Identification of a Novel Melon Transcription Factor CmNAC60 as a Potential Regulator of Leaf Senescence
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials and Dark Treatment
2.2. Expression of CmNAC60 in Different Tissues of Oriental Melon
2.3. Isolation and Bioinformatic Analysis of CmNAC60
2.4. Isolation and Bioinformatic Analysis of the CmNAC60 Promoter
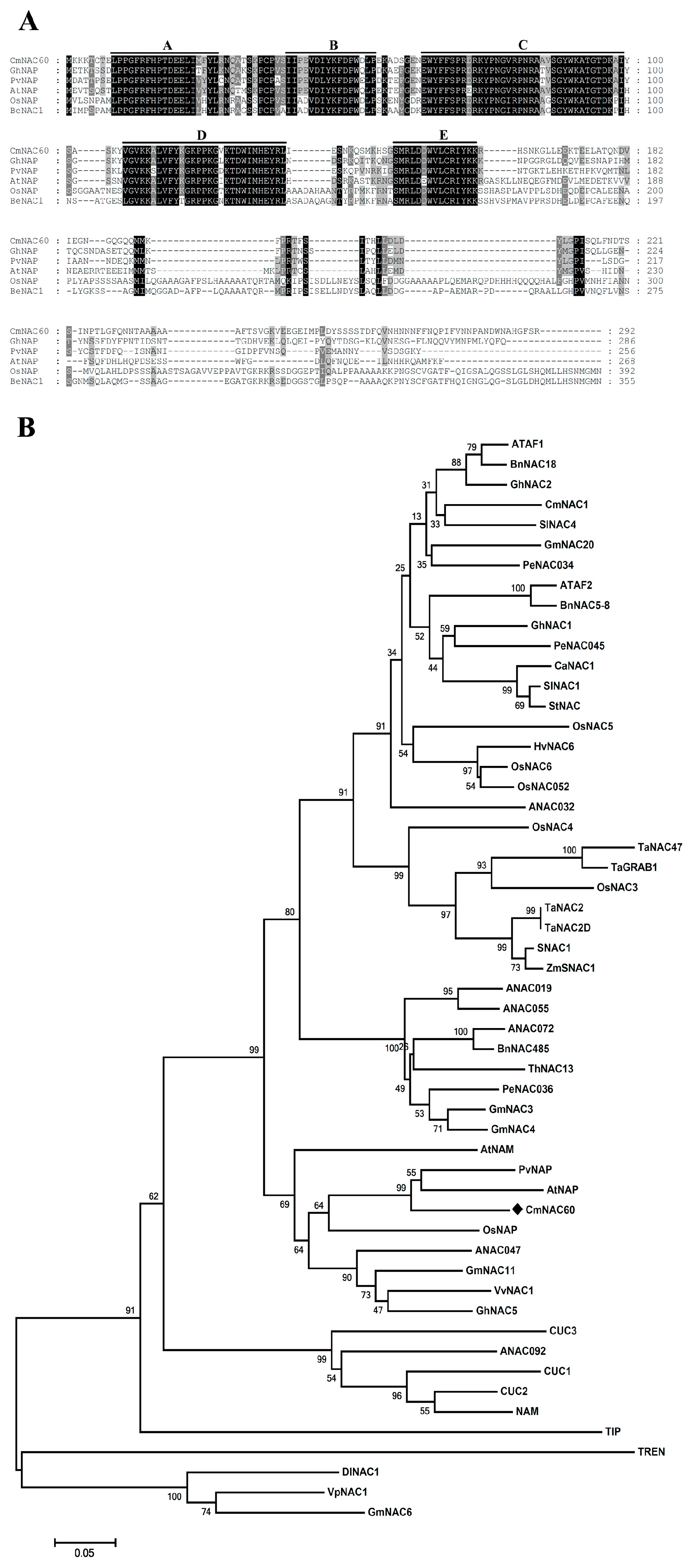
2.5. Multiple Sequence Alignment and Phylogenetic Analysis of CmNAC60
2.6. Subcellular Localization of CmNAC60
2.7. Transcriptional Activation Assay of CmNAC60
2.8. Tissue Specific Expression of CmNAC60
2.9. Generation of Overexpression Transgenic Arabidopsis Plants
2.10. RNA Isolation, qRT-PCR, and Measurement of Chlorophyll Content
2.11. Statistical Analysis
3. Results
3.1. Sequence Characterization of the CDS and Promoter of CmNAC60
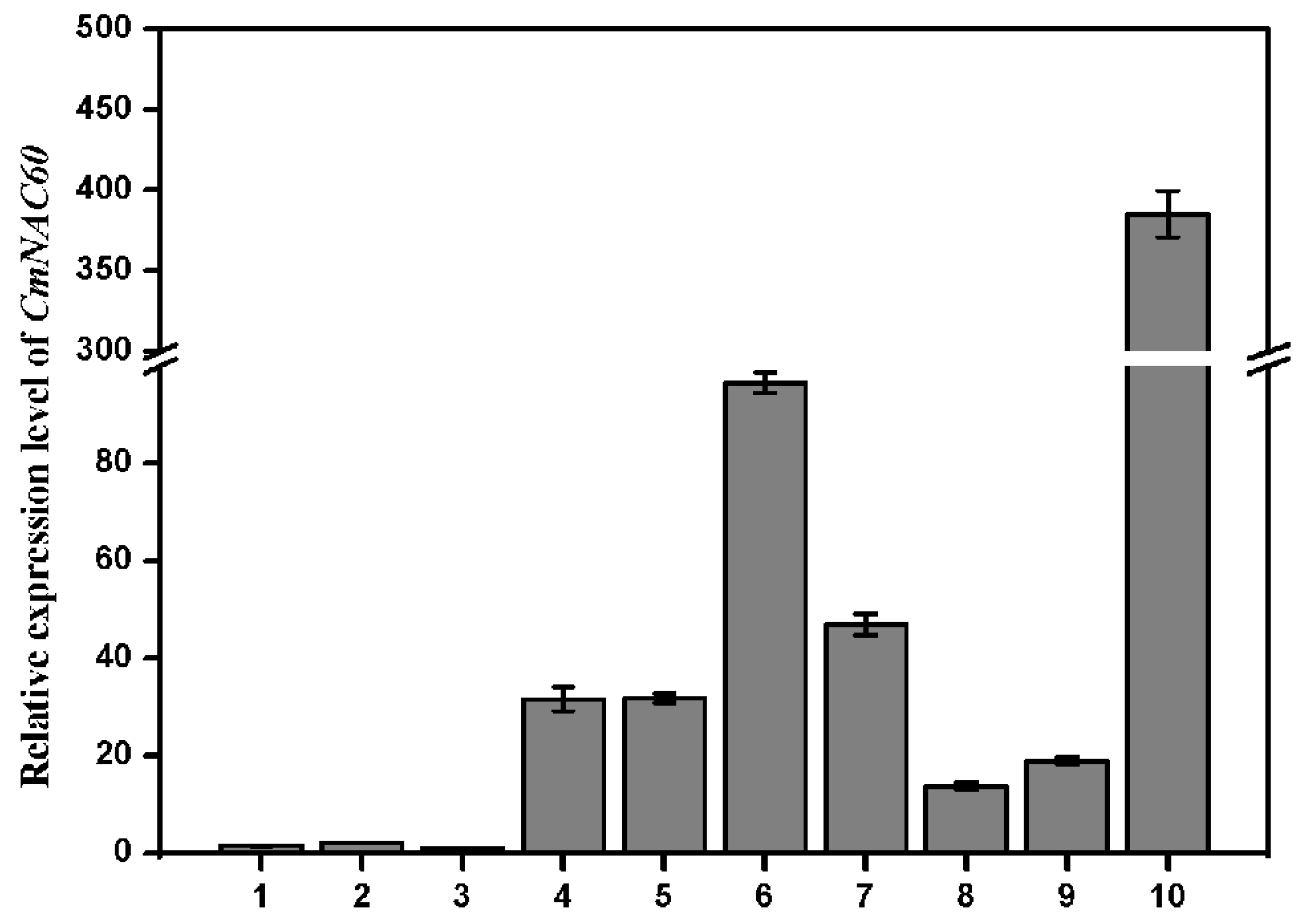
3.2. Expression of CmNAC60 in Different Tissues of Oriental Melon
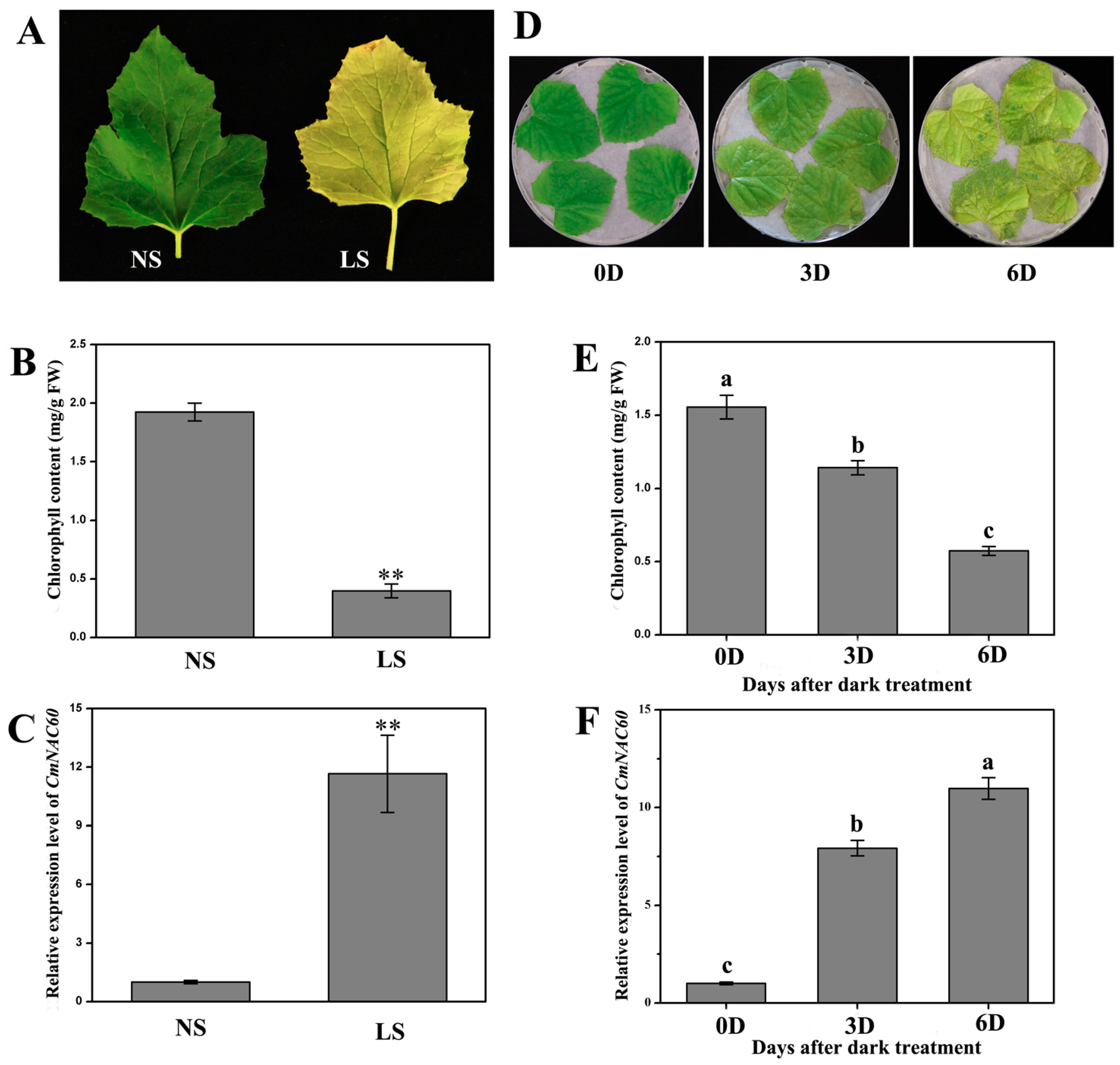
3.3. Relative Expression of CmNAC60 during Natural Senescence and Dark Treatment
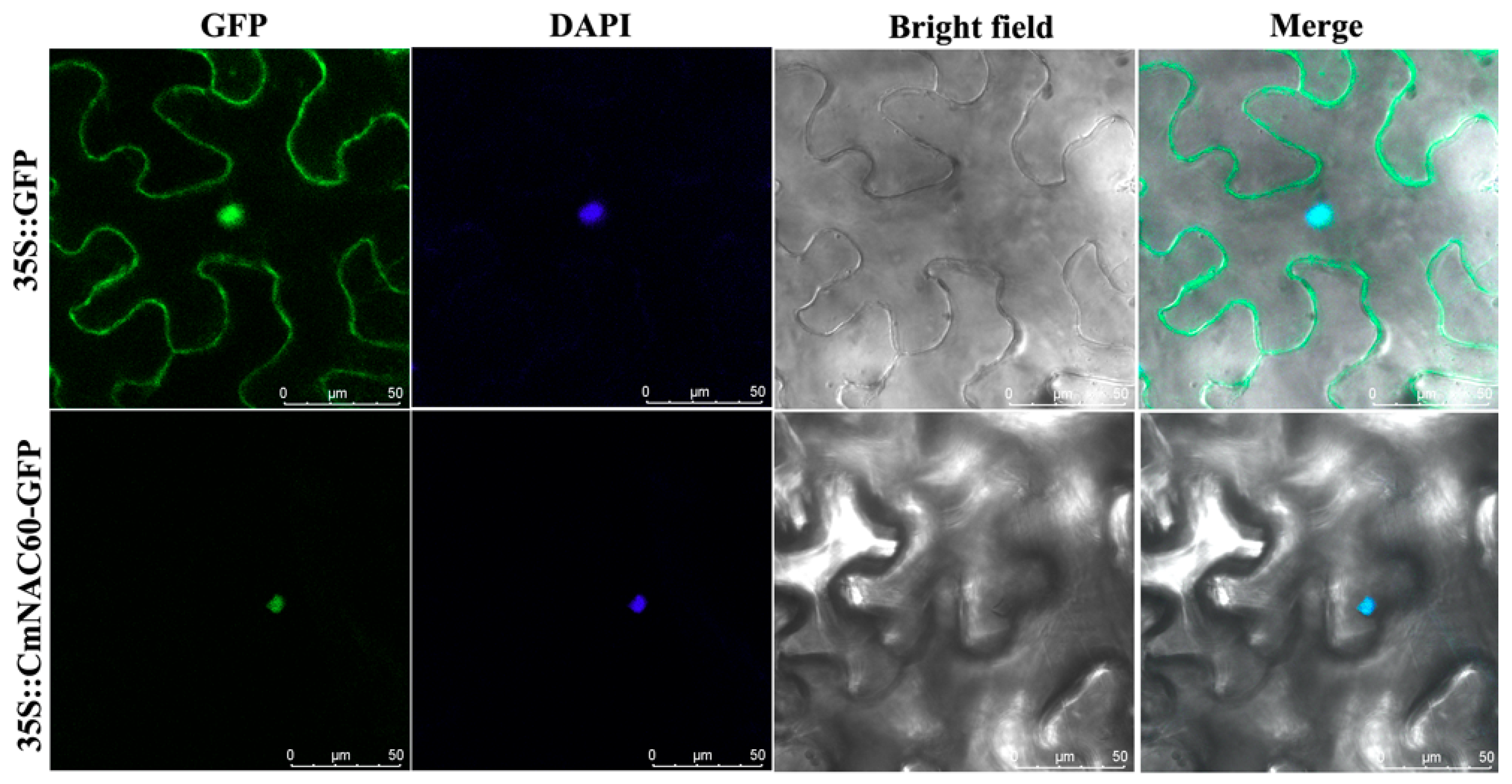
3.4. Subcellular Localization of CmNAC60
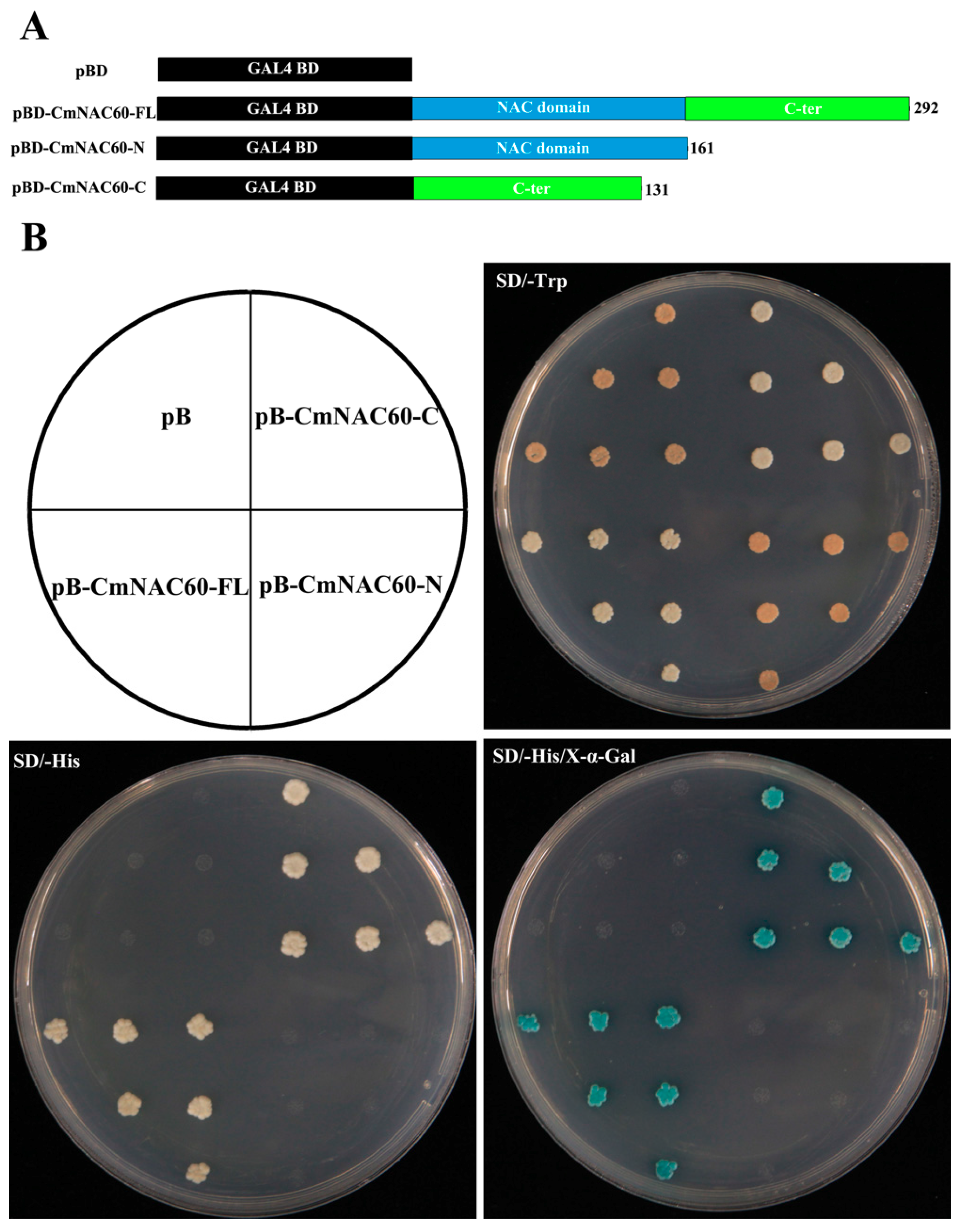
3.5. Transactivation Activity Assay
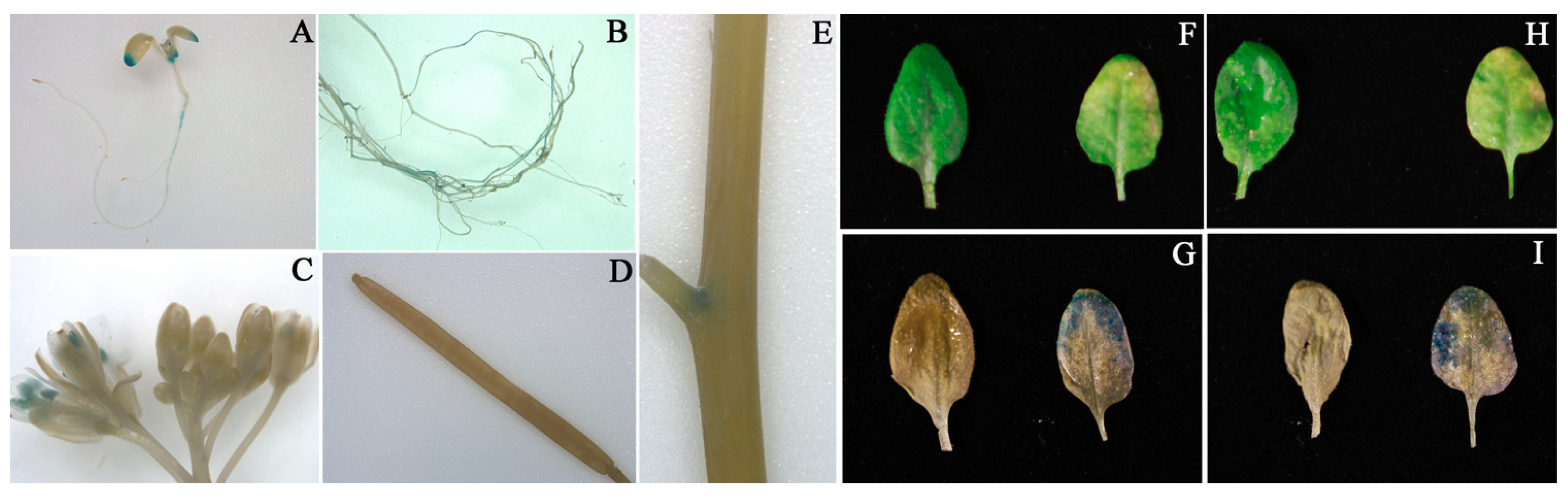
3.6. Tissue Specific Expression of CmNAC60
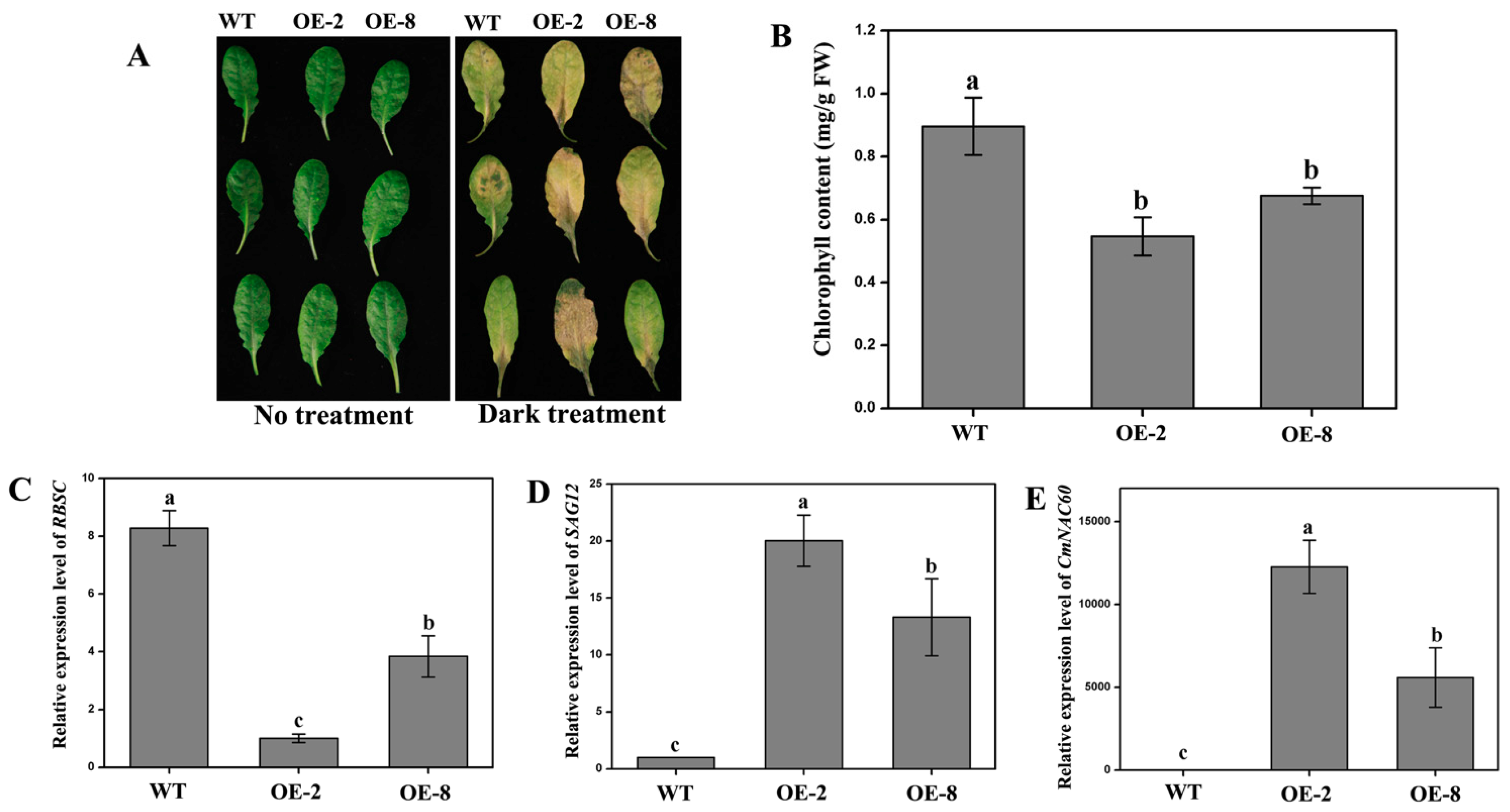
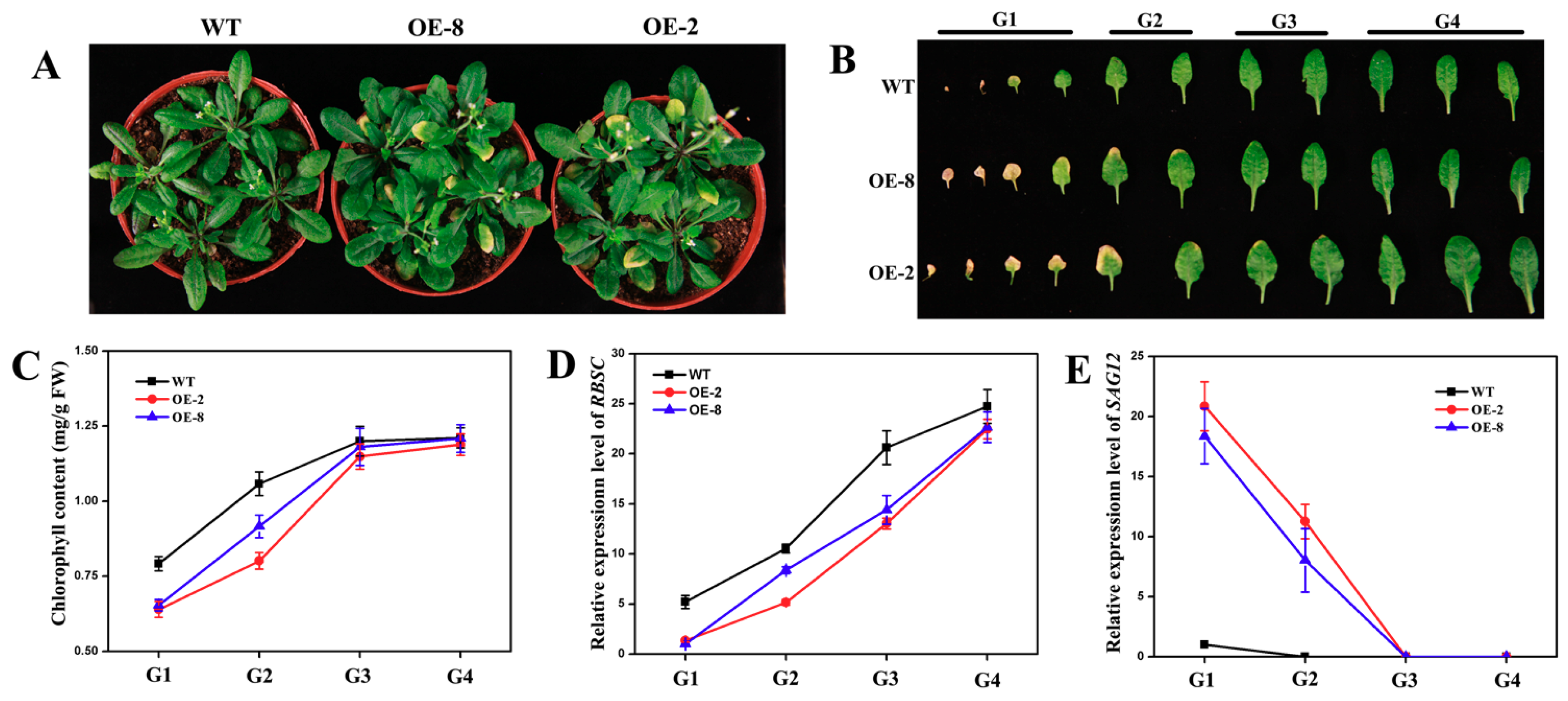
3.7. Overexpression of CmNAC60 in Arabidopsis Accelerates Leaf Senescence
4. Discussion
4.1. CmNAC60 Is Functionally Related to Leaf Senescence
4.2. The CmNAC60 Promoter May Contain Cis-Acting Regulatory Element Involved in Leaf Senescence
4.3. Relative Expression Difference of CmNAC60 between the Male and Female Flowers of Oriental Melon
4.4. CmNAC60 Is a Member of the Melon NAC Transcription Factors
4.5. CmNAC60 Overexpression Can Accelerate Leaf Senescence in Arabidopsis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lim, P.O.; Kim, H.J.; Gil Nam, H. Leaf Senescence. Annu. Rev. Plant Boil. 2007, 58, 115–136. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, X.-Y.; Kuai, B.-K.; Jia, J.-Z.; Jing, H.-C. Regulation of Leaf Senescence and Crop Genetic Improvement. J. Integr. Plant Boil. 2012, 54, 936–952. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y. Towards systems biological understanding of leaf senescence. Plant Mol. Biol. 2013, 82, 519–528. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Gao, X.; Guo, Y. Initiation, Progression, and Genetic Manipulation of Leaf Senescence. Methods Mol. Biol. 2018, 1744, 9–31. [Google Scholar] [PubMed]
- Breeze, E.; Harrison, E.; McHattie, S.; Hughes, L.; Hickman, R.; Hill, C.; Kiddle, S.; Kim, Y.-S.; Penfold, C.A.; Jenkins, D.; et al. High-Resolution Temporal Profiling of Transcripts during Arabidopsis Leaf Senescence Reveals a Distinct Chronology of Processes and Regulation. Plant Cell 2011, 23, 873–894. [Google Scholar] [CrossRef] [PubMed]
- Riechmann, J.L.; Heard, J.; Martin, G.; Reuber, L.; Jiang, C.-Z.; Keddie, J.; Adam, L.; Pineda, O.; Ratcliffe, O.J.; Samaha, R.R.; et al. Arabidopsis Transcription Factors: Genome-Wide Comparative Analysis Among Eukaryotes. Science 2000, 290, 2105–2110. [Google Scholar] [CrossRef] [PubMed]
- Buchanan-Wollaston, V.; Page, T.; Harrison, E.; Breeze, E.; Lim, P.O.; Gil Nam, H.; Lin, J.-F.; Wu, S.-H.; Swidzinski, J.; Ishizaki, K.; et al. Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J. 2005, 42, 567–585. [Google Scholar] [CrossRef]
- Guo, Y.; Gan, S.S. Convergence and divergence in gene expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant Cell Environ. 2012, 35, 644–655. [Google Scholar] [CrossRef]
- Balazadeh, S.; Riaño-Pachón, D.M.; Mueller-Roeber, B. Transcription factors regulating leaf senescence in Arabidopsis thaliana. Plant Boil. 2008, 10, 63–75. [Google Scholar] [CrossRef]
- Souer, E.; Van Houwelingen, A.; Kloos, D.; Mol, J.; Koes, R. The No Apical Meristem Gene of Petunia Is Required for Pattern Formation in Embryos and Flowers and Is Expressed at Meristem and Primordia Boundaries. Cell 1996, 85, 159–170. [Google Scholar] [CrossRef] [Green Version]
- Aida, M.; Ishida, T.; Fukaki, H.; Fujisawa, H.; Tasaka, M. Genes involved in organ separation in Arabidopsis: An analysis of the cup-shaped cotyledon mutant. Plant Cell 1997, 9, 841–857. [Google Scholar] [CrossRef]
- Olsen, A.N.; Ernst, H.A.; Leggio, L.L.; Skriver, K. NAC transcription factors: Structurally distinct, functionally diverse. Trends Plant Sci. 2005, 10, 79–87. [Google Scholar] [CrossRef]
- Nakashima, K.; Takasaki, H.; Mizoi, J.; Shinozaki, K.; Yamaguchi-Shinozaki, K. NAC transcription factors in plant abiotic stress responses. Biochim. Biophys. Acta BBA Bioenerg. 2012, 1819, 97–103. [Google Scholar] [CrossRef]
- Ooka, H.; Satoh, K.; Doi, K.; Nagata, T.; Otomo, Y.; Murakami, K.; Matsubara, K.; Osato, N.; Kawai, J.; Carninci, P.; et al. Comprehensive Analysis of NAC Family Genes in Oryza sativa and Arabidopsis thaliana. DNA Res. 2003, 10, 239–247. [Google Scholar] [CrossRef]
- Hu, R.; Qi, G.; Kong, Y.; Kong, D.; Gao, Q.; Zhou, G. Comprehensive Analysis of NAC Domain Transcription Factor Gene Family in Populus trichocarpa. BMC Plant Boil. 2010, 10, 145. [Google Scholar] [CrossRef]
- Le, D.T.; Nishiyama, R.; Watanabe, Y.; Mochida, K.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Tran, L.-S.P. Genome-Wide Survey and Expression Analysis of the Plant-Specific NAC Transcription Factor Family in Soybean During Development and Dehydration Stress. DNA Res. 2011, 18, 263–276. [Google Scholar] [CrossRef] [Green Version]
- Wang, N.; Zheng, Y.; Xin, H.; Fang, L.; Li, S. Comprehensive analysis of NAC domain transcription factor gene family in Vitis vinifera. Plant Cell Rep. 2013, 32, 61–75. [Google Scholar] [CrossRef]
- Su, H.; Zhang, S.; Yuan, X.; Chen, C.; Wang, X.-F.; Hao, Y.-J. Genome-wide analysis and identification of stress-responsive genes of the NAM-ATAF1,2-CUC2 transcription factor family in apple. Plant Physiol. Biochem. 2013, 71, 11–21. [Google Scholar] [CrossRef]
- Singh, A.K.; Sharma, V.; Pal, A.K.; Acharya, V.; Ahuja, P.S. Genome-Wide Organization and Expression Profiling of the NAC Transcription Factor Family in Potato (Solanum tuberosum L.). DNA Res. 2013, 20, 403–423. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.M.; Yu, H.J.; Sun, C.; Deng, J.; Zhang, X.; Liu, P.; Li, Y.Y.; Li, Q.; Jiang, W.J. Genome-wide characterization and expression profiling of the NAC genes under abiotic stresses in Cucumis sativus. Plant Physiol. Biochem. 2017, 113, 98–109. [Google Scholar] [CrossRef]
- Kim, Y.-S.; Kim, S.-G.; Park, J.-E.; Park, H.-Y.; Lim, M.-H.; Chua, N.-H.; Park, C.-M. A Membrane-Bound NAC Transcription Factor Regulates Cell Division in Arabidopsis. Plant Cell 2006, 18, 3132–3144. [Google Scholar] [CrossRef]
- Kim, J.H.; Woo, H.R.; Kim, J.; Lim, P.O.; Lee, I.C.; Choi, S.H.; Hwang, D.; Nam, H.G. Trifurcate Feed-Forward Regulation of Age-Dependent Cell Death Involving miR164 in Arabidopsis. Science 2009, 323, 1053–1057. [Google Scholar] [CrossRef]
- Zhong, R.; Lee, C.; Ye, Z.H. Functional characterization of poplar wood-associated NAC domain transcription factors. Plant Physiol. 2010, 152, 1044–1055. [Google Scholar] [CrossRef]
- Zhao, Y.; Sun, J.; Xu, P.; Zhang, R.; Li, L. Intron-Mediated Alternative Splicing of WOOD-ASSOCIATED NAC TRANSCRIPTION FACTOR1B Regulates Cell Wall Thickening during Fiber Development in Populus Species. Plant Physiol. 2014, 164, 765–776. [Google Scholar] [CrossRef]
- Vroemen, C.W.; Mordhorst, A.P.; Albrecht, C.; Kwaaitaal, M.A.C.J.; De Vries, S.C. The CUP-SHAPED COTYLEDON3 Gene Is Required for Boundary and Shoot Meristem Formation in Arabidopsis. Plant Cell 2003, 15, 1563–1577. [Google Scholar] [CrossRef]
- Hao, Y.-J.; Wei, W.; Song, Q.-X.; Chen, H.-W.; Zhang, Y.-Q.; Wang, F.; Zou, H.-F.; Lei, G.; Tian, A.-G.; Zhang, W.-K.; et al. Soybean NAC transcription factors promote abiotic stress tolerance and lateral root formation in transgenic plants. Plant J. 2011, 68, 302–313. [Google Scholar] [CrossRef]
- Kou, X.; Watkins, C.B.; Gan, S.-S. Arabidopsis AtNAP regulates fruit senescence. J. Exp. Bot. 2012, 63, 6139–6147. [Google Scholar] [CrossRef] [Green Version]
- Guo, Y.; Gan, S. AtNAP, a NAC family transcription factor, has an important role in leaf senescence. Plant J. 2006, 46, 601–612. [Google Scholar] [CrossRef]
- Balazadeh, S.; Kwasniewski, M.; Caldana, C.; Mehrnia, M.; Zanor, M.I.; Xue, G.-P.; Mueller-Roeber, B. ORS1, an H2O2-Responsive NAC Transcription Factor, Controls Senescence in Arabidopsis thaliana. Mol. Plant 2011, 4, 346–360. [Google Scholar] [CrossRef]
- Takasaki, H.; Maruyama, K.; Takahashi, F.; Fujita, M.; Yoshida, T.; Nakashima, K.; Myouga, F.; Toyooka, K.; Yamaguchi-Shinozaki, K.; Shinozaki, K. SNAC-As, stress-responsive NAC transcription factors, mediate ABA-inducible leaf senescence. Plant J. 2015, 84, 1114–1123. [Google Scholar] [CrossRef]
- Lee, S.; Lee, H.-J.; Huh, S.U.; Paek, K.-H.; Ha, J.-H.; Park, C.-M. The Arabidopsis NAC transcription factor NTL4 participates in a positive feedback loop that induces programmed cell death under heat stress conditions. Plant Sci. 2014, 227, 76–83. [Google Scholar] [CrossRef]
- Shao, H.; Wang, H.; Tang, X. NAC transcription factors in plant multiple abiotic stress responses: Progress and prospects. Front. Plant Sci. 2015, 6, 902. [Google Scholar] [CrossRef]
- Le Hénanff, G.; Profizi, C.; Courteaux, B.; Rabenoelina, F.; Gérard, C.; Clément, C.; Baillieul, F.; Cordelier, S.; Dhondt-Cordelier, S. Grapevine NAC1 transcription factor as a convergent node in developmental processes, abiotic stresses, and necrotrophic/biotrophic pathogen tolerance. J. Exp. Bot. 2013, 64, 4877–4893. [Google Scholar] [CrossRef] [Green Version]
- Fan, K.; Bibi, N.; Gan, S.; Li, F.; Yuan, S.; Ni, M.; Wang, M.; Shen, H.; Wang, X. A novel NAP member GhNAP is involved in leaf senescence in Gossypium hirsutum. J. Exp. Bot. 2015, 66, 4669–4682. [Google Scholar] [CrossRef]
- Chen, Y.; Qiu, K.; Kuai, B.; Ding, Y. Identification of an NAP-like transcription factor BeNAC1 regulating leaf senescence in bamboo (Bambusa emeiensis‘Viridiflavus’). Physiol. Plant. 2011, 142, 361–371. [Google Scholar] [CrossRef]
- Liang, C.; Wang, Y.; Zhu, Y.; Tang, J.; Hu, B.; Liu, L.; Ou, S.; Wu, H.; Sun, X.; Chu, J.; et al. OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc. Natl. Acad. Sci. USA 2014, 111, 10013–10018. [Google Scholar] [CrossRef] [Green Version]
- El Mannai, Y.; Akabane, K.; Hiratsu, K.; Satoh-Nagasawa, N.; Wabiko, H. The NAC Transcription Factor Gene OsY37 (ONAC011) Promotes Leaf Senescence and Accelerates Heading Time in Rice. Int. J. Mol. Sci. 2017, 18, 2165. [Google Scholar] [CrossRef]
- Zhu, Z.; Li, G.; Yan, C.; Liu, L.; Zhang, Q.; Han, Z.; Li, B. DRL1, Encoding A NAC Transcription Factor, Is Involved in Leaf Senescence in Grapevine. Int. J. Mol. Sci. 2019, 20, 2678. [Google Scholar] [CrossRef]
- Wei, S.; Gao, L.; Zhang, Y.; Zhang, F.; Yang, X.; Huang, D. Genome-wide investigation of the NAC transcription factor family in melon (Cucumis melo L.) and their expression analysis under salt stress. Plant Cell Rep. 2016, 35, 1827–1839. [Google Scholar] [CrossRef]
- Garcia-Mas, J.; Benjak, A.; Sanseverino, W.; Bourgeois, M.; Mir, G.; González, V.M.; Hénaff, E.; Câmara, F.; Cozzuto, L.; Lowy, E.; et al. The genome of melon (Cucumis melo L.). Proc. Natl. Acad. Sci. USA 2012, 109, 11872–11877. [Google Scholar] [CrossRef]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method forAgrobacterium-mediated transformation ofArabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef]
- Jefferson, R.; Kavanagh, T.; Bevan, M. GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [CrossRef]
- Zhang, Z.; Guo, Y. Hormone Treatments in Studying Leaf Senescence. Methods Mol. Biol. 2018, 1744, 125–132. [Google Scholar]
- Khan, M.; Rozhon, W.; Poppenberger, B. The Role of Hormones in the Aging of Plants—A Mini-Review. Gerontology 2014, 60, 49–55. [Google Scholar] [CrossRef]
- Reinbothe, C.; Springer, A.; Samol, I.; Reinbothe, S. Plant oxylipins: Role of jasmonic acid during programmed cell death, defence and leaf senescence. FEBS J. 2009, 276, 4666–4681. [Google Scholar] [CrossRef]
- Yang, J.; Worley, E.; Udvardi, M. A NAP-AAO3 Regulatory Module Promotes Chlorophyll Degradation via ABA Biosynthesis in Arabidopsis Leaves. Plant Cell 2014, 26, 4862–4874. [Google Scholar] [CrossRef]
- Balazadeh, S.; Siddiqui, H.; Allu, A.D.; Matallana-Ramirez, L.P.; Caldana, C.; Mehrnia, M.; Zanor, M.-I.; Köhler, B.; Mueller-Roeber, B. A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J. 2010, 62, 250–264. [Google Scholar] [CrossRef]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cao, S.; Zhang, Z.; Wang, C.; Li, X.; Guo, C.; Yang, L.; Guo, Y. Identification of a Novel Melon Transcription Factor CmNAC60 as a Potential Regulator of Leaf Senescence. Genes 2019, 10, 584. https://doi.org/10.3390/genes10080584
Cao S, Zhang Z, Wang C, Li X, Guo C, Yang L, Guo Y. Identification of a Novel Melon Transcription Factor CmNAC60 as a Potential Regulator of Leaf Senescence. Genes. 2019; 10(8):584. https://doi.org/10.3390/genes10080584
Chicago/Turabian StyleCao, Songxiao, Zhenbiao Zhang, Chenghui Wang, Xiaoxu Li, Cun Guo, Liyu Yang, and Yongfeng Guo. 2019. "Identification of a Novel Melon Transcription Factor CmNAC60 as a Potential Regulator of Leaf Senescence" Genes 10, no. 8: 584. https://doi.org/10.3390/genes10080584
APA StyleCao, S., Zhang, Z., Wang, C., Li, X., Guo, C., Yang, L., & Guo, Y. (2019). Identification of a Novel Melon Transcription Factor CmNAC60 as a Potential Regulator of Leaf Senescence. Genes, 10(8), 584. https://doi.org/10.3390/genes10080584




