Single-Nucleotide Polymorphisms Sequencing Identifies Candidate Functional Variants at Prostate Cancer Risk Loci
Abstract
1. Introduction
2. Materials and Methods
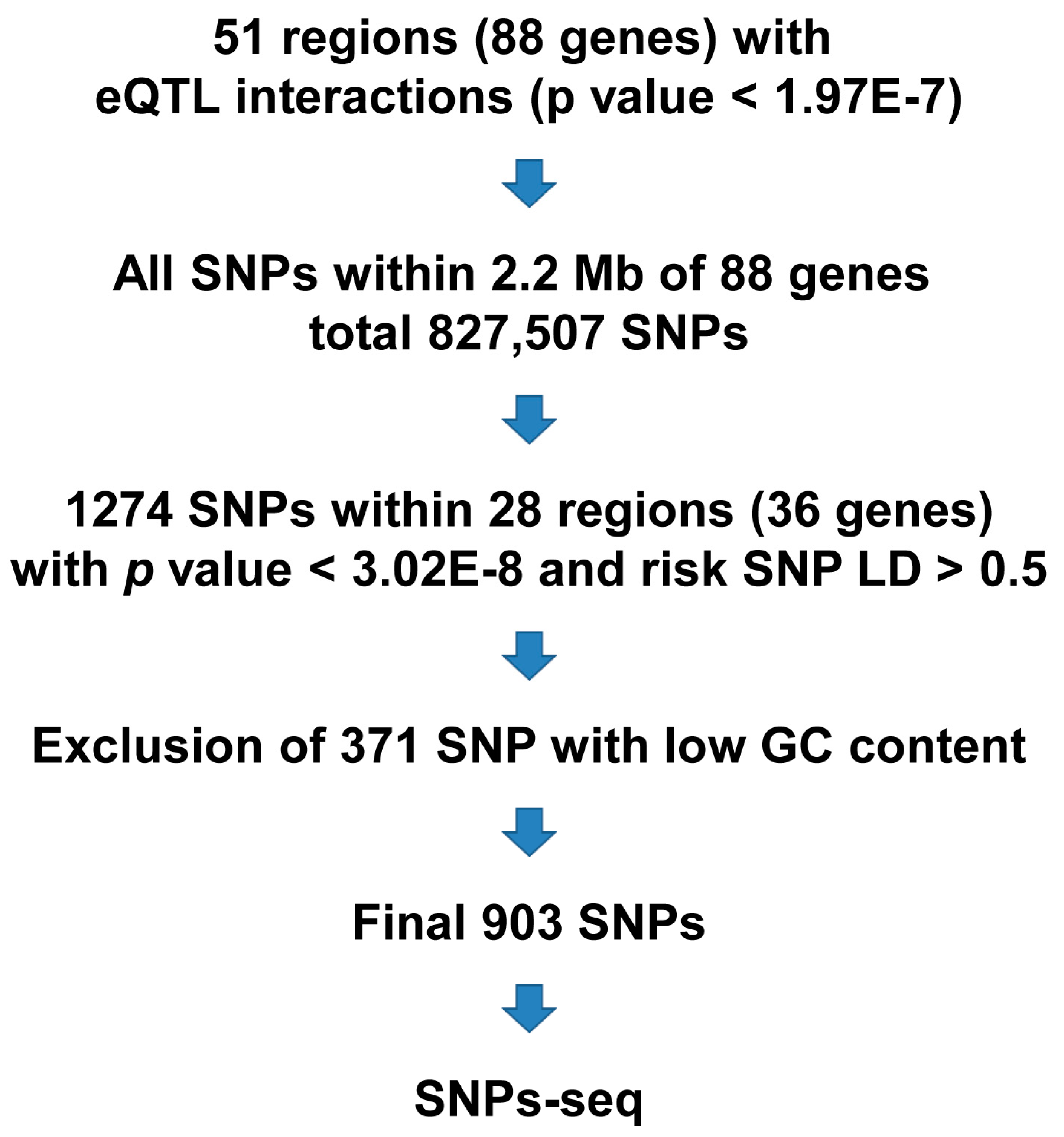
2.1. Selection of Candidate SNPs
2.2. Cell Culture and Nuclear Extraction
2.3. SNPs-seq
2.4. Database Search for Potential Functional SNPs
2.5. Electrophoretic Mobility Shift Assay (EMSA)
2.6. Clinical Association Analysis
3. Results
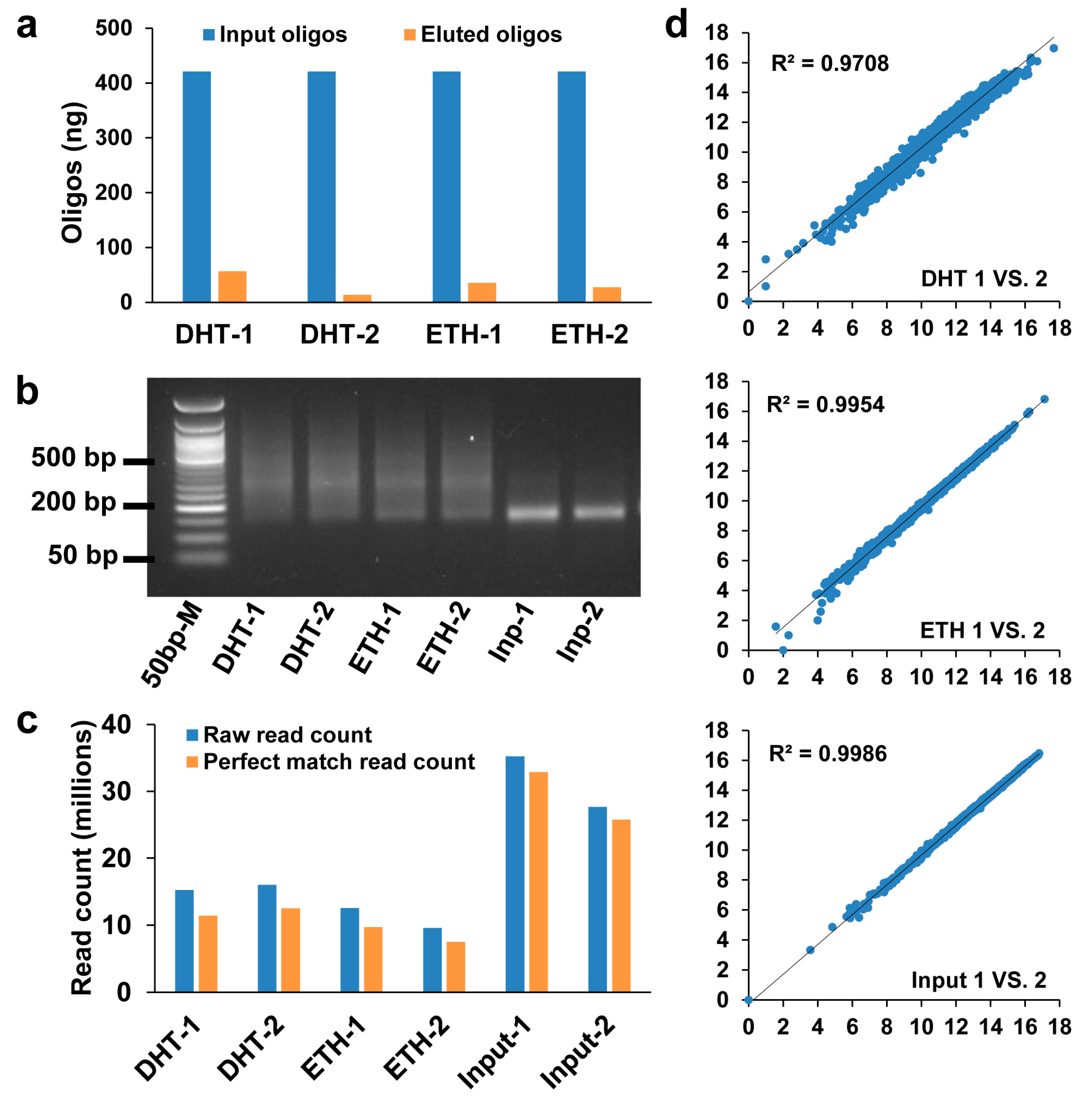
3.1. High-Quality SNPs-seq Libraries
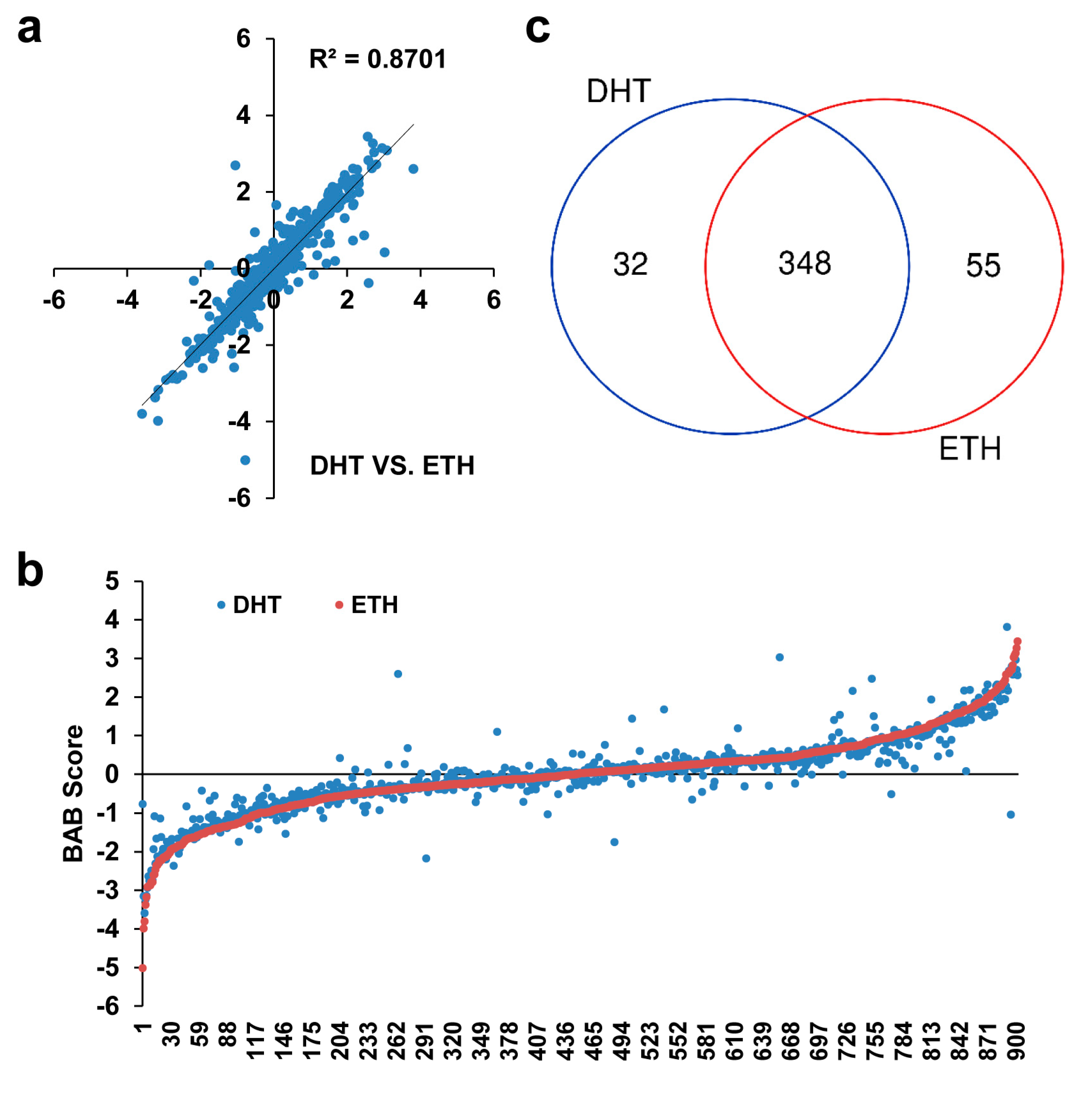
3.2. Candidate SNPs with Allele-Dependent Protein Binding
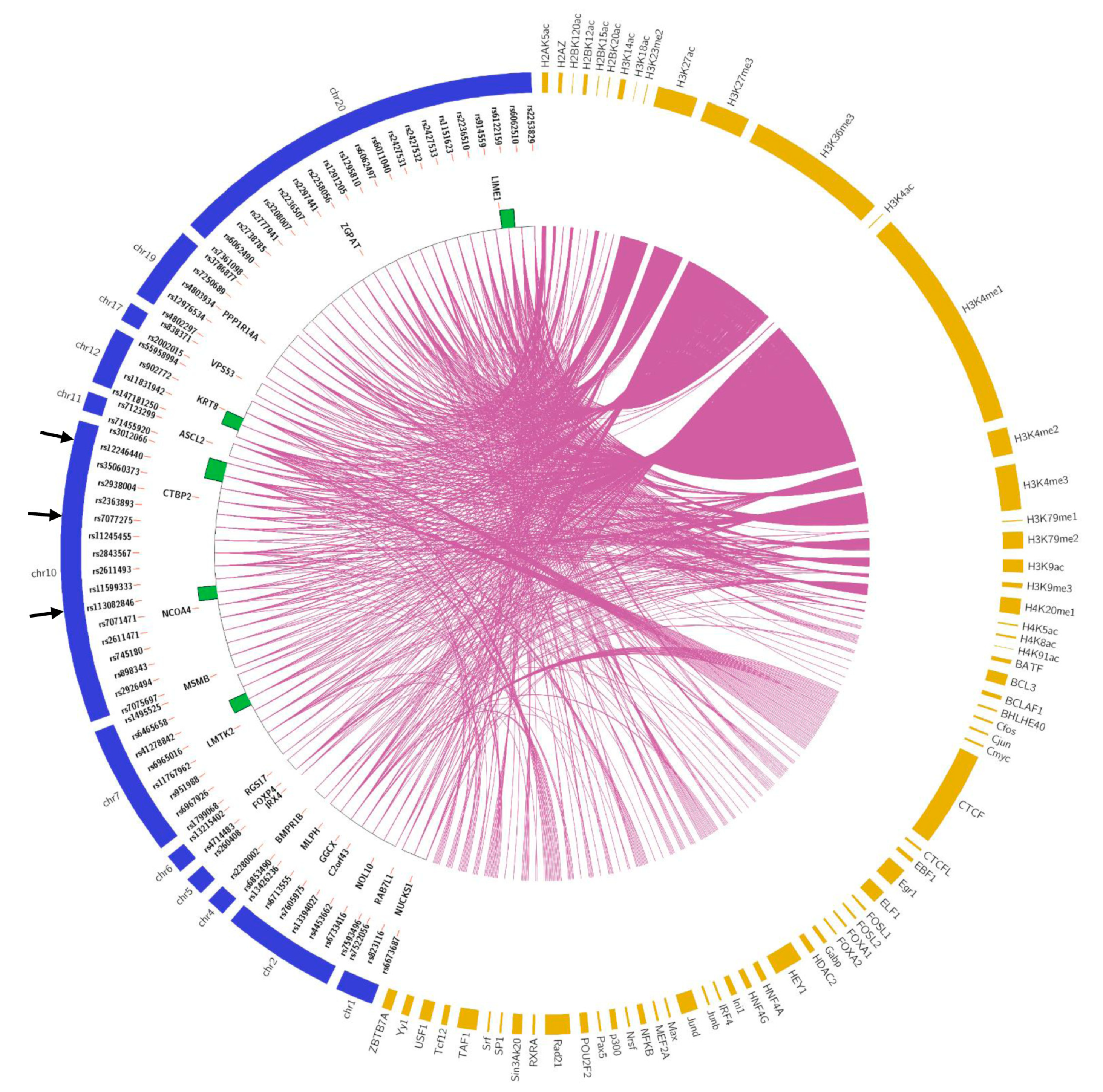
3.3. Candidate SNPs to Prioritize Functional Validation
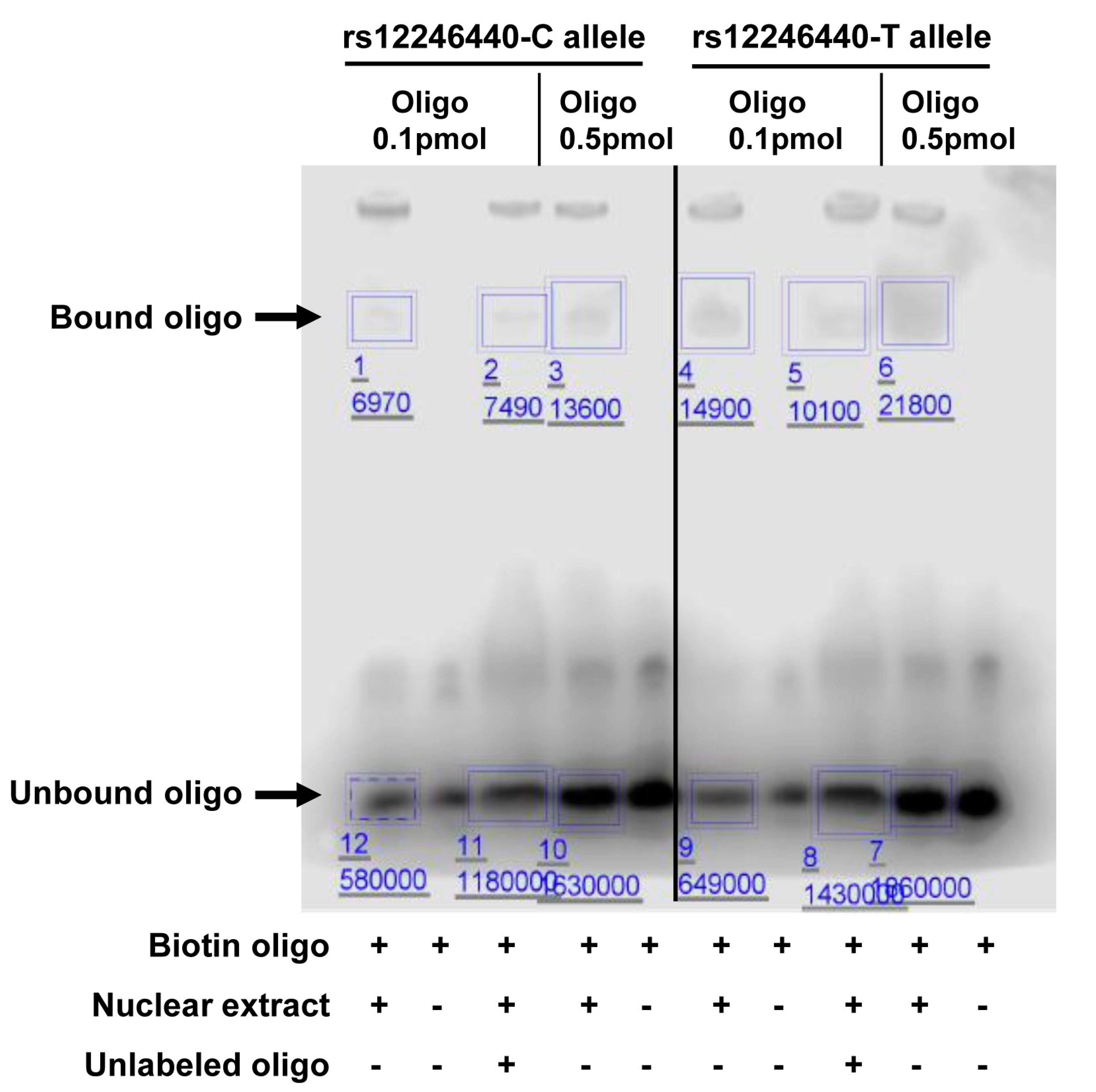
3.4. Validation of Selected Candidate SNPs
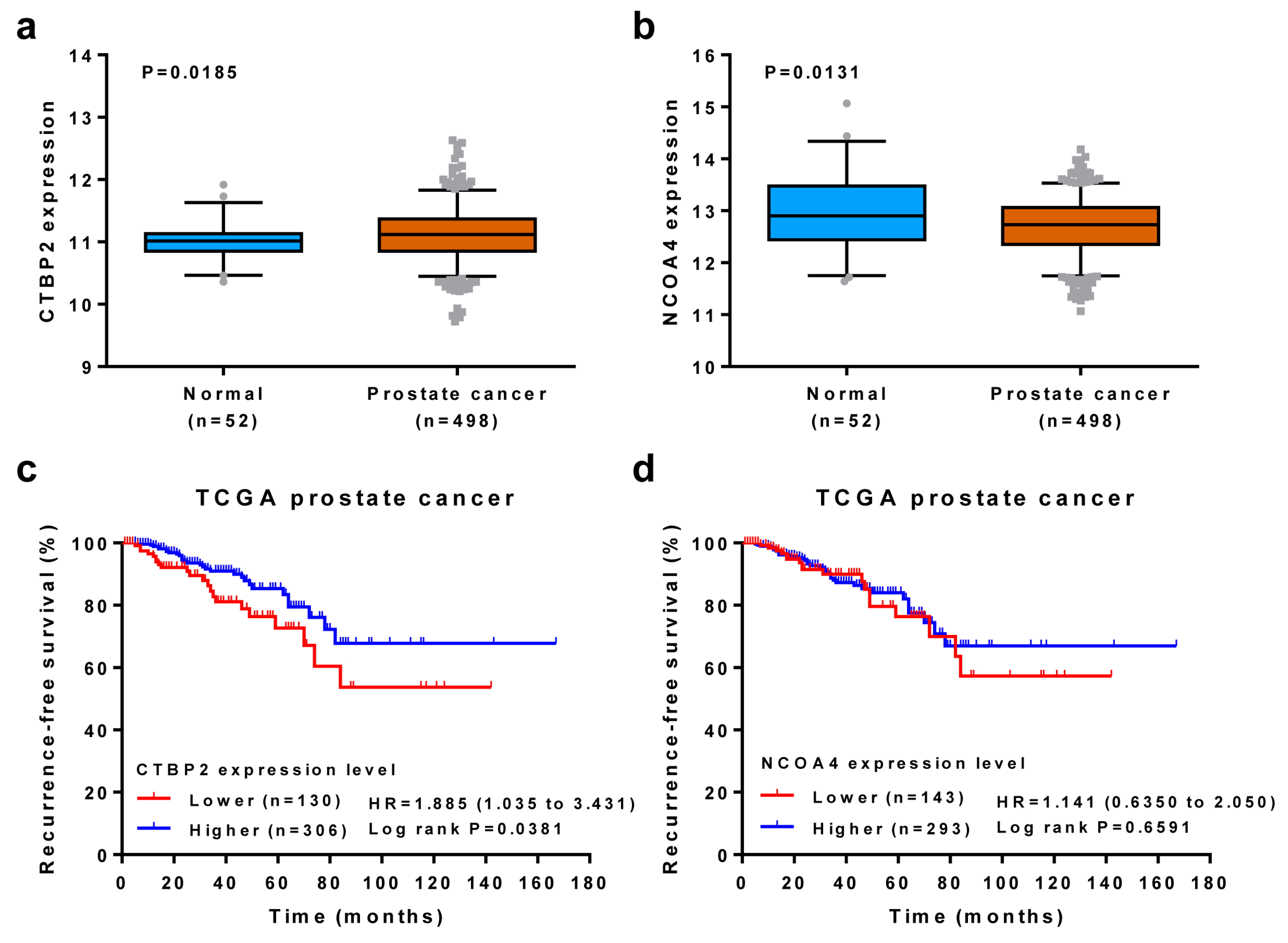
3.5. Association of Candidate Genes with Clinical Outcomes
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Attard, G.; Parker, C.; Eeles, R.A.; Schroder, F.; Tomlins, S.A.; Tannock, I.; Drake, C.G.; de Bono, J.S. Prostate cancer. Lancet 2016, 387, 70–82. [Google Scholar] [CrossRef]
- Wang, G.; Zhao, D.; Spring, D.J.; DePinho, R.A. Genetics and biology of prostate cancer. Genes Dev. 2018, 32, 1105–1140. [Google Scholar] [CrossRef] [PubMed]
- Neslund-Dudas, C.; Levin, A.M.; Rundle, A.; Beebe-Dimmer, J.; Bock, C.H.; Nock, N.L.; Jankowski, M.; Datta, I.; Krajenta, R.; Dou, Q.P.; et al. Case-only gene-environment interaction between ALAD tagSNPs and occupational lead exposure in prostate cancer. Prostate 2014, 74, 637–646. [Google Scholar] [CrossRef] [PubMed]
- Buniello, A.; MacArthur, J.A.L.; Cerezo, M.; Harris, L.W.; Hayhurst, J.; Malangone, C.; McMahon, A.; Morales, J.; Mountjoy, E.; Sollis, E.; et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 2019, 47, D1005–D1012. [Google Scholar] [CrossRef]
- Benafif, S.; Kote-Jarai, Z.; Eeles, R.A.; Consortium, P. A Review of Prostate Cancer Genome-Wide Association Studies (GWAS). Cancer Epidemiol. Biomark. Prev. 2018, 27, 845–857. [Google Scholar] [CrossRef] [PubMed]
- Farashi, S.; Kryza, T.; Clements, J.; Batra, J. Post-GWAS in prostate cancer: From genetic association to biological contribution. Nat. Rev. Cancer 2019, 19, 46–59. [Google Scholar] [CrossRef]
- Chen, H.; Yu, H.; Wang, J.; Zhang, Z.; Gao, Z.; Chen, Z.; Lu, Y.; Liu, W.; Jiang, D.; Zheng, S.L.; et al. Systematic enrichment analysis of potentially functional regions for 103 prostate cancer risk-associated loci. Prostate 2015, 75, 1264–1276. [Google Scholar] [CrossRef]
- Westra, H.J.; Franke, L. From genome to function by studying eQTLs. Biochim. Biophys. Acta 2014, 1842, 1896–1902. [Google Scholar] [CrossRef]
- Thibodeau, S.N.; French, A.J.; McDonnell, S.K.; Cheville, J.; Middha, S.; Tillmans, L.; Riska, S.; Baheti, S.; Larson, M.C.; Fogarty, Z.; et al. Identification of candidate genes for prostate cancer-risk SNPs utilizing a normal prostate tissue eQTL data set. Nat. Commun. 2015, 6, 8653. [Google Scholar] [CrossRef]
- Zhang, P.; Xia, J.H.; Zhu, J.; Gao, P.; Tian, Y.J.; Du, M.; Guo, Y.C.; Suleman, S.; Zhang, Q.; Kohli, M.; et al. High-throughput screening of prostate cancer risk loci by single nucleotide polymorphisms sequencing. Nat. Commun. 2018, 9, 2022. [Google Scholar] [CrossRef]
- Boyle, A.P.; Hong, E.L.; Hariharan, M.; Cheng, Y.; Schaub, M.A.; Kasowski, M.; Karczewski, K.J.; Park, J.; Hitz, B.C.; Weng, S.; et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012, 22, 1790–1797. [Google Scholar] [CrossRef]
- Ward, L.D.; Kellis, M. HaploReg: A resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 2012, 40, D930–D934. [Google Scholar] [CrossRef]
- Hazelett, D.J.; Rhie, S.K.; Gaddis, M.; Yan, C.; Lakeland, D.L.; Coetzee, S.G.; Ellipse, G.-O.N.C.; Practical, C.; Henderson, B.E.; Noushmehr, H.; et al. Comprehensive functional annotation of 77 prostate cancer risk loci. PLoS Genet. 2014, 10, e1004102. [Google Scholar] [CrossRef]
- Taberlay, P.C.; Statham, A.L.; Kelly, T.K.; Clark, S.J.; Jones, P.A. Reconfiguration of nucleosome-depleted regions at distal regulatory elements accompanies DNA methylation of enhancers and insulators in cancer. Genome Res. 2014, 24, 1421–1432. [Google Scholar] [CrossRef]
- Chelala, C.; Khan, A.; Lemoine, N.R. SNPnexus: A web database for functional annotation of newly discovered and public domain single nucleotide polymorphisms. Bioinformatics 2009, 25, 655–661. [Google Scholar] [CrossRef]
- Consortium, E.P. An integrated encyclopedia of DNA elements in the human genome. Nature 2012, 489, 57–74. [Google Scholar] [CrossRef]
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef]
- Kel, A.E.; Gossling, E.; Reuter, I.; Cheremushkin, E.; Kel-Margoulis, O.V.; Wingender, E. MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 2003, 31, 3576–3579. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Chang, B.L.; Cramer, S.D.; Wiklund, F.; Isaacs, S.D.; Stevens, V.L.; Sun, J.; Smith, S.; Pruett, K.; Romero, L.M.; Wiley, K.E.; et al. Fine mapping association study and functional analysis implicate a SNP in MSMB at 10q11 as a causal variant for prostate cancer risk. Hum. Mol. Genet. 2009, 18, 1368–1375. [Google Scholar] [CrossRef]
- McLean, C.Y.; Bristor, D.; Hiller, M.; Clarke, S.L.; Schaar, B.T.; Lowe, C.B.; Wenger, A.M.; Bejerano, G. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 2010, 28, 495–501. [Google Scholar] [CrossRef]
- Gao, T.; He, B.; Liu, S.; Zhu, H.; Tan, K.; Qian, J. EnhancerAtlas: A resource for enhancer annotation and analysis in 105 human cell/tissue types. Bioinformatics 2016, 32, 3543–3551. [Google Scholar] [CrossRef]
- Gallagher, M.D.; Chen-Plotkin, A.S. The Post-GWAS Era: From Association to Function. Am. J. Hum. Genet. 2018, 102, 717–730. [Google Scholar] [CrossRef]
- Chang, K.H.; Li, R.; Kuri, B.; Lotan, Y.; Roehrborn, C.G.; Liu, J.; Vessella, R.; Nelson, P.S.; Kapur, P.; Guo, X.; et al. A gain-of-function mutation in DHT synthesis in castration-resistant prostate cancer. Cell 2013, 154, 1074–1084. [Google Scholar] [CrossRef]
- Shafi, A.A.; Yen, A.E.; Weigel, N.L. Androgen receptors in hormone-dependent and castration-resistant prostate cancer. Pharmacol. Ther. 2013, 140, 223–238. [Google Scholar] [CrossRef]
- Bernstein, B.E.; Stamatoyannopoulos, J.A.; Costello, J.F.; Ren, B.; Milosavljevic, A.; Meissner, A.; Kellis, M.; Marra, M.A.; Beaudet, A.L.; Ecker, J.R.; et al. The NIH Roadmap Epigenomics Mapping Consortium. Nat. Biotechnol. 2010, 28, 1045–1048. [Google Scholar] [CrossRef]
- Hellman, L.M.; Fried, M.G. Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions. Nat. Protoc. 2007, 2, 1849–1861. [Google Scholar] [CrossRef]
- Park, P.J. ChIP-seq: Advantages and challenges of a maturing technology. Nat. Rev. Genet. 2009, 10, 669–680. [Google Scholar] [CrossRef]
- Stankiewicz, T.R.; Gray, J.J.; Winter, A.N.; Linseman, D.A. C-terminal binding proteins: Central players in development and disease. Biomol. Concepts 2014, 5, 489–511. [Google Scholar] [CrossRef]
- Thomas, G.; Jacobs, K.B.; Yeager, M.; Kraft, P.; Wacholder, S.; Orr, N.; Yu, K.; Chatterjee, N.; Welch, R.; Hutchinson, A.; et al. Multiple loci identified in a genome-wide association study of prostate cancer. Nat. Genet. 2008, 40, 310–315. [Google Scholar] [CrossRef]
- Zhang, C.; Li, S.; Qiao, B.; Yang, K.; Liu, R.; Ma, B.; Liu, Y.; Zhang, Z.; Xu, Y. CtBP2 overexpression is associated with tumorigenesis and poor clinical outcome of prostate cancer. Arch. Med. Sci. 2015, 11, 1318–1323. [Google Scholar] [CrossRef]
- Zhang, C.; Gao, C.; Xu, Y.; Zhang, Z. CtBP2 could promote prostate cancer cell proliferation through c-Myc signaling. Gene 2014, 546, 73–79. [Google Scholar] [CrossRef]
- Debiais-Delpech, C.; Godet, J.; Pedretti, N.; Bernard, F.X.; Irani, J.; Cathelineau, X.; Cussenot, O.; Fromont, G. Expression patterns of candidate susceptibility genes HNF1beta and CtBP2 in prostate cancer: Association with tumor progression. Urol. Oncol. 2014, 32, 426–432. [Google Scholar] [CrossRef]
- Takayama, K.; Suzuki, T.; Fujimura, T.; Urano, T.; Takahashi, S.; Homma, Y.; Inoue, S. CtBP2 modulates the androgen receptor to promote prostate cancer progression. Cancer Res. 2014, 74, 6542–6553. [Google Scholar] [CrossRef]
- Xuan, Q.; Zhong, X.; Li, W.; Mo, Z.; Huang, Y.; Hu, Y. CtBP2 is associated with angiogenesis and regulates the apoptosis of prostate cancer cells. Oncol. Rep. 2017, 38, 1259–1267. [Google Scholar] [CrossRef]
- Kollara, A.; Brown, T.J. Expression and function of nuclear receptor co-activator 4: Evidence of a potential role independent of co-activator activity. Cell. Mol. Life Sci. 2012, 69, 3895–3909. [Google Scholar] [CrossRef]
- Pomerantz, M.M.; Shrestha, Y.; Flavin, R.J.; Regan, M.M.; Penney, K.L.; Mucci, L.A.; Stampfer, M.J.; Hunter, D.J.; Chanock, S.J.; Schafer, E.J.; et al. Analysis of the 10q11 cancer risk locus implicates MSMB and NCOA4 in human prostate tumorigenesis. PLoS Genet. 2010, 6, e1001204. [Google Scholar] [CrossRef]
- Li, P.; Yu, X.; Ge, K.; Melamed, J.; Roeder, R.G.; Wang, Z. Heterogeneous expression and functions of androgen receptor co-factors in primary prostate cancer. Am. J. Pathol. 2002, 161, 1467–1474. [Google Scholar] [CrossRef]
- Mestayer, C.; Blanchere, M.; Jaubert, F.; Dufour, B.; Mowszowicz, I. Expression of androgen receptor coactivators in normal and cancer prostate tissues and cultured cell lines. Prostate 2003, 56, 192–200. [Google Scholar] [CrossRef]
- Hu, Y.C.; Yeh, S.; Yeh, S.D.; Sampson, E.R.; Huang, J.; Li, P.; Hsu, C.L.; Ting, H.J.; Lin, H.K.; Wang, L.; et al. Functional domain and motif analyses of androgen receptor coregulator ARA70 and its differential expression in prostate cancer. J. Biol. Chem. 2004, 279, 33438–33446. [Google Scholar] [CrossRef]
- Tsui, K.H.; Feng, T.H.; Chung, L.C.; Chao, C.H.; Chang, P.L.; Juang, H.H. Prostate specific antigen gene expression in androgen insensitive prostate carcinoma subculture cell line. Anticancer Res. 2008, 28, 1969–1976. [Google Scholar]
| Risk Regions | Genes | SNPs with eQTL Signal | SNPs LD >0.5 | SNPs Selected |
|---|---|---|---|---|
| chr1:205,657,824-205,857,824 | RAB7L1, NUCKS1 | 155 | 40 | 40 |
| chr2:10,610,730-10,810,730 ** | NOL10 | 368 | 286 | 19 |
| chr2:20,788,265-20,988,265 | C2orf43 | 74 | 13 | 13 |
| chr2:85,677,270-85,894,297 | GGCX | 19 | 19 | 19 |
| chr2:238,287,228-238,543,226 ** | MLPH | 383 | 255 | 9 |
| chr3:87,010,674-87,341,497 | CHMP2B | 73 | 17 | 17 |
| chr4:95,414,609-95,662,877 | BMPR1B | 225 | 166 | 166 |
| chr5:1,795,829-1,995,829 * | IRX4 | 71 | 12 | 71 |
| chr6:41,436,427-41,636,427 | FOXP4 | 54 | 43 | 43 |
| chr6:76,395,882-76,595,882 | MYO6 | 171 | 63 | 63 |
| chr6:153,341,079-153,541,079 | RGS17 | 330 | 76 | 76 |
| chr7:97,716,327-97,916,327 | BHLHA15, LMTK2 | 304 | 75 | 75 |
| chr10:51,424,971-51,649,496 | NCOA4, MSMB | 156 | 135 | 135 |
| chr10:126,596,872-126,796,872 * | CTBP2 | 99 | 25 | 99 |
| chr11:2,133,574-2,333,574 * | ASCL2 | 53 | 53 | 53 |
| chr11:102,301,661-102,501,661 | MMP7 | 69 | 3 | 3 |
| chr12:48,138,757-48,519,618 | COL2A1 | 15 | 15 | 15 |
| chr12:53,173,904-53,373,904 | KRT8, KRT18 | 62 | 49 | 49 |
| chr12:114,585,571-114,785,571 | TBX5 | 3 | 3 | 3 |
| chr14:61,022,526-61,222,526 ** | C14orf39 | 445 | 243 | 2 |
| chr17:518,965-718,965 | VPS53, FAM57A, GEMIN4 | 513 | 60 | 60 |
| chr17:35,974,979-36,201,156 * | HNF1B | 21 | 18 | 21 |
| chr19:38,635,613-38,835,613 | PPP1R14A | 278 | 17 | 17 |
| chr19:41,885,587-42,085,624 | PCAT19 | 50 | 20 | 20 |
| chr20:62,262,563-62,462,563 | LIME1, ZGPAT | 105 | 104 | 104 |
| chrX:9,714,135-9,914,135 | SHROOM2, GPR143 | 62 | 40 | 40 |
| chrX:51,110,057-51,341,672 ** | NUDT11 | 1226 | 362 | 2 |
| chrX:70,307,983-70,507,983 | GJB1 | 59 | 40 | 40 |
| Total SNPs | 5443 | 2252 | 1274 |
| Oligo Amount | Bound/Unbound oligo | C/T Ratio | ||
|---|---|---|---|---|
| C Allele | T Allele | |||
| EMSA | 0.1 pmol | 0.012 | 0.023 | 0.522 |
| 0.5 pmol | 0.008 | 0.012 | 0.667 | |
| SNPs-seq (DHT) | 0.400 | |||
| SNPs-seq (ETH) | 0.378 | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, P.; Tillmans, L.S.; Thibodeau, S.N.; Wang, L. Single-Nucleotide Polymorphisms Sequencing Identifies Candidate Functional Variants at Prostate Cancer Risk Loci. Genes 2019, 10, 547. https://doi.org/10.3390/genes10070547
Zhang P, Tillmans LS, Thibodeau SN, Wang L. Single-Nucleotide Polymorphisms Sequencing Identifies Candidate Functional Variants at Prostate Cancer Risk Loci. Genes. 2019; 10(7):547. https://doi.org/10.3390/genes10070547
Chicago/Turabian StyleZhang, Peng, Lori S. Tillmans, Stephen N. Thibodeau, and Liang Wang. 2019. "Single-Nucleotide Polymorphisms Sequencing Identifies Candidate Functional Variants at Prostate Cancer Risk Loci" Genes 10, no. 7: 547. https://doi.org/10.3390/genes10070547
APA StyleZhang, P., Tillmans, L. S., Thibodeau, S. N., & Wang, L. (2019). Single-Nucleotide Polymorphisms Sequencing Identifies Candidate Functional Variants at Prostate Cancer Risk Loci. Genes, 10(7), 547. https://doi.org/10.3390/genes10070547





