Nitrogen Supply Drives Senescence-Related Seed Storage Protein Expression in Rapeseed Leaves
Abstract
1. Introduction
2. Material and Methods
2.1. Plant Growth Conditions
2.2. H2O2 Measurements
2.3. Generation of Anti-SSP Antisera
2.4. Protein Extraction and Western-Blotting
2.5. RNA Extraction and qRT-PCR
2.6. Chlorophyll Measurements
2.7. Microarray Data Evaluation
2.8. Total N and δ15N-Measurements
2.9. Catalase Zymograms
3. Results
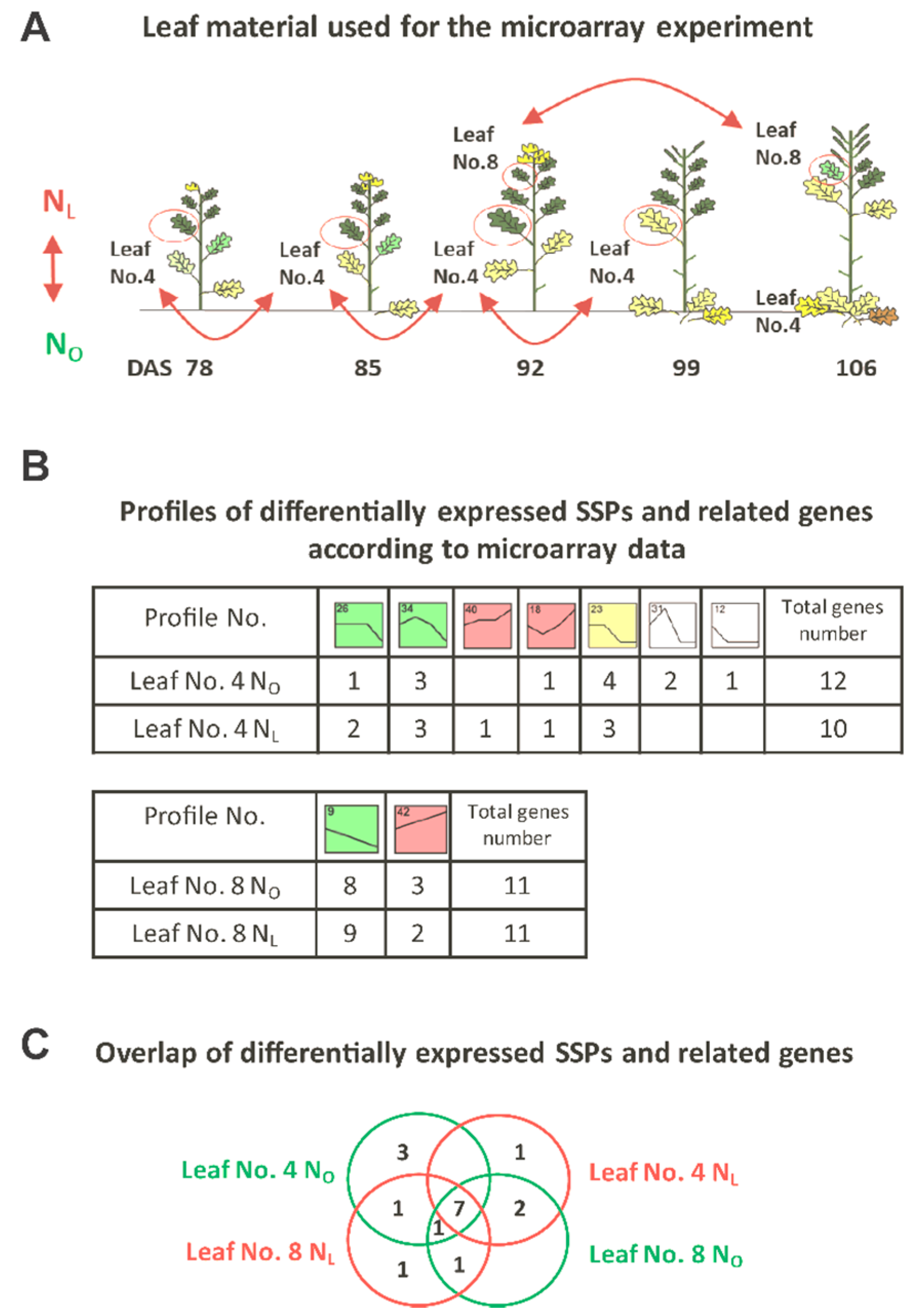
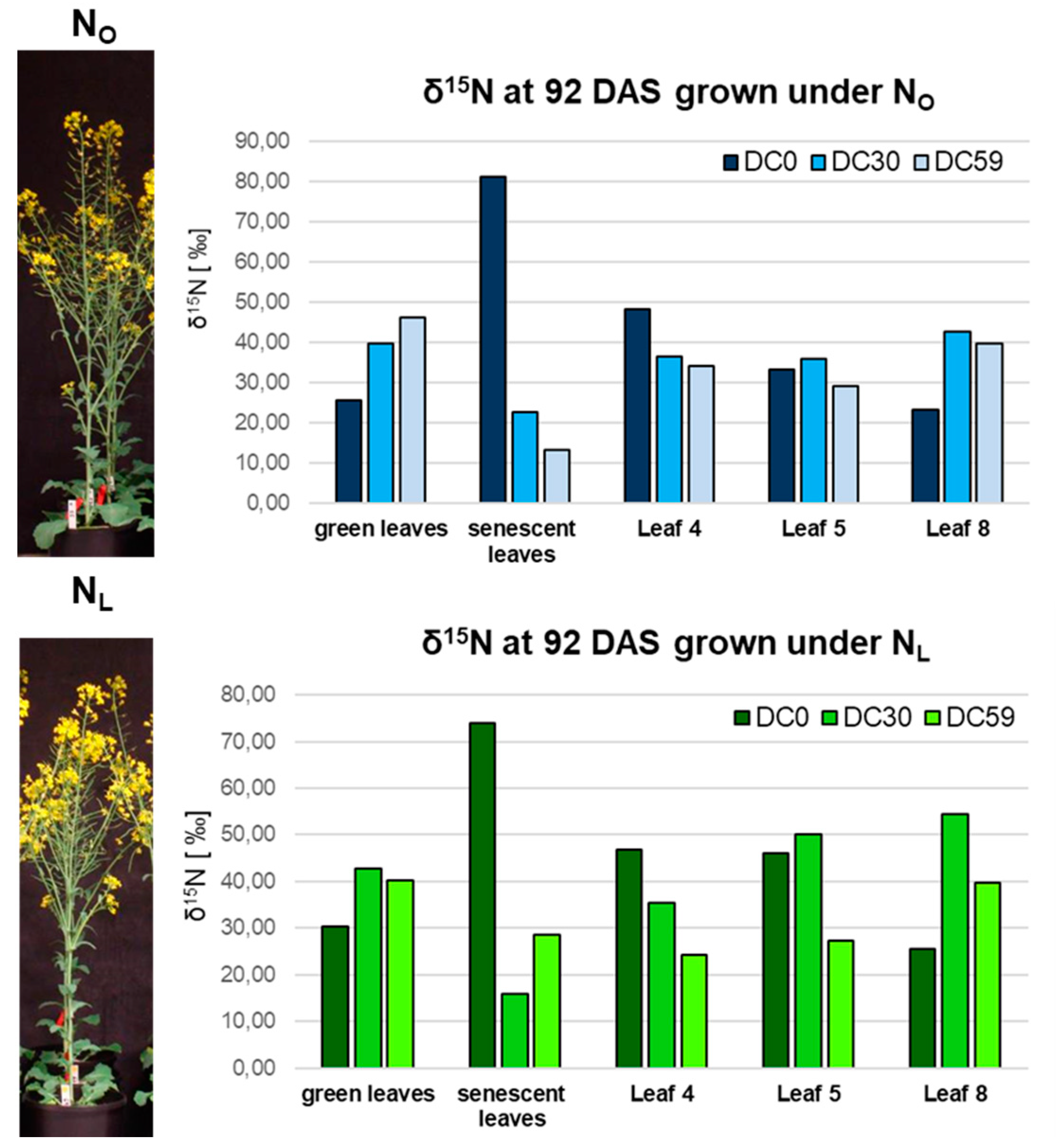
3.1. Transcriptome Analysis of OSR during Induction of Senescence under Two Different N-Supplies
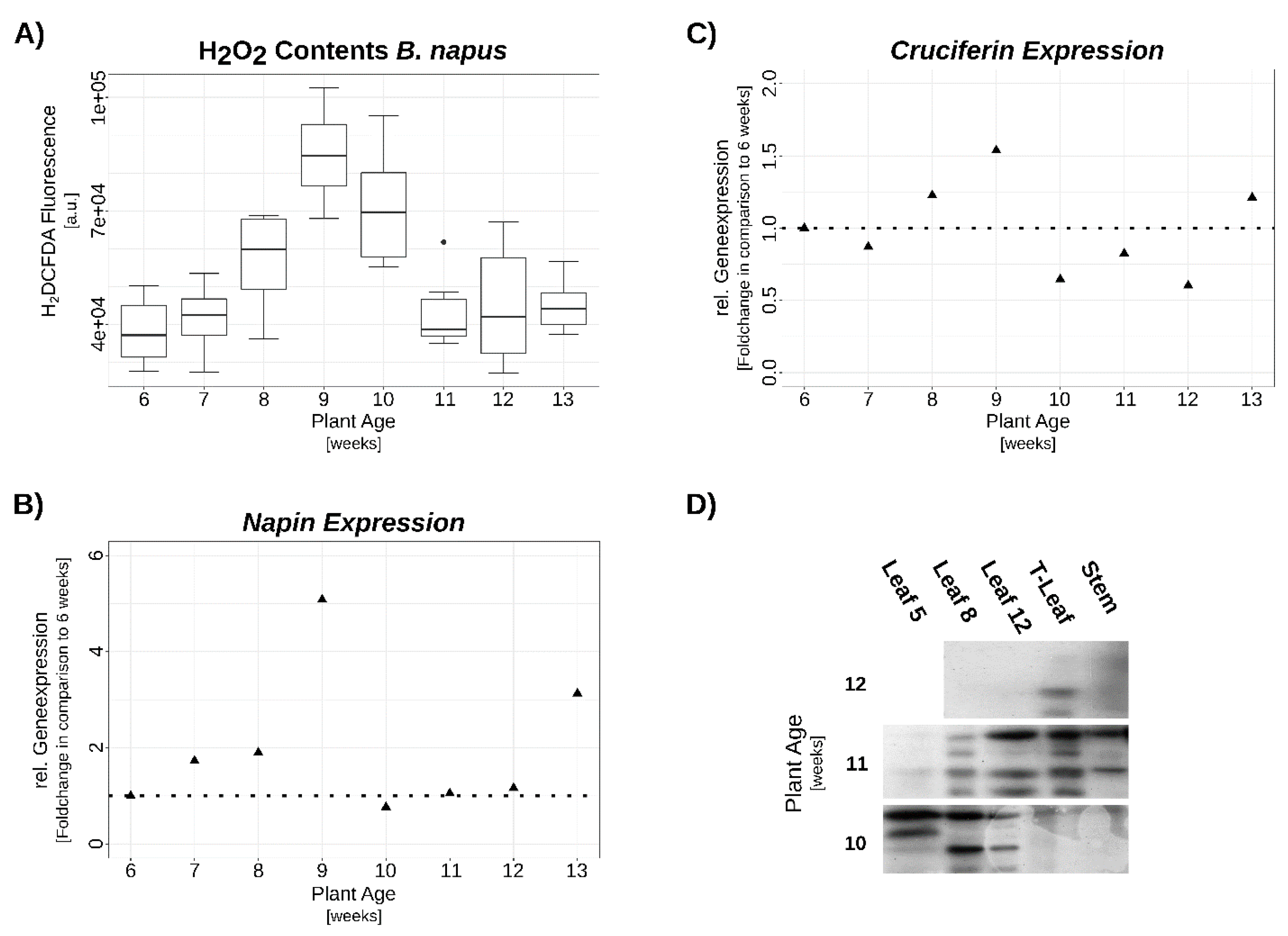
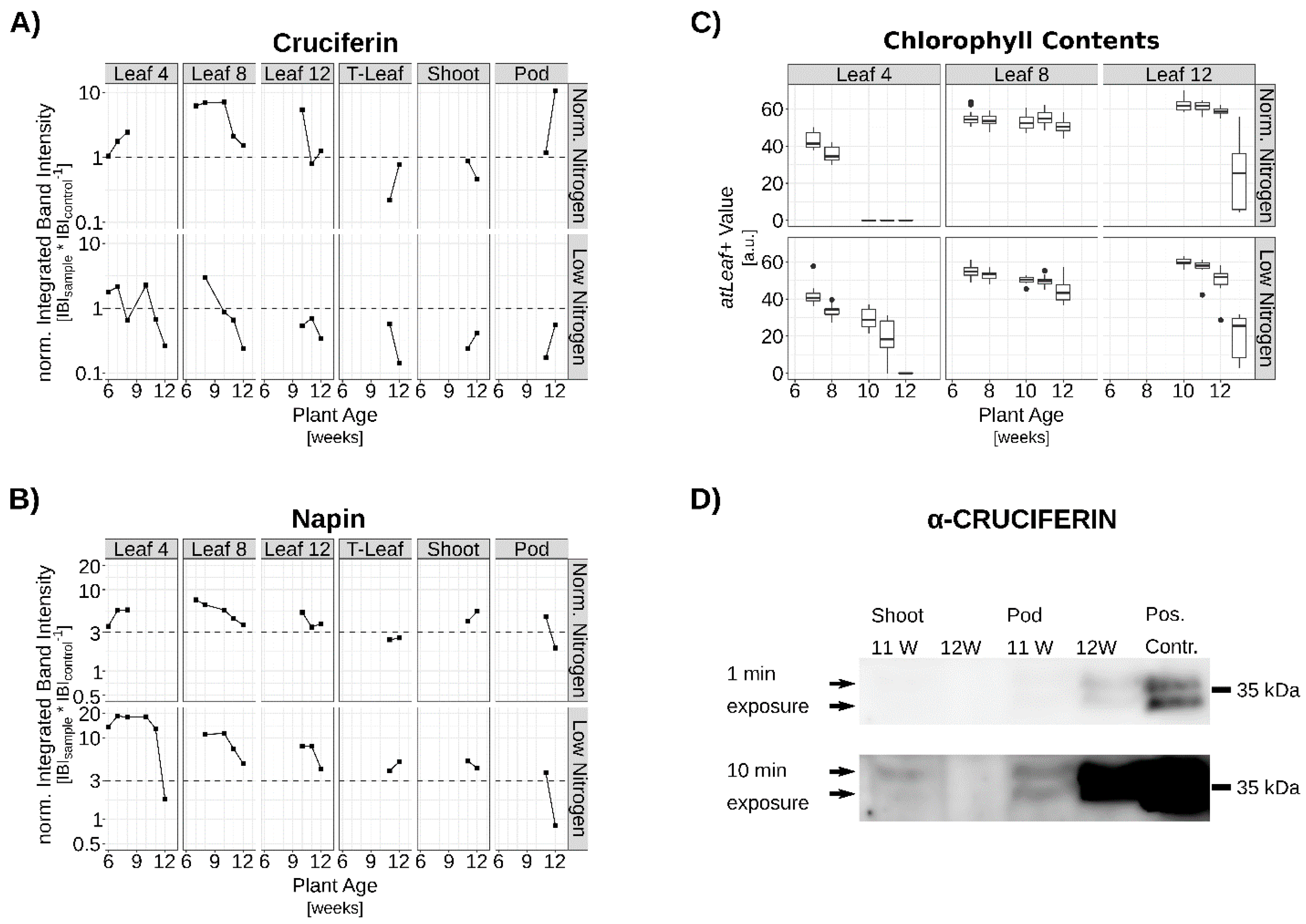
3.2. Verification of SSP Expression via qRT-PCR and Western-Blot
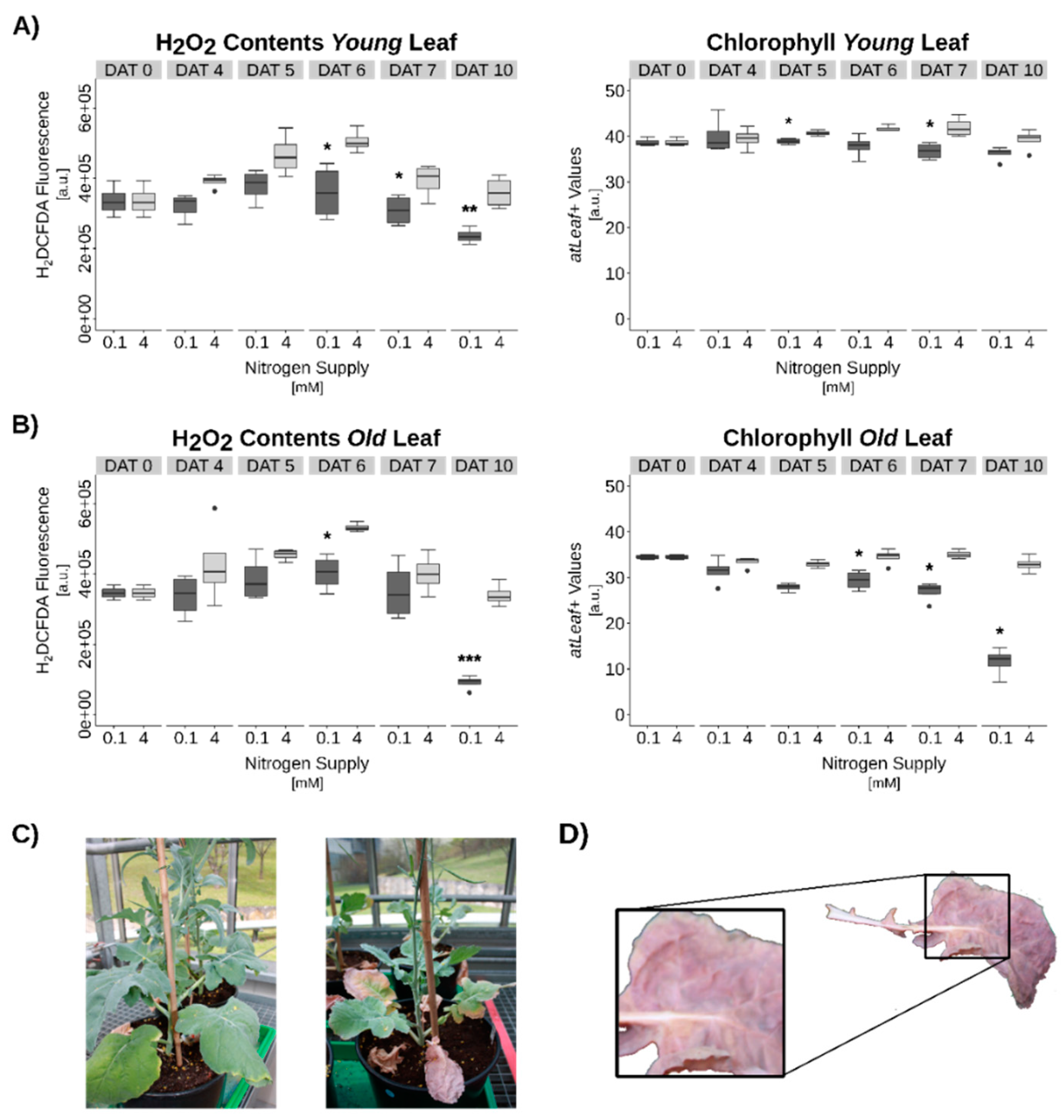
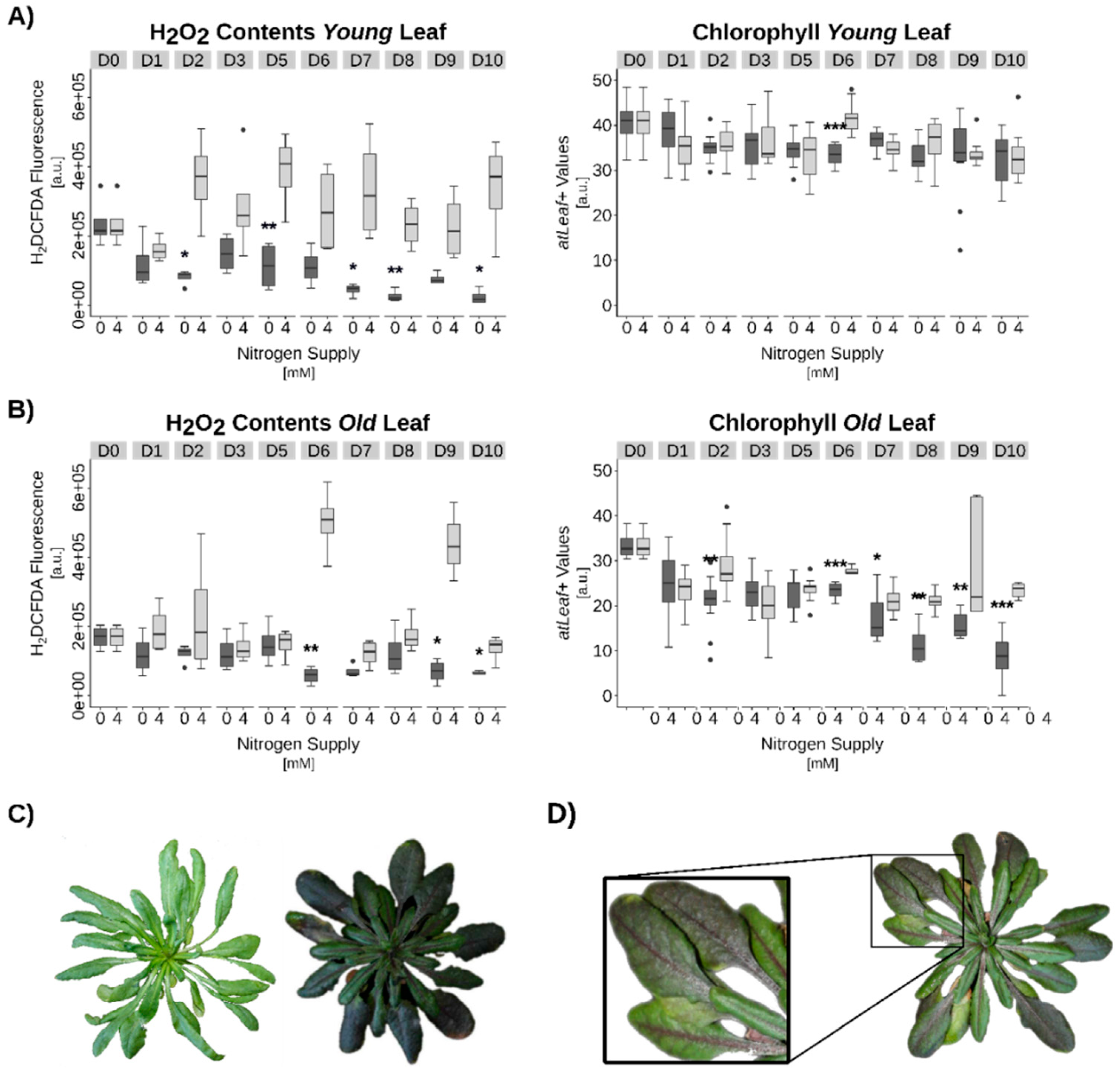
3.3. Hydrogen Peroxide Signaling under Complete N-Starvation
4. Discussions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brunel-Muguet, S.; D’Hooghe, P.; Bataille, M.P.; Larre, C.; Kim, T.H.; Trouverie, J.; Avice, J.C.; Etienne, P.; Durr, C. Heat stress during seed filling interferes with sulfur restriction on grain composition and seed germination in oilseed rape (Brassica napus L.). Front. Plant Sci. 2015, 6, 213. [Google Scholar] [CrossRef] [PubMed]
- Monsalve, R.I.; Lopez-Otin, C.; Villalba, M.; Rodriguez, R. A new distinct group of 2 S albumins from rapeseed. Amino acid sequence of two low molecular weight napins. FEBS Lett. 1991, 295, 207–210. [Google Scholar] [CrossRef]
- Schwenke, K.D.; Raab, B.; Linow, K.J.; Pahtz, W.; Uhlig, J. Isolation of the 12 S globulin from rapeseed (Brassica napus L.) and characterization as a “neutral” protein. On seed proteins. Part 13. Die Nahrung 1981, 25, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Perera, S.P.; McIntosh, T.C.; Wanasundara, J.P. Structural properties of cruciferin and napin of Brassica napus (Canola) show distinct responses to changes in pH and temperature. Plants 2016, 5, 36. [Google Scholar] [CrossRef] [PubMed]
- Gacek, K.; Bartkowiak-Broda, I.; Batley, J. Genetic and Molecular Regulation of Seed Storage Proteins (SSPs) to Improve Protein Nutritional Value of Oilseed Rape (Brassica napus L.) Seeds. Front. Plant Sci. 2018, 9, 890. [Google Scholar] [CrossRef] [PubMed]
- Chao, H.; Wang, H.; Wang, X.; Guo, L.; Gu, J.; Zhao, W.; Li, B.; Chen, D.; Raboanatahiry, N.; Li, M. Genetic dissection of seed oil and protein content and identification of networks associated with oil content in Brassica napus. Sci. Rep. 2017, 7, 46295. [Google Scholar] [CrossRef]
- Gu, J.; Chao, H.; Gan, L.; Guo, L.; Zhang, K.; Li, Y.; Wang, H.; Raboanatahiry, N.; Li, M. Proteomic Dissection of Seed Germination and Seedling Establishment in Brassica napus. Front. Plant Sci. 2016, 7, 1482. [Google Scholar] [CrossRef]
- Tilsner, J.; Kassner, N.; Struck, C.; Lohaus, G. Amino acid contents and transport in oilseed rape (Brassica napus L.) under different nitrogen conditions. Planta 2005, 221, 328–338. [Google Scholar] [CrossRef]
- Kichey, T.; Hirel, B.; Heumez, E.; Dubois, F.; Le Gouis, J. In winter wheat (Triticum aestivum L.), post-anthesis nitrogen uptake and remobilisation to the grain correlates with agronomic traits and nitrogen physiological markers. Field Crop. Res. 2007, 102, 22–32. [Google Scholar] [CrossRef]
- Breeze, E.; Harrison, E.; McHattie, S.; Hughes, L.; Hickman, R.; Hill, C.; Kiddle, S.; Kim, Y.S.; Penfold, C.A.; Jenkins, D.; et al. High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 2011, 23, 873–894. [Google Scholar] [CrossRef]
- Bieker, S.; Riester, L.; Stahl, M.; Franzaring, J.; Zentgraf, U. Senescence-specific alteration of hydrogen peroxide levels in Arabidopsis thaliana and oilseed rape spring variety Brassica napus L. cv. Mozart. J. Integr. Plant Biol. 2012, 54, 540–554. [Google Scholar] [CrossRef] [PubMed]
- Zimmermann, P.; Heinlein, C.; Orendi, G.; Zentgraf, U. Senescence-specific regulation of catalases in Arabidopsis thaliana (L.) Heynh. Plant Cell Environ. 2006, 29, 1049–1060. [Google Scholar] [CrossRef] [PubMed]
- Rossato, L.; Lainé, P.; Ourry, A. Nitrogen storage and remobilization in Brassica napus L. during the growth cycle: Nitrogen fluxes within the plant and changes in soluble protein patterns. J. Exp. Bot. 2001, 52, 1655–1663. [Google Scholar] [CrossRef] [PubMed]
- Oury, F.-X.; Godin, C. Yield and grain protein concentration in bread wheat: How to use the negative relationship between the two characters to identify favourable genotypes? Euphytica 2007, 157, 45–57. [Google Scholar] [CrossRef]
- Bogard, M.; Jourdan, M.; Allard, V.; Martre, P.; Perretant, M.R.; Ravel, C.; Heumez, E.; Orford, S.; Snape, J.; Griffiths, S.; et al. Anthesis date mainly explained correlations between post-anthesis leaf senescence, grain yield, and grain protein concentration in a winter wheat population segregating for flowering time QTLs. J. Exp. Bot. 2011, 62, 3621–3636. [Google Scholar] [CrossRef] [PubMed]
- Avice, J.C.; Etienne, P. Leaf senescence and nitrogen remobilization efficiency in oilseed rape (Brassica napus L.). J. Exp. Bot. 2014, 65, 3813–3824. [Google Scholar] [CrossRef]
- Desclos, M.; Dubousset, L.; Etienne, P.; Le Caherec, F.; Satoh, H.; Bonnefoy, J.; Ourry, A.; Avice, J.C. A proteomic profiling approach to reveal a novel role of Brassica napus drought 22 kD/water-soluble chlorophyll-binding protein in young leaves during nitrogen remobilization induced by stressful conditions. Plant Physiol. 2008, 147, 1830–1844. [Google Scholar] [CrossRef]
- Etienne, P.; Desclos, M.; Le Gou, L.; Gombert, J.; Bonnefoy, J.; Maurel, K.; Le Dily, F.; Ourry, A.; Avice, J.-C. N-protein mobilisation associated with the leaf senescence process in oilseed rape is concomitant with the disappearance of trypsin inhibitor activity. Funct. Plant Biol. 2007, 34, 895–906. [Google Scholar] [CrossRef]
- Safavi-Rizi, V.; Franzaring, J.; Fangmeier, A.; Kunze, R. Divergent N Deficiency-Dependent Senescence and Transcriptome Response in Developmentally Old and Young Brassica napus Leaves. Front. Plant Sci. 2018, 9, 48. [Google Scholar] [CrossRef]
- Franzaring, J.; Weller, S.; Schmid, I.; Fangmeier, A. Growth, senescence and water use efficiency of spring oilseed rape (Brassica napus L. cv. Mozart) grown in a factorial combination of nitrogen supply and elevated CO2. Environ. Exp. Bot. 2011, 72, 284–296. [Google Scholar] [CrossRef]
- Franzaring, J.; Gensheimer, G.; Weller, S.; Schmid, I.; Fangmeier, A. Allocation and remobilisation of nitrogen in spring oilseed rape (Brassica napus L. cv. Mozart) as affected by N supply and elevated CO2. Environ. Exp. Bot. 2012, 83, 12–22. [Google Scholar] [CrossRef]
- Koeslin-Findeklee, F.; Meyer, A.; Girke, A.; Beckmann, K.; Horst, W.J. The superior nitrogen efficiency of winter oilseed rape (Brassica napus L.) hybrids is not related to delayed nitrogen starvation-induced leaf senescence. Plant Soil 2014, 384, 347–362. [Google Scholar] [CrossRef]
- Cathcart, R.; Schwiers, E.; Ames, B.N. Detection of picomole levels of hydroperoxides using a fluorescent dichlorofluorescein assay. Anal. Biochem. 1983, 134, 111–116. [Google Scholar] [CrossRef]
- Arvidsson, S.; Kwasniewski, M.; Riano-Pachon, D.M.; Mueller-Roeber, B. QuantPrime-a flexible tool for reliable high-throughput primer design for quantitative PCR. BMC Bioinf. 2008, 9, 465. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucl. Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucl. Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Camacho, C.; Coulouris, G.; Avagyan, V.; Ma, N.; Papadopoulos, J.; Bealer, K.; Madden, T.L. BLAST+: Architecture and applications. BMC Bioinf. 2009, 10, 421. [Google Scholar] [CrossRef] [PubMed]
- Lamesch, P.; Berardini, T.Z.; Li, D.; Swarbreck, D.; Wilks, C.; Sasidharan, R.; Muller, R.; Dreher, K.; Alexander, D.L.; Garcia-Hernandez, M.; et al. The Arabidopsis Information Resource (TAIR): Improved gene annotation and new tools. Nucl. Acids Res. 2012, 40, D1202–D1210. [Google Scholar] [CrossRef]
- Li, L.; Shimada, T.; Takahashi, H.; Ueda, H.; Fukao, Y.; Kondo, M.; Nishimura, M.; Hara-Nishimura, I. MAIGO2 is involved in exit of seed storage proteins from the endoplasmic reticulum in Arabidopsis thaliana. Plant Cell 2006, 18, 3535–3547. [Google Scholar] [CrossRef]
- Li, L.; Shimada, T.; Takahashi, H.; Koumoto, Y.; Shirakawa, M.; Takagi, J.; Zhao, X.; Tu, B.; Jin, H.; et al. MAG2 and three MAG2-INTERACTING PROTEINs form an ER-localized complex to facilitate storage protein transport in Arabidopsis thaliana. Plant J. 2013, 76, 781–791. [Google Scholar] [CrossRef]
- Gironde, A.; Etienne, P.; Trouverie, J.; Bouchereau, A.; Le Caherec, F.; Leport, L.; Orsel, M.; Niogret, M.F.; Nesi, N.; Carole, D.; et al. The contrasting N management of two oilseed rape genotypes reveals the mechanisms of proteolysis associated with leaf N remobilization and the respective contributions of leaves and stems to N storage and remobilization during seed filling. BMC Plant Biol. 2015, 15, 59. [Google Scholar] [CrossRef] [PubMed]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.; Tang, H.; Wang, X.; et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950–953. [Google Scholar] [CrossRef] [PubMed]
- Solovyev, V.V.; Shahmuradov, I.A.; Salamov, A.A. Identification of promoter regions and regulatory sites. Methods Mol. Biol. 2010, 674, 57. [Google Scholar] [PubMed]
- Wang, X.; Larkins, B.A. Genetic analysis of amino acid accumulation in opaque-2 maize endosperm. Plant Physiol. 2001, 125, 1766–1777. [Google Scholar] [CrossRef]
- Fernandez, D.E.; Turner, F.R.; Crouch, M.L. In situ localization of storage protein mRNAs in developing meristems of Brassica napus embryos. Development 1991, 111, 299–313. [Google Scholar] [PubMed]
- Finkelstein, R.R.; Tenbarge, K.M.; Shumway, J.E.; Crouch, M.L. Role of ABA in Maturation of Rapeseed Embryos. Plant Physiol. 1985, 78, 630–636. [Google Scholar] [CrossRef]
- Rajjou, L.; Duval, M.; Gallardo, K.; Catusse, J.; Bally, J.; Job, C.; Job, D. Seed germination and vigor. Ann. Rev. Plant Biol. 2012, 63, 507–533. [Google Scholar] [CrossRef]
- Lim, P.O.; Kim, H.J.; Nam, H.G. Leaf senescence. Ann. Revi. Plant Biol. 2007, 58, 115–136. [Google Scholar] [CrossRef]
- Koeslin-Findeklee, F.; Becker, M.A.; van der Graaff, E.; Roitsch, T.; Horst, W.J. Differences between winter oilseed rape (Brassica napus L.) cultivars in nitrogen starvation-induced leaf senescence are governed by leaf-inherent rather than root-derived signals. J. Exp. Bot. 2015, 66, 3669–3681. [Google Scholar] [CrossRef]
- Suzuki, A.; Wu, C.Y.; Washida, H.; Takaiwa, F. Rice MYB protein OSMYB5 specifically binds to the AACA motif conserved among promoters of genes for storage protein glutelin. Plant Cell Physiol. 1998, 39, 555–559. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, R.J.; Burr, F.A.; Aukerman, M.J.; Burr, B. Maize regulatory gene opaque-2 encodes a protein with a “leucine-zipper” motif that binds to zein DNA. Proc. Natl. Acad. Sci. USA 1990, 87, 46–50. [Google Scholar] [CrossRef] [PubMed]
- Alonso, R.; Onate-Sanchez, L.; Weltmeier, F.; Ehlert, A.; Diaz, I.; Dietrich, K.; Vicente-Carbajosa, J.; Droge-Laser, W. A pivotal role of the basic leucine zipper transcription factor bZIP53 in the regulation of Arabidopsis seed maturation gene expression based on heterodimerization and protein complex formation. Plant Cell 2009, 21, 1747–1761. [Google Scholar] [CrossRef] [PubMed]
- Desikan, R.; S, A.H.-M.; Hancock, J.T.; Neill, S.J. Regulation of the Arabidopsis transcriptome by oxidative stress. Plant Physiol. 2001, 127, 159–172. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.; Bell, E.; Sadka, A.; Mullet, J.E. Arabidopsis thaliana Atvsp is homologous to soybean VspA and VspB, genes encoding vegetative storage protein acid phosphatases, and is regulated similarly by methyl jasmonate, wounding, sugars, light and phosphate. Plant Mol. Biol. 1995, 27, 933–942. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Ahn, J.E.; Datta, S.; Salzman, R.A.; Moon, J.; Huyghues-Despointes, B.; Pittendrigh, B.; Murdock, L.L.; Koiwa, H.; Zhu-Salzman, K. Arabidopsis vegetative storage protein is an anti-insect acid phosphatase. Plant Physiol. 2005, 139, 1545–1556. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Xu, G.; Tang, W.; Jing, Y.; Ji, Q.; Fei, Z.; Lin, R. Antagonistic basic helix-loop-helix/bZIP transcription factors form transcriptional modules that integrate light and reactive oxygen species signaling in Arabidopsis. Plant Cell 2013, 25, 1657–1673. [Google Scholar] [CrossRef] [PubMed]
- Peleg-Grossman, S.; Melamed-Book, N.; Cohen, G.; Levine, A. Cytoplasmic H2O2 prevents translocation of NPR1 to the nucleus and inhibits the induction of PR genes in Arabidopsis. Plant Signal Behav. 2010, 5, 1401–1406. [Google Scholar] [CrossRef]
- Job, C.; Rajjou, L.; Lovigny, Y.; Belghazi, M.; Job, D. Patterns of protein oxidation in Arabidopsis seeds and during germination. Plant Physiol. 2005, 138, 790–802. [Google Scholar] [CrossRef]
- Barba-Espin, G.; Diaz-Vivancos, P.; Job, D.; Belghazi, M.; Job, C.; Hernandez, J.A. Understanding the role of H2O2 during pea seed germination: A combined proteomic and hormone profiling approach. Plant Cell Environ. 2011, 34, 1907–1919. [Google Scholar] [CrossRef]
- El-Maarouf-Bouteau, H.; Meimoun, P.; Job, C.; Job, D.; Bailly, C. Role of protein and mRNA oxidation in seed dormancy and germination. Front. Plant Sci. 2013, 4, 77. [Google Scholar] [CrossRef]
- Nguyen, T.P.; Cueff, G.; Hegedus, D.D.; Rajjou, L.; Bentsink, L. A role for seed storage proteins in Arabidopsis seed longevity. J. Exp. Bot. 2015, 66, 6399–6413. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.; Wang, R.; Nacry, P.; Breton, G.; Kay, S.A.; Pruneda-Paz, J.L.; Davani, A.; Crawford, N.M. Nitrate foraging by Arabidopsis roots is mediated by the transcription factor TCP20 through the systemic signaling pathway. Proc. Natl. Acad. Sci. USA 2014, 111, 15267–15272. [Google Scholar] [CrossRef] [PubMed]
- Danisman, S.; van Dijk, A.D.; Bimbo, A.; van der Wal, F.; Hennig, L.; de Folter, S.; Angenent, G.C.; Immink, R.G. Analysis of functional redundancies within the Arabidopsis TCP transcription factor family. J. Exp. Bot. 2013, 64, 5673–5685. [Google Scholar] [CrossRef]
- Huang, L.; Zhang, H.; Zhang, H.; Deng, X.W.; Wei, N. HY5 regulates nitrite reductase 1 (NIR1) and ammonium transporter1;2 (AMT1;2) in Arabidopsis seedlings. Plant Sci. 2015, 238, 330–339. [Google Scholar] [CrossRef] [PubMed]
- Chai, T.; Zhou, J.; Liu, J.; Xing, D. LSD1 and HY5 antagonistically regulate red light induced-programmed cell death in Arabidopsis. Front. Plant Sci. 2015, 6, 292. [Google Scholar] [CrossRef] [PubMed]
- Tran, L.S.; Nakashima, K.; Sakuma, Y.; Simpson, S.D.; Fujita, Y.; Maruyama, K.; Fujita, M.; Seki, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. Isolation and functional analysis of Arabidopsis stress-inducible NAC transcription factors that bind to a drought-responsive cis-element in the early responsive to dehydration stress 1 promoter. Plant Cell 2004, 16, 2481–2498. [Google Scholar] [CrossRef] [PubMed]
- Zhu, X.; Chen, J.; Xie, Z.; Gao, J.; Ren, G.; Gao, S.; Zhou, X.; Kuai, B. Jasmonic acid promotes degreening via MYC2/3/4- and ANAC019/055/072-mediated regulation of major chlorophyll catabolic genes. Plant J. 2015, 84, 597–610. [Google Scholar] [CrossRef]
- Hickman, R.; Hill, C.; Penfold, C.A.; Breeze, E.; Bowden, L.; Moore, J.D.; Zhang, P.; Jackson, A.; Cooke, E.; Bewicke-Copley, F.; et al. A local regulatory network around three NAC transcription factors in stress responses and senescence in Arabidopsis leaves. Plant J. 2013, 75, 26–39. [Google Scholar] [CrossRef]
- Yan, J.; Tong, T.; Li, X.; Chen, Q.; Dai, M.; Niu, F.; et al. A Novel NAC-Type Transcription Factor, NAC87, from Oilseed Rape Modulates Reactive Oxygen Species Accumulation and Cell Death. Plant Cell Physiol. 2018, 59, 290–303. [Google Scholar] [CrossRef]
- Bi, Y.M.; Zhang, Y.; Signorelli, T.; Zhao, R.; Zhu, T.; Rothstein, S. Genetic analysis of Arabidopsis GATA transcription factor gene family reveals a nitrate-inducible member important for chlorophyll synthesis and glucose sensitivity. Plant J. 2005, 44, 680–692. [Google Scholar] [CrossRef]
- Fritz, C.; Palacios-Rojas, N.; Feil, R.; Stitt, M. Regulation of secondary metabolism by the carbon-nitrogen status in tobacco: Nitrate inhibits large sectors of phenylpropanoid metabolism. Plant J. 2006, 46, 533–548. [Google Scholar] [CrossRef] [PubMed]
- Peng, M.; Hudson, D.; Schofield, A.; Tsao, R.; Yang, R.; Gu, H.; Bi, Y.M.; Rothstein, S.J. Adaptation of Arabidopsis to nitrogen limitation involves induction of anthocyanin synthesis which is controlled by the NLA gene. J. Exp. Bot. 2008, 59, 2933–2944. [Google Scholar] [CrossRef] [PubMed]
- Soubeyrand, E.; Basteau, C.; Hilbert, G.; van Leeuwen, C.; Delrot, S.; Gomes, E. Nitrogen supply affects anthocyanin biosynthetic and regulatory genes in grapevine cv. Cabernet-Sauvignon berries. Phytochemistry 2014, 103, 38–49. [Google Scholar] [CrossRef]
- Misyura, M.; Colasanti, J.; Rothstein, S.J. Physiological and genetic analysis of Arabidopsis thaliana anthocyanin biosynthesis mutants under chronic adverse environmental conditions. J. Exp. Bot. 2013, 64, 229–240. [Google Scholar] [CrossRef] [PubMed]
- Clément, G.; Moison, M.; Soulay, F.; Reisdorf-Cren, M.; Masclaux-Daubresse, C. Metabolomics of laminae and midvein during leaf senescence and source-sink metabolite management in Brassica napus L. leaves. J. Exp. Bot. 2018, 69, 891–903. [Google Scholar] [CrossRef] [PubMed]
- Guo, Y.; Gan, S.S. Convergence and divergence in gene expression profiles induced by leaf senescence and 27 senescence-promoting hormonal, pathological and environmental stress treatments. Plant Cell Environ. 2012, 35, 644–655. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bieker, S.; Riester, L.; Doll, J.; Franzaring, J.; Fangmeier, A.; Zentgraf, U. Nitrogen Supply Drives Senescence-Related Seed Storage Protein Expression in Rapeseed Leaves. Genes 2019, 10, 72. https://doi.org/10.3390/genes10020072
Bieker S, Riester L, Doll J, Franzaring J, Fangmeier A, Zentgraf U. Nitrogen Supply Drives Senescence-Related Seed Storage Protein Expression in Rapeseed Leaves. Genes. 2019; 10(2):72. https://doi.org/10.3390/genes10020072
Chicago/Turabian StyleBieker, Stefan, Lena Riester, Jasmin Doll, Jürgen Franzaring, Andreas Fangmeier, and Ulrike Zentgraf. 2019. "Nitrogen Supply Drives Senescence-Related Seed Storage Protein Expression in Rapeseed Leaves" Genes 10, no. 2: 72. https://doi.org/10.3390/genes10020072
APA StyleBieker, S., Riester, L., Doll, J., Franzaring, J., Fangmeier, A., & Zentgraf, U. (2019). Nitrogen Supply Drives Senescence-Related Seed Storage Protein Expression in Rapeseed Leaves. Genes, 10(2), 72. https://doi.org/10.3390/genes10020072



