Expansion and Functional Divergence of Inositol Polyphosphate 5-Phosphatases in Angiosperms
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Sources and Homolog Searches
2.2. Phylogenetic Analyses
2.3. Motif and Synteny Analyses
2.4. Expression Analysis
2.5. Calculation of Ka/Ks-Values
3. Results
3.1. Identification of 5PTase Genes in Plants, Animals, and Fungi
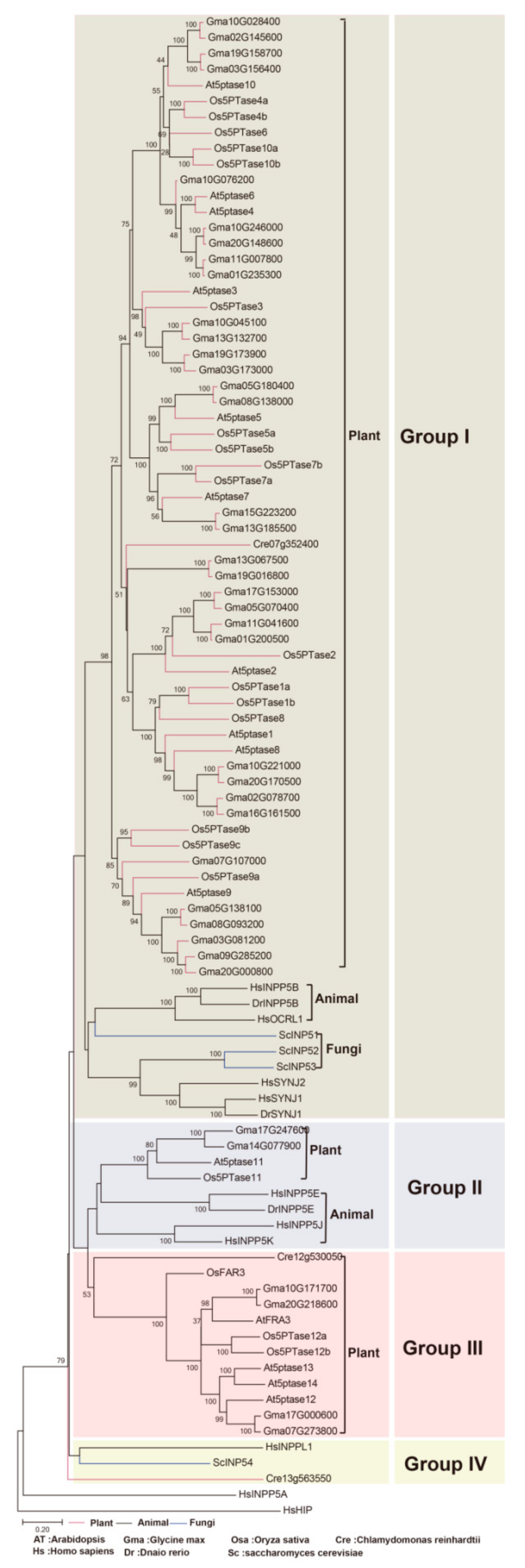
3.2. Phylogenetic Classification of 5PTase Genes into Four Groups
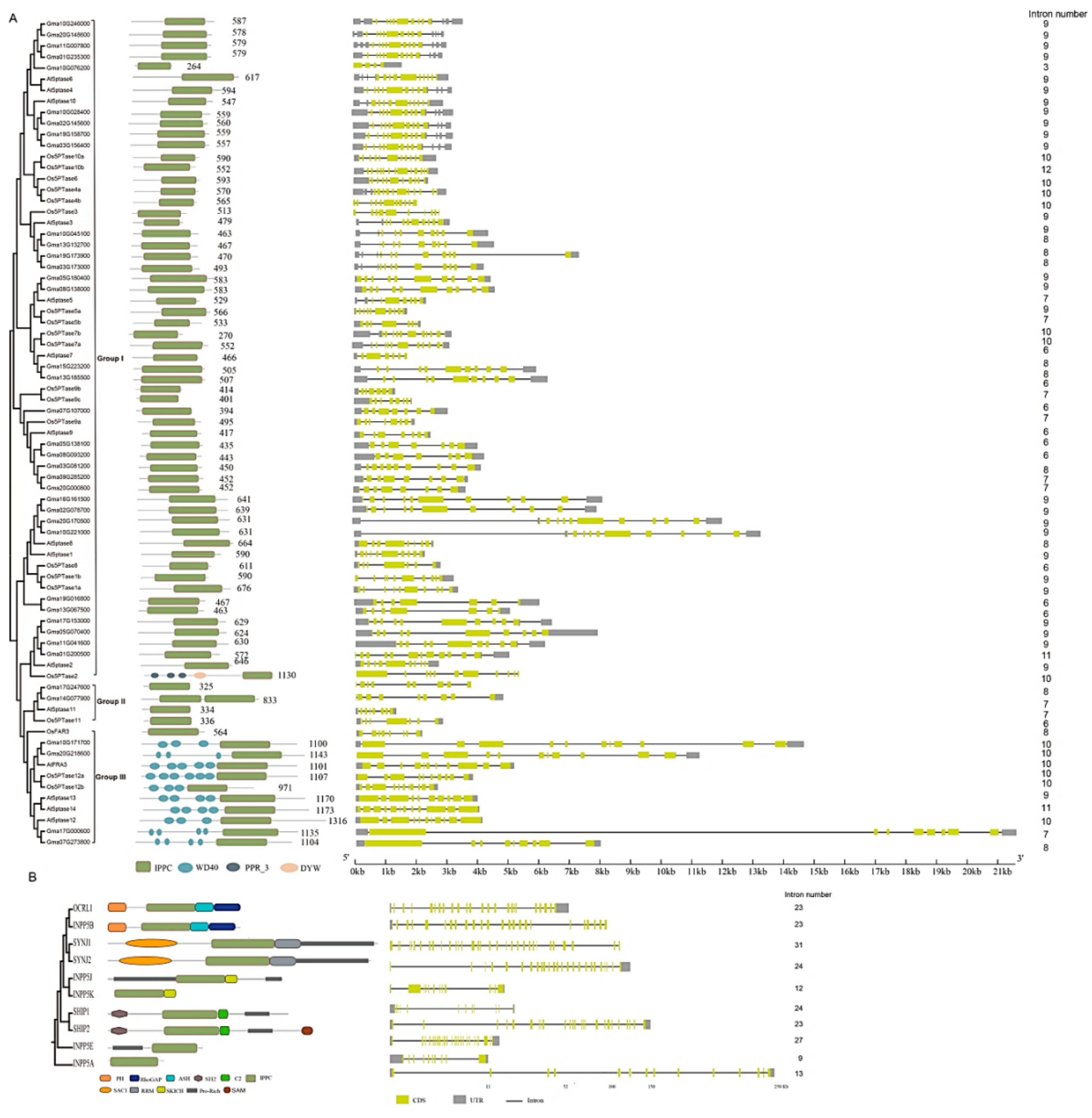
3.3. Domain Features of 5PTase Proteins
3.4. Structural Analysis of 5PTase Family Genes
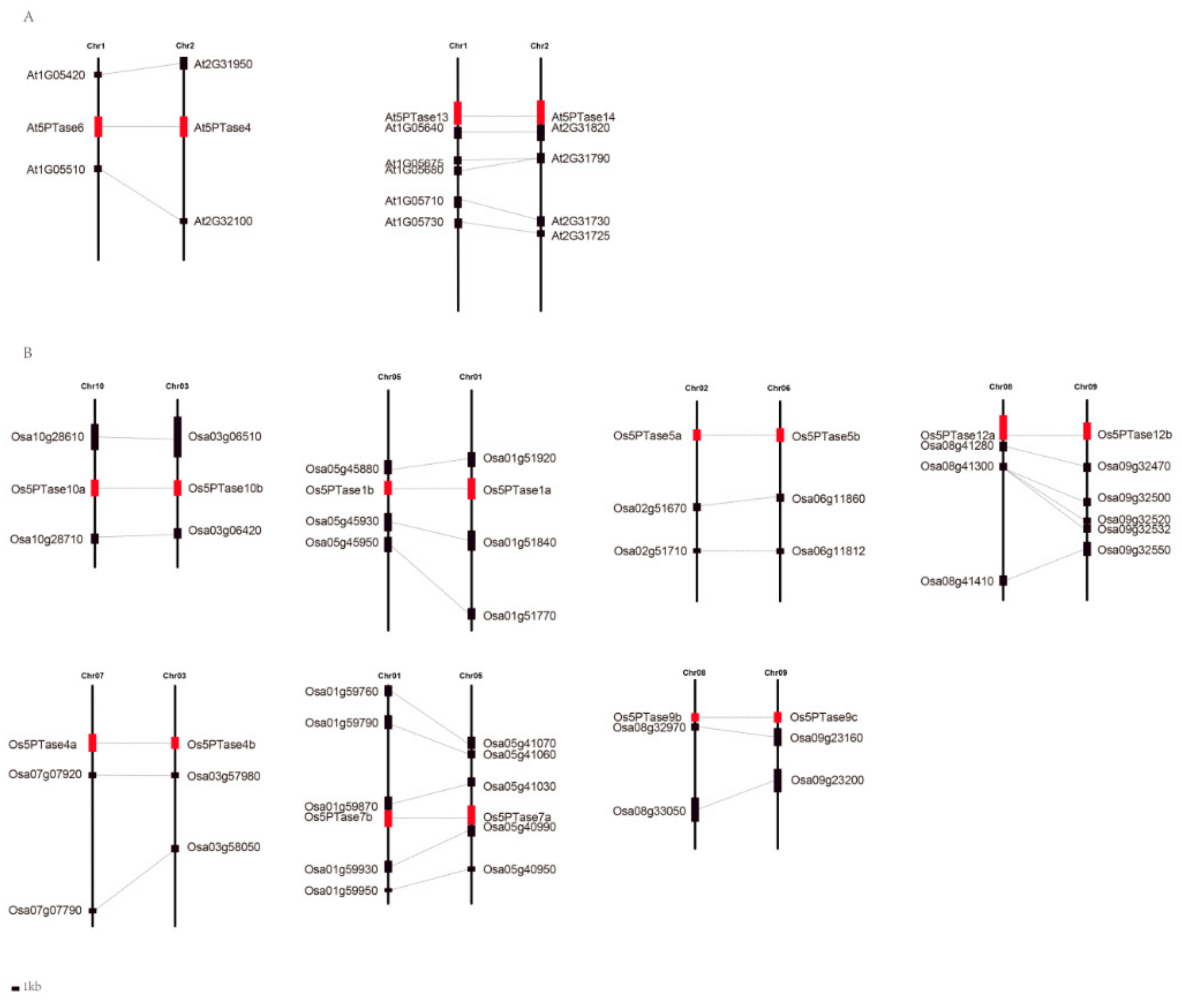
3.5. Identification of Multiple Duplication Events in Land Plant 5PTases
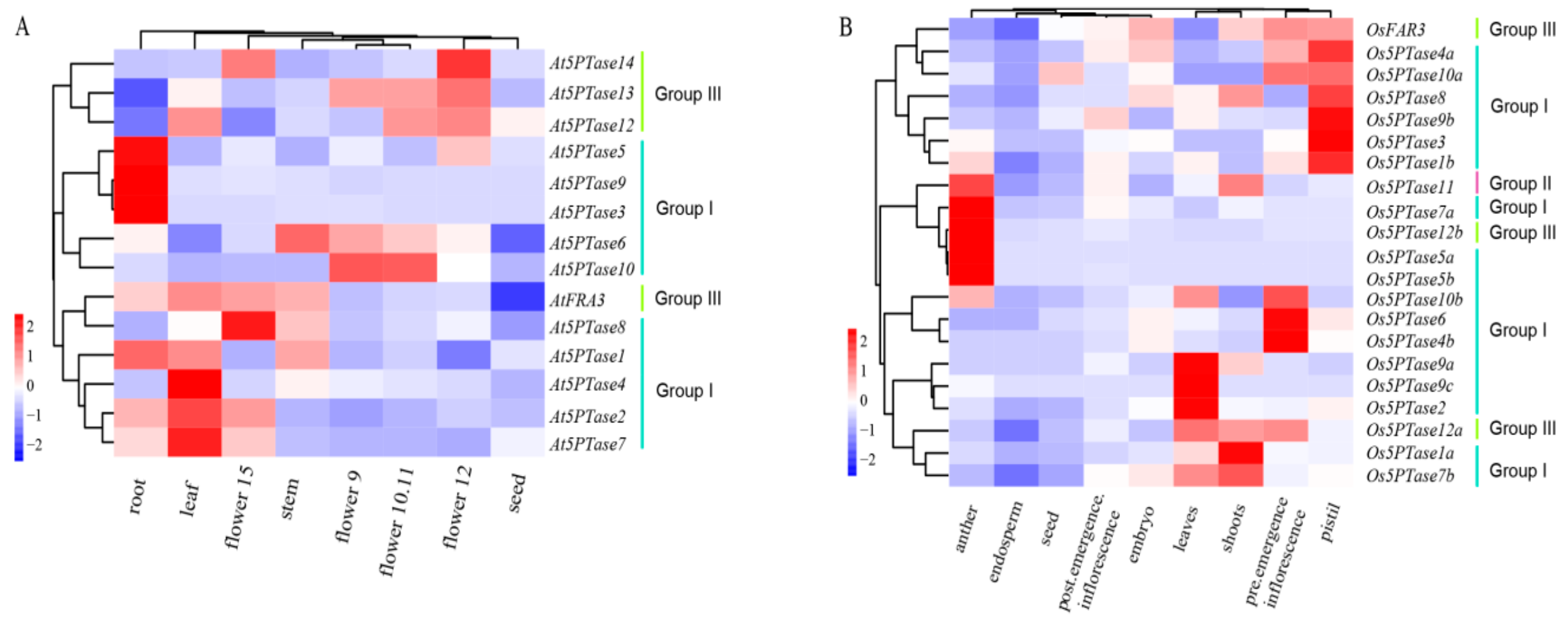
3.6. Expression and Functional Divergence of 5PTase
4. Discussion
4.1. 5PTases Genes Have Been Conserved during Speciation
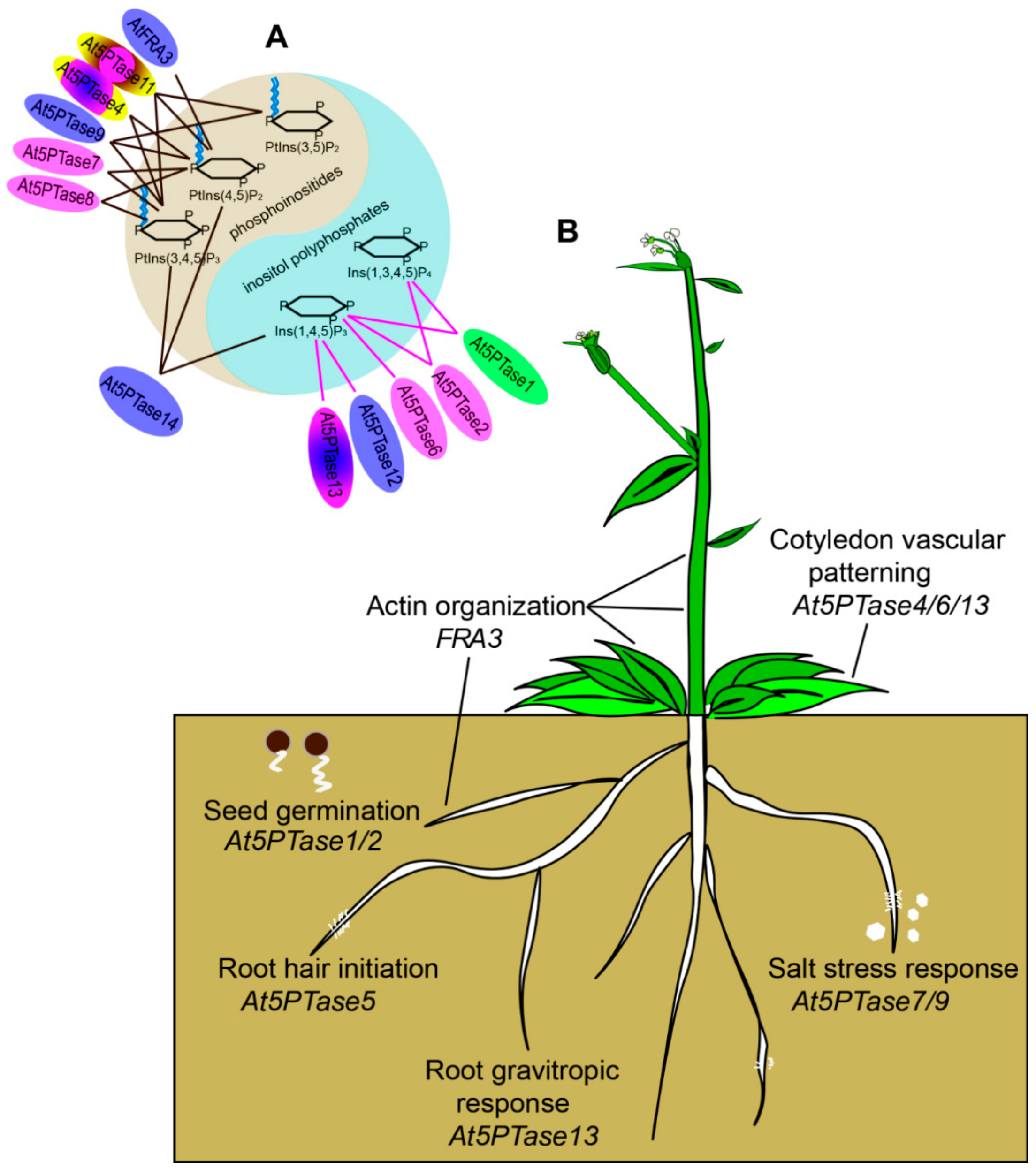
4.2. Functional Divergence in the Evolution of 5PTase Genes
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Erneux, C.; Govaerts, C.; Communi, D.; Pesesse, X. The diversity and possible functions of the inositol polyphosphate 5-phosphatases. Biochim. Biophys. Acta 1998, 1436, 185–199. [Google Scholar] [CrossRef]
- Majerus, P.W.; Kisseleva, M.V.; Norris, F.A. The role of phosphatases in inositol signaling reactions. J. Biol. Chem. 1999, 274, 10669–10672. [Google Scholar] [CrossRef] [PubMed]
- Munnik, T.; Testerink, C. Plant phospholipid signaling: “in a nutshell”. J. Lipid Res. 2009, 50, S260–S265. [Google Scholar] [CrossRef]
- Kerr, W.G.; Colucci, F. Inositol phospholipid signaling and the biology of natural killer cells. J. Innate Immun. 2011, 3, 249–257. [Google Scholar] [CrossRef]
- Tsujishita, Y.; Guo, S.; Stolz, L.E.; York, J.D.; Hurley, J.H. Specificity Determinants in Phosphoinositide Dephosphorylation: Crystal Structure of an Archetypal Inositol Polyphosphate 5-Phosphatase. Cell 2001, 105, 379–389. [Google Scholar] [CrossRef]
- Stevenson, J.M.; Perera, I.Y.; Heilmann, I.I.; Persson, S.; Boss, W.F. Inositol signaling and plant growth. Trends Plant Sci. 2000, 5, 252–258. [Google Scholar] [CrossRef]
- Astle, M.V.; Seaton, G.; Davies, E.M.; Fedele, C.G.; Rahman, P.; Arsala, L.; Mitchell, C.A. Regulation of phosphoinositide signaling by the inositol polyphosphate 5-phosphatases. IUBMB Life 2006, 58, 451–456. [Google Scholar] [CrossRef] [PubMed]
- Xue, H.; Chen, X.; Li, G. Involvement of phospholipid signaling in plant growth and hormone effects. Curr. Opin. Plant Biol. 2007, 10, 483–489. [Google Scholar] [CrossRef]
- Xue, H.W.; Chen, X.; Mei, Y. Function and regulation of phospholipid signalling in plants. Biochem. J. 2009, 421, 145–156. [Google Scholar] [CrossRef]
- Pirruccello, M.; De Camilli, P. Inositol 5-phosphatases: Insights from the Lowe syndrome protein OCRL. Trends Biochem. Sci. 2012, 37, 134–143. [Google Scholar] [CrossRef] [PubMed]
- Berdy, S.E.; Kudla, J.; Gruissem, W.; Gillaspy, G.E. Molecular characterization of At5PTase1, an inositol phosphatase capable of terminating inositol trisphosphate signaling. Plant Physiol. 2001, 126, 801–810. [Google Scholar] [CrossRef] [PubMed]
- Majerus, P.W. Inositols do it all. Genes Dev. 1996, 10, 1051–1053. [Google Scholar] [CrossRef] [PubMed]
- Stolz, L.E.; Huynh, C.V.; Thorner, J.; York, J.D. Identification and characterization of an essential family of inositol polyphosphate 5-phosphatases (INP51, INP52 and INP53 gene products) in the yeast Saccharomyces cerevisiae. Genetics 1998, 148, 1715–1729. [Google Scholar]
- Sarbassov, D.D.; Guertin, D.A.; Ali, S.M.; Sabatini, D.M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 2005, 307, 1098–1101. [Google Scholar] [CrossRef]
- Ai, J.; Maturu, A.; Johnson, W.; Wang, Y.; Marsh, C.B.; Tridandapani, S. The inositol phosphatase SHIP-2 down-regulates FcgammaR-mediated phagocytosis in murine macrophages independently of SHIP-1. Blood 2006, 107, 813–820. [Google Scholar] [CrossRef]
- Majerus, P.W.; York, J.D. Phosphoinositide phosphatases and disease. J. Lipid Res. 2009, 50, S249–S254. [Google Scholar] [CrossRef]
- Zhong, R.; Ye, Z.H. The SAC domain-containing protein gene family in Arabidopsis. Plant Physiol. 2003, 132, 544–555. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Ye, Z.H. Molecular and biochemical characterization of three WD-repeat-domain-containing inositol polyphosphate 5-phosphatases in Arabidopsis thaliana. Plant Cell Physiol. 2004, 45, 1720–1728. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Lee, S.H.; Kim, H.; Jin, J.B.; Kim, D.H.; Hwang, I. A WD40 repeat protein, Arabidopsis Sec13 homolog 1, may play a role in vacuolar trafficking by controlling the membrane association of AtDRP2A. Mol. Cells 2006, 22, 210–219. [Google Scholar]
- Wiradjaja, F.; Ooms, L.M.; Whisstock, J.C.; McColl, B.; Helfenbaum, L.; Sambrook, J.F.; Gething, M.J.; Mitchell, C.A. The yeast inositol polyphosphate 5-phosphatase Inp54p localizes to the endoplasmic reticulum via a C-terminal hydrophobic anchoring tail: Regulation of secretion from the endoplasmic reticulum. J. Biol. Chem. 2001, 276, 7643–7653. [Google Scholar] [CrossRef] [PubMed]
- Bottcher, C.; Wicky, S.; Schwarz, H.; Singer-Kruger, B. Sjl2p is specifically involved in early steps of endocytosis intimately linked to actin dynamics via the Ark1p/Prk1p kinases. Febs Lett. 2006, 580, 633–641. [Google Scholar] [CrossRef]
- Ha, S.A.; Bunch, J.T.; Hama, H.; DeWald, D.B.; Nothwehr, S.F. A novel mechanism for localizing membrane proteins to yeast trans-Golgi network requires function of synaptojanin-like protein. Mol. Biol. Cell 2001, 12, 3175–3190. [Google Scholar] [CrossRef]
- Roth, M.G. Phosphoinositides in constitutive membrane traffic. Physiol. Rev. 2004, 84, 699–730. [Google Scholar] [CrossRef]
- Ooms, L.M.; Horan, K.A.; Rahman, P.; Seaton, G.; Gurung, R.; Kethesparan, D.S.; Mitchell, C.A. The role of the inositol polyphosphate 5-phosphatases in cellular function and human disease. Biochem. J. 2009, 419, 29–49. [Google Scholar] [CrossRef] [PubMed]
- Attree, O.; Olivos, I.M.; Okabe, I.; Bailey, L.C.; Nelson, D.L.; Lewis, R.A.; McInnes, R.R.; Nussbaum, R.L. The Lowe’s oculocerebrorenal syndrome gene encodes a protein highly homologous to inositol polyphosphate-5-phosphatase. Nature 1992, 358, 239–242. [Google Scholar] [CrossRef] [PubMed]
- Hoopes, R.R., Jr.; Shrimpton, A.E.; Knohl, S.J.; Hueber, P.; Hoppe, B.; Matyus, J.; Simckes, A.; Tasic, V.; Toenshoff, B.; Suchy, S.F.; et al. Dent Disease with mutations in OCRL1. Am. J. Hum. Genet. 2005, 76, 260–267. [Google Scholar] [CrossRef]
- Helgason, C.D.; Damen, J.E.; Rosten, P.; Grewal, R.; Sorensen, P.; Chappel, S.M.; Borowski, A.; Jirik, F.; Krystal, G.; Humphries, R.K. Targeted disruption of SHIP leads to hemopoietic perturbations, lung pathology, and a shortened life span. Genes Dev. 1998, 12, 1610–1620. [Google Scholar] [CrossRef] [PubMed]
- Paraiso, K.H.; Ghansah, T.; Costello, A.; Engelman, R.W.; Kerr, W.G. Induced SHIP deficiency expands myeloid regulatory cells and abrogates graft-versus-host disease. J. Immunol. 2007, 178, 2893–2900. [Google Scholar] [CrossRef] [PubMed]
- Suwa, A.; Kurama, T.; Shimokawa, T. SHIP2 and its involvement in various diseases. Expert Opin. Ther. Targets 2010, 14, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Sleeman, M.W.; Wortley, K.E.; Lai, K.M.; Gowen, L.C.; Kintner, J.; Kline, W.O.; Garcia, K.; Stitt, T.N.; Yancopoulos, G.D.; Wiegand, S.J.; et al. Absence of the lipid phosphatase SHIP2 confers resistance to dietary obesity. Nat. Med. 2005, 11, 199–205. [Google Scholar] [CrossRef]
- Clement, S.; Krause, U.; Desmedt, F.; Tanti, J.F.; Behrends, J.; Pesesse, X.; Sasaki, T.; Penninger, J.; Doherty, M.; Malaisse, W.; et al. The lipid phosphatase SHIP2 controls insulin sensitivity. Nature 2001, 409, 92–97. [Google Scholar] [CrossRef]
- Bielas, S.L.; Silhavy, J.L.; Brancati, F.; Kisseleva, M.V.; Al-Gazali, L.; Sztriha, L.; Bayoumi, R.A.; Zaki, M.S.; Abdel-Aleem, A.; Rosti, R.O.; et al. Mutations in INPP5E, encoding inositol polyphosphate-5-phosphatase E, link phosphatidyl inositol signaling to the ciliopathies. Nat. Genet. 2009, 41, 1032–1036. [Google Scholar] [CrossRef]
- Gunesekera, B.; Torabinejad, J.; Robinson, J.; Gillaspy, G.E. Inositol polyphosphate 5-phosphatases 1 and 2 are required for regulating seedling growth. Plant Physiol. 2007, 143, 1408–1417. [Google Scholar] [CrossRef]
- Zhong, R.; Burk, D.H.; Morrison, W.H., 3rd; Ye, Z.H. FRAGILE FIBER3, an Arabidopsis gene encoding a type II inositol polyphosphate 5-phosphatase, is required for secondary wall synthesis and actin organization in fiber cells. Plant Cell 2004, 16, 3242–3259. [Google Scholar] [CrossRef]
- Jones, M.A.; Raymond, M.J.; Smirnoff, N. Analysis of the root-hair morphogenesis transcriptome reveals the molecular identity of six genes with roles in root-hair development in Arabidopsis. Plant J. Cell Mol. Biol. 2006, 45, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Carland, F.M.; Nelson, T. Cotyledon vascular pattern2-mediated inositol (1,4,5) triphosphate signal transduction is essential for closed venation patterns of Arabidopsis foliar organs. Plant Cell 2004, 16, 1263–1275. [Google Scholar] [CrossRef] [PubMed]
- Carland, F.; Nelson, T. CVP2- and CVL1-mediated phosphoinositide signaling as a regulator of the ARF GAP SFC/VAN3 in establishment of foliar vein patterns. Plant J. Cell Mol. Biol. 2009, 59, 895–907. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.H.; Wang, Y.; Mueller-Roeber, B.; Brearley, C.A.; Xu, Z.H.; Xue, H.W. At5PTase13 modulates cotyledon vein development through regulating auxin homeostasis. Plant Physiol. 2005, 139, 1677–1691. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Lin, W.H.; Chen, X.; Xue, H.W. The role of Arabidopsis 5PTase13 in root gravitropism through modulation of vesicle trafficking. Cell Res. 2009, 19, 1191–1204. [Google Scholar] [CrossRef]
- Golani, Y.; Kaye, Y.; Gilhar, O.; Ercetin, M.; Gillaspy, G.; Levine, A. Inositol polyphosphate phosphatidylinositol 5-phosphatase9 (At5ptase9) controls plant salt tolerance by regulating endocytosis. Mol. Plant 2013, 6, 1781–1794. [Google Scholar] [CrossRef]
- Kaye, Y.; Golani, Y.; Singer, Y.; Leshem, Y.; Cohen, G.; Ercetin, M.; Gillaspy, G.; Levine, A. Inositol polyphosphate 5-phosphatase7 regulates the production of reactive oxygen species and salt tolerance in Arabidopsis. Plant Physiol. 2011, 157, 229–241. [Google Scholar] [CrossRef] [PubMed]
- Avila, L.M.; Cerrudo, D.; Swanton, C.; Lukens, L. Brevis plant1, a putative inositol polyphosphate 5-phosphatase, is required for internode elongation in maize. J. Exp. Bot. 2016, 67, 1577–1588. [Google Scholar] [CrossRef]
- Eddy, S.R. Profile hidden Markov models. Bioinformatics 1998, 14, 755–763. [Google Scholar] [CrossRef]
- Finn, R.D.; Miller, B.L.; Clements, J.; Bateman, A. iPfam: A database of protein family and domain interactions found in the Protein Data Bank. Nucleic Acids Res. 2014, 42, D364–D373. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Doerks, T.; Bork, P. SMART 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, D302–D305. [Google Scholar] [CrossRef] [PubMed]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R.; et al. CDD: A Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 2006, 22, 2688–2690. [Google Scholar] [CrossRef]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar]
- Tang, H.; Bowers, J.E.; Wang, X.; Ming, R.; Alam, M.; Paterson, A.H. Synteny and collinearity in plant genomes. Science 2008, 320, 486–488. [Google Scholar] [CrossRef]
- Catchen, J.M.; Conery, J.S.; Postlethwait, J.H. Automated identification of conserved synteny after whole-genome duplication. Genome Res. 2009, 19, 1497–1505. [Google Scholar] [CrossRef] [PubMed]
- Hruz, T.; Laule, O.; Szabo, G.; Wessendorp, F.; Bleuler, S.; Oertle, L.; Widmayer, P.; Gruissem, W.; Zimmermann, P. Genevestigator v3: A reference expression database for the meta-analysis of transcriptomes. Adv. Bioinform. 2008, 2008, 420747. [Google Scholar] [CrossRef]
- Shen, Y.; Zhou, Z.; Wang, Z.; Li, W.; Fang, C.; Wu, M.; Ma, Y.; Liu, T.; Kong, L.A.; Peng, D.L.; et al. Global dissection of alternative splicing in paleopolyploid soybean. Plant Cell 2014, 26, 996–1008. [Google Scholar] [CrossRef]
- McPherson, P.S.; Garcia, E.P.; Slepnev, V.I.; David, C.; Zhang, X.; Grabs, D.; Sossin, W.S.; Bauerfeind, R.; Nemoto, Y.; De Camilli, P. A presynaptic inositol-5-phosphatase. Nature 1996, 379, 353–357. [Google Scholar] [CrossRef]
- Zeng, L.; Zhang, N.; Zhang, Q.; Endress, P.K.; Huang, J.; Ma, H. Resolution of deep eudicot phylogeny and their temporal diversification using nuclear genes from transcriptomic. New Phytol. 2017, 214, 1338–1354. [Google Scholar] [CrossRef] [PubMed]
- Zeng, L.; Zhang, Q.; Sun, R.; Kong, H.; Zhang, N.; Ma, H. Resolution of deep angiosperm phylogeny using conserved nuclear genes and estimates of early divergence times. Nat. Commun. 2014, 5, 4956. [Google Scholar] [CrossRef]
- Huang, C.H.; Sun, R.; Hu, Y.; Zeng, L.; Ma, H. Resolution of Brassicaceae Phylogeny Using Nuclear Genes Uncovers Nested Radiations and Supports Convergent Morphological Evolution. Mol. Biol. Evol. 2015, 33, 394–412. [Google Scholar] [CrossRef] [PubMed]
- Voronov, S.V.; Frere, S.G.; Giovedi, S.; Pollina, E.A.; Borel, C.; Zhang, H.; Schmidt, C.; Akeson, E.C.; Wenk, M.R.; Cimasoni, L.; et al. Synaptojanin 1-linked phosphoinositide dyshomeostasis and cognitive deficits in mouse models of Down’s syndrome. Proc. Natl. Acad. Sci. USA 2008, 105, 9415–9420. [Google Scholar] [CrossRef]
- Suchy, S.F.; Olivos-Glander, I.M.; Nussabaum, R.L. Lowe syndrome, a deficiency of phosphatidylinositol 4,5-bisphosphate 5-phosphatase in the Golgi apparatus. Hum. Mol. Genet. 1995, 4, 2245–2250. [Google Scholar] [CrossRef]
- Zhang, X.; Hartz, P.A.; Philip, E.; Racusen, L.C.; Majerus, P.W. Cell lines from kidney proximal tubules of a patient with Lowe syndrome lack OCRL inositol polyphosphate 5-phosphatase and accumulate phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 1998, 273, 1574–1582. [Google Scholar] [CrossRef]
- Roy, S.W.; Gilbert, W. Rates of intron loss and gain: Implications for early eukaryotic evolution. Proc. Natl. Acad. Sci. USA 2005, 102, 5773–5778. [Google Scholar] [CrossRef]
- Schmutz, J.; Cannon, S.B.; Schlueter, J.; Ma, J.; Mitros, T.; Nelson, W.; Hyten, D.L.; Song, Q.; Thelen, J.J.; Cheng, J.; et al. Genome sequence of the palaeopolyploid soybean. Nature 2010, 463, 178–183. [Google Scholar] [CrossRef]
- Cannon, S.B.; McKain, M.R.; Harkess, A.; Nelson, M.N.; Dash, S.; Deyholos, M.K.; Peng, Y.; Joyce, B.; Stewart, C.N., Jr.; Rolf, M.; et al. Multiple polyploidy events in the early radiation of nodulating and nonnodulating legumes. Mol. Biol. Evol. 2015, 32, 193–210. [Google Scholar] [CrossRef]
- Maere, S.; De Bodt, S.; Raes, J.; Casneuf, T.; Van Montagu, M.; Kuiper, M.; Van de Peer, Y. Modeling gene and genome duplications in eukaryotes. Proc. Natl. Acad. Sci. USA 2005, 102, 5454–5459. [Google Scholar] [CrossRef] [PubMed]
- Van de Peer, Y.; Maere, S.; Meyer, A. The evolutionary significance of ancient genome duplications. Nat. Rev. Genet. 2009, 10, 725–732. [Google Scholar] [CrossRef]
- Lei, L.; Zhou, S.L.; Ma, H.; Zhang, L.S. Expansion and diversification of the SET domain gene family following whole-genome duplications in Populus trichocarpa. BMC Evol. Biol. 2012, 12, 51. [Google Scholar] [CrossRef]
- Lynch, M.; Conery, J.S. The evolutionary fate and consequences of duplicate genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef]
- Ganko, E.W.; Meyers, B.C.; Vision, T.J. Divergence in expression between duplicated genes in Arabidopsis. Mol. Biol. Evol. 2007, 24, 2298–2309. [Google Scholar] [CrossRef] [PubMed]
- Lan, T.; Yang, Z.L.; Yang, X.; Liu, Y.J.; Wang, X.R.; Zeng, Q.Y. Extensive functional diversification of the Populus glutathione S-transferase supergene family. Plant Cell 2009, 21, 3749–3766. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.L.; Liu, H.J.; Wang, X.R.; Zeng, Q.Y. Molecular evolution and expression divergence of the Populus polygalacturonase supergene family shed light on the evolution of increasingly complex organs in plants. New Phytol. 2013, 197, 1353–1365. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Liu, B.; Yu, L.; Feng, D.; Wang, H.; Wang, J. Phylogenetic analysis, structural evolution and functional divergence of the 12-oxo-phytodienoate acid reductase gene family in plants. BMC Evol. Biol. 2009, 9, 90. [Google Scholar] [CrossRef] [PubMed]


| Paralog Pair | Ka | Ks | Ka/Ks | Duplication Date (mya) | Duplication Type | |
|---|---|---|---|---|---|---|
| At5PTase4 | At5PTase6 | 0.06 | 1.01 | 0.06 | 82.75 | WGD or segmental |
| At5PTase13 | At5PTase14 | 0.16 | 0.70 | 0.22 | 57.08 | WGD or segmental |
| Os5PTase1a | Os5PTase1b | 0.41 | 0.22 | 1.87 | 18.03 | WGD or segmental |
| Os5PTase4a | Os5PTase4b | 0.13 | 1.06 | 0.12 | 87.07 | WGD or segmental |
| Os5PTase5a | Os5PTase5b | 0.21 | 2.17 | 0.10 | 178.04 | WGD or segmental |
| Os5PTase7a | Os5PTase7b | 0.20 | 2.10 | 0.09 | 172.52 | WGD or segmental |
| Os5PTase9b | Os5PTase9c | 0.32 | 0.32 | 1.01 | 26.23 | WGD or segmental |
| Os5PTase10a | Os5PTase10b | 0.12 | 1.16 | 0.11 | 95.09 | WGD or segmental |
| Os5PTase12a | Os5PTase12b | 0.15 | 0.80 | 0.19 | 65.88 | WGD or segmental |
| Gma11G041600 | Gma01G200500 | 0.03 | 0.17 | 0.20 | 14.25 | WGD or segmental |
| Gma17G153000 | Gma05G070400 | 0.03 | 0.12 | 0.27 | 9.97 | WGD or segmental |
| Gma02G078700 | Gma16G161500 | 0.03 | 0.08 | 0.34 | 6.41 | WGD or segmental |
| Gma10G221000 | Gma20G170500 | 0.02 | 0.13 | 0.19 | 10.76 | WGD or segmental |
| Gma20G148600 | Gma10G246000 | 0.01 | 0.07 | 0.14 | 6.11 | WGD or segmental |
| Gma11G007800 | Gma10G076200 | 0.01 | 0.13 | 0.07 | 10.46 | WGD or segmental |
| Gma10G028400 | Gma02G145600 | 0.02 | 0.12 | 0.15 | 9.95 | WGD or segmental |
| Gma19G158700 | Gma03G156400 | 0.02 | 0.11 | 0.17 | 9.20 | WGD or segmental |
| Gma15G223200 | Gma13G185500 | 0.02 | 0.09 | 0.18 | 7.57 | WGD or segmental |
| Gma05G180400 | Gma08G138000 | 0.02 | 0.08 | 0.22 | 6.49 | WGD or segmental |
| Gma19G173900 | Gma03G173000 | 0.04 | 0.13 | 0.30 | 10.85 | WGD or segmental |
| Gma10G045100 | Gma13G132700 | 0.03 | 0.14 | 0.21 | 11.49 | WGD or segmental |
| Gma20G000800 | Gma03G081200 | 0.08 | 0.19 | 0.39 | 15.68 | WGD or segmental |
| Gma05G138100 | Gma08G093200 | 0.02 | 0.08 | 0.28 | 6.42 | WGD or segmental |
| Gma13G067500 | Gma19G016800 | 0.01 | 0.12 | 0.11 | 9.68 | WGD or segmental |
| Gma17G000600 | Gma07G273800 | 0.03 | 0.09 | 0.29 | 7.70 | WGD or segmental |
| Gma10G171700 | Gma20G218600 | 0.02 | 0.09 | 0.21 | 7.18 | WGD or segmental |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Z.; Li, Y.; Luo, Z.; Kong, S.; Zhao, Y.; Zhang, C.; Zhang, W.; Yuan, H.; Cheng, L. Expansion and Functional Divergence of Inositol Polyphosphate 5-Phosphatases in Angiosperms. Genes 2019, 10, 393. https://doi.org/10.3390/genes10050393
Zhang Z, Li Y, Luo Z, Kong S, Zhao Y, Zhang C, Zhang W, Yuan H, Cheng L. Expansion and Functional Divergence of Inositol Polyphosphate 5-Phosphatases in Angiosperms. Genes. 2019; 10(5):393. https://doi.org/10.3390/genes10050393
Chicago/Turabian StyleZhang, Zaibao, Yuting Li, Zhaoyi Luo, Shuwei Kong, Yilin Zhao, Chi Zhang, Wei Zhang, Hongyu Yuan, and Lin Cheng. 2019. "Expansion and Functional Divergence of Inositol Polyphosphate 5-Phosphatases in Angiosperms" Genes 10, no. 5: 393. https://doi.org/10.3390/genes10050393
APA StyleZhang, Z., Li, Y., Luo, Z., Kong, S., Zhao, Y., Zhang, C., Zhang, W., Yuan, H., & Cheng, L. (2019). Expansion and Functional Divergence of Inositol Polyphosphate 5-Phosphatases in Angiosperms. Genes, 10(5), 393. https://doi.org/10.3390/genes10050393




