Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae
Abstract
1. Introduction
2. Effect of High Salt Concentration on Green Algae
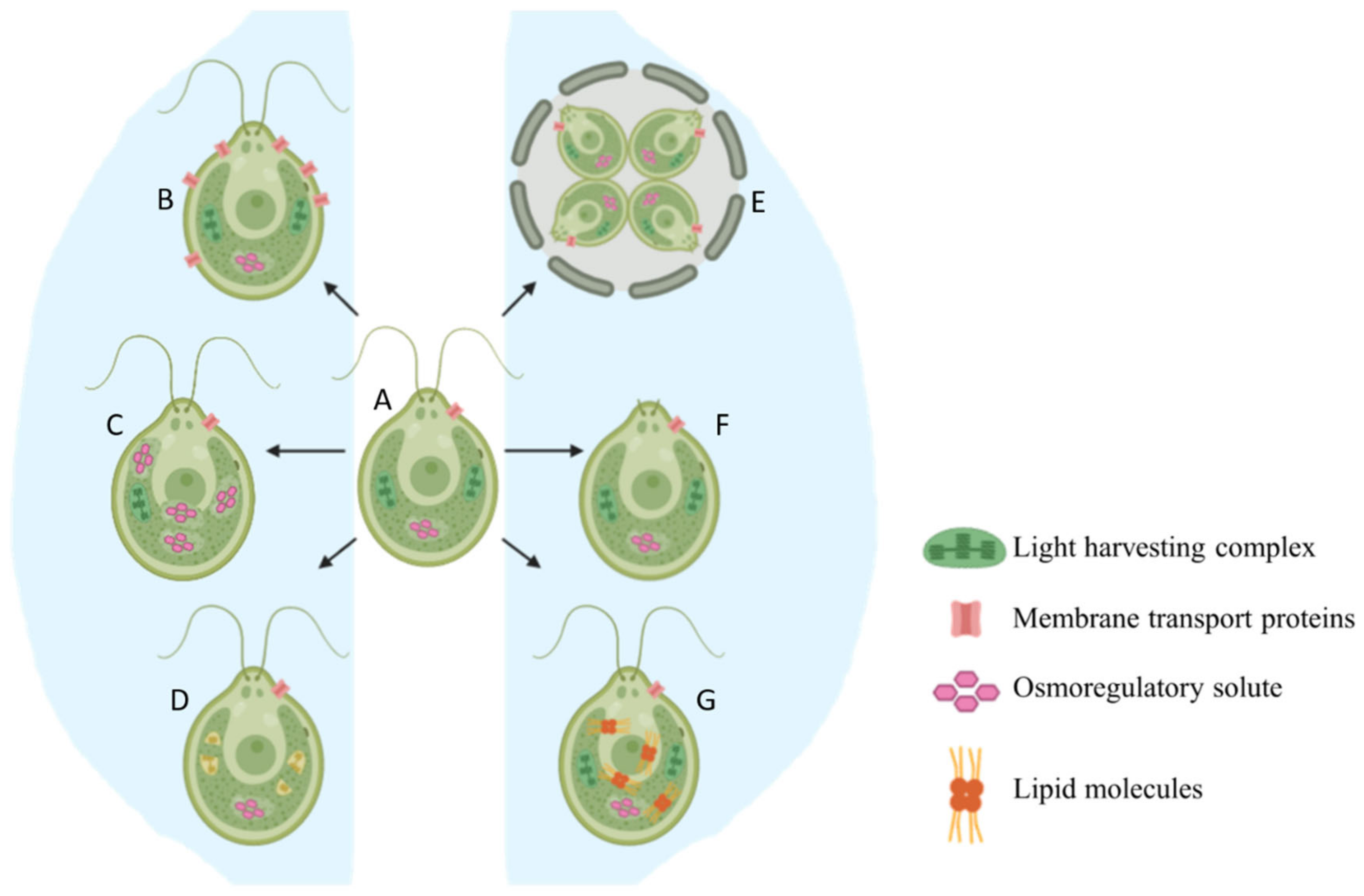
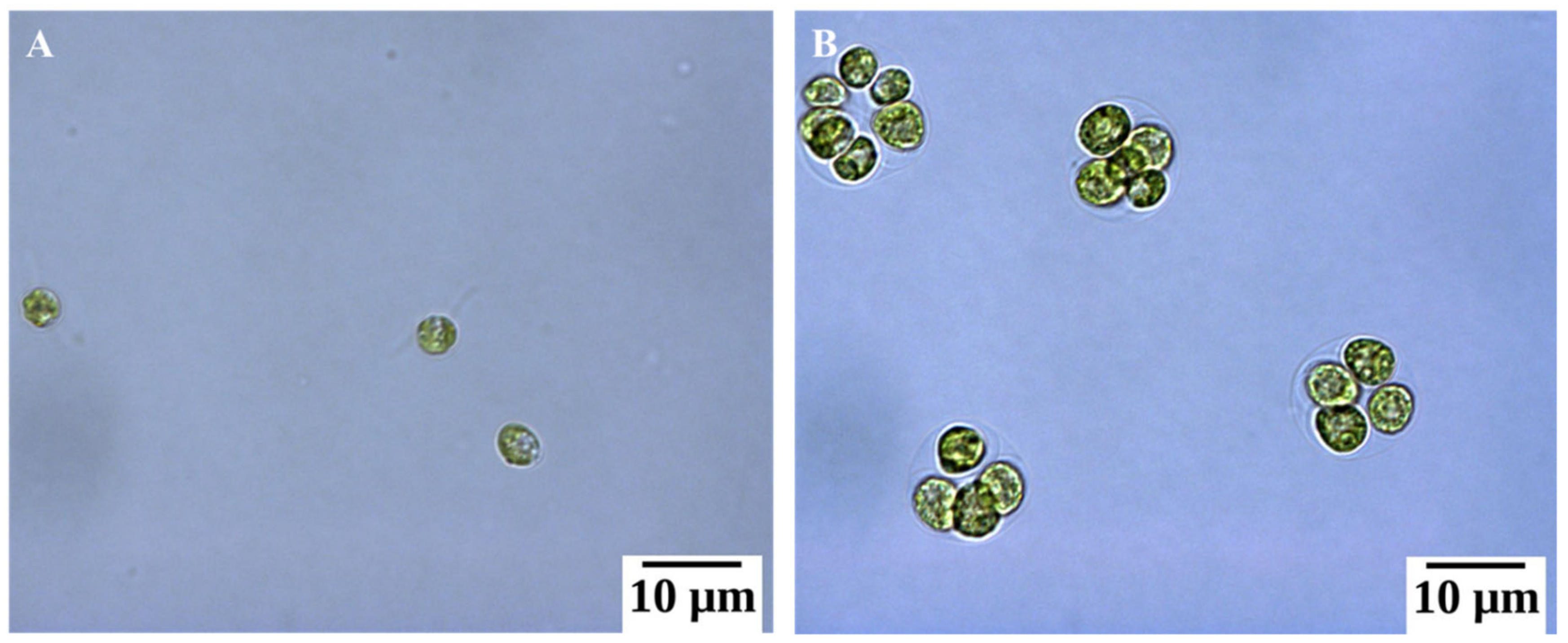
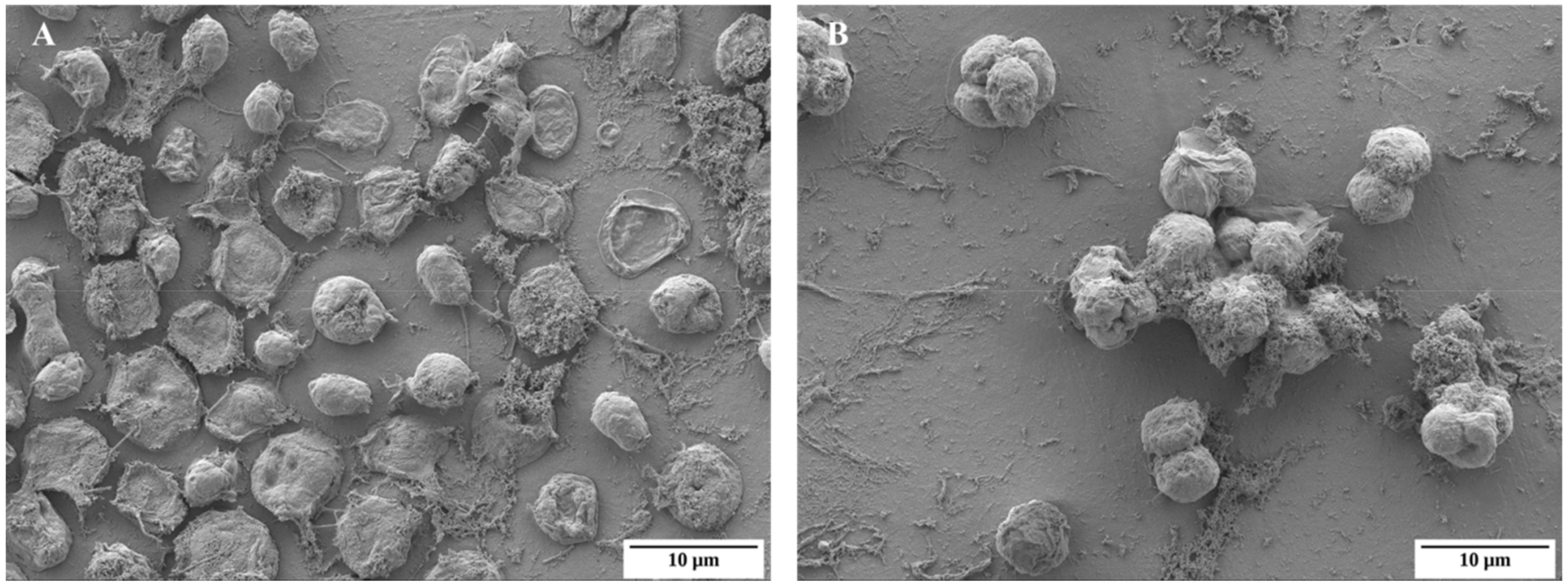
2.1. Basic Morphological Changes
2.2. Production of Osmoregulatory Solutes
2.3. Reorganization of Membrane Transport Proteins
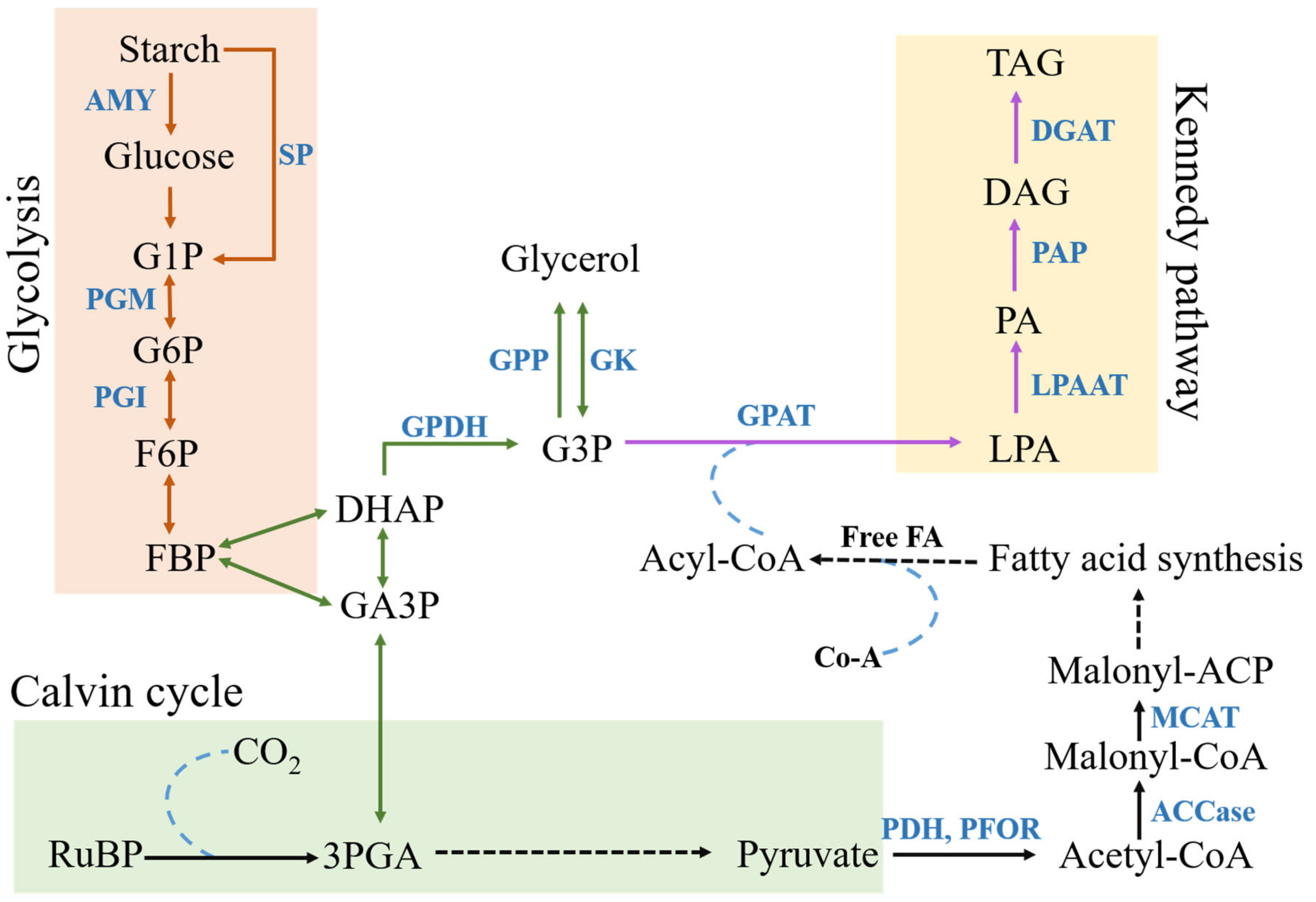
2.4. Lipid Accumulation
2.5. Impact of High Salt Stress on Photosynthesis
2.6. Glycolysis
2.7. Role of Acetate
3. Development of Salt Tolerant Freshwater Algae
4. Perspectives of Salt Stress Utilization in Biotechnological Applications
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brock, T.D. Salinity and the ecology of Dunaliella from Great Salt Lake. Microbiology 1975, 89, 285–292. [Google Scholar] [CrossRef]
- Costelloe, J.F.; Powling, J.; Reid, J.R.W.; Shiel, R.J.; Hudson, P. Algal diversity and assemblages in arid zone rivers of the Lake Eyre Basin, Australia. River Res. Appl. 2005, 21, 337–349. [Google Scholar] [CrossRef]
- Ahmad, I.; Hellebust, J.A. The role of glycerol and inorganic ions in osmoregulatory responses of the euryhaline flagellate Chlamydomonas pulsatilla Wollenweber. Plant Physiol. 1986, 82, 406–410. [Google Scholar] [CrossRef]
- Hellebust, J.A.; Le Gresley, S.M. Growth characteristics of the marine rock pool flagellate Chlamydomonas pulsatilla Wollenweber (Chlorophyta). Phycologia 1985, 24, 225–229. [Google Scholar] [CrossRef]
- Miyasaka, H.; Ohnishi, Y.; Akano, T.; Fukatsu, K.; Mizoguchi, T.; Yagi, K.; Isamu, M.; Yoshiaki, I.; Hiroyo, M.; Norio, S.; et al. Excretion of glycerol by the marine Chlamydomonas sp. strain W-80 in high CO2 cultures. J. Ferment. Bioeng. 1998, 85, 122–124. [Google Scholar] [CrossRef]
- Lee, H.N.; Kim, S.H.; Han, Y.J.; Im, S.; Jeong, W.J.; Park, E.J.; Mi, S.H.; Choi, D.W. PsCYP1 of marine red alga Pyropia seriata (Bangiales, Rhodophyta) confers salt and heat tolerance in Chlamydomonas. J. Appl. Phycol. 2017, 29, 617–625. [Google Scholar] [CrossRef]
- Huang, L.F.; Lin, J.Y.; Pan, K.Y.; Huang, C.K.; Chu, Y.K. Overexpressing ferredoxins in Chlamydomonas reinhardtii increase starch and oil yields and enhance electric power production in a photo microbial fuel cell. Int. J. Mol. Sci. 2015, 16, 19308–19325. [Google Scholar] [CrossRef]
- Tanaka, S.; Suda, Y.; Ikeda, K.; Ono, M.; Miyasaka, H.; Watanabe, M.; Ken, S.; Hirata, K. A novel gene with antisalt and anticadmium stress activities from a halotolerant marine green alga Chlamydomonas sp. W80. FEMS Microbiol. Lett. 2007, 271, 48–52. [Google Scholar] [CrossRef][Green Version]
- Miyasaka, H.; Kanaboshi, H.; Ikeda, K. Isolation of several anti-stress genes from the halotolerant green alga Chlamydomonas by simple functional expression screening with Escherichia coli. World J. Microbiol. Biotechnol. 2000, 16, 23–29. [Google Scholar] [CrossRef]
- Tanaka, S.; Ikeda, K.; Miyasaka, H. Enhanced tolerance against salt-stress and freezing-stress of Escherichia coli cells expressing algal bbc1 gene. Curr. Microbiol. 2001, 42, 173–177. [Google Scholar] [CrossRef]
- Yoshimura, K.; Miyao, K.; Gaber, A.; Takeda, T.; Kanaboshi, H.; Miyasaka, H.; Shigeoka, S. Enhancement of stress tolerance in transgenic tobacco plants overexpressing Chlamydomonas glutathione peroxidase in chloroplasts or cytosol. Plant J. 2004, 37, 21–33. [Google Scholar] [CrossRef]
- Tanaka, S.; Ikeda, K.; Miyasaka, H. Isolation of a new member of group 3 late embryogenesis abundant protein gene from a halotorelant green alga by a functional expression screening with cyanobacterial cells. FEMS Microbiol. Lett. 2004, 236, 41–45. [Google Scholar] [CrossRef][Green Version]
- Hema, R.; Senthil-Kumar, M.; Shivakumar, S.; Reddy, P.C.; Udayakumar, M. Chlamydomonas reinhardtii, a model system for functional validation of abiotic stress responsive genes. Planta 2007, 226, 655–670. [Google Scholar] [CrossRef]
- Khona, D.K.; Shirolikar, S.M.; Gawde, K.K.; Hom, E.; Deodhar, M.A.; D’Souza, J.S. Characterization of salt stress-induced palmelloids in the green alga, Chlamydomonas reinhardtii. Algal Res. 2016, 16, 434–448. [Google Scholar] [CrossRef]
- Nakamura, K.A.Z.U.O.; Bray, D.F.; Wagenaar, E.B. Ultrastructure of Chlamydomonas eugametos palmelloids induced by chloroplatinic acid treatment. J. Bacteriol. 1975, 121, 338–343. [Google Scholar]
- Neelam, S.; Subramanyam, R. Alteration of photochemistry and protein degradation of photosystem II from Chlamydomonas reinhardtii under high salt grown cells. J. Photochem. Photobiol. B Biol. 2013, 124, 63–70. [Google Scholar] [CrossRef]
- Talebi, A.F.; Tabatabaei, M.; Mohtashami, S.K.; Tohidfar, M.; Moradi, F. Comparative salt stress study on intracellular ion concentration in marine and salt-adapted freshwater strains of microalgae. Not. Sci. Biol. 2013, 5, 309–315. [Google Scholar] [CrossRef]
- Demetriou, G.; Neonaki, C.; Navakoudis, E.; Kotzabasis, K. Salt stress impact on the molecular structure and function of the photosynthetic apparatus—The protective role of polyamines. Biochim. Biophys. Acta (BBA)-Bioenerg. 2007, 1767, 272–280. [Google Scholar] [CrossRef]
- Tordai, H.; Bányai, L.; Patthy, L. The PAN module: The N-terminal domains of plasminogen and hepatocyte growth factor are homologous with the apple domains of the prekallikrein family and with a novel domain found in numerous nematode proteins. FEBS Lett. 1999, 461, 63–67. [Google Scholar] [CrossRef]
- Verma, R.P.; Hansch, C. Matrix metalloproteinases (MMPs): Chemical–biological functions and (Q) SARs. Bioorg. Med. Chem. 2007, 15, 2223–2268. [Google Scholar] [CrossRef]
- Ahmad, I.; Hellebust, J.A. Osmoregulation in the extremely euryhaline marine micro-alga Chlorella autotrophica. Plant Physiol. 1984, 74, 1010–1015. [Google Scholar] [CrossRef] [PubMed]
- Kaçka, A.; Dönmez, G. Isolation of Dunaliella spp. from a hypersaline lake and their ability to accumulate glycerol. Bioresour. Technol. 2008, 99, 8348–8352. [Google Scholar] [PubMed]
- Fu, W.; Paglia, G.; Magnúsdóttir, M.; Steinarsdóttir, E.A.; Gudmundsson, S.; Palsson, B.Ø.; Brynjólfsson, S. Effects of abiotic stressors on lutein production in the green microalga Dunaliella salina. Microb. Cell Factories 2014, 13, 3. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.D.; Simpson, J.R. Water relations of sugar-tolerant yeasts: The role of intracellular polyols. Microbiology 1972, 72, 589–591. [Google Scholar] [CrossRef]
- Läuchli, A.; Lüttge, U. Salinity: Environment-Plants-Molecules; Kluwer Academic Publishers: Dordrecht, The Netherlands, 2002. [Google Scholar]
- Miyasaka, H.; Ikeda, K. Osmoregulating mechanism of the halotolerant green alga Chlamydomonas, strain HS-5. Plant Sci. 1997, 127, 91–96. [Google Scholar]
- León, R.; Galván, F. Halotolerance studies on Chlamydomonas reinhardtii: Glycerol excretion by free and immobilized cells. J. Appl. Phycol. 1994, 6, 13–20. [Google Scholar] [CrossRef]
- Salama, E.S.; Kim, H.C.; Abou-Shanab, R.A.; Ji, M.K.; Oh, Y.K.; Kim, S.H.; Jeon, B.H. Biomass, lipid content, and fatty acid composition of freshwater Chlamydomonas mexicana and Scenedesmus obliquus grown under salt stress. Bioprocess Biosyst. Eng. 2013, 36, 827–833. [Google Scholar] [CrossRef]
- Ho, S.H.; Nakanishi, A.; Kato, Y.; Yamasaki, H.; Chang, J.S.; Misawa, N.; Yuu, H.; Jun, M.; Tomohisda, H.; Kondo, A. Dynamic metabolic profiling together with transcription analysis reveals salinity-induced starch-to-lipid biosynthesis in alga Chlamydomonas sp. JSC4. Sci. Rep. 2017, 7, 45471. [Google Scholar] [CrossRef]
- Ben-Amotz, A.; Avron, M. The role of glycerol in the osmotic regulation of the halophilic alga Dunaliella parva. Plant Physiol. 1973, 51, 875–878. [Google Scholar] [CrossRef]
- Hellebust, J.A.; Lin, Y.-H. Regulation of glycerol and starch metabolism in Chlamydomonas pulsatilla in response to changes in salinity. Plant Cell Environ. 1989, 12, 621–627. [Google Scholar] [CrossRef]
- Chen, H.; Lu, Y.; Jiang, J.G. Comparative analysis on the key enzymes of the glycerol cycle metabolic pathway in Dunaliella salina under osmotic stresses. PLoS ONE 2012, 7, e37578. [Google Scholar]
- Driver, T.; Trivedi, D.K.; McIntosh, O.A.; Dean, A.P.; Goodacre, R.; Pittman, J.K. Two glycerol-3-phosphate dehydrogenases from Chlamydomonas have distinct roles in lipid metabolism. Plant Physiol. 2017, 174, 2083–2097. [Google Scholar] [CrossRef] [PubMed]
- Casais-Molina, M.L.; Peraza-Echeverria, S.; Echevarría-Machado, I.; Herrera-Valencia, V.A. Expression of Chlamydomonas reinhardtii CrGPDH2 and CrGPDH3 cDNAs in yeast reveals that they encode functional glycerol-3-phosphate dehydrogenases involved in glycerol production and osmotic stress tolerance. J. Appl. Phycol. 2016, 28, 219–226. [Google Scholar] [CrossRef]
- Cohu, C.M.; Muller, O.; Stewart, J.J.; Demmig-Adams, B.; Adams III, W.W. Association between minor loading vein architecture and light-and CO2-saturated rates of photosynthetic oxygen evolution among Arabidopsis thaliana ecotypes from different latitudes. Front. Plant Sci. 2013, 4, 264. [Google Scholar] [CrossRef] [PubMed]
- Adams, W.W., III; Cohu, C.M.; Muller, O.; Demmig-Adams, B. Foliar phloem infrastructure in support of photosynthesis. Front. Plant Sci. 2013, 4, 194. [Google Scholar] [CrossRef]
- Burch, T.A.; Adams, W.W.; Degrenne, B.L.; Englert, C.H.; Mines, B.R.; Nash, P.C.; Emma, C.B.; Demmig-Adams, B. Environmental manipulation of growth and energy carrier release from freshwater and marine Chlamydomonas species. J. Appl. Phycol. 2015, 27, 1127–1136. [Google Scholar] [CrossRef]
- Grizeau, D.; Navarro, J.M. Glycerol production by Dunaliella tertiolecta immobilized within Ca-alginate beads. Biotechnol. Lett. 1986, 8, 261–264. [Google Scholar] [CrossRef]
- Liu, J.; Zhu, J.K. Proline accumulation and salt-stress-induced gene expression in a salt-hypersensitive mutant of Arabidopsis. Plant Physiol. 1997, 114, 591–596. [Google Scholar] [CrossRef]
- Brown, L.M.; Hellebust, J.A. Sorbitol and proline as intracellular osmotic solutes in the green alga Stichococcus bacillaris. Can. J. Bot. 1978, 56, 676–679. [Google Scholar] [CrossRef]
- Reynoso, G.T.; De Gamboa, B.A. Salt tolerance in the freshwater algae Chlamydomonas reinhardii: Effect of proline and taurine. Comp. Biochem. Physiol. Part A Physiol. 1982, 73, 95–99. [Google Scholar] [CrossRef]
- Aggarwal, M.; Sharma, S.; Kaur, N.; Pathania, D.; Bhandhari, K.; Kaushal, N.; Ramanpreet, K.; Kamaljit, S.; Alok, S.; Nayyar, H. Exogenous proline application reduces phytotoxic effects of selenium by minimising oxidative stress and improves growth in bean (Phaseolus vulgaris L.) seedlings. Biol. Trace Elem. Res. 2011, 140, 354–367. [Google Scholar] [CrossRef]
- Henley, W.J.; Hironaka, J.L.; Guillou, L.; Buchheim, M.A.; Buchheim, J.A.; Fawley, M.W.; Fawley, K.P. Phylogenetic analysis of the ‘Nannochloris-like’algae and diagnoses of Picochlorum oklahomensis gen. et sp. nov. (Trebouxiophyceae, Chlorophyta). Phycologia 2004, 43, 641–652. [Google Scholar] [CrossRef]
- Foflonker, F.; Ananyev, G.; Qiu, H.; Morrison, A.; Palenik, B.; Dismukes, G.C.; Bhattacharya, D. The unexpected extremophile: Tolerance to fluctuating salinity in the green alga Picochlorum. Algal Res. 2016, 16, 465–472. [Google Scholar] [CrossRef]
- Goyal, A. Osmoregulation in Dunaliella, Part II: Photosynthesis and starch contribute carbon for glycerol synthesis during a salt stress in Dunaliella tertiolecta. Plant Physiol. Biochem. 2007, 45, 705–710. [Google Scholar] [CrossRef] [PubMed]
- Xia, B.B.; Wang, S.H.; Duan, J.B.; Bai, L.H. The relationship of glycerol and glycolysis metabolism pathway under hyperosmotic stress in Dunaliella salina. Cent. Eur. J. Biol. 2014, 9, 901–908. [Google Scholar]
- Kaushik, J.K.; Bhat, R. Why is trehalose an exceptional protein stabilizer? An analysis of the thermal stability of proteins in the presence of the compatible osmolyte trehalose. J. Biol. Chem. 2003, 278, 26458–26465. [Google Scholar] [CrossRef]
- Wang, N.; Qian, Z.; Luo, M.; Fan, S.; Zhang, X.; Zhang, L. Identification of salt stress responding genes using transcriptome analysis in green alga Chlamydomonas reinhardtii. Int. J. Mol. Sci. 2018, 19, 3359. [Google Scholar] [CrossRef]
- del Pilar Bremauntz, M.; Torres-Bustillos, L.G.; Cañizares-Villanueva, R.O.; Duran-Paramo, E.; Fernández-Linares, L. Trehalose and sucrose osmolytes accumulated by algae as potential raw material for bioethanol. Nat. Resour. 2011, 2, 173. [Google Scholar]
- Zeid, I.M. Trehalose as osmoprotectant for maize under salinity-induced stress. Res. J. Agric. Biol. Sci. 2009, 5, 613–622. [Google Scholar]
- Hellebust, J.A. Effect of salinity on photosynthesis and mannitol synthesis in the green flagellate Platymonas suecica. Can. J. Bot. 1976, 54, 1735–1741. [Google Scholar] [CrossRef]
- Brown, L.M.; Hellebust, J.A. Some new taxonomic characteristics applied to Stichococcus bacillaris (Chlorophyceae). Can. J. Bot. 1980, 58, 1405–1411. [Google Scholar] [CrossRef]
- Chakraborty, K.; Bose, J.; Shabala, L.; Shabala, S. Difference in root K+ retention ability and reduced sensitivity of K+-permeable channels to reactive oxygen species confer differential salt tolerance in three Brassica species. J. Exp. Bot. 2016, 67, 4611–4625. [Google Scholar] [CrossRef] [PubMed]
- Shabala, S. Regulation of potassium transport in leaves: From molecular to tissue level. Ann. Bot. 2003, 92, 627–634. [Google Scholar] [CrossRef] [PubMed]
- Sithtisarn, S.; Yokthongwattana, K.; Mahong, B.; Roytrakul, S.; Paemanee, A.; Phaonakrop, N.; Yokthongwattana, C. Comparative proteomic analysis of Chlamydomonas reinhardtii control and a salinity-tolerant strain revealed a differential protein expression pattern. Planta 2017, 246, 843–856. [Google Scholar] [CrossRef] [PubMed]
- Pick, U.; Karni, L.; Avron, M. Determination of ion content and ion fluxes in the halotolerant alga Dunaliella salina. Plant Physiol. 1986, 81, 92–96. [Google Scholar] [CrossRef]
- Fisher, M.; Gokhman, I.; Pick, U.; Zamir, A. A salt-resistant plasma membrane carbonic anhydrase is induced by salt in Dunaliella salina. J. Biol. Chem. 1996, 271, 17718–17723. [Google Scholar] [CrossRef]
- Fisher, M.; Gokhman, I.; Pick, U.; Zamir, A. A structurally novel transferrin-like protein accumulates in the plasma membrane of the unicellular green alga Dunaliella salina grown in high salinities. J. Biol. Chem. 1997, 272, 1565–1570. [Google Scholar] [CrossRef]
- Sun, X.; Wang, W.; Shen, J.; Xu, N. Transcriptome sequencing of the marine microalga, Chlorella pyrenoidosa (Chlorophyta), and analysis of carbonic anhydrase expression under salt stress. Bot. Mar. 2014, 57, 403–412. [Google Scholar] [CrossRef]
- Moroney, J.V.; Ma, Y.; Frey, W.D.; Fusilier, K.A.; Pham, T.T.; Simms, T.A.; Robert, J.D.; Jing, Y.; Bratati, M. The carbonic anhydrase isoforms of Chlamydomonas reinhardtii: Intracellular location, expression, and physiological roles. Photosynth. Res. 2011, 109, 133–149. [Google Scholar] [CrossRef]
- Goncalves, E.C.; Wilkie, A.C.; Kirst, M.; Rathinasabapathi, B. Metabolic regulation of triacylglycerol accumulation in the green algae: Identification of potential targets for engineering to improve oil yield. Plant Biotechnol. J. 2016, 14, 1649–1660. [Google Scholar] [CrossRef]
- Ho, S.H.; Nakanishi, A.; Ye, X.; Chang, J.S.; Hara, K.; Hasunuma, T.; Kondo, A. Optimizing biodiesel production in marine Chlamydomonas sp. JSC4 through metabolic profiling and an innovative salinity-gradient strategy. Biotechnol. Biofuels 2014, 7, 97. [Google Scholar] [CrossRef]
- de Jaeger, L.; Carreres, B.M.; Springer, J.; Schaap, P.J.; Eggink, G.; Dos Santos, V.A.M.; Rene, H.W.; Dirk, E.M. Neochloris oleoabundans is worth its salt: Transcriptomic analysis under salt and nitrogen stress. PLoS ONE 2018, 13, e0194834. [Google Scholar] [CrossRef] [PubMed]
- Perrineau, M.M.; Zelzion, E.; Gross, J.; Price, D.C.; Boyd, J.; Bhattacharya, D. Evolution of salt tolerance in a laboratory reared population of Chlamydomonas reinhardtii. Environ. Microbiol. 2014, 16, 1755–1766. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yuan, Y.; Cheng, D.; Gao, J.; Kong, L.; Zhao, Q.; Wei, W.; Sun, Y. Exploring stress tolerance mechanism of evolved freshwater strain Chlorella sp. S30 under 30 g/L salt. Bioresour. Technol. 2018, 250, 495–504. [Google Scholar] [CrossRef] [PubMed]
- Kato, Y.; Ho, S.H.; Vavricka, C.J.; Chang, J.S.; Hasunuma, T.; Kondo, A. Evolutionary engineering of salt-resistant Chlamydomonas sp. strains reveals salinity stress-activated starch-to-lipid biosynthesis switching. Bioresour. Technol. 2017, 245, 1484–1490. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.M.; Ren, L.J.; Bi, Z.Q.; Ji, X.J.; Zhao, Q.Y.; Huang, H. Adaptive evolution of microalgae Schizochytrium sp. under high salinity stress to alleviate oxidative damage and improve lipid biosynthesis. Bioresour. Technol. 2018, 267, 438–444. [Google Scholar] [CrossRef]
- Sudhir, P.; Murthy, S.D.S. Effects of salt stress on basic processes of photosynthesis. Photosynthetica 2004, 42, 481–486. [Google Scholar] [CrossRef]
- Subramanyam, R.; Jolley, C.; Thangaraj, B.; Nellaepalli, S.; Webber, A.N.; Fromme, P. Structural and functional changes of PSI-LHCI supercomplexes of Chlamydomonas reinhardtii cells grown under high salt conditions. Planta 2010, 231, 913–922. [Google Scholar] [CrossRef]
- Ye, Z.-W.; Jiang, J.G.; Wu, G.H. Biosynthesis and regulation of carotenoids in Dunaliella: Progresses and prospects. Biotechnol. Adv. 2008, 26, 352–360. [Google Scholar] [CrossRef]
- Mishra, A.; Mandoli, A.; Jha, B. Physiological characterization and stress-induced metabolic responses of Dunaliella salina isolated from salt pan. J. Ind. Microbiol. Biotechnol. 2008, 35, 1093. [Google Scholar] [CrossRef]
- Hiremath, S.; Mathad, P. Impact of salinity on the physiological and biochemical traits of Chlorella vulgaris Beijerinck. J. Algal Biomass Utln 2010, 1, 51–59. [Google Scholar]
- Plaxton, W.C. The organization and regulation of plant glycolysis. Annu. Rev. Plant Biol. 1996, 47, 185–214. [Google Scholar] [CrossRef] [PubMed]
- Capriotti, A.L.; Borrelli, G.M.; Colapicchioni, V.; Papa, R.; Piovesana, S.; Samperi, R.; Serena, S.; Aldo, L. Proteomic study of a tolerant genotype of durum wheat under salt-stress conditions. Anal. Bioanal. Chem. 2014, 406, 1423–1435. [Google Scholar] [CrossRef] [PubMed]
- Espartero, J.; Sánchez-Aguayo, I.; Pardo, J.M. Molecular characterization of glyoxalase-I from a higher plant; upregulation by stress. Plant Mol. Biol. 1995, 29, 1223–1233. [Google Scholar] [CrossRef] [PubMed]
- Heifetz, P.B.; Förster, B.; Osmond, C.B.; Giles, L.J.; Boynton, J.E. Effects of Acetate on Facultative Autotrophy in Chlamydomonas reinhardtii Assessed by Photosynthetic Measurements and Stable Isotope Analyses. Plant Physiol. 2000, 122, 1439–1446. [Google Scholar] [CrossRef] [PubMed]
- Talarski, A.; Manning, S.R.; La Claire, J.W. Transcriptome analysis of the euryhaline alga, Prymnesium parvum (Prymnesiophyceae): Effects of salinity on differential gene expression. Phycologia 2016, 55, 33–44. [Google Scholar] [CrossRef]
- Tanaka, A.; Shikazono, N.; Hase, Y. Studies on biological effects of ion beams on lethality, molecular nature of mutation, mutation rate, and spectrum of mutation phenotype for mutation breeding in higher plants. J. Radiat. Res. 2010, 51, 223–233. [Google Scholar] [CrossRef]
- Su, H.Y.; Chou, H.H.; Chow, T.J.; Lee, T.M.; Chang, J.S.; Huang, W.L.; Chen, H.J. Improvement of outdoor culture efficiency of cyanobacteria by over-expression of stress tolerance genes and its implication as bio-refinery feedstock. Bioresour. Technol. 2017, 244, 1294–1303. [Google Scholar] [CrossRef]
- Takouridis, S.J.; Tribe, D.E.; Gras, S.L.; Martin, G.J. The selective breeding of the freshwater microalga Chlamydomonas reinhardtii for growth in salinity. Bioresour. Technol. 2015, 184, 18–22. [Google Scholar] [CrossRef]
- Trainor, F.R. Reproduction in Scenedesmus. Algae Korean J. Phycol. 1996, 11, 183–201. [Google Scholar]
- Adair, W.S.; Hwang, C.; Goodenough, U.W. Identification and visualization of the sexual agglutinin from the mating-type plus flagellar membrane of Chlamydomonas. Cell 1983, 33, 183–193. [Google Scholar] [CrossRef]
- Oren, A. A hundred years of Dunaliella research: 1905–2005. Saline Syst. 2005, 1, 2. [Google Scholar] [CrossRef] [PubMed]
- Pisal, D.S.; Lele, S.S. Carotenoid production from microalga, Dunaliella salina. Indian J. Biotechnol. 2005, 4, 476–483. [Google Scholar]
- Afify, A.E.M.M.; Shalaby, E.A.; Shanab, S.M. Enhancement of biodiesel production from different species of algae. Grasas Y Aceites 2010, 61, 416–422. [Google Scholar]
- von Alvensleben, N.; Stookey, K.; Magnusson, M.; Heimann, K. Salinity tolerance of Picochlorum atomus and the use of salinity for contamination control by the freshwater cyanobacterium Pseudanabaena limnetica. PLoS ONE 2013, 8, e63569. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Z.J.; Zhu, Y.R.; Bai, Y.L.; Wang, Y. Volatile communication between Chlamydomonas reinhardtii cells under salt stress. Biochem. Syst. Ecol. 2012, 40, 19–24. [Google Scholar] [CrossRef]

| Gene Name | Organism From | Organism Transformed | Effect Observed | Reference |
|---|---|---|---|---|
| Cyclophilins (CYP) PsCYP1 | Pyropia seriata (marine red algae) | Chlamydomonas sp. | Heat and salt tolerance | [6] |
| Photosynthetic ferredoxin (PETF) and ferredoxin-5 gene (FDX5) | Chlamydomonas sp. | Chlamydomonas sp. | Salt tolerance | [7] |
| Gene with unknown function | Chlamydomonas W80 | E. coli | Salt and cadmium tolerance | [8] |
| Anti-stress genes | Chlamydomonas W80 | E. coli | Salt stress protection | [9] |
| Breast basic conserved gene (bbc1) | Chlamydomonas W80 | E. coli | Salt stress protection | [10] |
| Glutathione peroxidase | Chlamydomonas W80 | Tobacco plant | Salt stress protection | [11] |
| Group-3 late embryogenesis abundant protein gene (cw80lea3) | Chlamydomonas W80 | Synechococcus PCC7942 | Salt and cold stress protection | [12] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shetty, P.; Gitau, M.M.; Maróti, G. Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae. Cells 2019, 8, 1657. https://doi.org/10.3390/cells8121657
Shetty P, Gitau MM, Maróti G. Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae. Cells. 2019; 8(12):1657. https://doi.org/10.3390/cells8121657
Chicago/Turabian StyleShetty, Prateek, Margaret Mukami Gitau, and Gergely Maróti. 2019. "Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae" Cells 8, no. 12: 1657. https://doi.org/10.3390/cells8121657
APA StyleShetty, P., Gitau, M. M., & Maróti, G. (2019). Salinity Stress Responses and Adaptation Mechanisms in Eukaryotic Green Microalgae. Cells, 8(12), 1657. https://doi.org/10.3390/cells8121657





