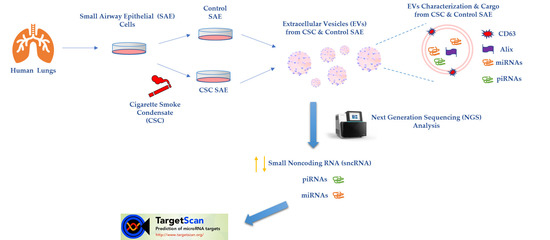
Cigarette Smoke Condensate Exposure Changes RNA Content of Extracellular Vesicles Released from Small Airway Epithelial Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Small Airway Epithelial Cultures and Stimulation with Cigarette Smoke Condensate
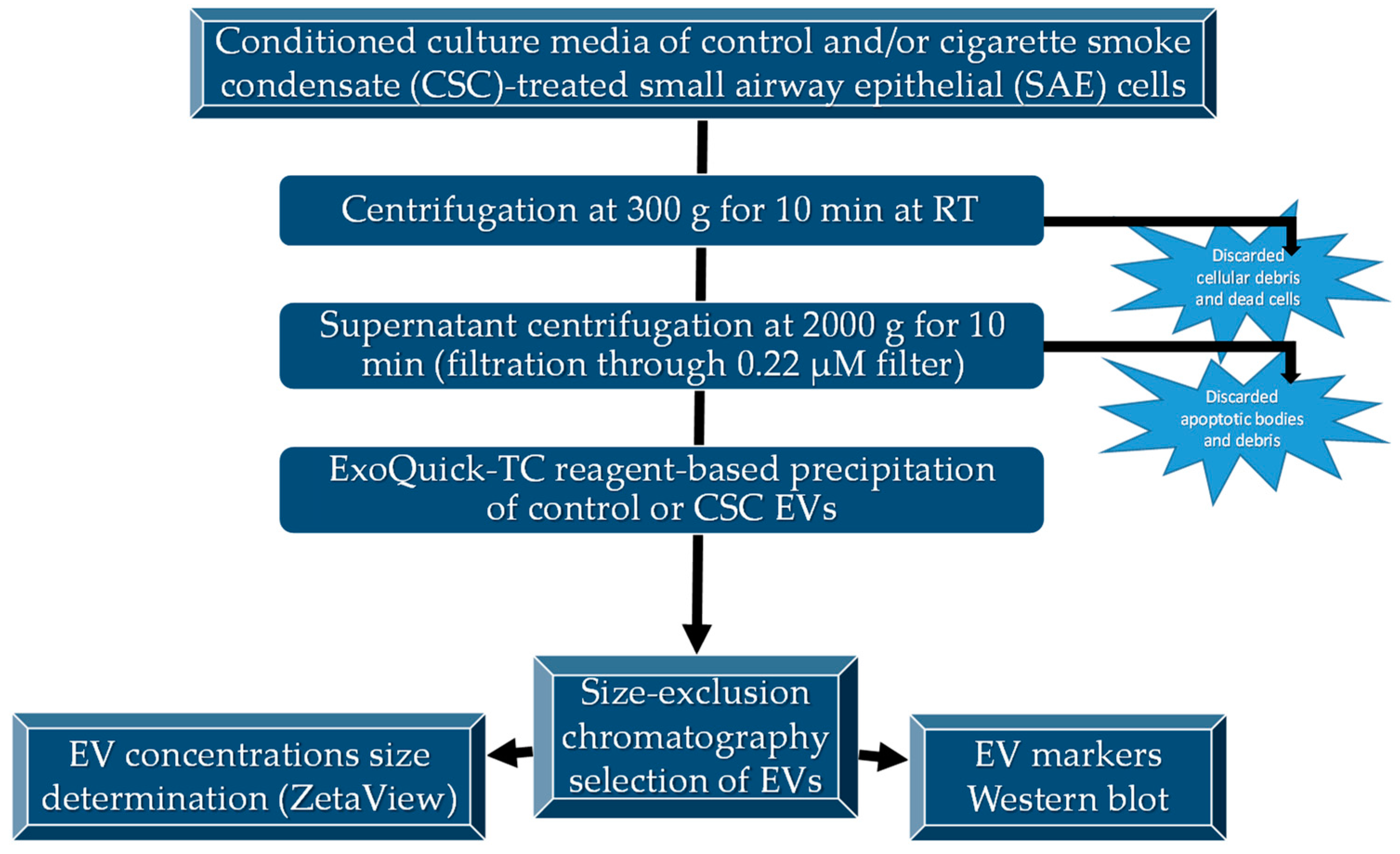
2.2. Extracellular Vesicles Purification
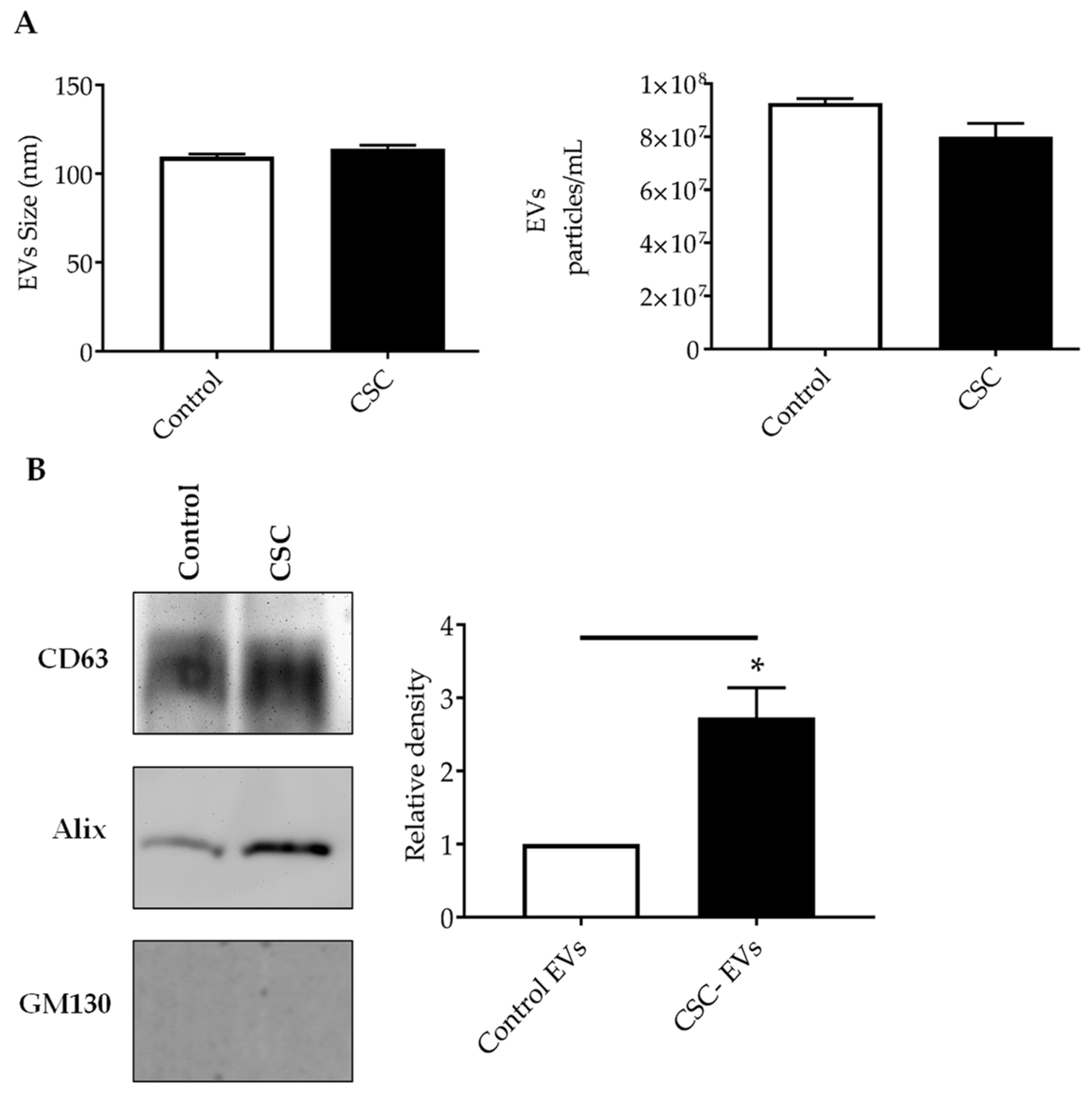
2.3. Nanoparticle Tracking Analysis with ZetaView®
2.4. Western Blot Analysis for EVs Markers
2.5. Extraction of EVs RNA and Next Generation Sequencing (NGS)
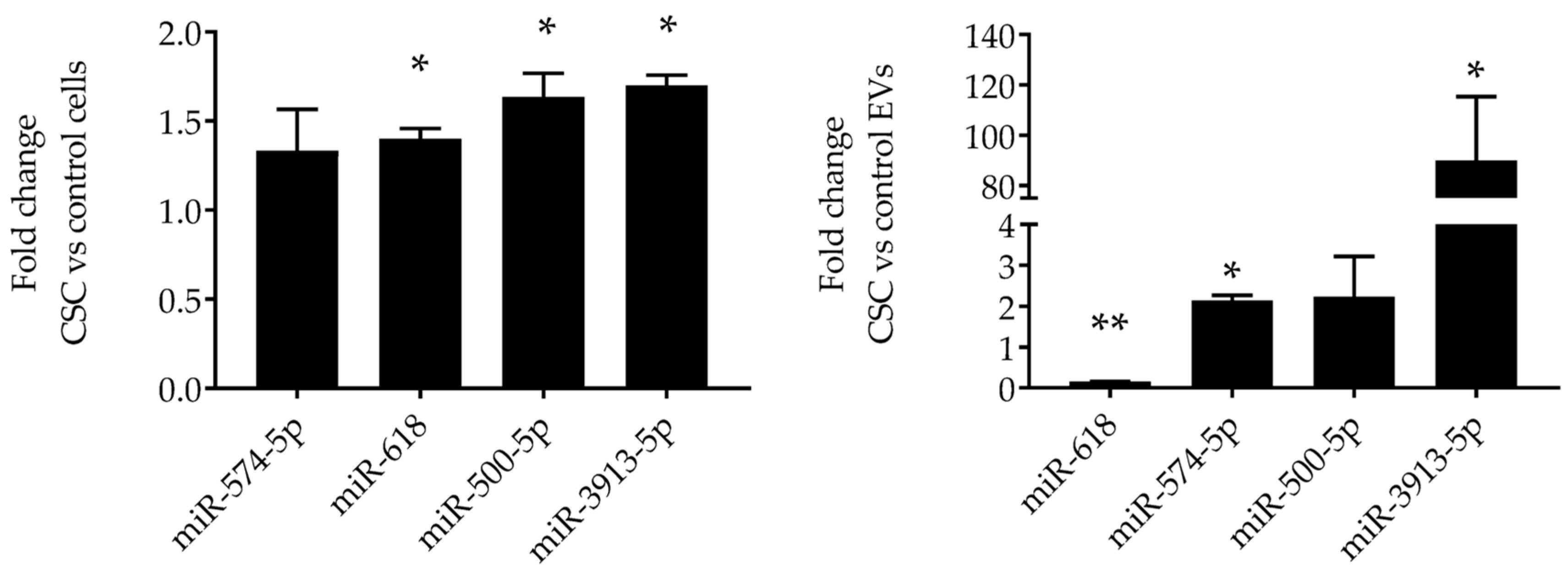
2.6. Reverse Transcription (RT)-PCR
2.7. Predicted Targets of Hsa-miR-3913-5p and Functional Analysis
2.8. Statistical Analysis
3. Results
3.1. Characteristics of EVs Released by SAE Cells under Control and CSC Conditions
3.2. EVs from Control and CSC-Treated SAE Cells Contain Small RNAs
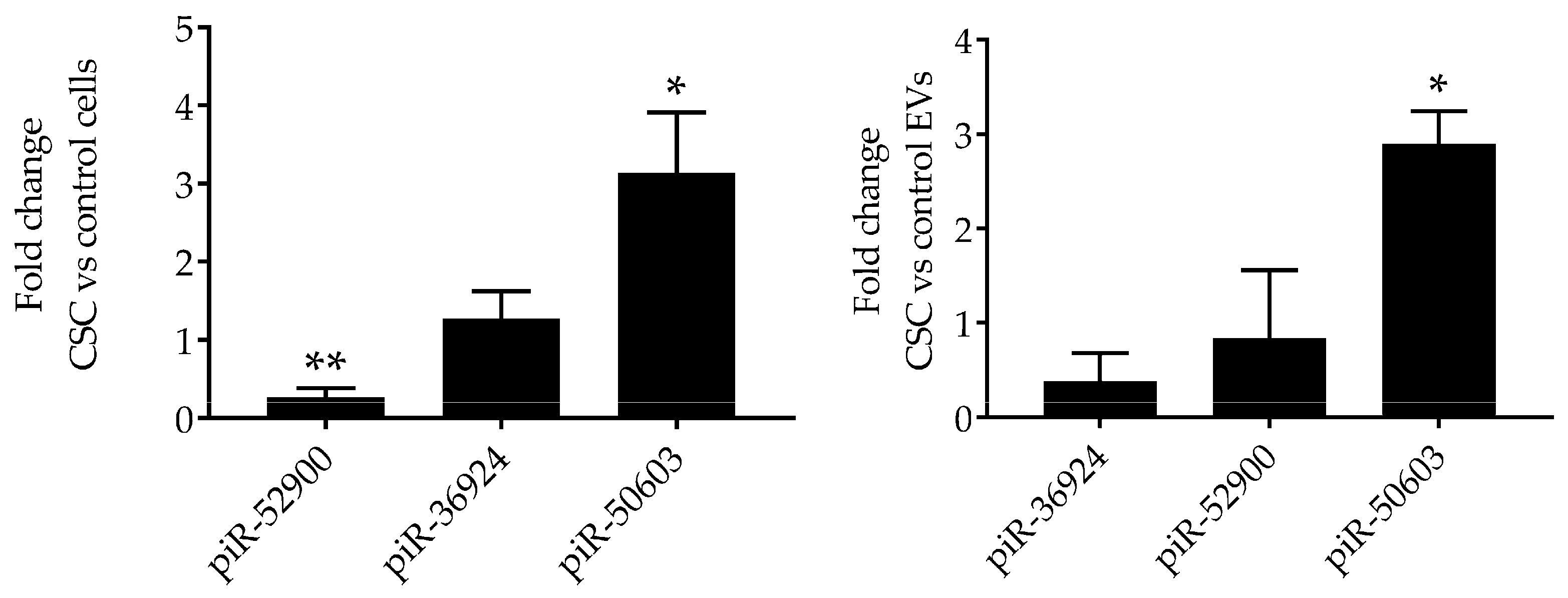
3.3. CSC Induces Changes in miRNAs Expression Profile in EVs from CSC-Treated SAE Cells
3.4. Potential Targets and Gene Ontology Prediction of the Upregulated Hsa-miR-3913-5p in CSC EVs
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Margolis, L.; Sadovsky, Y. The biology of extracellular vesicles: The known unknowns. PLoS Biol. 2019, 17, e3000363. [Google Scholar] [CrossRef] [PubMed]
- Chiang, C.-Y.; Chen, C. Toward characterizing extracellular vesicles at a single-particle level. J. Biomed. Sci. 2019, 26, 9. [Google Scholar] [CrossRef] [PubMed]
- Tkach, M.; Thery, C. Communication by Extracellular Vesicles: Where We Are and Where We Need to Go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef] [PubMed]
- Skalnikova, H.K.; Bohuslavova, B.; Turnovcova, K.; Juhasova, J.; Juhas, S.; Rodinova, M.; Vodicka, P. Isolation and Characterization of Small Extracellular Vesicles from Porcine Blood Plasma, Cerebrospinal Fluid, and Seminal Plasma. Proteomes 2019, 7, 17. [Google Scholar] [CrossRef] [PubMed]
- Wang, X. Isolation of Extracellular Vesicles from Breast Milk. Methods Mol. Biol. 2017, 1660, 351–353. [Google Scholar] [CrossRef] [PubMed]
- Parimon, T.; Garrett Iii, N.E.; Chen, P.; Antes, T.J. Isolation of Extracellular Vesicles from Murine Bronchoalveolar Lavage Fluid Using an Ultrafiltration Centrifugation Technique. JoVE 2018. [Google Scholar] [CrossRef]
- Iwai, K.; Yamamoto, S.; Yoshida, M.; Shiba, K. Isolation of Extracellular Vesicles in Saliva Using Density Gradient Ultracentrifugation. Methods Mol. Biol. 2017, 1660, 343–350. [Google Scholar] [CrossRef]
- Markowska, A.; Pendergrast, R.S.; Pendergrast, J.S.; Pendergrast, P.S. A novel method for the isolation of extracellular vesicles and RNA from urine. J. Circ. Biomark. 2017, 6. [Google Scholar] [CrossRef]
- Kosanovic, M.; Milutinovic, B.; Goc, S.; Mitic, N.; Jankovic, M. Ion-exchange chromatography purification of extracellular vesicles. Biotechniques 2017, 63, 65–71. [Google Scholar] [CrossRef]
- Thery, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Lotvall, J.; Hill, A.F.; Hochberg, F.; Buzas, E.I.; Di Vizio, D.; Gardiner, C.; Gho, Y.S.; Kurochkin, I.V.; Mathivanan, S.; Quesenberry, P.; et al. Minimal experimental requirements for definition of extracellular vesicles and their functions: A position statement from the International Society for Extracellular Vesicles. J. Extracell. Vesicles 2014, 3, 26913. [Google Scholar] [CrossRef] [PubMed]
- Yanez-Mo, M.; Siljander, P.R.; Andreu, Z.; Zavec, A.B.; Borras, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef] [PubMed]
- Pitt, J.M.; Kroemer, G.; Zitvogel, L. Extracellular vesicles: Masters of intercellular communication and potential clinical interventions. J. Clin. Investig. 2016, 126, 1139–1143. [Google Scholar] [CrossRef] [PubMed]
- Paolicelli, R.C.; Bergamini, G.; Rajendran, L. Cell-to-cell Communication by Extracellular Vesicles: Focus on Microglia. Neuroscience 2019, 405, 148–157. [Google Scholar] [CrossRef]
- Candelario, K.M.; Steindler, D.A. The role of extracellular vesicles in the progression of neurodegenerative disease and cancer. Trends Mol. Med. 2014, 20, 368–374. [Google Scholar] [CrossRef]
- Han, L.; Lam, E.W.; Sun, Y. Extracellular vesicles in the tumor microenvironment: Old stories, but new tales. Mol. Cancer 2019, 18, 59. [Google Scholar] [CrossRef]
- Hadley, E.E.; Sheller-Miller, S.; Saade, G.; Salomon, C.; Mesiano, S.; Taylor, R.N.; Taylor, B.D.; Menon, R. Amnion epithelial cell-derived exosomes induce inflammatory changes in uterine cells. Am. J. Obstet. Gynecol. 2018, 219, 478.e1–478.e21. [Google Scholar] [CrossRef]
- Thery, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef]
- Todorova, D.; Simoncini, S.; Lacroix, R.; Sabatier, F.; Dignat-George, F. Extracellular Vesicles in Angiogenesis. Circ. Res. 2017, 120, 1658–1673. [Google Scholar] [CrossRef]
- Chahar, H.S.; Corsello, T.; Kudlicki, A.S.; Komaravelli, N.; Casola, A. Respiratory Syncytial Virus Infection Changes Cargo Composition of Exosome Released from Airway Epithelial Cells. Sci. Rep. 2018, 8, 387. [Google Scholar] [CrossRef]
- Vader, P.; Mol, E.A.; Pasterkamp, G.; Schiffelers, R.M. Extracellular vesicles for drug delivery. Adv. Drug Deliv. Rev. 2016, 106, 148–156. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Kim, H.S. Engineering of extracellular vesicles as drug delivery vehicles. Stem. Cell Investig. 2017, 4, 74. [Google Scholar] [CrossRef] [PubMed]
- Di Leva, G.; Garofalo, M.; Croce, C.M. MicroRNAs in cancer. Annu. Rev. Pathol. 2014, 9, 287–314. [Google Scholar] [CrossRef] [PubMed]
- Bartel, S.; La Grutta, S.; Cilluffo, G.; Perconti, G.; Bongiovanni, A.; Giallongo, A.; Behrends, J.; Kruppa, J.; Hermann, S.; Chiang, D.; et al. Human airway epithelial extracellular vesicle miRNA signature is altered upon asthma development. Allergy 2019. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Ling, M.; Xue, J.; Dai, X.; Sun, Q.; Chen, C.; Liu, Y.; Zhou, L.; Liu, J.; Luo, F.; et al. Exosomal microRNA-21 derived from bronchial epithelial cells is involved in aberrant epithelium-fibroblast cross-talk in COPD induced by cigarette smoking. Theranostics 2018, 8, 5419–5433. [Google Scholar] [CrossRef] [PubMed]
- Stassen, F.R.M.; van Eijck, P.H.; Savelkoul, P.H.M.; Wouters, E.F.M.; Rohde, G.G.U.; Briede, J.J.; Reynaert, N.L.; de Kok, T.M.; Benedikter, B.J. Cell Type- and Exposure-Specific Modulation of CD63/CD81-Positive and Tissue Factor-Positive Extracellular Vesicle Release in response to Respiratory Toxicants. Oxid. Med. Cell Longev. 2019, 2019, 5204218. [Google Scholar] [CrossRef]
- Benedikter, B.J.; Bouwman, F.G.; Heinzmann, A.C.A.; Vajen, T.; Mariman, E.C.; Wouters, E.F.M.; Savelkoul, P.H.M.; Koenen, R.R.; Rohde, G.G.U.; van Oerle, R.; et al. Proteomic analysis reveals procoagulant properties of cigarette smoke-induced extracellular vesicles. J. Extracell. Vesicles 2019, 8, 1585163. [Google Scholar] [CrossRef]
- Castro, S.M.; Kolli, D.; Guerrero-Plata, A.; Garofalo, R.P.; Casola, A. Cigarette smoke condensate enhances respiratory syncytial virus-induced chemokine release by modulating NF-kappa B and interferon regulatory factor activation. Toxicol. Sci. 2008, 106, 509–518. [Google Scholar] [CrossRef]
- Kuksa, P.P.; Amlie-Wolf, A.; Katanic, Z.; Valladares, O.; Wang, L.S.; Leung, Y.Y. DASHR 2.0: Integrated database of human small non-coding RNA genes and mature products. Bioinformatics 2019, 35, 1033–1039. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Shivdasani, R.A. MicroRNAs: Regulators of gene expression and cell differentiation. Blood 2006, 108, 3646–3653. [Google Scholar] [CrossRef] [PubMed]
- Catalanotto, C.; Cogoni, C.; Zardo, G. MicroRNA in Control of Gene Expression: An Overview of Nuclear Functions. Int. J. Mol. Sci. 2016, 17, 1712. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Rios, P.; Simonelig, M. piRNAs and PIWI proteins: Regulators of gene expression in development and stem cells. Development 2018, 145. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef] [PubMed]
- Available online: www.targetscan.org (accessed on 1 December 2019).
- Lu, X.; Kazmierczak, K.; Jiang, X.; Jones, M.; Watt, J.; Helfman, D.M.; Moore, J.R.; Szczesna-Cordary, D.; Lossos, I.S. Germinal center-specific protein human germinal center associated lymphoma directly interacts with both myosin and actin and increases the binding of myosin to actin. FEBS J. 2011, 278, 1922–1931. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Fu, C.; Turck, C.W.; Kurosaki, T.; Chan, A.C. BLNK: A central linker protein in B cell activation. Immunity 1998, 9, 93–103. [Google Scholar] [CrossRef]
- Moscat, J.; Diaz-Meco, M.T.; Rennert, P. NF-kappaB activation by protein kinase C isoforms and B-cell function. EMBO Rep. 2003, 4, 31–36. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef]
- Mi, H.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2019, 47, D419–D426. [Google Scholar] [CrossRef]
- Luppi, F.; Aarbiou, J.; van Wetering, S.; Rahman, I.; de Boer, W.I.; Rabe, K.F.; Hiemstra, P.S. Effects of cigarette smoke condensate on proliferation and wound closure of bronchial epithelial cells in vitro: Role of glutathione. Respir. Res. 2005, 6, 140. [Google Scholar] [CrossRef]
- Benedikter, B.J.; Volgers, C.; van Eijck, P.H.; Wouters, E.F.M.; Savelkoul, P.H.M.; Reynaert, N.L.; Haenen, G.; Rohde, G.G.U.; Weseler, A.R.; Stassen, F.R.M. Cigarette smoke extract induced exosome release is mediated by depletion of exofacial thiols and can be inhibited by thiol-antioxidants. Free Radic. Biol. Med. 2017, 108, 334–344. [Google Scholar] [CrossRef] [PubMed]
- Davis, C.N.; Phillips, H.; Tomes, J.J.; Swain, M.T.; Wilkinson, T.J.; Brophy, P.M.; Morphew, R.M. The importance of extracellular vesicle purification for downstream analysis: A comparison of differential centrifugation and size exclusion chromatography for helminth pathogens. PLoS Negl. Trop. Dis. 2019, 13, e0007191. [Google Scholar] [CrossRef] [PubMed]
- Haque, S.; Sinha, N.; Ranjit, S.; Midde, N.M.; Kashanchi, F.; Kumar, S. Monocyte-derived exosomes upon exposure to cigarette smoke condensate alter their characteristics and show protective effect against cytotoxicity and HIV-1 replication. Sci. Rep. 2017, 7, 16120. [Google Scholar] [CrossRef] [PubMed]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Izzotti, A.; Calin, G.A.; Arrigo, P.; Steele, V.E.; Croce, C.M.; De Flora, S. Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke. FASEB J. 2009, 23, 806–812. [Google Scholar] [CrossRef]
- Fujita, Y.; Araya, J.; Ito, S.; Kobayashi, K.; Kosaka, N.; Yoshioka, Y.; Kadota, T.; Hara, H.; Kuwano, K.; Ochiya, T. Suppression of autophagy by extracellular vesicles promotes myofibroblast differentiation in COPD pathogenesis. J. Extracell. Vesicles 2015, 4, 28388. [Google Scholar] [CrossRef]
- He, S.; Chen, D.; Hu, M.; Zhang, L.; Liu, C.; Traini, D.; Grau, G.E.; Zeng, Z.; Lu, J.; Zhou, G.; et al. Bronchial epithelial cell extracellular vesicles ameliorate epithelial-mesenchymal transition in COPD pathogenesis by alleviating M2 macrophage polarization. Nanomedicine 2019, 18, 259–271. [Google Scholar] [CrossRef]
- Baskoro, H.; Sato, T.; Karasutani, K.; Suzuki, Y.; Mitsui, A.; Arano, N.; Nurwidya, F.; Kato, M.; Takahashi, F.; Kodama, Y.; et al. Regional heterogeneity in response of airway epithelial cells to cigarette smoke. BMC Pulm. Med. 2018, 18, 148. [Google Scholar] [CrossRef]
- Morissette, M.C.; Shen, P.; Thayaparan, D.; Stampfli, M.R. Disruption of pulmonary lipid homeostasis drives cigarette smoke-induced lung inflammation in mice. Eur. Respir. J. 2015, 46, 1451–1460. [Google Scholar] [CrossRef]
- Haslam, R.J.; Koide, H.B.; Hemmings, B.A. Pleckstrin domain homology. Nature 1993, 363, 309–310. [Google Scholar] [CrossRef]
- Kim, C.; Lee, J.M.; Park, S.W.; Kim, K.S.; Lee, M.W.; Paik, S.; Jang, A.S.; Kim, D.J.; Uh, S.; Kim, Y.; et al. Attenuation of Cigarette Smoke-Induced Emphysema in Mice by Apolipoprotein A-1 Overexpression. Am. J. Respir. Cell Mol. Biol. 2016, 54, 91–102. [Google Scholar] [CrossRef] [PubMed]
| RNA Type | % Control EVs | % CSC EVs | p Value |
|---|---|---|---|
| protein coding | 2.9 ± 1.3 | 3 ± 1.4 | 0.4 |
| processed transcript | 1.5 ± 0.7 | 1.3 ± 0.7 | 0.4 |
| miRNA | 5.0 ± 0.7 | 7.5 ± 0.8 | 0.04 |
| tRNA | 9.1 ± 4.0 | 12.5 ± 6.2 | 0.3 |
| piRNA | 0.4 ± 0.01 | 0.38 ± 0.02 | 0.3 |
| snoRNA | 6.1± 3.0 | 5.3 ± 2.9 | 0.4 |
| rRNA | 8.1 ± 0.3 | 8.4 ± 0.9 | 0.4 |
| miscRNA | 1.4 ± 0.6 | 2.3 ± 1.0 | 0.2 |
| snRNA | 0.8 ± 0.4 | 0.9 ± 0.4 | 0.4 |
| unannotated | 63.6 ± 9.3 | 57 ± 11.3 | 0.3 |
| miRNAs | Log2 Fold Change | p Value | Read Count (Avg) | |
|---|---|---|---|---|
| CSC EVs | Control EVs | |||
| hsa-miR-3913-5p | 10.28 | 0.0003 | 356.00 | 1.62 |
| hsa-miR-574-5p | 10.25 | 0.0006 | 183.57 | 0 |
| hsa-miR-656-5p | 9.76 | 0.0014 | 126.40 | 0 |
| hsa-miR-3180-5p | 9.21 | 0.0022 | 39.98 | 0 |
| hsa-miR-500a-5p | 8.91 | 0.0033 | 84.5 | 0 |
| hsa-miR-618 | −9.25 | 0.00241 | 0 | 76.053 |
| hsa-miR-222-5p | −8.78 | 0.00472 | 0 | 55.44 |
| hsa-miR-130b-5p | −8.20 | 0.005 | 0 | 38.35 |
| piRNAs | Log2 Fold Change | p Value | Read Count (Avg) | |
|---|---|---|---|---|
| CSC EVs | Control EVs | |||
| piR-36705 | −10.19 | 0.002 | 0 | 191.02 |
| piR-37183 | −10.19 | 0.002 | 0 | 191.02 |
| piR-59260 | −10.19 | 0.002 | 0 | 191.02 |
| piR-36924 | −10.24 | 0.002 | 0.32 | 358.4 |
| piR-52900 | −10.28 | 0.004 | 0 | 211.63 |
| piR-31985 | 10.52 | 0.004 | 498.48 | 0 |
| piR-50603 | 9.78 | 0.001 | 276.51 | 0 |
| Target Gene | Cumulative Weighted Context ++ Score | Target Gene | Cumulative Weighted Context ++ Score |
|---|---|---|---|
| PLEKHS1 | −1.39 | AC005477.1 | −0.58 |
| GCSAM | −1 | FCGR2A | −0.58 |
| BLNK | −0.75 | ETNK1 | −0.55 |
| ACMSD | −0.74 | MYF5 | −0.55 |
| AP000708.1 | −0.72 | NME4 | −0.54 |
| KCNAB2 | −0.7 | APOA4 | −0.54 |
| IQCK | −0.7 | KRTAP4-6 | −0.53 |
| RHAG | −0.65 | AL049747.1 | −0.53 |
| S100A10 | −0.65 | ZNF740 | −0.53 |
| NHLH1 | −0.61 | SEPT3 | −0.53 |
| OR51F2 | −0.61 | ITGAE | −0.53 |
| VSNL1 | −0.6 | PRKCDBP | −0.52 |
| ERGIC1 | −0.6 | TPMT | −0.52 |
| PYY | −0.59 | C21ORF33 | −0.5 |
| YWHAZ | −0.59 | RP11-321F6.1 | −0.5 |
| PSMC4 | −0.59 |
| Term | Genes | Enrichment |
|---|---|---|
| Biological process | ||
| lipid transport | NME4, APOA4 | 19.21 |
| regulation of mRNA stability | YWHAZ, PSMC4 | 14.17 |
| extracellular matrix organization | ITGAE, MYF5 | 7.44 |
| RNA polymerase II promoter | MYF5, NHLH1 | 2.84 |
| Cellular component | ||
| cytosol | APOA4, YWHAZ, KCNAB2, PSMC4, VSNL1, ACMSD, ETNK1, TPMT, BLNK | 2.06 |
| membrane | KCNAB2, PSMC4, VSNL1, ETNK1, RHAG, ERGIC1 | 2.07 |
| blood microparticle | APOA4, YWHAZ | 9.99 |
| plasma membrane | KCNAB2, ITGAE, OR51F2, ETNK1, GCSAM, FCGR2A, RHAG, BLNK | 1.47 |
| extracellular exosome | APOA4, YWHAZ, ACMSD, S100A10, FCGR2A, TPMT | 1.62 |
| cytoskeleton | SEPT3, KCNAB2 | 4.09 |
| cell junction | SEPT3, KCNAB2 | 3.3 |
| integral component of membrane | KCNAB2, ACMSD, OR51F2, ETNK1, S100A10, FCGR2A, RHAG, ERGIC1 | 1.17 |
| Molecular function | ||
| lipid binding | NME4, APOA4, S100A10 | 13.97 |
| protein binding | YWHAZ, SEPT3, MYF5, ACMSD, GCSAM, S100A10, PRKCDBP, ERGIC1, APOA4, NME4, PSMC4, VSNL1, ETNK1, FCGR2A, PYY, PLEKHS1, BLNK | 1.36 |
| transcriptional activation only | MYF5, NHLH1 | 14.80 |
| hydrolase activity | PSMC4, ACMSD | 7.36 |
| protein kinase binding | YWHAZ, GCSAM | 3.74 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Corsello, T.; Kudlicki, A.S.; Garofalo, R.P.; Casola, A. Cigarette Smoke Condensate Exposure Changes RNA Content of Extracellular Vesicles Released from Small Airway Epithelial Cells. Cells 2019, 8, 1652. https://doi.org/10.3390/cells8121652
Corsello T, Kudlicki AS, Garofalo RP, Casola A. Cigarette Smoke Condensate Exposure Changes RNA Content of Extracellular Vesicles Released from Small Airway Epithelial Cells. Cells. 2019; 8(12):1652. https://doi.org/10.3390/cells8121652
Chicago/Turabian StyleCorsello, Tiziana, Andrzej S. Kudlicki, Roberto P. Garofalo, and Antonella Casola. 2019. "Cigarette Smoke Condensate Exposure Changes RNA Content of Extracellular Vesicles Released from Small Airway Epithelial Cells" Cells 8, no. 12: 1652. https://doi.org/10.3390/cells8121652
APA StyleCorsello, T., Kudlicki, A. S., Garofalo, R. P., & Casola, A. (2019). Cigarette Smoke Condensate Exposure Changes RNA Content of Extracellular Vesicles Released from Small Airway Epithelial Cells. Cells, 8(12), 1652. https://doi.org/10.3390/cells8121652