Regulation of KIR3DL3 Expression via miRNA
Abstract
1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Bioinformatic Prediction of Potential miRNAs for KIR3DL3
2.3. Quantitative Real-Time Reverse Transcription Polymerase Chain Reaction (qRT-PCR)
2.4. Reporter Plasmid Construction and Luciferase Reporter Assay
2.5. Targeted miRNA Inhibition
2.6. Flow Cytometry
2.7. Statistical Analysis
3. Results
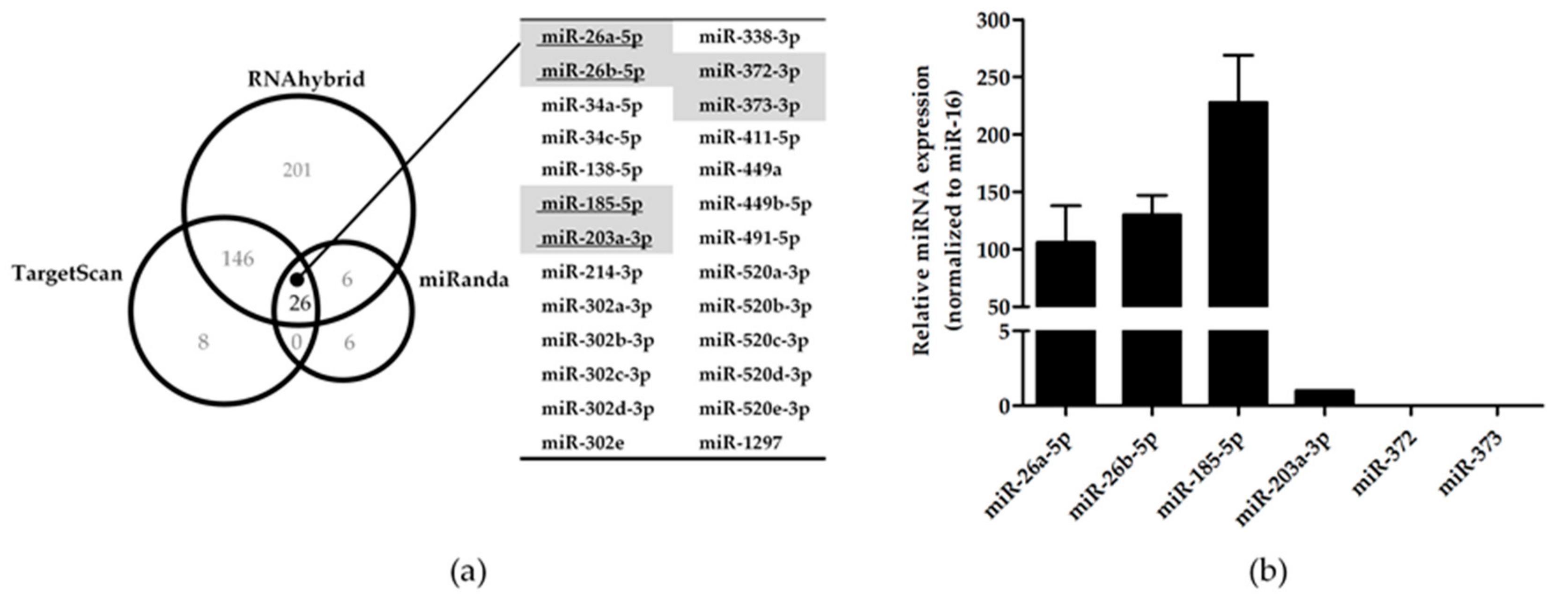
3.1. Bioinformatic Prediction of Potential miRNA Candidates for KIR3DL3
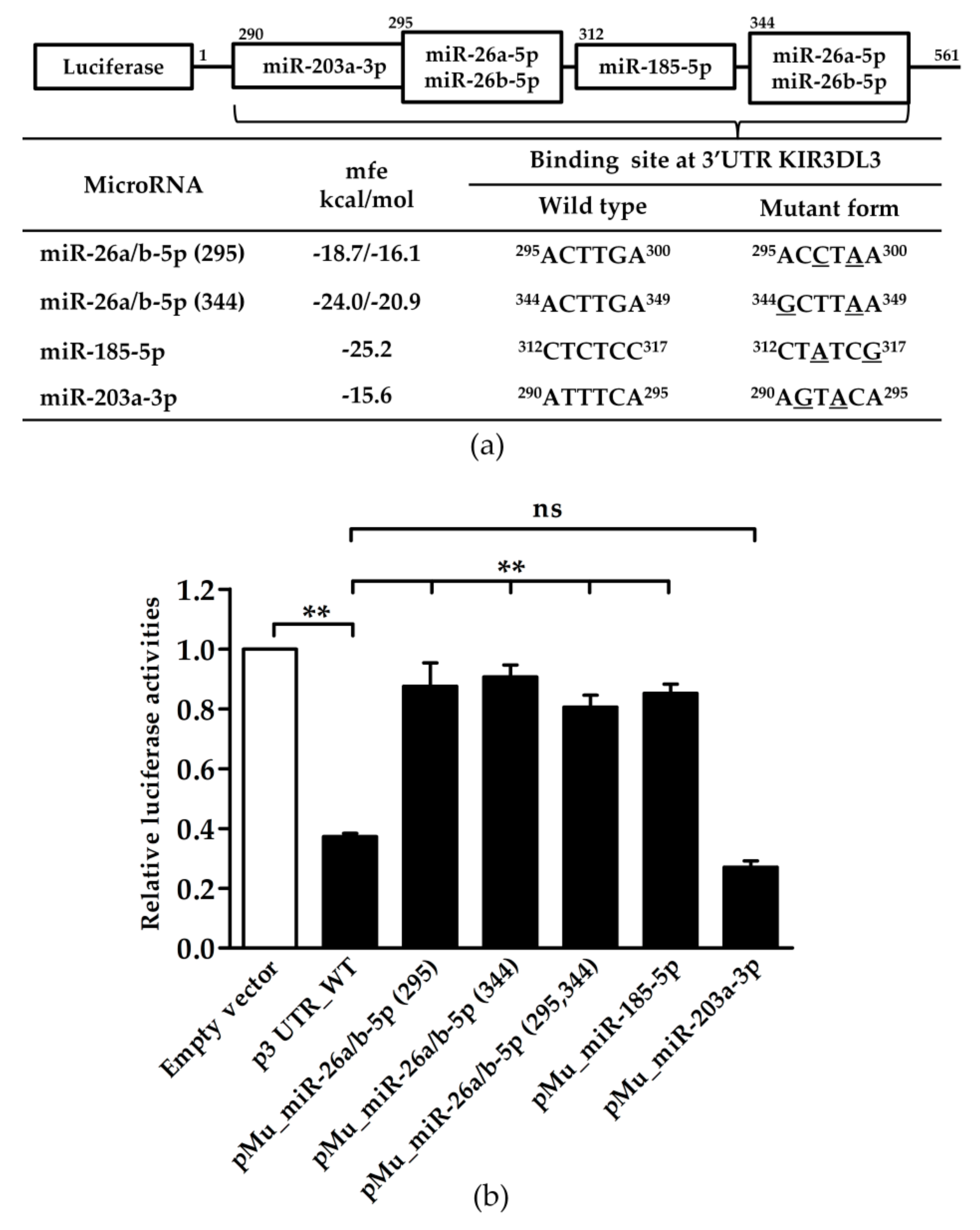
3.2. Potential miRNA Candidates Targeting KIR3DL3 Through the 3’UTR
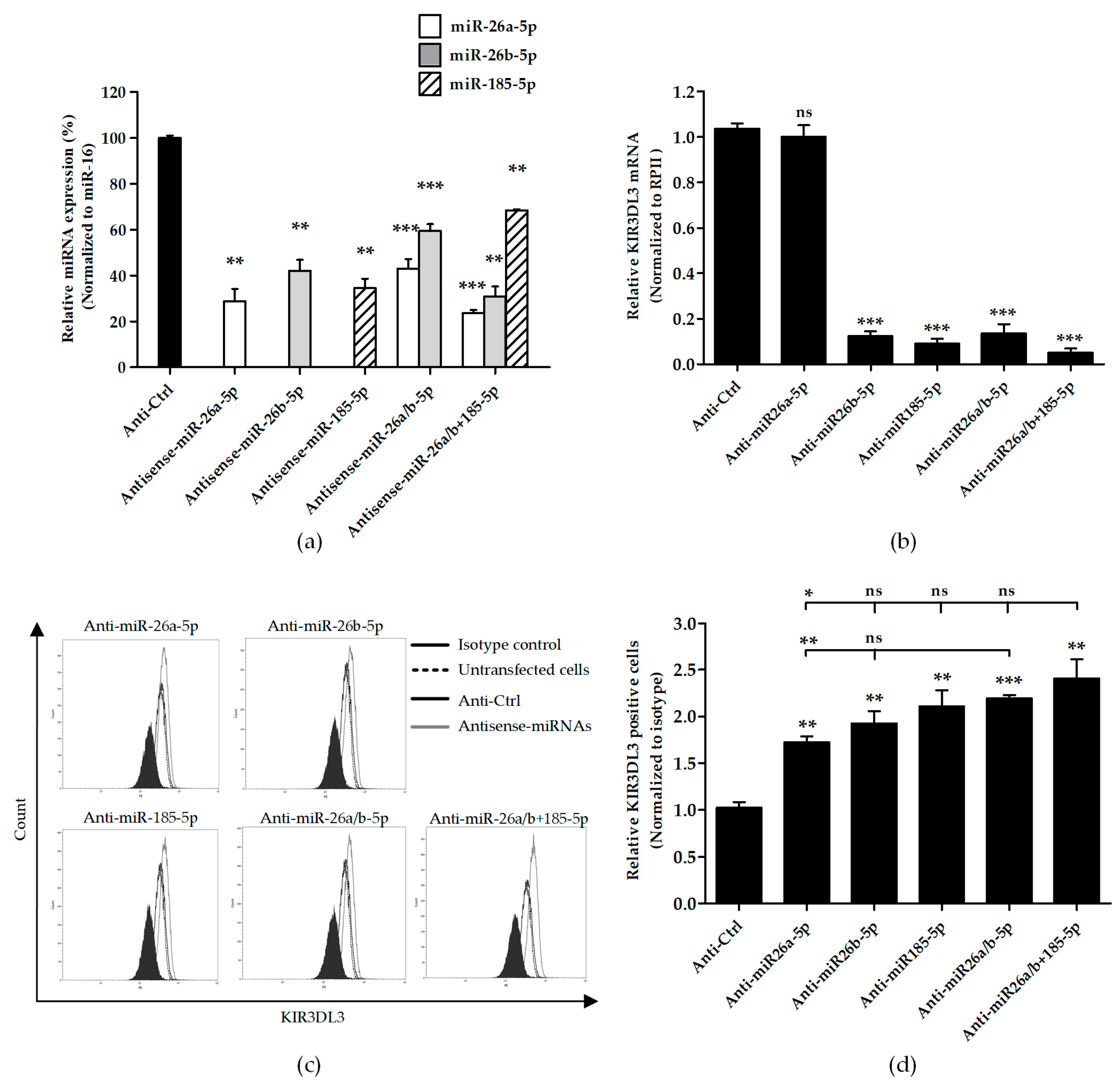
3.3. Knocking Down Candidate miRNAs Resulted in Higher Cell Surface KIR3DL3 Expression
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Vilches, C.; Parham, P. KIR: diverse, rapidly evolving receptors of innate and adaptive immunity. Annu. Rev. Immunol. 2002, 20, 217–251. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.H.; Chen, M.J.; Chen, H.F.; Lee, T.H.; Ho, H.N.; Yang, Y.S. Decreased expression of killer cell inhibitory receptors on natural killer cells in eutopic endometrium in women with adenomyosis. Hum. Reprod. 2004, 19, 1974–1978. [Google Scholar] [CrossRef] [PubMed]
- Ljunggren, H.G.; Karre, K. In search of the ‘missing self’: MHC molecules and NK cell recognition. Immunol. Today 1990, 11, 237–244. [Google Scholar] [CrossRef]
- Valiante, N.M.; Uhrberg, M.; Shilling, H.G.; Lienert-Weidenbach, K.; Arnett, K.L.; D’Andrea, A.; Phillips, J.H.; Lanier, L.L.; Parham, P. Functionally and structurally distinct NK cell receptor repertoires in the peripheral blood of two human donors. Immunity 1997, 7, 739–751. [Google Scholar] [CrossRef]
- Husain, Z.; Alper, C.A.; Yunis, E.J.; Dubey, D.P. Complex expression of natural killer receptor genes in single natural killer cells. Immunology 2002, 106, 373–380. [Google Scholar] [CrossRef] [PubMed]
- Hsu, K.C.; Chida, S.; Geraghty, D.E.; Dupont, B. The killer cell immunoglobulin-like receptor (KIR) genomic region: gene-order, haplotypes and allelic polymorphism. Immunol. Rev. 2002, 190, 40–52. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.J.; Torkar, M.; Haude, A.; Milne, S.; Jones, T.; Sheer, D.; Beck, S.; Trowsdale, J. Plasticity in the organization and sequences of human KIR/ILT gene families. Proc. Natl. Acad. Sci. USA 2000, 97, 4778–4783. [Google Scholar] [CrossRef] [PubMed]
- Rajagopalan, S.; Long, E.O. A human histocompatibility leukocyte antigen (HLA)-G-specific receptor expressed on all natural killer cells. J. Exp. Med. 1999, 189, 1093–1100. [Google Scholar] [CrossRef]
- Chan, H.W.; Miller, J.S.; Moore, M.B.; Lutz, C.T. Epigenetic control of highly homologous killer Ig-like receptor gene alleles. J. Immunol. 2005, 175, 5966–5974. [Google Scholar] [CrossRef]
- Santourlidis, S.; Trompeter, H.I.; Weinhold, S.; Eisermann, B.; Meyer, K.L.; Wernet, P.; Uhrberg, M. Crucial role of DNA methylation in determination of clonally distributed killer cell Ig-like receptor expression patterns in NK cells. J. Immunol. 2002, 169, 4253–4261. [Google Scholar] [CrossRef]
- Uhrberg, M. Shaping the human NK cell repertoire: an epigenetic glance at KIR gene regulation. Mol. Immunol. 2005, 42, 471–475. [Google Scholar] [CrossRef] [PubMed]
- Torkar, M.; Norgate, Z.; Colonna, M.; Trowsdale, J.; Wilson, M.J. Isotypic variation of novel immunoglobulin-like transcript/killer cell inhibitory receptor loci in the leukocyte receptor complex. Eur. J. Immunol. 1998, 28, 3959–3967. [Google Scholar] [CrossRef]
- Vilches, C.; Rajalingam, R.; Uhrberg, M.; Gardiner, C.M.; Young, N.T.; Parham, P. KIR2DL5, a novel killer-cell receptor with a D0-D2 configuration of Ig-like domains. J. Immunol. 2000, 164, 5797–5804. [Google Scholar] [CrossRef] [PubMed]
- Long, E.O.; Barber, D.F.; Burshtyn, D.N.; Faure, M.; Peterson, M.; Rajagopalan, S.; Renard, V.; Sandusky, M.; Stebbins, C.C.; Wagtmann, N.; et al. Inhibition of natural killer cell activation signals by killer cell immunoglobulin-like receptors (CD158). Immunol. Rev. 2001, 181, 223–233. [Google Scholar] [CrossRef] [PubMed]
- Leaton, L.A.; Shortt, J.; Kichula, K.M.; Tao, S.; Nemat-Gorgani, N.; Mentzer, A.J.; Oppenheimer, S.J.; Deng, Z.; Hollenbach, J.A.; Gignoux, C.R.; et al. Conservation, Extensive Heterozygosity, and Convergence of Signaling Potential All Indicate a Critical Role for KIR3DL3 in Higher Primates. Front. Immunol. 2019, 10, 24. [Google Scholar] [CrossRef] [PubMed]
- Trundley, A.E.; Hiby, S.E.; Chang, C.; Sharkey, A.M.; Santourlidis, S.; Uhrberg, M.; Trowsdale, J.; Moffett, A. Molecular characterization of KIR3DL3. Immunogenetics 2006, 57, 904–916. [Google Scholar] [CrossRef] [PubMed]
- Trompeter, H.I.; Gomez-Lozano, N.; Santourlidis, S.; Eisermann, B.; Wernet, P.; Vilches, C.; Uhrberg, M. Three structurally and functionally divergent kinds of promoters regulate expression of clonally distributed killer cell Ig-like receptors (KIR), of KIR2DL4, and of KIR3DL3. J. Immunol. 2005, 174, 4135–4143. [Google Scholar] [CrossRef] [PubMed]
- Pesce, S.; Squillario, M.; Greppi, M.; Loiacono, F.; Moretta, L.; Moretta, A.; Sivori, S.; Castagnola, P.; Barla, A.; Candiani, S.; et al. New miRNA Signature Heralds Human NK Cell Subsets at Different Maturation Steps: Involvement of miR-146a-5p in the Regulation of KIR Expression. Front. Immunol. 2018, 9, 2360. [Google Scholar] [CrossRef]
- Beaulieu, A.M.; Bezman, N.A.; Lee, J.E.; Matloubian, M.; Sun, J.C.; Lanier, L.L. MicroRNA function in NK-cell biology. Immunol. Rev. 2013, 253, 40–52. [Google Scholar] [CrossRef]
- Ni, F.; Guo, C.; Sun, R.; Fu, B.; Yang, Y.; Wu, L.; Ren, S.; Tian, Z.; Wei, H. MicroRNA transcriptomes of distinct human NK cell populations identify miR-362-5p as an essential regulator of NK cell function. Sci. Rep. 2015, 5, 9993. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Baek, D.; Villen, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- Kruger, J.; Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 2006, 34, W451–W454. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. Targetscanhuman: Prediction of microRNA Targets. Available online: http://www.targetscan.org/vert_61/ (accessed on 30 June 2019).
- Betel, D. Miranda: A Comprehensive Resource of microRNA Target Predictions and Expression Profiles. Available online: http://www.microrna.org/microrna/getDownloads.do (accessed on 30 June 2019).
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar] [CrossRef] [PubMed]
- Varkonyi-Gasic, E.; Wu, R.; Wood, M.; Walton, E.F.; Hellens, R.P. Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 2007, 3, 12. [Google Scholar] [CrossRef]
- Tajik, N.; Shahsavar, F.; Nasiri, M.; Radjabzadeh, M.F. Compound KIR-HLA genotype analyses in the Iranian population by a novel PCR-SSP assay. Int. J. Immunogenet. 2010, 37, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Stern-Ginossar, N.; Gur, C.; Biton, M.; Horwitz, E.; Elboim, M.; Stanietsky, N.; Mandelboim, M.; Mandelboim, O. Human microRNAs regulate stress-induced immune responses mediated by the receptor NKG2D. Nat. Immunol. 2008, 9, 1065–1073. [Google Scholar] [CrossRef]
- Bezman, N.A.; Cedars, E.; Steiner, D.F.; Blelloch, R.; Hesslein, D.G.; Lanier, L.L. Distinct requirements of microRNAs in NK cell activation, survival, and function. J. Immunol. 2010, 185, 3835–3846. [Google Scholar] [CrossRef]
- Bezman, N.A.; Chakraborty, T.; Bender, T.; Lanier, L.L. miR-150 regulates the development of NK and iNKT cells. J. Exp. Med. 2011, 208, 2717–2731. [Google Scholar] [CrossRef]
- Cichocki, F.; Felices, M.; McCullar, V.; Presnell, S.R.; Al-Attar, A.; Lutz, C.T.; Miller, J.S. Cutting edge: microRNA-181 promotes human NK cell development by regulating Notch signaling. J. Immunol. 2011, 187, 6171–6175. [Google Scholar] [CrossRef] [PubMed]
- Espinoza, J.L.; Takami, A.; Yoshioka, K.; Nakata, K.; Sato, T.; Kasahara, Y.; Nakao, S. Human microRNA-1245 down-regulates the NKG2D receptor in natural killer cells and impairs NKG2D-mediated functions. Haematologica 2012, 97, 1295–1303. [Google Scholar] [CrossRef] [PubMed]
- Chan, H.W.; Kurago, Z.B.; Stewart, C.A.; Wilson, M.J.; Martin, M.P.; Mace, B.E.; Carrington, M.; Trowsdale, J.; Lutz, C.T. DNA methylation maintains allele-specific KIR gene expression in human natural killer cells. J. Exp. Med. 2003, 197, 245–255. [Google Scholar] [CrossRef] [PubMed]
- Kelley, J.; Walter, L.; Trowsdale, J. Comparative genomics of natural killer cell receptor gene clusters. PLoS Genet. 2005, 1, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Wu, X.; Liu, B.; Wang, C.; Liu, Y.; Zhou, Q.; Xu, K. MiR-26a enhances metastasis potential of lung cancer cells via AKT pathway by targeting PTEN. Biochim. Biophys. Acta. 2012, 1822, 1692–1704. [Google Scholar] [CrossRef] [PubMed]
- Ostadrahimi, S.; Abedi Valugerdi, M.; Hassan, M.; Haddad, G.; Fayaz, S.; Parvizhamidi, M.; Mahdian, R.; Fard Esfahani, P. miR-1266-5p and miR-185-5p Promote Cell Apoptosis in Human Prostate Cancer Cell Lines. Asian. Pac. J. Cancer Prev. 2018, 19, 2305–2311. [Google Scholar]
- Rizzo, M.; Berti, G.; Russo, F.; Fazio, S.; Evangelista, M.; D’Aurizio, R.; Pellegrini, M.; Rainaldi, G. Discovering the miR-26a-5p Targetome in Prostate Cancer Cells. J. Cancer 2017, 8, 2729–2739. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jin, F.; Wang, Y.; Li, M.; Zhu, Y.; Liang, H.; Wang, C.; Wang, F.; Zhang, C.Y.; Zen, K.; Li, L. MiR-26 enhances chemosensitivity and promotes apoptosis of hepatocellular carcinoma cells through inhibiting autophagy. Cell Death Dis. 2017, 8, e2540. [Google Scholar] [CrossRef]
| Plasmid Names | Information |
|---|---|
| p3’UTR_WT | Wild type 3’UTR of KIR3DL3 |
| pMu_miR-26a/b-5p (295) | Mutated miR-26a/b-5p binding site 3’UTR at positions 297 and 299 |
| pMu_miR-26a/b-5p (344) | Mutated miR-26a/b-5p binding site 3’UTR at positions 344 and 348 |
| pMu_miR-26a/b-5p (295,344) | Mutated both miR-26a-5p and miR-26b-5p binding sites |
| pMu_miR-185-5p | Mutated miR-185-5p binding site |
| pMu_miR-203a-3p | Mutated miR-203a-3p binding site |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nutalai, R.; Gaudieri, S.; Jumnainsong, A.; Leelayuwat, C. Regulation of KIR3DL3 Expression via miRNA. Genes 2019, 10, 603. https://doi.org/10.3390/genes10080603
Nutalai R, Gaudieri S, Jumnainsong A, Leelayuwat C. Regulation of KIR3DL3 Expression via miRNA. Genes. 2019; 10(8):603. https://doi.org/10.3390/genes10080603
Chicago/Turabian StyleNutalai, Rungtiwa, Silvana Gaudieri, Amonrat Jumnainsong, and Chanvit Leelayuwat. 2019. "Regulation of KIR3DL3 Expression via miRNA" Genes 10, no. 8: 603. https://doi.org/10.3390/genes10080603
APA StyleNutalai, R., Gaudieri, S., Jumnainsong, A., & Leelayuwat, C. (2019). Regulation of KIR3DL3 Expression via miRNA. Genes, 10(8), 603. https://doi.org/10.3390/genes10080603






