 1979, 15, 361–367. [Google Scholar] [CrossRef]
- Habib el, S.E.; Scarsdale, J.N.; Reynolds, K.A. Biosynthetic origin of hygromycin A. Antimicrob. Agents Chemother. 2003, 47, 2065–2071. [Google Scholar] [CrossRef]
- Omura, S.; Otoguro, K.; Nishikiori, T.; Oiwa, R.; Iwai, Y. Setamycin, a new antibiotic. J. Antibiot. 1981, 34, 1253–1256. [Google Scholar] [CrossRef]
- Takahashi, Y.; Iwai, Y.; S, O. Two new species of the genus Kitasatosporia, Kitasatosporia phosalacinea sp. nov. and Kitasatosporia griseola sp. nov. J. Gen. Appl. Microbiol. 1984, 30, 377–387. [Google Scholar] [CrossRef]
- Omura, S.; Murata, M.; Hanaki, H.; Hinotozawa, K.; Oiwa, R.; Tanaka, H. Phosalacine, a new herbicidal antibiotic containing phosphinothricin. Fermentation, isolation, biological activity and mechanism of action. J. Antibiot. 1984, 37, 829–835. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; van Wezel, G.P.; Hae Choi, Y. Identification of novel endophenaside antibiotics produced by Kitasatospora sp. MBT66. J. Antibiot. 2015, 68, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Book, A.J.; Lewin, G.R.; McDonald, B.R.; Takasuka, T.E.; Wendt-Pienkowski, E.; Doering, D.T.; Suh, S.; Raffa, K.F.; Fox, B.G.; Currie, C.R. Evolution of high cellulolytic activity in symbiotic Streptomyces through selection of expanded gene content and coordinated gene expression. PLoS Biol. 2016, 14, e1002475. [Google Scholar] [CrossRef] [PubMed]
- Lacombe-Harvey, M.E.; Brzezinski, R.; Beaulieu, C. Chitinolytic functions in actinobacteria: Ecology, enzymes, and evolution. Appl. Microbiol. Biotechnol. 2018, 102, 7219–7230. [Google Scholar] [CrossRef]
- Sawaguchi, A.; Ono, S.; Oomura, M.; Inami, K.; Kumeta, Y.; Honda, K.; Sameshima-Saito, R.; Sakamoto, K.; Ando, A.; Saito, A. Chitosan degradation and associated changes in bacterial community structures in two contrasting soils. Soil Sci. Plant Nutr. 2015, 61, 471–480. [Google Scholar] [CrossRef]
- Zitouni, M.; Viens, P.; Ghinet, M.G.; Brzezinski, R. Diversity of family GH46 chitosanases in Kitasatospora setae KM-6054. Appl. Microbiol. Biotechnol. 2017, 101, 7877–7888. [Google Scholar] [CrossRef]
- Hwang, S.; Yun, Y.; Choi, W.H.; Kim, S.B.; Shin, J.; Lee, M.J.; Oh, D.C. Acidiphilamides A-E, modified peptides as autophagy inhibitors from an acidophilic actinobacterium, Streptacidiphilus rugosus. J. Nat. Prod. 2019, 82, 341–348. [Google Scholar] [CrossRef]
- Newman, D. Screening and identification of novel biologically active natural compounds. F1000Research 2017, 6, 783. [Google Scholar] [CrossRef]
- Omura, S.; Ikeda, H.; Ishikawa, J.; Hanamoto, A.; Takahashi, C.; Shinose, M.; Takahashi, Y.; Horikawa, H.; Nakazawa, H.; Osonoe, T.; et al. Genome sequence of an industrial microorganism Streptomyces avermitilis: Deducing the ability of producing secondary metabolites. Proc. Natl. Acad. Sci. USA 2001, 98, 12215–12220. [Google Scholar] [CrossRef]
- Udwary, D.W.; Zeigler, L.; Asolkar, R.N.; Singan, V.; Lapidus, A.; Fenical, W.; Jensen, P.R.; Moore, B.S. Genome sequencing reveals complex secondary metabolome in the marine actinomycete Salinispora tropica. Proc. Natl. Acad. Sci. USA 2007, 104, 10376–10381. [Google Scholar] [CrossRef]
- Cho, S.H.; Han, J.H.; Seong, C.N.; Kim, S.B. Phylogenetic diversity of acidophilic sporoactinobacteria isolated from various soils. J. Microbiol. 2006, 44, 600–606. [Google Scholar] [PubMed]
- Cho, S.H.; Han, J.H.; Ko, H.Y.; Kim, S.B. Streptacidiphilus anmyonensis sp. nov., Streptacidiphilus rugosus sp. nov. and Streptacidiphilus melanogenes sp. nov., acidophilic actinobacteria isolated from Pinus soils. Int. J. Syst. Evol. Microbiol. 2008, 58, 1566–1570. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.T.; Khan, M.R. Antibiotics—A soil microbiologist’s viewpoint. Postepy Hig. I Med. Dosw. 1974, 28, 395–408. [Google Scholar]
- Williams, S.T.; Robinson, C.S. The role of streptomycetes in decomposition of chitin in acidic soils. J. Gen. Microbiol. 1981, 127, 55–63. [Google Scholar] [CrossRef]
- Iuv, A.; Zenova, G.M. Antagonistic activity of soil acidophilic actinomycetes. Izv. Akad. Nauk. Seriia Biol. 2007, 4, 402–405. [Google Scholar]
- Poomthongdee, N.; Duangmal, K.; Pathom-aree, W. Acidophilic actinomycetes from rhizosphere soil: Diversity and properties beneficial to plants. J. Antibiot. 2015, 68, 106–114. [Google Scholar] [CrossRef]
- Kim, Y.R.; Park, S.; Kim, T.S.; Kim, M.K.; Han, J.H.; Joung, Y.; Kim, S.B. Draft genome sequence of Streptacidiphilus oryzae TH49T, an acidophilic actinobacterium isolated from soil. Genome Announc. 2015, 3. [Google Scholar] [CrossRef]
- Komaki, H.; Ichikawa, N.; Oguchi, A.; Hamada, M.; Tamura, T.; Fujita, N. Draft genome sequences of six type strains of the genus Streptacidiphilus. Genome Announc. 2015, 3. [Google Scholar] [CrossRef]
- Arthur, R.A.; Gulvik, C.A.; Humrighouse, B.W.; Lasker, B.A.; Batra, D.; Rowe, L.A.; Igual, J.M.; Nouioui, I.; Klenk, H.P.; McQuiston, J.R. Complete genome sequence of Streptacidiphilus sp. strain 15-057A, obtained from bronchial lavage fluid. Microbiol. Resour. Announc. 2018, 7. [Google Scholar] [CrossRef]
- Eid, J.; Fehr, A.; Gray, J.; Luong, K.; Lyle, J.; Otto, G.; Peluso, P.; Rank, D.; Baybayan, P.; Bettman, B.; et al. Real-time DNA sequencing from single polymerase molecules. Science 2009, 323, 133–138. [Google Scholar] [CrossRef]
- Chin, C.S.; Alexander, D.H.; Marks, P.; Klammer, A.A.; Drake, J.; Heiner, C.; Clum, A.; Copeland, A.; Huddleston, J.; Eichler, E.E.; et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat. Methods 2013, 10, 563–569. [Google Scholar] [CrossRef] [PubMed]
- Hyatt, D.; Chen, G.L.; Locascio, P.F.; Land, M.L.; Larimer, F.W.; Hauser, L.J. Prodigal: Prokaryotic gene recognition and translation initiation site identification. BMC Bioinform. 2010, 11, 119. [Google Scholar] [CrossRef] [PubMed]
- Huerta-Cepas, J.; Forslund, K.; Coelho, L.P.; Szklarczyk, D.; Jensen, L.J.; von Mering, C.; Bork, P. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Bio. Evol. 2017, 34, 2115–2122. [Google Scholar] [CrossRef] [PubMed]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.C.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef]
- Blin, K.; Wolf, T.; Chevrette, M.G.; Lu, X.; Schwalen, C.J.; Kautsar, S.A.; Suarez Duran, H.G.; de Los Santos, E.L.C.; Kim, H.U.; Nave, M.; et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res. 2017, 45, W36–W41. [Google Scholar] [CrossRef]
- Zhang, H.; Yohe, T.; Huang, L.; Entwistle, S.; Wu, P.; Yang, Z.; Busk, P.K.; Xu, Y.; Yin, Y. dbCAN2: A meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 2018, 46, W95–W101. [Google Scholar] [CrossRef]
- Lombard, V.; Golaconda Ramulu, H.; Drula, E.; Coutinho, P.M.; Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 2014, 42, D490–D495. [Google Scholar] [CrossRef]
- Gerlt, J.A.; Bouvier, J.T.; Davidson, D.B.; Imker, H.J.; Sadkhin, B.; Slater, D.R.; Whalen, K.L. Enzyme Function Initiative-Enzyme Similarity Tool (EFI-EST): A web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 2015, 1854, 1019–1037. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.H.; Ha, S.M.; Lim, J.; Kwon, S.; Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie Leeuwenhoek 2017, 110, 1281–1286. [Google Scholar] [CrossRef] [PubMed]
- Chaudhari, N.M.; Gupta, V.K.; Dutta, C. BPGA—An ultra-fast pan-genome analysis pipeline. Sci. Rep. 2016, 6, 24373. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Jia, X.; Yang, J.; Ling, Y.; Zhang, Z.; Yu, J.; Wu, J.; Xiao, J. PanGP: A tool for quickly analyzing bacterial pan-genome profile. Bioinformatics 2014, 30, 1297–1299. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Kong, F.; Zhou, S.; Huang, D.; Zheng, J.; Zhu, W. Streptomyces tirandamycinicus sp. nov., a novel marine sponge-derived actinobacterium with antibacterial potential against Streptococcus agalactiae. Front. Microbiol. 2019, 10, 482. [Google Scholar] [CrossRef] [PubMed]
- Kuo, C.H.; Ochman, H. The extinction dynamics of bacterial pseudogenes. PLoS Genet. 2010, 6, e1001050. [Google Scholar] [CrossRef] [PubMed]
- Labeda, D.P.; Dunlap, C.A.; Rong, X.; Huang, Y.; Doroghazi, J.R.; Ju, K.S.; Metcalf, W.W. Phylogenetic relationships in the family Streptomycetaceae using multi-locus sequence analysis. Antonie Van Leeuwenhoek. 2017, 110, 563–583. [Google Scholar] [CrossRef]
- Nouioui, I.; Klenk, H.-P.; Igual, J.M.; Gulvik, C.A.; Lasker, B.A.; McQuiston, J.R. Streptacidiphilus bronchialis sp. nov., a ciprofloxacin-resistant bacterium from a human clinical specimen; reclassification of Streptomyces griseoplanus as Streptacidiphilus griseoplanus comb. nov. and emended description of the genus Streptacidiphilus. Int. J. Syst. Evol. Microbiol. 2019, 69, 1047–1056. [Google Scholar] [CrossRef]
- Huang, Y.; Cui, Q.; Wang, L.; Rodriguez, C.; Quintana, E.; Goodfellow, M.; Liu, Z. Streptacidiphilus jiangxiensis sp. nov., a novel actinomycete isolated from acidic rhizosphere soil in China. Antonie Van Leeuwenhoek 2004, 86, 159–165. [Google Scholar] [CrossRef]
- Wang, L.; Huang, Y.; Liu, Z.; Goodfellow, M.; Rodríguez, C. Streptacidiphilus oryzae sp. nov., an actinomycete isolated from rice-field soil in Thailand. Int. J. Syst. Evol. Microbiol. 2006, 56, 1257–1261. [Google Scholar] [CrossRef]
- Nouioui, I.; Carro, L.; García-López, M.; Meier-Kolthoff, J.P.; Woyke, T.; Kyrpides, N.C.; Pukall, R.; Klenk, H.P.; Goodfellow, M.; Göker, M. Streptomyces albus (Rossi Doria 1891) Waksman and Henrici 1943 emend. Genome-Based Taxonomic Classification of the Phylum Actinobacteria. Front. Microbiol. 2018, 9, 2007. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.B.; Goodfellow, M. Streptomyces avermitilis sp. nov., nom. rev., a taxonomic home for the avermectin-producing streptomycetes. Int. J. Syst. Evol. Microbiol. 2002, 52, 2011–2014. [Google Scholar] [CrossRef] [PubMed]
- Hodgson, D.A. Carbohydrate Utilisation in Streptomyces coelicolor A3(2). Ph.D. Thesis, University of East Anglia, Norwich, UK, 1980. Available online: https://ethos.bl.uk/OrderDetails.do?uin=uk.bl.ethos.258081 (accessed on 17 July 2020).
- Zhang, Z.; Wang, Y.; Ruan, J. A proposal to revive the genus Kitasatospora (Omura, Takahashi, Iwai, and Tanaka 1982). Int. J. Syst. Bacteriol. 1997, 47, 1048–1054. [Google Scholar] [CrossRef] [PubMed]
- Labeda, D.P. Kitasatosporia mediocidica sp. nov. Int. J. Syst. Bacteriol. 1988, 38, 287–290. [Google Scholar] [CrossRef]
- Omura, S.; Takahashi, Y.; Iwai, Y.; Tanaka, H. Kitasatosporia, a new genus of the order Actinomycetales. J. Antibiot. 1982, 35, 1013–1019. [Google Scholar] [CrossRef]
- Vigil Stenman, C.T. Effects and Dynamics of Insertion Sequences in the Evolution of Cyanobacteria. Ph.D. Thesis, Stockholm University, Stockholm, Sweden, 2015. [Google Scholar]
- Vigil-Stenman, T.; Ininbergs, K.; Bergman, B.; Ekman, M. High abundance and expression of transposases in bacteria from the Baltic Sea. ISME J. 2017, 11, 2611–2623. [Google Scholar] [CrossRef]
- Zhang, M.M.; Wong, F.T.; Wang, Y.; Luo, S.; Lim, Y.H.; Heng, E.; Yeo, W.L.; Cobb, R.E.; Enghiad, B.; Ang, E.L.; et al. CRISPR-Cas9 strategy for activation of silent Streptomyces biosynthetic gene clusters. Nat. Chem. Biol. 2017. [Google Scholar] [CrossRef]
- Tao, W.; Yang, A.; Deng, Z.; Sun, Y. CRISPR/Cas9-based editing of Streptomyces for discovery, characterization, and production of natural products. Front. Microbiol. 2018, 9, 1660. [Google Scholar] [CrossRef]
- Wilkens, S. Structure and mechanism of ABC transporters. F1000Prime Rep. 2015, 7, 14. [Google Scholar] [CrossRef]
- Kim, J.N.; Kim, Y.; Jeong, Y.; Roe, J.H.; Kim, B.G.; Cho, B.K. Comparative genomics reveals the core and accessory genomes of Streptomyces species. J. Microbiol. Biotechnol. 2015, 25, 1599–1605. [Google Scholar] [CrossRef]
- Medini, D.; Donati, C.; Tettelin, H.; Masignani, V.; Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet. Dev. 2005, 15, 589–594. [Google Scholar] [CrossRef] [PubMed]
- De Maayer, P.; Chan, W.Y.; Rubagotti, E.; Venter, S.N.; Toth, I.K.; Birch, P.R.; Coutinho, T.A. Analysis of the Pantoea ananatis pan-genome reveals factors underlying its ability to colonize and interact with plant, insect and vertebrate hosts. BMC Genom. 2014, 15, 404. [Google Scholar] [CrossRef] [PubMed]
- Callister, S.J.; McCue, L.A.; Turse, J.E.; Monroe, M.E.; Auberry, K.J.; Smith, R.D.; Adkins, J.N.; Lipton, M.S. Comparative bacterial proteomics: Analysis of the core genome concept. PLoS ONE 2008, 3, e1542. [Google Scholar] [CrossRef] [PubMed]
- Bibb, M.J. Regulation of secondary metabolism in streptomycetes. Curr. Opin. Microbiol. 2005, 8, 208–215. [Google Scholar] [CrossRef] [PubMed]
- Méndez, C.; Salas, J.A. The role of ABC transporters in antibiotic-producing organisms: Drug secretion and resistance mechanisms. Res. Microbiol. 2001, 152, 341–350. [Google Scholar] [CrossRef]
- Campbell, J.W.; Cronan, J.E., Jr. Bacterial fatty acid biosynthesis: Targets for antibacterial drug discovery. Annu. Rev. Microbiol. 2001, 55, 305–332. [Google Scholar] [CrossRef]
- Manuse, S.; Fleurie, A.; Zucchini, L.; Lesterlin, C.; Grangeasse, C. Role of eukaryotic-like serine/threonine kinases in bacterial cell division and morphogenesis. FEMS Microbiol. Rev. 2016, 40, 41–56. [Google Scholar] [CrossRef] [PubMed]
- Petříčková, K.; Petříček, M. Eukaryotic-type protein kinases in Streptomyces coelicolor: Variations on a common theme. Microbiology 2003, 149, 1609–1621. [Google Scholar] [CrossRef]
- Claessen, D.; de Jong, W.; Dijkhuizen, L.; Wosten, H.A. Regulation of Streptomyces development: Reach for the sky! Trends Microbiol. 2006, 14, 313–319. [Google Scholar] [CrossRef]
- Den Hengst, C.D.; Tran, N.T.; Bibb, M.J.; Chandra, G.; Leskiw, B.K.; Buttner, M.J. Genes essential for morphological development and antibiotic production in Streptomyces coelicolor are targets of BldD during vegetative growth. Mol. Microbiol. 2010, 78, 361–379. [Google Scholar] [CrossRef]
- El-Gebali, S.; Mistry, J.; Bateman, A.; Eddy, S.R.; Luciani, A.; Potter, S.C.; Qureshi, M.; Richardson, L.J.; Salazar, G.A.; Smart, A.; et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019, 47, D427–D432. [Google Scholar] [CrossRef] [PubMed]
- Chater, K.F.; Chandra, G. The evolution of development in Streptomyces analysed by genome comparisons. FEMS Microbiol. Rev. 2006, 30, 651–672. [Google Scholar] [CrossRef] [PubMed]
- Flardh, K.; Buttner, M.J. Streptomyces morphogenetics: Dissecting differentiation in a filamentous bacterium. Nat. Rev. Microbiol. 2009, 7, 36–49. [Google Scholar] [CrossRef] [PubMed]
- McCormick, J.R.; Flardh, K. Signals and regulators that govern Streptomyces development. FEMS Microbiol. Rev. 2012, 36, 206–231. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Li, N.; Kudo, H.; Zhang, Z.; Li, J.; Wang, L.; Zhang, W.; Takechi, K.; Takano, H. Genes sufficient for synthesizing peptidoglycan are retained in Gymnosperm genomes, and MurE from Larix gmelinii can rescue the albino phenotype of Arabidopsis MurE mutation. Plant Cell Physiol. 2017, 58, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Richaud, C.; Higgins, W.; Mengin-Lecreulx, D.; Stragier, P. Molecular cloning, characterization, and chromosomal localization of dapF, the Escherichia coli gene for diaminopimelate epimerase. J. Bacteriol. 1987, 169, 1454–1459. [Google Scholar] [CrossRef]
- Ichikawa, N.; Oguchi, A.; Ikeda, H.; Ishikawa, J.; Kitani, S.; Watanabe, Y.; Nakamura, S.; Katano, Y.; Kishi, E.; Sasagawa, M.; et al. Genome sequence of Kitasatospora setae NBRC 14216T: An evolutionary snapshot of the family Streptomycetaceae. DNA Res. 2010, 17, 393–406. [Google Scholar] [CrossRef]
- Hwang, J.Y.; Kim, S.H.; Oh, H.R.; Kwon, E.; Nam, D.H. Analysis of a draft genome sequence of Kitasatospora cheerisanensis KCTC 2395 producing bafilomycin antibiotics. J. Microbiol. 2015, 53, 84–89. [Google Scholar] [CrossRef]
- Blin, K.; Pascal Andreu, V.; de Los Santos, E.L.C.; Del Carratore, F.; Lee, S.Y.; Medema, M.H.; Weber, T. The antiSMASH database version 2: A comprehensive resource on secondary metabolite biosynthetic gene clusters. Nucleic Acids Res. 2019, 47, D625–D630. [Google Scholar] [CrossRef]
- Rascher, A.; Hu, Z.; Viswanathan, N.; Schirmer, A.; Reid, R.; Nierman, W.C.; Lewis, M.; Hutchinson, C.R. Cloning and characterization of a gene cluster for geldanamycin production in Streptomyces hygroscopicus NRRL 3602. FEMS Microbiol. Lett. 2003, 218, 223–230. [Google Scholar] [CrossRef]
- Kautsar, S.A.; Blin, K.; Shaw, S.; Navarro-Muñoz, J.C.; Terlouw, B.R.; van der Hooft, J.J.J.; van Santen, J.A.; Tracanna, V.; Suarez Duran, H.G.; Pascal Andreu, V.; et al. MIBiG 2.0: A repository for biosynthetic gene clusters of known function. Nucleic Acids Res. 2020, 48, D454–D458. [Google Scholar] [CrossRef] [PubMed]
- McAlpine, J.B.; Bachmann, B.O.; Piraee, M.; Tremblay, S.; Alarco, A.M.; Zazopoulos, E.; Farnet, C.M. Microbial genomics as a guide to drug discovery and structural elucidation: ECO-02301, a novel antifungal agent, as an example. J. Nat. Prod. 2005, 68, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Wyatt, M.A.; Ahilan, Y.; Argyropoulos, P.; Boddy, C.N.; Magarvey, N.A.; Harrison, P.H. Biosynthesis of ebelactone A: Isotopic tracer, advanced precursor and genetic studies reveal a thioesterase-independent cyclization to give a polyketide β-lactone. J. Antibiot. 2013, 66, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Rounge, T.B.; Rohrlack, T.; Tooming-Klunderud, A.; Kristensen, T.; Jakobsen, K.S. Comparison of cyanopeptolin genes in Planktothrix, Microcystis, and Anabaena strains: Evidence for independent evolution within each genus. Appl. Environ. Microbiol. 2007, 73, 7322–7330. [Google Scholar] [CrossRef]
- Kodani, S.; Bicz, J.; Song, L.; Deeth, R.J.; Ohnishi-Kameyama, M.; Yoshida, M.; Ochi, K.; Challis, G.L. Structure and biosynthesis of scabichelin, a novel tris-hydroxamate siderophore produced by the plant pathogen Streptomyces scabies 87.22. Org. Biomol. Chem. 2013, 11, 4686–4694. [Google Scholar] [CrossRef]
- Izawa, M.; Kawasaki, T.; Hayakawa, Y. Cloning and heterologous expression of the thioviridamide biosynthesis gene cluster from Streptomyces olivoviridis. Appl. Environ. Microbiol. 2013, 79, 7110–7113. [Google Scholar] [CrossRef]
- Santos, C.L.; Correia-Neves, M.; Moradas-Ferreira, P.; Mendes, M.V. A walk into the LuxR regulators of Actinobacteria: Phylogenomic distribution and functional diversity. PLoS ONE 2012, 7, e46758. [Google Scholar] [CrossRef]
- Seipke, R.F.; Hutchings, M.I. The regulation and biosynthesis of antimycins. Beilstein J. Org. Chem. 2013, 9, 2556–2563. [Google Scholar] [CrossRef]
- Perez, M.; Schleissner, C.; Fernandez, R.; Rodriguez, P.; Reyes, F.; Zuniga, P.; de la Calle, F.; Cuevas, C. PM100117 and PM100118, new antitumor macrolides produced by a marine Streptomyces caniferus GUA-06-05-006A. J. Antibiot. 2016, 69, 388–394. [Google Scholar] [CrossRef]
- Zhang, Q.; Yu, Y.; Velasquez, J.E.; van der Donk, W.A. Evolution of lanthipeptide synthetases. Proc. Natl. Acad. Sci. USA 2012, 109, 18361–18366. [Google Scholar] [CrossRef]
- Repka, L.M.; Chekan, J.R.; Nair, S.K.; van der Donk, W.A. Mechanistic understanding of lanthipeptide biosynthetic enzymes. Chem. Rev. 2017, 117, 5457–5520. [Google Scholar] [CrossRef] [PubMed]
- Oldfield, E.; Lin, F.Y. Terpene biosynthesis: Modularity rules. Angew. Chem. Int. Ed. Engl. 2012, 51, 1124–1137. [Google Scholar] [CrossRef] [PubMed]
- Hay, R. Investigation of the Role of Gene Clusters in Terpene Biosynthesis in Sorghum Bicolor. Theses 2018. Available online: https://irl.umsl.edu/thesis/320 (accessed on 3 February 2020).
- Durr, C.; Schnell, H.J.; Luzhetskyy, A.; Murillo, R.; Weber, M.; Welzel, K.; Vente, A.; Bechthold, A. Biosynthesis of the terpene phenalinolactone in Streptomyces sp. Tu6071: Analysis of the gene cluster and generation of derivatives. Chem. Biol. 2006, 13, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Yamada, Y.; Kuzuyama, T.; Komatsu, M.; Shin-Ya, K.; Omura, S.; Cane, D.E.; Ikeda, H. Terpene synthases are widely distributed in bacteria. 2015. Proc. Natl. Acad. Sci. USA 2015, 112, 857–862. [Google Scholar] [CrossRef]
- Dairi, T.; Hamano, Y.; Kuzuyama, T.; Itoh, N.; Furihata, K.; Seto, H. Eubacterial diterpene cyclase genes essential for production of the isoprenoid antibiotic terpentecin. J. Bacteriol. 2001, 183, 6085–6094. [Google Scholar] [CrossRef]
- Hayashi, Y.; Matsuura, N.; Toshima, H.; Itoh, N.; Ishikawa, J.; Mikami, Y.; Dairi, T. Cloning of the gene cluster responsible for the biosynthesis of brasilicardin A, a unique diterpenoid. J. Antibiot. 2008, 61, 164–174. [Google Scholar] [CrossRef]
- Kannenberg, E.L.; Poralla, K. Hopanoid biosynthesis and function in bacteria. Naturwissenschaften 1999, 86, 168–176. [Google Scholar] [CrossRef]
- Ward, A.C.; Allenby, N.E. Genome mining for the search and discovery of bioactive compounds: The Streptomyces paradigm. FEMS Microbiol. Lett. 2018, 365. [Google Scholar] [CrossRef]
- Sadeghi, A.; Soltani, B.M.; Nekouei, M.K.; Jouzani, G.S.; Mirzaei, H.H.; Sadeghizadeh, M. Diversity of the ectoines biosynthesis genes in the salt tolerant Streptomyces and evidence for inductive effect of ectoines on their accumulation. Microbiol. Res. 2014, 169, 699–708. [Google Scholar] [CrossRef]
- Smith, W.C.; Xiang, L.; Shen, B. Genetic localization and molecular characterization of the nonS gene required for macrotetrolide biosynthesis in Streptomyces griseus DSM40695. Antimicrob. Agents Chemother. 2000, 44, 1809–1817. [Google Scholar] [CrossRef]
- Zhu, J.; Chen, W.; Li, Y.Y.; Deng, J.J.; Zhu, D.Y.; Duan, J.; Liu, Y.; Shi, G.Y.; Xie, C.; Wang, H.X.; et al. Identification and catalytic characterization of a nonribosomal peptide synthetase-like (NRPS-like) enzyme involved in the biosynthesis of echosides from Streptomyces sp. LZ35. Gene 2014, 546, 352–358. [Google Scholar] [CrossRef] [PubMed]
- Andre, I.; Potocki-Veronese, G.; Barbe, S.; Moulis, C.; Remaud-Simeon, M. CAZyme discovery and design for sweet dreams. Curr. Opin. Chem. Biol. 2014, 19, 17–24. [Google Scholar] [CrossRef] [PubMed]
- Adams, A.S.; Jordan, M.S.; Adams, S.M.; Suen, G.; Goodwin, L.A.; Davenport, K.W.; Currie, C.R.; Raffa, K.F. Cellulose-degrading bacteria associated with the invasive woodwasp Sirex noctilio. ISME J. 2011, 5, 1323–1331. [Google Scholar] [CrossRef] [PubMed]
- Lakhundi, S.; Siddiqui, R.; Khan, N.A. Cellulose degradation: A therapeutic strategy in the improved treatment of Acanthamoeba infections. Parasit. Vectors 2015, 8, 23. [Google Scholar] [CrossRef]
- Lopez-Mondejar, R.; Zuhlke, D.; Becher, D.; Riedel, K.; Baldrian, P. Cellulose and hemicellulose decomposition by forest soil bacteria proceeds by the action of structurally variable enzymatic systems. Sci. Rep. 2016, 6, 25279. [Google Scholar] [CrossRef]
- Koeck, D.E.; Pechtl, A.; Zverlov, V.V.; Schwarz, W.H. Genomics of cellulolytic bacteria. Curr. Opin. Biotechnol. 2014, 29, 171–183. [Google Scholar] [CrossRef]
- Swiontek Brzezinska, M.; Jankiewicz, U.; Burkowska, A.; Walczak, M. Chitinolytic microorganisms and their possible application in environmental protection. Curr. Microbiol. 2014, 68, 71–81. [Google Scholar] [CrossRef]
- Synstad, B.; Gaseidnes, S.; Van Aalten, D.M.; Vriend, G.; Nielsen, J.E.; Eijsink, V.G. Mutational and computational analysis of the role of conserved residues in the active site of a family 18 chitinase. Eur. J. Biochem. 2004, 271, 253–262. [Google Scholar] [CrossRef]
- Collin, M.; Olsén, A. Effect of SpeB and EndoS from Streptococcus pyogenes on human immunoglobulins. Infect. Immun. 2001, 69, 7187–7189. [Google Scholar] [CrossRef]
- Nakamura, T.; Mine, S.; Hagihara, Y.; Ishikawa, K.; Uegaki, K. Structure of the catalytic domain of the hyperthermophilic chitinase from Pyrococcus furiosus. Acta Cryst. Sect. F Struct. Biol. Cryst. Commun. 2007, 63, 7–11. [Google Scholar] [CrossRef]
- Manjeet, K.; Purushotham, P.; Neeraja, C.; Podile, A.R. Bacterial chitin binding proteins show differential substrate binding and synergy with chitinases. Microbiol. Res. 2013, 168, 461–468. [Google Scholar] [CrossRef] [PubMed]
| Species | BioProject Accession | Size (Mb) | No. of Contigs | GC Content (%) | CDS | Pseudogenes | tRNA (rRNA) | Accessory Genes | Unique Genes | No. of Biosynthetic Gene Clusters (BGCs) | Reference |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | PRJNA234786 | 9.91 | 1 | 71.8 | 8,166 | 366 | 84 (27) | 3,458 | 2,066 | 35 | [5] |
| 2 | PRJDB3297 | 9.38 | 153 | 72 | 7941 | 453 | 73 (3) | 4678 | 759 | 30 | [23] |
| 3 | PRJNA438532 | 7.09 | 1 | 72.70 | 5681 | 247 | 66 (24) | 1915 | 1665 | 19 | [49] |
| 4 | PRJDB3297 | 8.45 | 139 | 71.3 | 7062 | 454 | 72 (3) | 3591 | 1162 | 32 | [5] |
| 5 | PRJNA238534 | 9.15 | 144 | 71.5 | 7769 | 257 | 73 (19) | 3990 | 1376 | 18 | - |
| 6 | PRJDB3297 | 9.52 | 96 | 72 | 8285 | 76 | 69 (5) | 4545 | 1247 | 30 | [50] |
| 7 | PRJDB3297 | 8.77 | 107 | 71.9 | 7481 | 390 | 74 (3) | 4547 | 554 | 31 | [23] |
| 8 | PRJDB3297 | 8.41 | 184 | 71.9 | 7119 | 459 | 71 (3) | 4193 | 567 | 38 | [5] |
| 9 | PRJNA234788 | 7.81 | 6 | 73.4 | 6526 | 272 | 63 (23) | 2447 | 1884 | 15 | [51] |
| 10 | PRJNA475452 | 8.43 | 281 | 71.8 | 7440 | 311 | 68 (8) | 4137 | 846 | 35 | [4] |
| 11 | PRJNA234778 | 9 | 4 | 71.8 | 7750 | 356 | 74 (27) | 4101 | 1171 | 22 | [23] |
| 12 | PRJNA271625 | 8.38 | 1 | 72.60 | 7330 | 58 | 65 (18) | NA | NA | 35 | [52] |
| 13 | PRJNA189 | 9.11 | 2 | 70.68 | 7446 | 0 | 68 (18) | NA | NA | 37 | [53] |
| 14 | PRJNA242 | 9.05 | 1 | 72 | 8152 | 60 | 65 (18) | NA | NA | 29 | [54] |
| 15 | PRJNA234862 | 8.27 | 3 | 71.6 | 6844 | 292 | 74 (29) | NA | NA | 30 | [55] |
| 16 | PRJNA234781 | 8.68 | 7 | 71.9 | 7103 | 254 | 71 (33) | NA | NA | 32 | [56] |
| 17 | PRJDA19951 | 8.78 | 1 | 74.20 | 7182 | 0 | 74 (27) | NA | NA | 38 | [57] |
| Cluster Type/Species | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | 13 | 14 | 15 | 16 | 17 | Total |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Terpene | 3 | 4 | 3 | 6 | 5 | 6 | 4 | 4 | 8 | 5 | 5 | 2 | 7 | 5 | 3 | 3 | 5 | 78 |
| T1PKS | 0 | 6 | 0 | 8 | 1 | 1 | 6 | 11 | 0 | 3 | 0 | 3 | 4 | 1 | 2 | 5 | 1 | 52 |
| NRPS | 4 | 2 | 1 | 4 | 3 | 5 | 1 | 2 | 0 | 4 | 2 | 2 | 4 | 3 | 0 | 3 | 2 | 42 |
| Siderophore | 2 | 2 | 1 | 2 | 2 | 2 | 3 | 2 | 1 | 2 | 2 | 2 | 4 | 3 | 2 | 2 | 2 | 36 |
| Lantipeptide | 7 | 1 | 3 | 2 | 2 | 1 | 0 | 2 | 1 | 1 | 1 | 2 | 0 | 3 | 3 | 1 | 4 | 34 |
| Bacteriocin | 0 | 1 | 1 | 2 | 1 | 0 | 1 | 1 | 2 | 3 | 1 | 1 | 1 | 2 | 3 | 0 | 3 | 23 |
| Butyrolactone | 2 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 0 | 0 | 2 | 1 | 6 | 22 |
| T2PKS | 0 | 2 | 0 | 0 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 2 | 2 | 0 | 1 | 12 |
| T3PKS | 0 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 0 | 2 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 12 |
| Lassopeptide | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 1 | 3 | 0 | 0 | 0 | 2 | 11 |
| T1PKS_NRPS | 1 | 0 | 3 | 1 | 0 | 1 | 0 | 1 | 0 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 0 | 10 |
| Thiopeptide | 2 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 0 | 0 | 2 | 0 | 0 | 9 |
| T1PKS_OtherKS | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 1 | 2 | 1 | 1 | 1 | 8 |
| Butyrolactone_OtherKS | 0 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 6 |
| Other hybrids | 9 | 6 | 4 | 1 | 1 | 5 | 6 | 6 | 0 | 2 | 1 | 8 | 5 | 3 | 4 | 8 | 6 | 75 |
| Others | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 1 | 2 | 5 | 3 | 3 | 1 | 3 | 2 | 23 |
| Total | 35 | 30 | 19 | 32 | 18 | 30 | 31 | 38 | 15 | 35 | 22 | 35 | 37 | 29 | 30 | 32 | 38 | 506 |
| Known BGC | Organism Name | Cluster No. | Cluster Type | Similarity (%) | MIBiG Reference |
|---|---|---|---|---|---|
| Cacibiocin | S. pinicola KCTC 49008T | 10 | Aminocoumarin | 64 | BGC0001154 |
| Cyanopeptin | S. albus JL83T | 30 | NRPS | 50 | BGC0000331 |
| Eicoseicosapentaenoic acid | S. bronchialis DSM 106435T | 18 | Otherks | 10 | BGC0000865 |
| Erythrochelin | S. rugosus AM-16T | 18 | NRPS | 28 | BGC0000349 |
| Frenolicin | S. jiangxiensis NBRC 100920T | 9 | T2PKS | 50 | BGC0000225 |
| Lankacidin | S. pinicola KCTC 49008T | 25 | Other | 26 | BGC0001100 |
| Micromonolactam | S. neutrinimicus NBRC 100921T | 2 | T1PKS | 100 | BGC0000095 |
| Organism Name | Genes (%) | GH | GT | CE | PL | CBM | AA |
|---|---|---|---|---|---|---|---|
| S. albus JL83T | 366 (4.48) | 145 (46) | 84 (14) | 76 (7) | 9 (6) | 120 (16) | 15 (5) |
| S. anmyonensis NBRC 103185T | 339 (4.27) | 134 (42) | 79 (14) | 71 (7) | 4 (2) | 108 (15) | 24 (6) |
| S. bronchialis DSM 106435T | 268 (4.72) | 132 (49) | 56 (13) | 60 (8) | 4 (3) | 86 (20) | 11 (4) |
| S. carbonis NBRC 100919T | 422 (5.98) | 235 (69) | 77 (13) | 77 (8) | 9 (5) | 152 (26) | 15 (6) |
| S. jeojiense NRRL B-24555T | 440 (5.66) | 236 (60) | 93 (12) | 74 (8) | 17 (5) | 129 (21) | 22 (6) |
| S. jiangxiensis NBRC 100920T | 370 (4.47) | 171 (56) | 88 (14) | 70 (8) | 1 (1) | 100 (18) | 29 (6) |
| S. melanogenes NBRC 103184T | 329 (4.40) | 133 (43) | 80 (14) | 64 (8) | 4 (3) | 96 (16) | 27 (6) |
| S. neutrinimicus NBRC 100921T | 313 (4.40) | 130 (47) | 75 (14) | 62 (7) | 5 (4) | 83 (16) | 25 (6) |
| S. oryzae TH49T | 326 (5.00) | 154 (53) | 83 (15) | 64 (8) | 4 (4) | 48 (16) | 20 (5) |
| S. pinicola KCTC 49008T | 390 (5.24) | 190 (61) | 87 (13) | 59 (9) | 9 (7) | 131 (20) | 24 (5) |
| S. rugosus AM-16T | 334 (4.31) | 147 (47) | 81 (14) | 66 (8) | 4 (4) | 99 (16) | 25 (6) |
| Str. albus DSM 41398T | 262 (3.57) | 120 (48) | 55 (11) | 54 (8) | 3 (3) | 30 (12) | 21 (7) |
| Str. avermitilis MA-4680T | 347 (4.52) | 169 (55) | 75 (13) | 60 (9) | 12 (9) | 75 (18) | 25 (7) |
| Str. coelicolor A3(2) | 383 (4.70) | 187 (60) | 68 (14) | 71 (9) | 15 (11) | 98 (19) | 23 (7) |
| Streptomyces sp. sirexAA | 297 (4.67) | 137 (50) | 63 (13) | 63 (9) | 8 (5) | 80 (21) | 17 (5) |
| K. azatica KCTC 9699T | 364 (5.34) | 172 (56) | 70 (15) | 67 (8) | 2 (2) | 160 (21) | 18 (5) |
| K. mediocidica KCTC 9733T | 333 (4.69) | 128 (48) | 84 (14) | 71 (9) | 8 (2) | 96 (18) | 15 (4) |
| K. setae KM-6054T | 308 (4.29) | 122 (41) | 78 (13) | 62 (8) | 3 (2) | 100 (19) | 21 (5) |
| Enzyme Class | CAZy Family | Main Activity | S. albus JL83T | S. anmyonensis NBRC 103185T | S. carbonis NBRC 100919T | S. jeojiense NRRL B-24555T | S. jiangxiensis NBRC 100920T | S. melanogenes NBRC 103184T | S. neutrinimicus NBRC 100921T | S. oryzae TH49T | S. pinicola KCTC 49008T | S. rugosus AM-16T | S. bronchialis DSM 106435T | Str. albus DSM 41398T | Str. avermitilis MA-4680T | Str. coelicolor A3(2) | Streptomyces sp. SirexAA | K. azatica KCTC 9699T | K. mediocidica KCTC 9733T | K. setae KM-6054T |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cellulose degrading CAZymes | GH1 | β-Glucosidase, β-galactosidase | 4 | 3 | 5 | 3 | 3 | 6 | 3 | 3 | 4 | 6 | 4 | 3 | 3 | 5 | 6 | 5 | 2 | 4 |
| GH3 | β-Glucosidases, β-D-xylopyranosidase | 8 | 7 | 8 | 9 | 14 | 10 | 8 | 7 | 11 | 9 | 7 | 6 | 5 | 9 | 7 | 7 | 2 | 4 | |
| GH5 | Endo-β-1,4-glucanase, β-mannosidase | 2 | 5 | 5 | 11 | 5 | 7 | 5 | 6 | 1 | 9 | 5 | 1 | 3 | 5 | 3 | 3 | 6 | 2 | |
| GH6 | Cellobiohydrolase, endo-β-1,4-glucanase | 3 | 1 | 4 | 1 | 4 | 1 | 1 | 1 | 1 | 1 | 2 | 2 | 4 | 3 | 1 | 3 | 2 | 3 | |
| GH8 | Endo-β-1,4-glucanase | 1 | 2 | 0 | 0 | 1 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | |
| GH9 | Endo-β-1,4-glucanase | 2 | 0 | 2 | 1 | 1 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 2 | 1 | 0 | 1 | 2 | |
| GH12 | Endo-β-1,4-glucanase | 0 | 0 | 1 | 2 | 2 | 1 | 0 | 0 | 0 | 1 | 1 | 0 | 3 | 2 | 1 | 1 | 0 | 0 | |
| GH44 | Endoglucanase | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | |
| GH48 | Cellobiohydrolase | 1 | 0 | 1 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 1 | 1 | 1 | 1 | 1 | |
| GH74 | Endoglucanase | 1 | 3 | 1 | 4 | 4 | 2 | 2 | 2 | 3 | 3 | 4 | 0 | 4 | 3 | 1 | 2 | 1 | 1 | |
| Subtotal | 22 | 21 | 28 | 32 | 35 | 31 | 22 | 22 | 21 | 33 | 26 | 12 | 23 | 30 | 21 | 22 | 16 | 18 | ||
| Hemicellulose degrading CAZymes | GH2 | β-Galactosidase, β-glucuronidase | 1 | 1 | 2 | 4 | 4 | 1 | 1 | 0 | 6 | 2 | 2 | 4 | 6 | 7 | 4 | 1 | 1 | 0 |
| GH10 | Endo-β-1,4-xylanase | 0 | 0 | 3 | 3 | 1 | 1 | 0 | 2 | 3 | 2 | 1 | 0 | 2 | 2 | 1 | 3 | 2 | 3 | |
| GH11 | Endo-β-1,4-xylanase | 0 | 0 | 2 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 0 | 1 | |
| GH16 | Xyloglycosyltransferase | 10 | 5 | 3 | 8 | 9 | 5 | 8 | 4 | 3 | 6 | 7 | 4 | 3 | 5 | 6 | 9 | 10 | 7 | |
| GH26 | Endo-β-1,4-mannanase | 0 | 2 | 0 | 3 | 3 | 5 | 3 | 3 | 0 | 3 | 0 | 1 | 1 | 1 | 1 | 1 | 0 | 0 | |
| GH30 | Endo-β-1,4-xylanase | 0 | 3 | 3 | 3 | 2 | 2 | 3 | 2 | 1 | 2 | 0 | 2 | 3 | 1 | 1 | 2 | 1 | 1 | |
| GH31 | α-Glucosidases, α-xylosidase | 0 | 0 | 1 | 4 | 4 | 1 | 0 | 0 | 5 | 1 | 0 | 2 | 2 | 2 | 4 | 4 | 0 | 0 | |
| GH39 | α-L-iduronidase, β-xylosidase | 3 | 0 | 0 | 2 | 2 | 1 | 1 | 1 | 2 | 0 | 2 | 0 | 2 | 0 | 0 | 1 | 0 | 0 | |
| GH42 | β-Galactosidase | 0 | 2 | 2 | 6 | 4 | 5 | 2 | 1 | 2 | 6 | 2 | 0 | 4 | 2 | 3 | 2 | 1 | 0 | |
| GH43 | α-l-arabinofuranosidase, β-xylosidase | 1 | 1 | 3 | 4 | 5 | 1 | 0 | 0 | 4 | 2 | 3 | 2 | 6 | 6 | 6 | 6 | 1 | 2 | |
| GH53 | Endo-β-1,4-galactanase | 1 | 0 | 0 | 7 | 4 | 3 | 2 | 4 | 3 | 8 | 1 | 0 | 0 | 0 | 0 | 5 | 4 | 0 | |
| Subtotal | 16 | 14 | 19 | 45 | 38 | 25 | 20 | 17 | 30 | 32 | 18 | 15 | 29 | 28 | 27 | 35 | 20 | 14 | ||
| Chitin and chitosan degrading CAZymes | GH18 | Chitinase | 14 | 12 | 6 | 12 | 17 | 16 | 12 | 10 | 7 | 12 | 14 | 5 | 8 | 12 | 10 | 14 | 12 | 19 |
| GH19 | Chitinase | 1 | 2 | 2 | 1 | 2 | 2 | 1 | 1 | 0 | 1 | 1 | 1 | 0 | 2 | 3 | 0 | 1 | 2 | |
| GH20 | Exo-β-N-acetylglucosaminidase | 2 | 4 | 2 | 3 | 3 | 5 | 4 | 2 | 4 | 4 | 4 | 5 | 3 | 4 | 2 | 2 | 2 | 2 | |
| GH46 | Endo-β-1,4-chitosanase | 1 | 0 | 1 | 1 | 1 | 0 | 0 | 0 | 1 | 0 | 0 | 2 | 2 | 2 | 3 | 1 | 0 | 3 | |
| Subtotal | 18 | 18 | 11 | 17 | 23 | 23 | 17 | 13 | 12 | 17 | 19 | 13 | 13 | 20 | 18 | 17 | 15 | 26 | ||
| Oxidative enzymes | AA10 | Lytic polysaccharide monooxygenase (LPMO) | 0 | 0 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 4 | 7 | 6 | 0 | 0 | 5 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Malik, A.; Kim, Y.R.; Kim, S.B. Genome Mining of the Genus Streptacidiphilus for Biosynthetic and Biodegradation Potential. Genes 2020, 11, 1166. https://doi.org/10.3390/genes11101166
Malik A, Kim YR, Kim SB. Genome Mining of the Genus Streptacidiphilus for Biosynthetic and Biodegradation Potential. Genes. 2020; 11(10):1166. https://doi.org/10.3390/genes11101166
Chicago/Turabian StyleMalik, Adeel, Yu Ri Kim, and Seung Bum Kim. 2020. "Genome Mining of the Genus Streptacidiphilus for Biosynthetic and Biodegradation Potential" Genes 11, no. 10: 1166. https://doi.org/10.3390/genes11101166
APA StyleMalik, A., Kim, Y. R., & Kim, S. B. (2020). Genome Mining of the Genus Streptacidiphilus for Biosynthetic and Biodegradation Potential. Genes, 11(10), 1166. https://doi.org/10.3390/genes11101166






