Fluorescent Light Incites a Conserved Immune and Inflammatory Genetic Response within Vertebrate Organs (Danio rerio, Oryzias latipes and Mus musculus)
Abstract
:1. Introduction
2. Materials and Methods
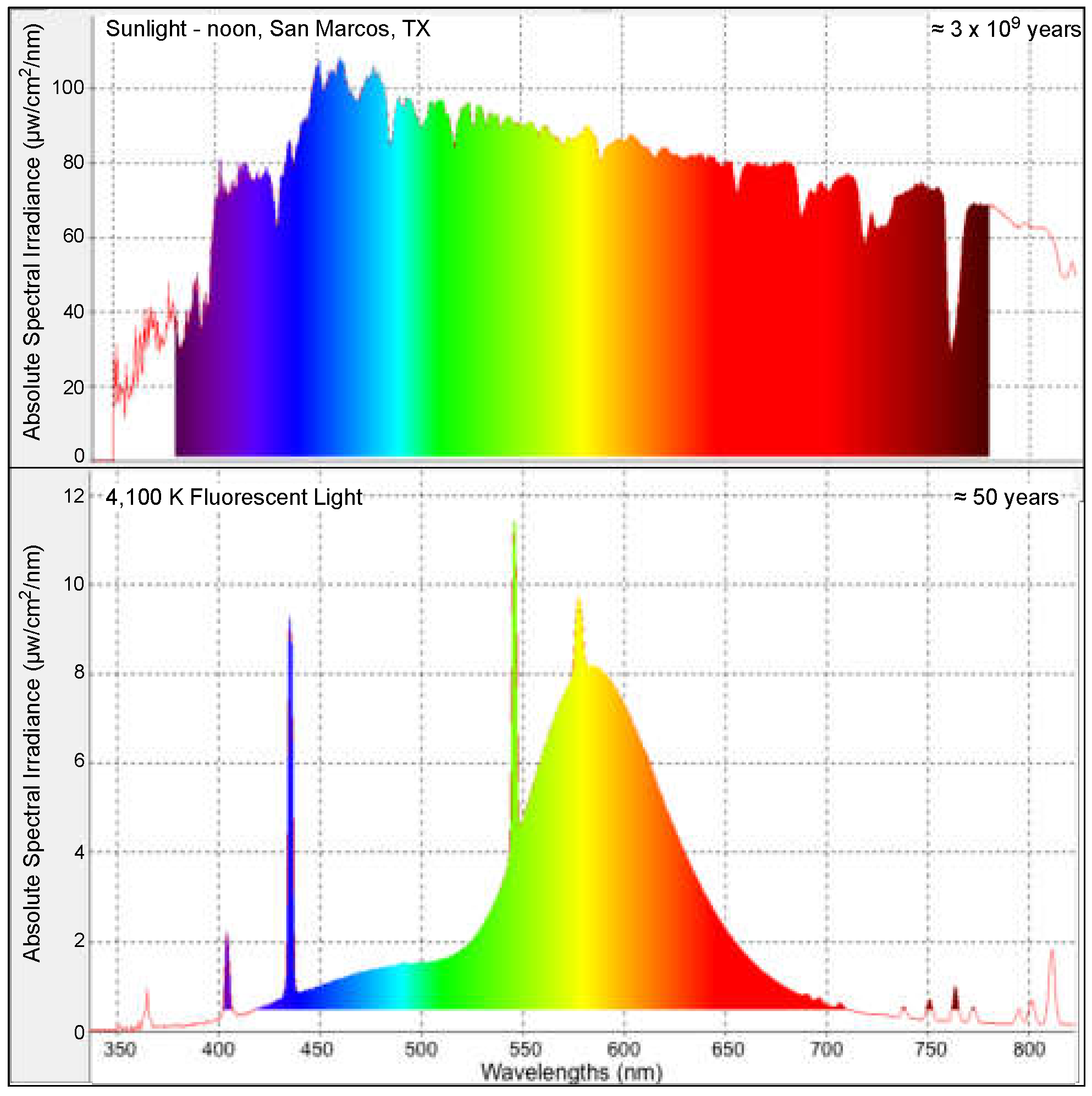
2.1. Fish Utilized and Fluorescent Light Exposure
2.2. Mice Utilized and Fluorescent Light Exposure
2.3. Statement of Animal Use
2.4. RNA Isolation and Sequencing
2.5. Differentially Expressed Gene (DEG) Analysis
2.6. Validation of RNA-Seq Gene Expression Results
3. Results
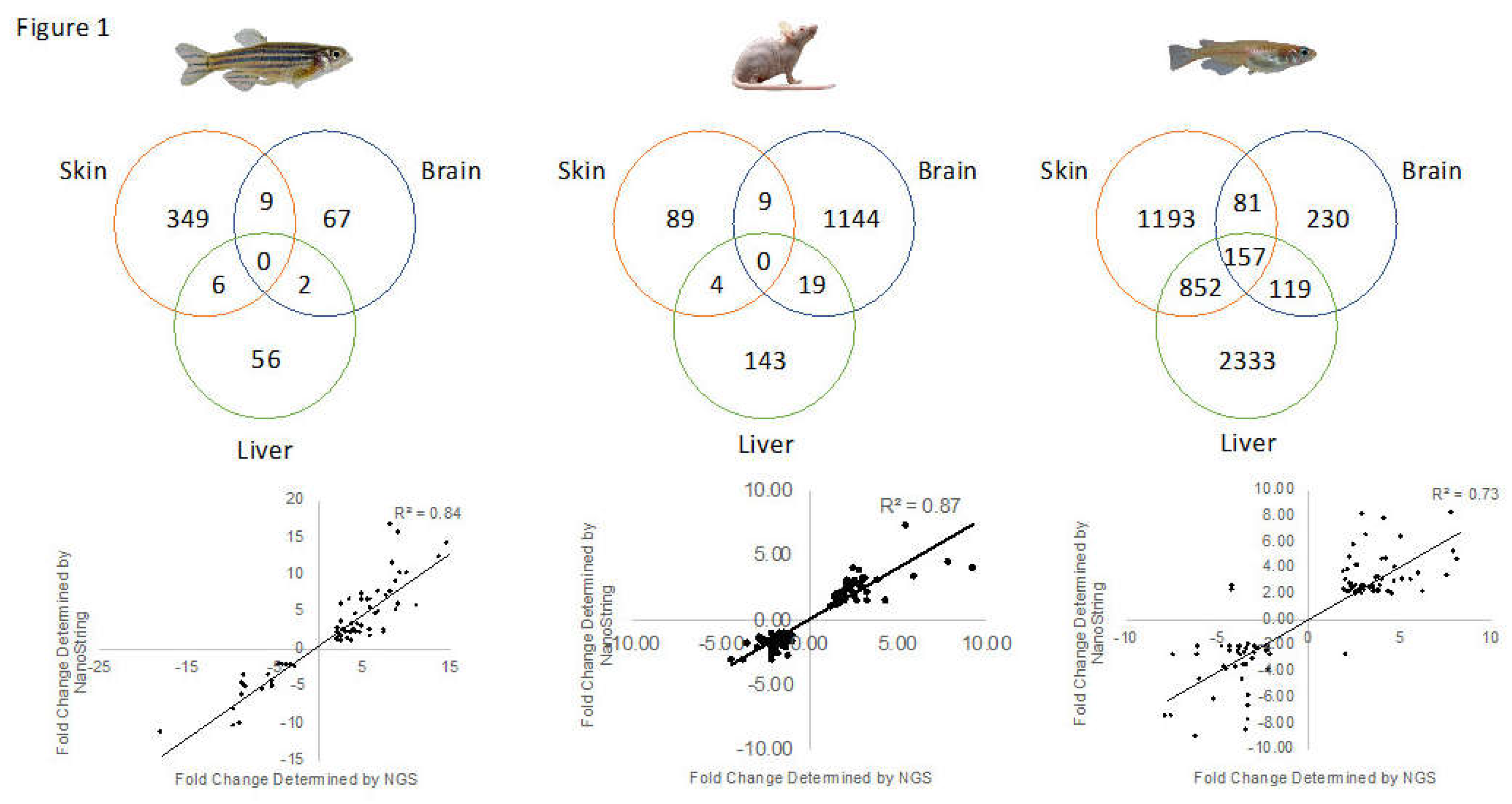
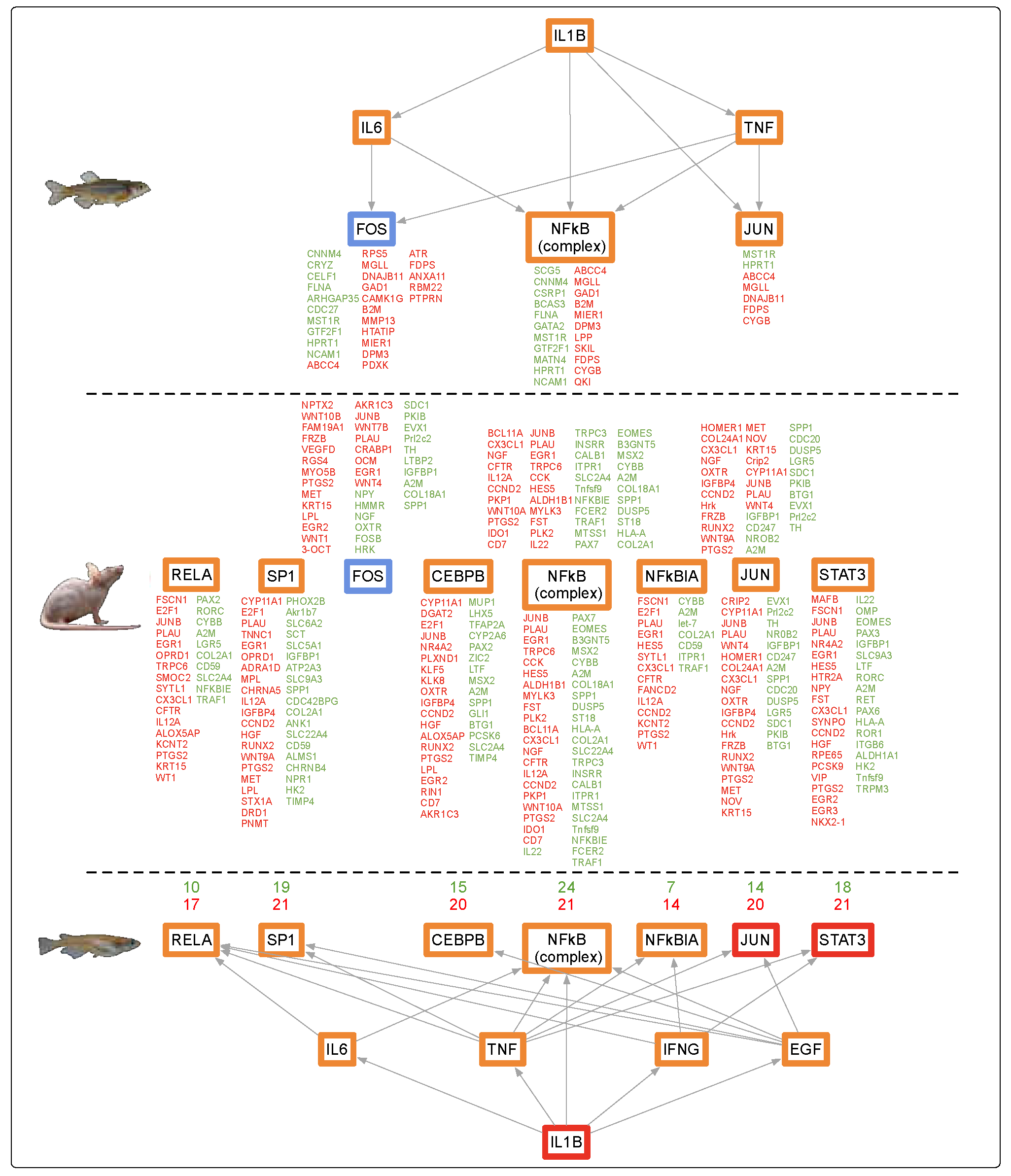
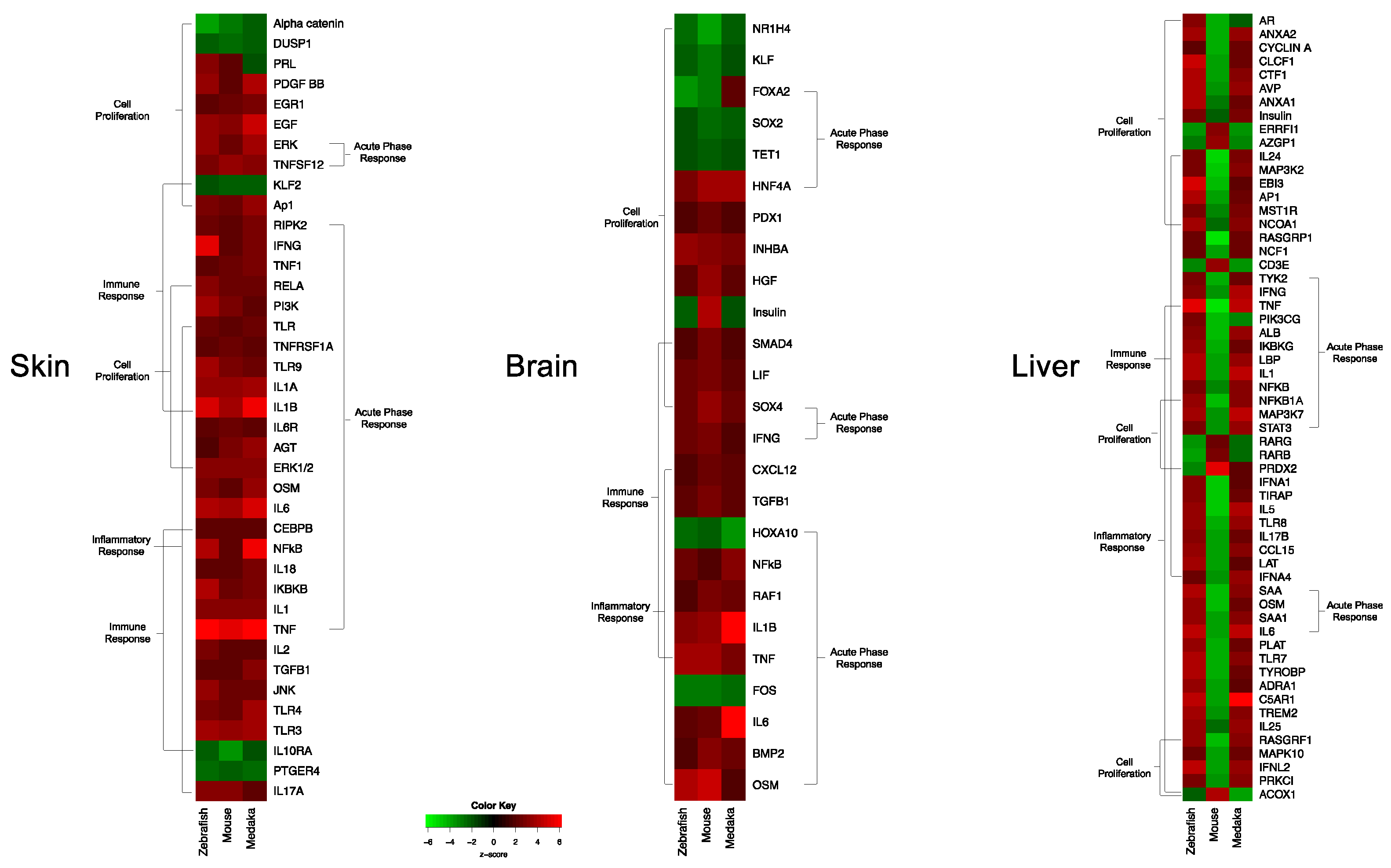
3.1. Genetic Response of Skin to Fluorescent Light
3.2. Genetic Response of Brain to Fluorescent Light
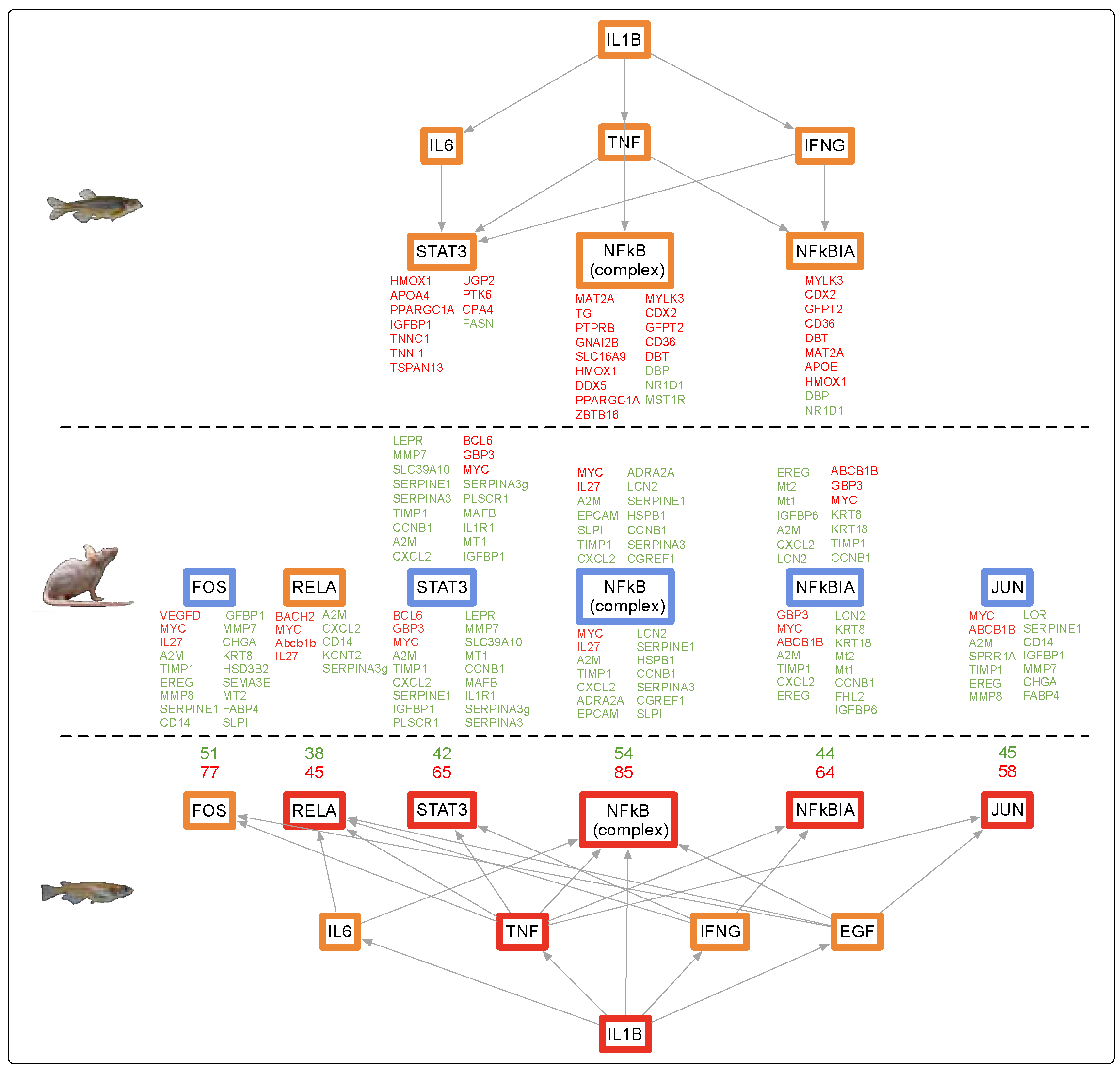
3.3. Genetic Response of Liver to Fluorescent Light
3.4. Conservation Among Fluorescent Light Responses
4. Discussion
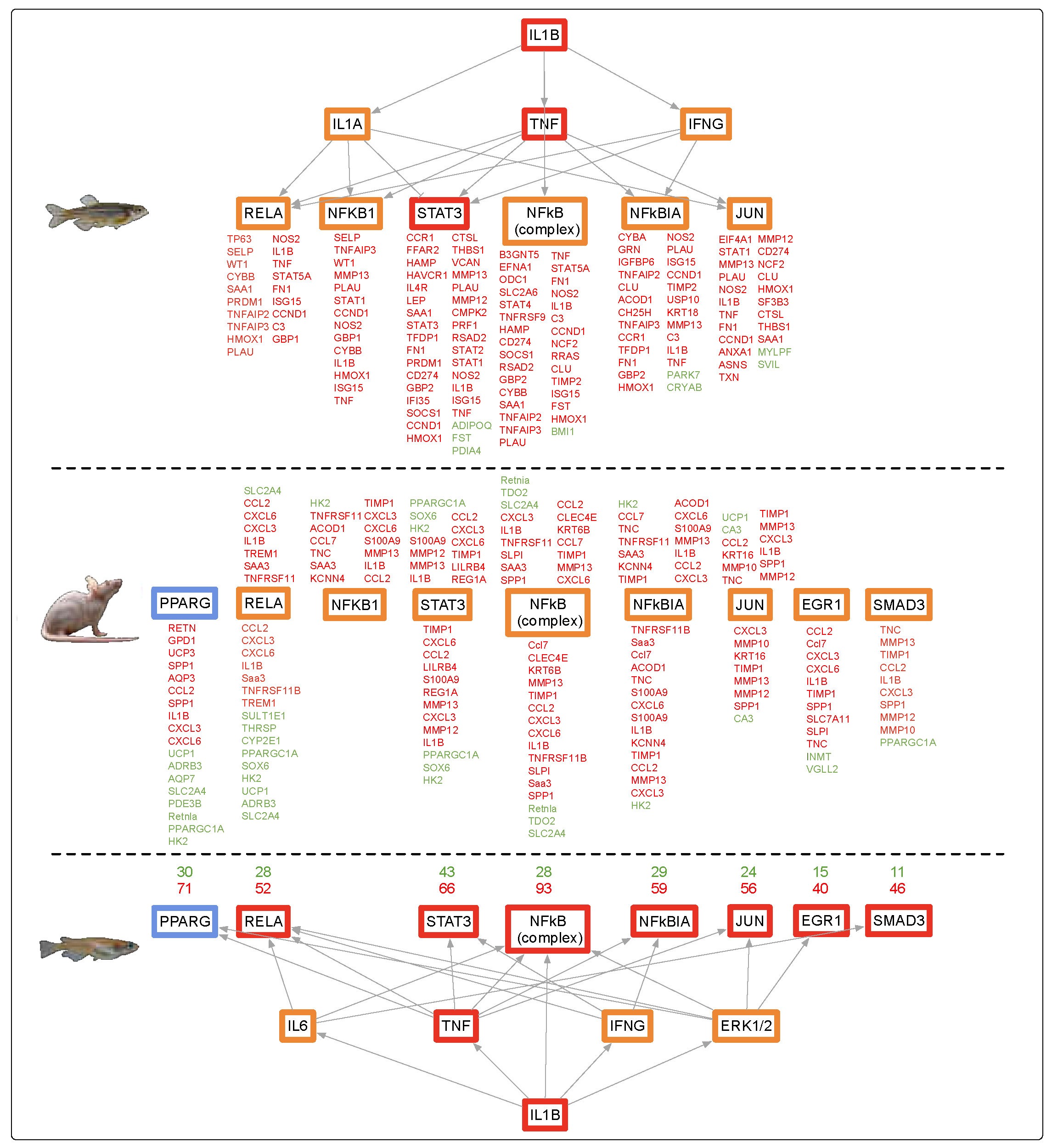
4.1. Alternate Pathway Analysis and Cross Validation
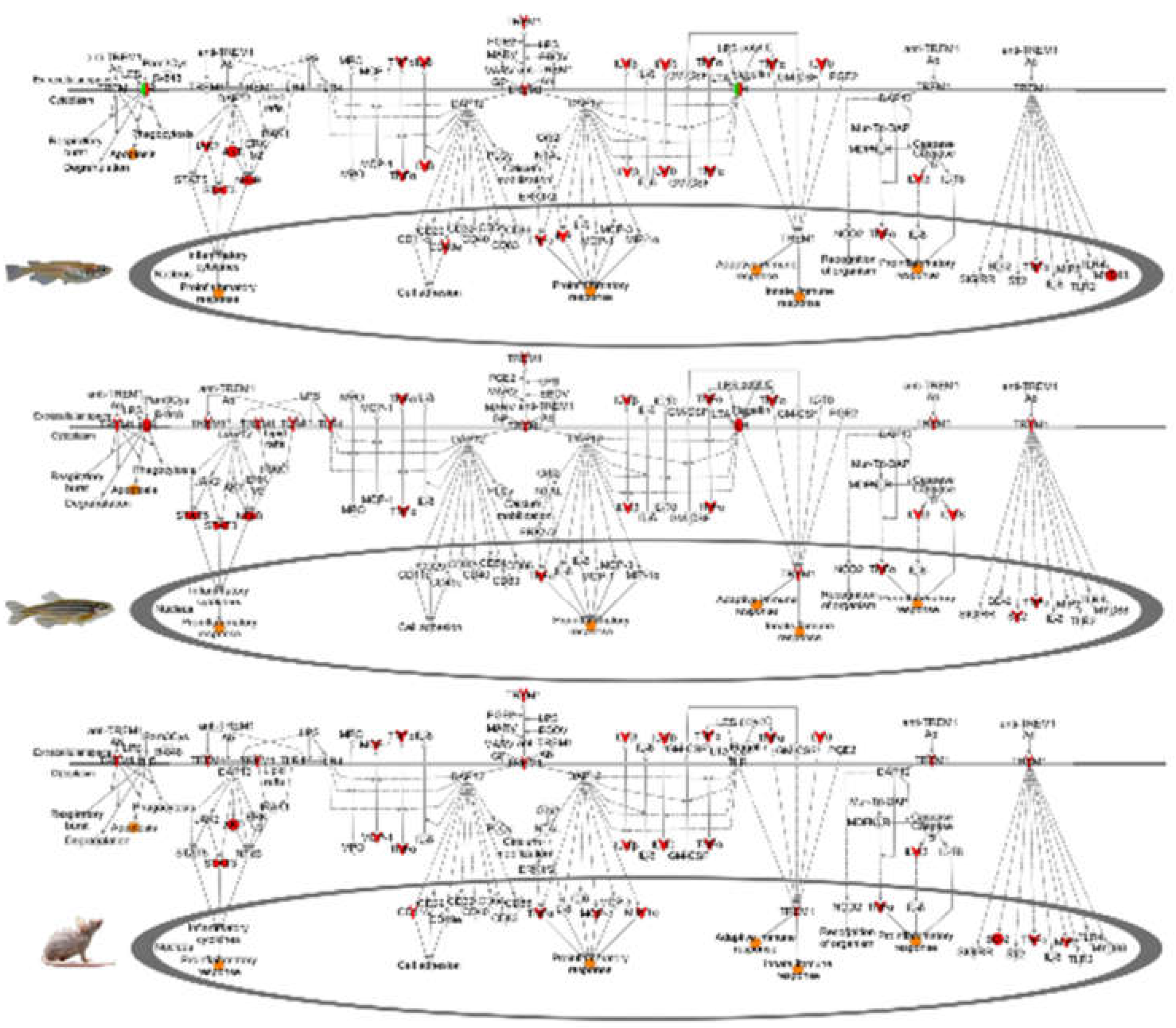
4.2. Fluorescent Light Genetic Response Conservation in the Skin of Vertebrates
4.3. Fluorescent Light Genetic Response Conservation in the Brain of Vertebrates
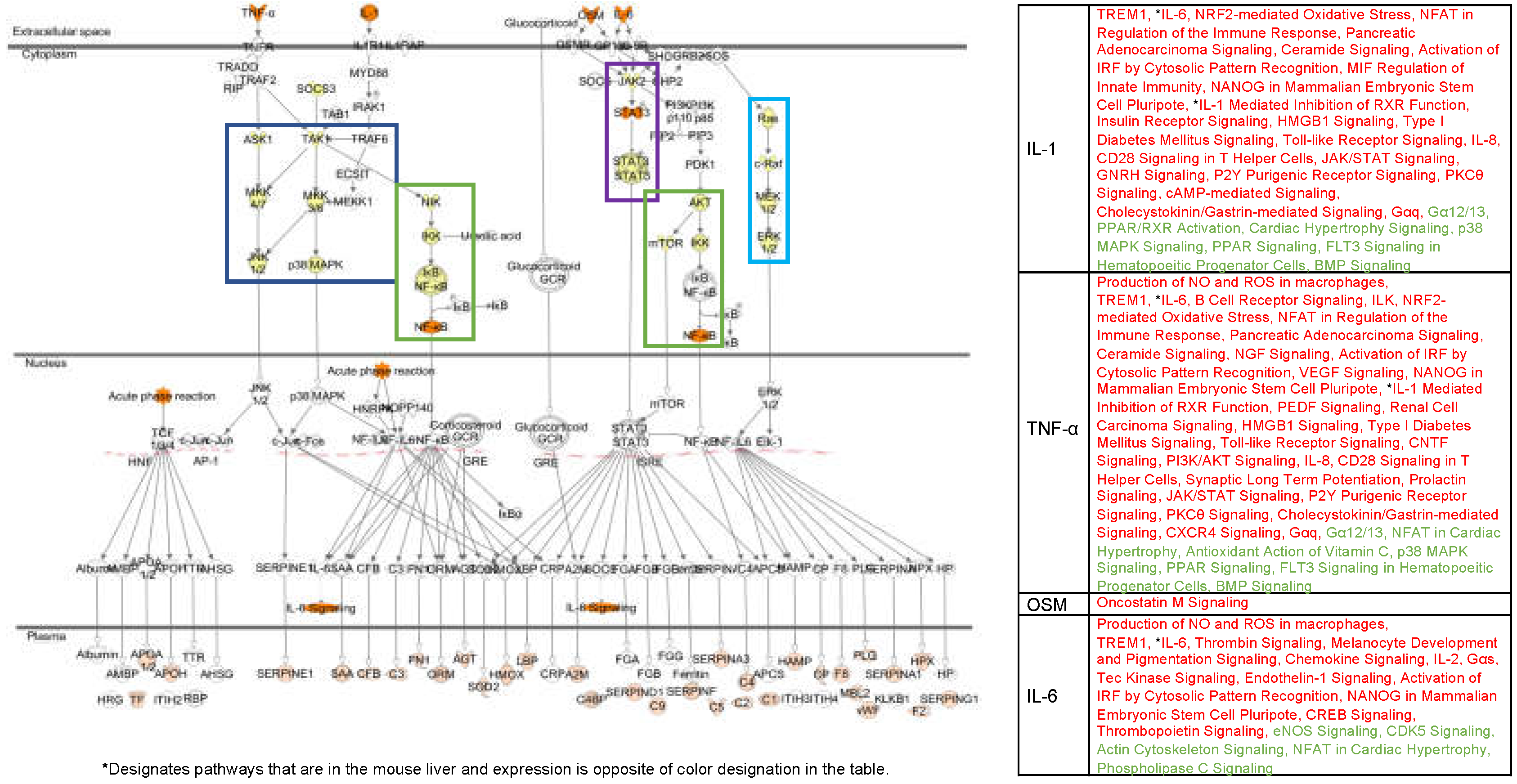
4.4. Fluorescent Light Genetic Response Conservation in the Liver of Vertebrates
4.5. Evolutionary Comparisons Highlighting Transcriptional Activation of the APR
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Boswell, M.; Boswell, W.; Lu, Y.; Savage, M.; Mazurek, Z.; Chang, J.; Muster, J.; Walter, R. The transcriptional response of skin to fluorescent light exposure in viviparous (Xiphophorus) and oviparous (Danio, Oryzias) fishes. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2018, 208, 77–86. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Boswell, M.; Walter, D.J.; Downs, K.P.; Gaston-Pravia, K.; Garcia, T.; Shen, Y.; Mitchell, D.L.; Walter, R.B. UVB-induced gene expression in the skin of Xiphophorus maculatus Jp 163 B. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2014, 163, 86–94. [Google Scholar] [CrossRef]
- Walter, R.B.; Boswell, M.; Chang, J.; Boswell, W.T.; Lu, Y.; Navarro, K.; Walter, S.M.; Walter, D.J.; Salinas, R.; Savage, M. Waveband specific transcriptional control of select genetic pathways in vertebrate skin (Xiphophorus maculatus). BMC Genom. 2018, 19, 355. [Google Scholar] [CrossRef]
- Badia, P.; Myers, B.; Boecker, M.; Culpepper, J.; Harsh, J.R. Bright light effects on body temperature, alertness, EEG and behavior. Physiol. Behav. 1991, 50, 583–588. [Google Scholar] [CrossRef]
- Veitch, J.A.; McColl, S.L. A critical examination of perceptual and cognitive effects attributed to full-spectrum fluorescent lighting. Ergonomics 2001, 44, 255–279. [Google Scholar] [CrossRef]
- Borisuit, A.; Linhart, F.; Scartezzini, J.-L.; Münch, M. Effects of realistic office daylighting and electric lighting conditions on visual comfort, alertness and mood. Light. Res. Technol. 2014, 47, 192–209. [Google Scholar] [CrossRef]
- McColl, S.L.; Veitch, J.A. Full-spectrum fluorescent lighting: A review of its effects on physiology and health. Psychol. Med. 2001, 31, 949–964. [Google Scholar] [CrossRef]
- Münch, M.; Kobialka, S.; Steiner, R.; Oelhafen, P.; Wirz-Justice, A.; Cajochen, C. Wavelength-dependent effects of evening light exposure on sleep architecture and sleep EEG power density in men. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2006, 290, R1421–R1428. [Google Scholar] [CrossRef] [PubMed]
- Scheer, F.A.J.L.; van Doornen, L.J.P.; Buijs, R.M. Light and Diurnal Cycle Affect Human Heart Rate: Possible Role for the Circadian Pacemaker. J. Biol. Rhythms 1999, 14, 202–212. [Google Scholar] [CrossRef] [PubMed]
- Lewy, A.J.; Wehr, T.A.; Goodwin, F.K.; Newsome, D.A.; Markey, S.P. Light suppresses melatonin secretion in humans. Science 1980, 210, 1267–1269. [Google Scholar] [CrossRef] [PubMed]
- Rybnikova, N.A.; Haim, A.; Portnov, B.A. Does artificial light-at-night exposure contribute to the worldwide obesity pandemic? Int. J. Obes. 2016, 40, 815–823. [Google Scholar] [CrossRef]
- Boswell, W.; Boswell, M.; Titus, J.; Savage, M.; Lu, Y.; Shen, J.; Walter, R.B. Sex-specific molecular genetic response to UVB exposure in Xiphophorus maculatus skin. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2015, 178, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, T.J.; Lu, Y.; Boswell, M.; Boswell, W.; Medrano, G.; Walter, S.; Ellis, S.; Savage, M.; Varga, Z.M.; Lawrence, C.; et al. Fluorescent light exposure incites acute and prolonged immune responses in zebrafish (Danio rerio) skin. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2018, 208, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Garcia, T.I.; Shen, Y.; Crawford, D.; Oleksiak, M.F.; Whitehead, A.; Walter, R.B. RNA-Seq reveals complex genetic response to Deepwater Horizon oil release in Fundulus grandis. BMC Genom. 2012, 13, 474. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Oliveros, J.C. VENNY. An Interactive Tool for Comparing Lists with Venn Diagrams. BioinfoGP of CNB-CSIC. 2007. Available online: http://bioinfogp.cnnb.csic.es/tools/venny/index.ht (accessed on 5 December 2017).
- Lu, Y.; Boswell, W.; Boswell, M.; Klotz, B.; Kneitz, S.; Regneri, J.; Savage, M.; Mendoza, C.; Postlethwait, J.; Warren, W.C.; et al. Application of the Transcriptional Disease Signature (TDSs) to Screen Melanoma-Effective Compounds in a Small Fish Model. Sci. Rep. 2019, 9, 530. [Google Scholar] [CrossRef]
- Geiss, G.K.; Bumgarner, R.E.; Birditt, B.; Dahl, T.; Dowidar, N.; Dunaway, D.L.; Fell, H.P.; Ferree, S.; George, R.D.; Grogan, T.; et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 2008, 26, 317–325. [Google Scholar] [CrossRef]
- Boswell, W.T.; Boswell, M.; Walter, D.J.; Navarro, K.L.; Chang, J.; Lu, Y.; Savage, M.G.; Shen, J.; Walter, R.B. Exposure to 4100 K fluorescent light elicits sex specific transcriptional responses in Xiphophorus maculatus skin. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2018, 208, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Walter, R.B.; Walter, D.J.; Boswell, W.T.; Caballero, K.L.; Boswell, M.; Lu, Y.; Chang, J.; Savage, M.G. Exposure to fluorescent light triggers down regulation of genes involved with mitotic progression in Xiphophorus skin. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2015, 178, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Lu, Y.; Boswell, W.T.; Boswell, M.; Caballero, K.L.; Walter, R.B. Molecular genetic response to varied wavelengths of light in Xiphophorus maculatus skin. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2015, 178, 104–115. [Google Scholar] [CrossRef]
- Lu, Y.; Reyes, J.; Walter, S.; Gonzalez, T.; Medrano, G.; Boswell, M.; Boswell, W.; Savage, M.; Walter, R. Characterization of basal gene expression trends over a diurnal cycle in Xiphophorus maculatus skin, brain and liver. Comp. Biochem. Physiol. Part C Toxicol. Pharmacol. 2018, 208, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, K.M.; Saunders, L.B.; Simmons, J.K.; Leon, E.; Calabrese, D.R.; Zhang, S.; Michalowski, A.; Gareiss, P.; Mock, B.A.; Schneekloth, J.S., Jr. Small Molecule Microarrays Enable the Identification of a Selective, Quadruplex-Binding Inhibitor of MYC Expression. ACS Chem. Biol. 2016, 11, 139–148. [Google Scholar] [CrossRef]
- Cray, C.; Zaias, J.; Altman, N.H. Acute phase response in animals: A review. Comp. Med. 2009, 59, 517–526. [Google Scholar]
- Forn-Cuní, G.; Varela, M.; Pereiro, P.; Novoa, B.; Figueras, A. Conserved gene regulation during acute inflammation between zebrafish and mammals. Sci. Rep. 2017, 7, 41905. [Google Scholar] [CrossRef]
- Kleczkowski, M.; Kluciński, W.; Czerski, M.; Kudyba, E. Association between acute phase response, oxidative status and mastitis in cows. Vet. Stanica 2017, 48, 177–186. [Google Scholar]
- Johar, D.; Roth, J.C.; Bay, G.H.; Walker, J.N.; Kroczak, T.J.; Los, M. Inflammatory response, reactive oxygen species, programmed (necrotic-like and apoptotic) cell death and cancer. Roczniki Akademii Medycznej w Bialymstoku 2004, 49, 31–39. [Google Scholar]
- Furuntani-Seiki, M.; Wittbrodt, J. Medaka and zebrafish, an evolutionary twin study. Mech. Dev. 2004, 121, 629–637. [Google Scholar] [CrossRef]
- Tena, J.J.; González-Aguilera, C.; Fernández-Miñán1, A.; Vázquez-Marín, J.; Parra-Acero, H.; Cross, J.W.; Rigby, P.W.J.; Carvajal, J.J.; Wittbrodt, J.J.; Gómez-Skarmeta, J.L.; et al. Comparative epigenomics in distantly related teleost species identifies conserved cis-regulatory nodes active during the vertebrate phylotypic period. Genome Res. 2014, 24, 1075–1085. [Google Scholar] [CrossRef]
- Shima, A.; Himmelbauer, H.; Mitani, H.; Furutani-Seiki, M.; Wittbrodt, J.; Schartl, M. Fish genomes flying. EMBO Rep. 2013, 4, 121–125. [Google Scholar] [CrossRef]
- Peterson, K.J.; Lyons, J.B.; Nowak, K.S.; Takacs, C.M.; Wargo, M.J.; McPeek, M.A. Estimating metazoan divergence times with a molecular clock. Proc. Natl. Acad. Sci. USA 2004, 101, 6536–6541. [Google Scholar] [CrossRef]
- Nobraga, M.A.; Pennacchio, L.A. Comparative genomic analysis as a tool for biological discovery. J. Physiol. 2003, 554, 31–39. [Google Scholar] [CrossRef]
- Kamburov, A.; Wierling, C.; Lehrach, H.; Herwig, R. ConsensusPathDB—A database for integrating human functional interaction networks. Nucleic Acids Res. 2009, 37, D623–D628. [Google Scholar] [CrossRef]
- Kamburov, A.; Pentchev, K.; Galicka, H.; Wierling, C.; Lehrach, H.; Herwig, R. ConsensusPathDB: Toward a more complete picture of cell biology. Nucleic Acids Res. 2011, 39, 12–17. [Google Scholar] [CrossRef]
- Pang, X.; Zhang, K.; Huang, J.; Wang, H.; Gao, L.; Wang, T.; Sun, Y.; Chen, L.; Wang, J. Decryption of Active Constituents and Action Mechanism of the Traditional Uighur Prescription (BXXTR) Alleviating IMQ-Induced Psoriasis-Like Skin Inflammation in BALB/c Mice. Int. J. Mol. Sci. 2018, 19, 1822. [Google Scholar] [CrossRef]
- Yoo, J.-Y.; Desiderio, S. Innate and acquired immunity intersect in a global view of the acute-phase response. Proc. Natl. Acad. Sci. USA 2003, 100, 1157–1162. [Google Scholar] [CrossRef]
- Yang, D.; Liu, Q.; Yang, M.; Wu, H.; Wang, Q.; Xiao, J.; Zhang, Y. RNA-seq liver transcriptome analysis reveals an activated MHC-I pathway and an inhibited MHC-II pathway at the early stage of vaccine immunization in zebrafish. BMC Genomics 2012, 13, 319. [Google Scholar] [CrossRef]
- Flutter, B.; Nestle, F.O. TLRs to cytokines: Mechanistic insights from the imiquimod mouse model of psoriasis. Eur. J. Immunol. 2013, 43, 3138–3146. [Google Scholar] [CrossRef]
- Wada, Y.; Cardinale, I.; Khatcherian, A.; Chu, J.; Kantor, A.B.; Gottlieb, A.B.; Tatsuta, N.; Jacobson, E.; Barsoum, J.; Krueger, J.G. Apilimod inhibits the production of IL-12 and IL-23 and reduces dendritic cell infiltration in psoriasis. PLoS ONE 2012, 7, e35069. [Google Scholar] [CrossRef]
- Beigel, F.; Friedrich, M.; Probst, C.; Sotlar, K.; Göke, B.; Diegelmann, J.; Brand, S. Oncostatin M mediates STAT3-dependent intestinal epithelial restitution via increased cell proliferation, decreased apoptosis and upregulation of SERPIN family members. PLoS ONE 2014, 9, e93498. [Google Scholar] [CrossRef]
- Matthews, V.B.; Knight, B.; Tirnitz-Parker, J.E.E.; Boon, J.; Olynyk, J.K.; Yeoh, G.C.T. Oncostatin M induces an acute phase response but does not modulate the growth or maturation-status of liver progenitor (oval) cells in culture. Exp. Cell Res. 2005, 306, 252–263. [Google Scholar] [CrossRef]
- Pohin, M.; Guesdon, W.; Mekouo, A.A.T.; Rabeony, H.; Paris, I.; Atanassov, H.; Favot, L.; Mcheik, J.; Bernard, F.X.; Richards, C.D.; et al. Oncostatin M overexpression induces skin inflammation but is not required in the mouse model of imiquimod-induced psoriasis-like inflammation. Eur. J. Immunol. 2016, 46, 1737–1751. [Google Scholar] [CrossRef] [PubMed]
- Böing, I.; Stross, C.; Radtke, S.; Lippok, B.E.; Heinrich, P.C.; Hermanns, H.M. Oncostatin M-induced activation of stress-activated MAP kinases depends on tyrosine 861 in the OSM receptor and requires Jak1 but not Src kinases. Cell Signal. 2006, 18, 50–61. [Google Scholar] [CrossRef]
- Frost, R.A.; Nystrom, G.J.; Lang, C.H. Lipopolysaccharide regulates proinflammatory cytokine expression in mouse myoblasts and skeletal muscle. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2002, 283, R698–R709. [Google Scholar] [CrossRef] [PubMed]
- Van Der Bruggen, T.; Nijenhuis, S.; Van Raaij, E.; Verhoef, J.; Van Asbeck, B.S. Lipopolysaccharide-induced tumor necrosis factor α production by human monocytes involves the Raf-1/MEK1-MEK2/ERK1-ERK2 pathway. Infect. Immun. 1999, 67, 3824–3829. [Google Scholar] [PubMed]
- Rizzo, R.R.; Musella, A.; De Vito, F.; Fresegna, D.; Bullitta, S.; Vanni, V.; Guadalupi, L.; Bassi, M.S.; Buttari, F.; Mandolesi, G.; et al. Tumor necrosis factor and interleukin-1B modulate synaptic plasticity during neuroinflammation. Neural Plast. 2018, 2018, 8430123. [Google Scholar] [CrossRef] [PubMed]
- Willard, S.S.; Koochekpour, S. Glutamate, glutamate receptors, and downstream signaling pathways. Int. J. Biol. Sci. 2013, 9, 948–959. [Google Scholar] [CrossRef]
- Hadley, D.; Wu, Z.; Kao, C.; Kini, A.; Mohamed-Hadley, A.; Thomas, K.; Vazquez, L.; Qiu, H.; Mentch, F.; Pellegrino, R.; et al. The impact of the metabotropic glutamate receptor and other gene family interaction networks on autism. Nat. Commun. 2014, 5, 4074. [Google Scholar] [CrossRef]
- Lüscher, C.; Malenka, R.C. NMDA receptor-dependent long-term potentiation and long-term depression (LTP/LTD). Cold Spring Harb. Perspect. Biol. 2012, 4, a005710. [Google Scholar] [CrossRef] [PubMed]
- Byrnes, K.R.; Stoica, B.; Loane, D.J.; Riccio, A.; Davis, M.I.; Faden, A.I. MGLUR5 activation inhibits micobial associated inflammation and neurotoxicity. Glia 2010, 57, 550–560. [Google Scholar] [CrossRef]
- Morrison, R.S.; Kinoshita, Y.; Johnson, M.D.; Ghatan, S.; Ho, J.T.; Garden, G. Neuronal survival and cell death signaling pathways. Adv. Exp. Med. Biol. 2002, 513, 41–86. [Google Scholar]
- Dong, X.X.; Wang, Y.; Qin, Z.H. Molecular mechanisms of excitotoxicity and their relevance to pathogenesis of neurodegenerative diseases. Acta Pharmacol. Sin. 2009, 30, 379–387. [Google Scholar] [CrossRef]
- Fonken, L.K.; Nelson, R.J. The effects of light at night on circadian clocks and metabolism. Endocr. Rev. 2014, 35, 648–670. [Google Scholar] [CrossRef]
- Delezie, J.; Challet, E. Interactions between metabolism and circadian clocks: Reciprocal disturbances. Ann. N. Y. Acad. Sci. 2011, 1243, 30–46. [Google Scholar] [CrossRef]
- Gnocchi, D.; Pedrelli, M.; Hurt-Camejo, E.; Parini, P. Lipids around the Clock: Focus on Circadian Rhythms and Lipid Metabolism. Biology 2015, 4, 104–132. [Google Scholar] [CrossRef]
- Shuboni, D.D.; Cramm, S.; Yan, L.; Nunez, A.A.; Smale, L. Acute behavioral responses to light and darkness in nocturnal Mus musculus and diurnal Arvicanthis niloticus. J. Biol. Rhythms 2012, 27, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Young, M.E.; McGinnis, G.R.; Young, M.E. Circadian regulation of metabolic homeostasis: Causes and consequences. Nat. Sci. Sleep 2016, 8, 163–180. [Google Scholar] [CrossRef] [PubMed]
- Eckel-Mahan, K.; Sassone-Corsi, P. Metabolism and the Circadian Clock Converge. Physiol. Rev. 2013, 93, 107–135. [Google Scholar] [CrossRef]
- Kalsbeek, A.; La Fleur, S.; Fliers, E. Circadian control of glucose metabolism. Mol. Metab. 2014, 3, 372–383. [Google Scholar] [CrossRef] [PubMed]
- Kurose, T.; Yabe, D.; Inagaki, N. Circadian rhythms and diabetes. J. Diabetes Investig. 2011, 2, 176–177. [Google Scholar] [CrossRef]
- Newton, K.; Dixit, V.M. Signaling in innate immunity and inflammation. Cold Spring Harb. Perspect. Biol. 2012, 4, a006049. [Google Scholar] [CrossRef] [PubMed]
- Neniskyte, U.; Vilalta, A.; Brown, G.C. Tumour necrosis factor α-induced neuronal loss is mediated by microglial phagocytosis. FEBS Lett. 2014, 588, 2952–2956. [Google Scholar] [CrossRef] [PubMed]
- Griffiths, M.R.; Gasque, P.; Neal, J.W. The multiple roles of the innate immune system in the regulation of apoptosis and inflammation in the brain. J. Neuropathol. Exp. Neurol. 2009, 68, 217–226. [Google Scholar] [CrossRef]
- Cerón, J.J.; Eckersall, P.D.; Martínez-Subiela, S. Acute phase proteins in dogs and cats: Current knowledge and future perspectives. Vet. Clin. Pathol. 2005, 34, 85–99. [Google Scholar] [CrossRef]
- Murata, H.; Shimada, N.; Yoshioka, M. Current research on acute phase proteins in veterinary diagnosis: An overview. Vet. J. 2004, 168, 28–40. [Google Scholar] [CrossRef]
- Petersen, H.H.; Nielsen, J.P.; Heegaard, P.M.H. Application of acute phase protein measurements in veterinary clinical chemistry. Vet. Res. 2004, 35, 163–187. [Google Scholar] [CrossRef]
| Species | Organ | Total Modulated | Up-Modulated | Down-Modulated | HUGO IDs | Mapped by IPA |
|---|---|---|---|---|---|---|
| Zebrafish | Skin | 523 | 336 | 187 | 482 | 364 |
| Brain | 94 | 49 | 45 | 89 | 78 | |
| Liver | 83 | 56 | 27 | 79 | 64 | |
| Medaka | Skin | 2304 | 1229 | 1075 | 2298 | 2284 |
| Brain | 592 | 433 | 159 | 589 | 587 | |
| Liver | 3492 | 1684 | 1808 | 3483 | 3464 | |
| Mouse | Skin | 107 | 49 | 58 | N/A | 102 |
| Brain | 1174 | 699 | 475 | N/A | 1172 | |
| Liver | 182 | 51 | 131 | N/A | 166 |
| Zebrafish Skin | ||||
| Ingenuity Canonical Pathways | z-score | p-value | Gene | Category |
| Acute Phase Response Signaling | 4.12 | 4.27E-02 | 9 | Im, If, AP, CP |
| Production of Nitric Oxide and ROS in Macrophages | 4.11 | 2.29E-02 | 11 | Im, AP |
| Oncostatin M Signaling | 3.19 | 3.09E-03 | 10 | Im, If, AP, CP |
| Leukocyte Extravasation Signaling | 3.07 | 2.88E-03 | 14 | Im, If |
| TREM1 Signaling | 3.00 | 2.69E-02 | 12 | Im, If, AP |
| mTOR Signaling | 2.90 | 2.09E-02 | 12 | CP, CS |
| Complement System | 2.71 | 2.02E-02 | 6 | Im, If, AP |
| IL-6 Signaling | 2.45 | 2.96E-02 | 11 | Im, If, AP |
| B Cell Receptor Signaling | 2.00 | 3.98E-02 | 10 | Im, AP |
| ILK Signaling | 2.00 | 4.27E-03 | 13 | Im, If, AP, CP |
| Nitric Oxide Signaling in the Cardiovascular System | 2.00 | 4.16E-02 | 6 | If |
| NRF2-mediated Oxidative Stress Response | 2.00 | 1.02E-02 | 9 | If, AP |
| Role of NFAT in Regulation of the Immune Response | 2.00 | 4.33E-02 | 6 | Im, AP |
| Thrombin Signaling | 2.00 | 1.28E-02 | 9 | If, AP |
| Gα12/13 Signaling | −2.00 | 4.17E-03 | 10 | If, AP, CP, CS |
| PPARα/RXRα Activation | −2.37 | 2.46E-01 | 7 | If, AP |
| Cardiac Hypertrophy Signaling | −2.87 | 7.76E-03 | 14 | If, AP, CP, CS |
| eNOS Signaling | −2.97 | 4.79E-02 | 8 | If, CP |
| CDK5 Signaling | −3.56 | 2.09E-03 | 9 | If, AP, CP |
| Calcium Signaling | −3.58 | 7.94E-12 | 26 | CP, CS |
| Actin Cytoskeleton Signaling | −2.47 | 2.40E-03 | 15 | AP, CP, CM, CS |
| Medaka Skin | ||||
| Ingenuity Canonical Pathways | z-score | p-value | Genes | |
| Acute Phase Response Signaling | 3.67 | 3.39E-09 | 45 | Im, If, AP, CP |
| Gα12/13 Signaling | 2.79 | 4.68E-03 | 24 | If, AP, CP |
| Pancreatic Adenocarcinoma Signaling | 2.79 | 1.10E-05 | 29 | Im, AP, CP |
| RhoA Signaling | 2.79 | 4.27E-03 | 23 | If, CP, CM |
| Eicosanoid Signaling | 2.73 | 9.33E-03 | 14 | If |
| IL-6 Signaling | 2.68 | 1.82E-05 | 30 | Im, If, AP |
| ILK Signaling | 2.49 | 7.59E-05 | 39 | Im, If, AP, CP |
| TREM1 Signaling | 2.43 | 5.01E-02 | 13 | Im, If, AP |
| Endothelin-1 Signaling | 2.41 | 5.75E-04 | 35 | AP, CP, CM |
| Ceramide Signaling | 2.37 | 7.76E-05 | 23 | If, AP, CP |
| ERK5 Signaling | 2.22 | 1.35E-02 | 13 | CP |
| NGF Signaling | 2.20 | 5.25E-03 | 22 | AP, CP |
| p38 MAPK Signaling | 2.20 | 2.45E-03 | 23 | Im, If |
| Activation of IRF by Cytosolic Pattern Recogn. Recep. | 2.16 | 2.05E-02 | 9 | Im, AP |
| MIF Regulation of Innate Immunity | 2.16 | 2.51E-02 | 9 | Im, AP |
| Cardiac β-adrenergic Signaling | 2.10 | 1.45E-02 | 23 | CS |
| VEGF Signaling | 2.10 | 7.41E-02 | 16 | AP, CP, CM |
| B Cell Receptor Signaling | 2.06 | 1.86E-03 | 33 | Im, AP |
| NANOG in Mammalian Embryonic Stem Cell Pluripotency | 2.00 | 1.51E-04 | 27 | AP, CP |
| ATM Signaling | −2.00 | 3.98E-02 | 14 | If, CP |
| Wnt/GSK-3β Signaling in the Pathos of Influenza | −2.00 | 1.26E-02 | 15 | Im |
| NFAT in Cardiac Hypertrophy | −2.11 | 6.31E-03 | 32 | Im, AP |
| PPARα/RXRα Activation | −2.68 | 1.70E-02 | 17 | If, AP |
| Antioxidant Action of Vitamin C | −2.82 | 4.17E-02 | 17 | If, AP, CP |
| Mouse Skin | ||||
| Ingenuity Canonical Pathways | z-score | p-value | Genes | |
| TREM1 Signaling | 4.20 | 2.00E-05 | 12 | Im, If, AP |
| Role of IL-17F Inflammatory | 4.00 | 3.98E-05 | 5 | If |
| Leukocyte Extravasation Signaling | 3.00 | 1.26E-02 | 5 | Im, If |
| Hepatic Stellate Cell Activation | 2.30 | 1.58E-05 | 7 | If |
| AMPK Signaling | 2.20 | 1.00E-02 | 5 | CP |
| Granulocyte Adhesion and Diapedesis | 2.20 | 2.00E-11 | 12 | If |
| IL-1 Mediated Inhibition of RXR Function | 2.20 | 1.58E-02 | 5 | Im, If, AP |
| PPARα/RXRα Activation | 2.20 | 7.94E-03 | 5 | If, AP |
| Acute Phase Response | 2.10 | 3.98E-03 | 17 | Im, If, AP, CP |
| Inhibition of Matrix Metalloproteases | 2.10 | 2.51E-05 | 5 | CM |
| LXR/RXR Activation | 2.10 | 2.00E-03 | 5 | Im, If |
| Agranulocyte Adhesion and Diapedesis | 2.00 | 1.00E-08 | 10 | Im, If |
| Zebrafish Brain | ||||
|---|---|---|---|---|
| Ingenuity Canonical Pathways | z-score | p-value | Gene | Category |
| Synaptic Long-Term Potentiation | 2.60 | 8.71E-03 | 13 | Im, If, AP, CP |
| Intrinsic Prothrombin Activation Pathway | 2.20 | 1.17E-08 | 6 | If |
| Coagulation System | 2.10 | 1.66E-08 | 6 | If |
| Acute Phase Response | 2.00 | 7.24E-04 | 17 | Im, If, AP, CP |
| Medaka Brain | ||||
| Acute Phase Response Signaling | 3.67 | 3.98E-08 | 20 | Im, If, AP, CP |
| Endothelin-1 Signaling | 3.36 | 2.00E-04 | 15 | AP, CP, CM |
| B Cell Receptor Signaling | 2.71 | 1.29E-02 | 11 | Im, AP |
| Prolactin Signaling | 2.65 | 1.91E-03 | 8 | Im, AP, CP |
| ERK/MAPK Signaling | 2.50 | 1.17E-03 | 14 | CP |
| IL-6 Signaling | 2.50 | 4.47E-05 | 13 | Im, If, AP |
| CREB Signaling in Neurons | 2.45 | 3.02E-02 | 10 | AP, CP, CS |
| JAK/Stat Signaling | 2.45 | 2.57E-02 | 6 | Im, AP, CP |
| Thrombopoietin Signaling | 2.45 | 8.51E-03 | 6 | If, AP, CP |
| P2Y Purigenic Receptor Signaling Pathway | 2.33 | 1.02E-02 | 9 | If, AP, CP |
| GNRH Signaling | 2.33 | 8.91E-03 | 9 | If, AP, CP, CM |
| Melanocyte Development and Pigmentation Signaling | 2.24 | 4.57E-02 | 6 | AP, CP, CS |
| ERK5 Signaling | 2.24 | 2.88E-02 | 5 | CP |
| Hypoxia Signaling in the Cardiovascular System | 2.24 | 1.91E-03 | 7 | If, CP |
| NFAT in Regulation of the Immune Response | 2.12 | 1.29E-02 | 11 | Im, AP |
| PKCθ Signaling in T Lymphocytes | 2.12 | 2.82E-02 | 8 | Im, AP |
| Ephrin Receptor Signaling | 2.12 | 3.02E-03 | 12 | CP, CM, CS |
| Chemokine Signaling | 2.12 | 6.92E-04 | 8 | If, AP, CP, CM |
| cAMP-mediated signaling | 2.00 | 3.98E-04 | 16 | If, AP, CS |
| IL-2 Signaling | 2.00 | 3.02E-02 | 5 | Im, AP, CP, CS |
| PPAR Signaling | −2.83 | 3.89E-03 | 8 | Im, AP |
| Synaptic Long-Term Potentiation | −3.23 | 3.98E-04 | 34 | Im, If, AP, CP |
| Mouse Brain | ||||
| Actin Cytoskeleton Signaling | 3.50 | 1.66E-03 | 22 | CP, CM |
| Gαs Signaling | 3.40 | 2.57E-03 | 11 | If, AP, CS |
| RhoA Signaling | 3.10 | 1.82E-02 | 12 | If, CS |
| VEGF Signaling | 2.60 | 4.68E-02 | 10 | CP, CM |
| Nitric Oxide Signaling | 2.50 | 1.35E-03 | 11 | If, CS |
| Cyclins and Cell Cycle Regulation | 2.40 | 4.37E-02 | 8 | CP |
| Gα12/13 Signaling | 2.30 | 2.24E-02 | 13 | If, AP, CS |
| Cholecystokinin/Gastrin-mediated Signaling | 2.10 | 3.89E-03 | 10 | AP, CP, CM |
| CXCR4 Signaling | 2.10 | 2.09E-02 | 16 | Im, AP, CP, CM |
| B Cell Receptor Signaling | 2.10 | 3.16E-02 | 18 | If, Im, AP, CM |
| Acute Phase Response Signaling | 2.10 | 3.80E-02 | 19 | If, Im, AP, CP |
| Gαq Signaling | 2.10 | 1.45E-03 | 15 | If, AP, CM, CS |
| IL-6 Signaling | 2.00 | 4.79E-02 | 21 | If, Im, AP |
| Synaptic Long-Term Potentiation | 2.00 | 1.95E-03 | 12 | If, AP, CS |
| ILK Signaling | 2.00 | 8.32E-04 | 19 | If, Im, AP, CP, CM |
| Tec Kinase Signaling | 2.00 | 4.79E-02 | 10 | AP, CP, CM |
| Calcium-induced T Lymphocyte Apoptosis | −2.00 | 1.70E-02 | 6 | Im |
| Glutamate Receptor Signaling | −2.00 | 4.37E-06 | 11 | If, CS |
| FLT3 Signaling in Hematopoietic Progenitor Cells | −2.00 | 4.90E-02 | 9 | Im, AP, CP |
| BMP signaling pathway | −2.10 | 2.09E-03 | 8 | AP, CP |
| Zebrafish Liver | ||||
| Ingenuity Canonical Pathways | z-score | p-value | Gene | Category |
| Acute Phase Response | 2.5 | 3.89E-02 | 18 | Im, If, AP, CP |
| Complement Signaling | 2.3 | 5.62E-03 | 9 | Im, If, AP |
| IL-1 Mediated Inhibition of RXR | 2.0 | 1.35E-02 | 22 | Im, If, AP |
| ILK Signaling | 2.2 | 4.79E-02 | 19 | If, Im, AP, CP, CM |
| Calcium Signaling | 2.0 | 2.24E-02 | 17 | CP, CS |
| Medaka Liver | ||||
| z-score | p-value | Gene # | Category | |
| Acute Phase Response Signaling | 3.2 | 9.77E-07 | 53 | Im, If, AP, CP |
| Tec Kinase Signaling | 2.9 | 1.02E-02 | 39 | AP, CP, CM |
| mTOR Signaling | 2.8 | 1.32E-02 | 44 | CP, CS |
| PEDF Signaling | 2.7 | 2.40E-06 | 31 | AP, CP, CM |
| Insulin Receptor Signaling | 2.6 | 1.70E-06 | 45 | If, AP, CP |
| IL-1 Signaling | 2.5 | 1.82E-06 | 33 | If, AP, CS |
| Renal Cell Carcinoma Signaling | 2.4 | 6.76E-03 | 22 | Im, AP, CP, CM |
| HMGB1 Signaling | 2.4 | 7.08E-04 | 36 | If, Im, AP |
| NANOG in Mammal Embry Stem Cell Pluripotency | 2.2 | 4.57E-03 | 31 | AP, CP |
| Type I Diabetes Mellitus Signaling | 2.2 | 3.80E-02 | 25 | If, Im, AP |
| Toll-like Receptor Signaling | 2.2 | 2.04E-03 | 22 | If, Im, AP |
| UVB-Induced MAPK Signaling | 2.2 | 2.69E-02 | 17 | If, AP, CS |
| CNTF Signaling | 2.2 | 1.74E-02 | 17 | Im, AP, CP, CS |
| Pancreatic Adenocarcinoma Signaling | 2.2 | 5.25E-05 | 36 | If, AP, CP |
| IL-1 Mediated Inhibition of RXR | 2.1 | 3.31E-03 | 51 | Im, If, AP |
| PI3K/AKT Signaling | 2.1 | 4.68E-06 | 40 | If, Im, AP, CP |
| IL-8 Signaling | 2.1 | 5.25E-07 | 59 | If, Im, AP |
| Macropinocytosis Signaling | 2.1 | 2.69E-02 | 20 | Im |
| CD28 Signaling in T Helper Cells | 2.1 | 2.14E-03 | 34 | Im, AP |
| ERK5 Signaling | 2.0 | 1.74E-04 | 22 | CP |
| G-βγ Signaling | 2.0 | 2.14E-03 | 25 | If, AP, CP |
| MIF Regulation of Innate Immunity | 2.0 | 4.07E-02 | 11 | Im, AP |
| Chemokine Signaling | −2.0 | 2.75E-03 | 21 | If, AP, CP, CM |
| Ephrin B Signaling | −2.1 | 2.57E-04 | 24 | AP, CP, CM |
| Phospholipase C Signaling | −2.1 | 2.04E-02 | 50 | AP, CP, CM |
| LXR/RXR Activation | −2.2 | 2.40E-06 | 40 | Im, If |
| p53 Signaling | −2.2 | 7.59E-03 | 28 | CP |
| CREB Signaling in Neurons | −2.3 | 2.69E-04 | 48 | AP, CP, CS |
| Wnt/Ca+ pathway | −2.3 | 3.02E-05 | 22 | CP |
| Sonic Hedgehog Signaling | −2.4 | 1.45E-02 | 10 | CP |
| Dopamine-DARPP32 Feedback in cAMP Signaling | −2.7 | 8.71E-05 | 45 | CS |
| Basal Cell Carcinoma Signaling | −2.9 | 7.41E-03 | 20 | CP, CM, CS |
| Calcium Signaling | −3.2 | 8.32E-04 | 45 | CP, CS |
| Mouse Liver | ||||
| z-score | p-value | Gene # | Category | |
| IL-1 Mediated Inhibition of RXR | −2.0 | 6.92E-04 | 23 | Im, If, AP |
| LXR/RXR Activation | −2.0 | 8.51E-03 | 13 | Im, If |
| VDR/RXR Activation | −2.0 | 1.48E-02 | 8 | Im, CP |
| Granulocyte Adhesion and Diapedesis | −2.1 | 1.15E-03 | 17 | If |
| Coagulation System | −2.1 | 1.58E-03 | 5 | If |
| STAT3 Pathway | −2.3 | 1.26E-02 | 7 | If, Im, AP |
| p38 MAPK Signaling | −3.0 | 7.59E-03 | 12 | Im, If |
| Acute Phase Response Signaling IL-6 Signaling | −4.0 −4.0 | 2.57E-02 1.55E-03 | 19 26 | I Im, If, AP, CP m, If, AP |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boswell, M.; Lu, Y.; Boswell, W.; Savage, M.; Hildreth, K.; Salinas, R.; Walter, C.A.; Walter, R.B. Fluorescent Light Incites a Conserved Immune and Inflammatory Genetic Response within Vertebrate Organs (Danio rerio, Oryzias latipes and Mus musculus). Genes 2019, 10, 271. https://doi.org/10.3390/genes10040271
Boswell M, Lu Y, Boswell W, Savage M, Hildreth K, Salinas R, Walter CA, Walter RB. Fluorescent Light Incites a Conserved Immune and Inflammatory Genetic Response within Vertebrate Organs (Danio rerio, Oryzias latipes and Mus musculus). Genes. 2019; 10(4):271. https://doi.org/10.3390/genes10040271
Chicago/Turabian StyleBoswell, Mikki, Yuan Lu, William Boswell, Markita Savage, Kim Hildreth, Raquel Salinas, Christi A. Walter, and Ronald B. Walter. 2019. "Fluorescent Light Incites a Conserved Immune and Inflammatory Genetic Response within Vertebrate Organs (Danio rerio, Oryzias latipes and Mus musculus)" Genes 10, no. 4: 271. https://doi.org/10.3390/genes10040271
APA StyleBoswell, M., Lu, Y., Boswell, W., Savage, M., Hildreth, K., Salinas, R., Walter, C. A., & Walter, R. B. (2019). Fluorescent Light Incites a Conserved Immune and Inflammatory Genetic Response within Vertebrate Organs (Danio rerio, Oryzias latipes and Mus musculus). Genes, 10(4), 271. https://doi.org/10.3390/genes10040271




