Intratumor Microbiome in Neuroendocrine Neoplasms: A New Partner of Tumor Microenvironment? A Pilot Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
2.2. Patients
2.3. Fluorescent In Situ Hybridization (FISH)
2.4. Confocal Microscopy
2.5. Statistical Analysis
3. Results
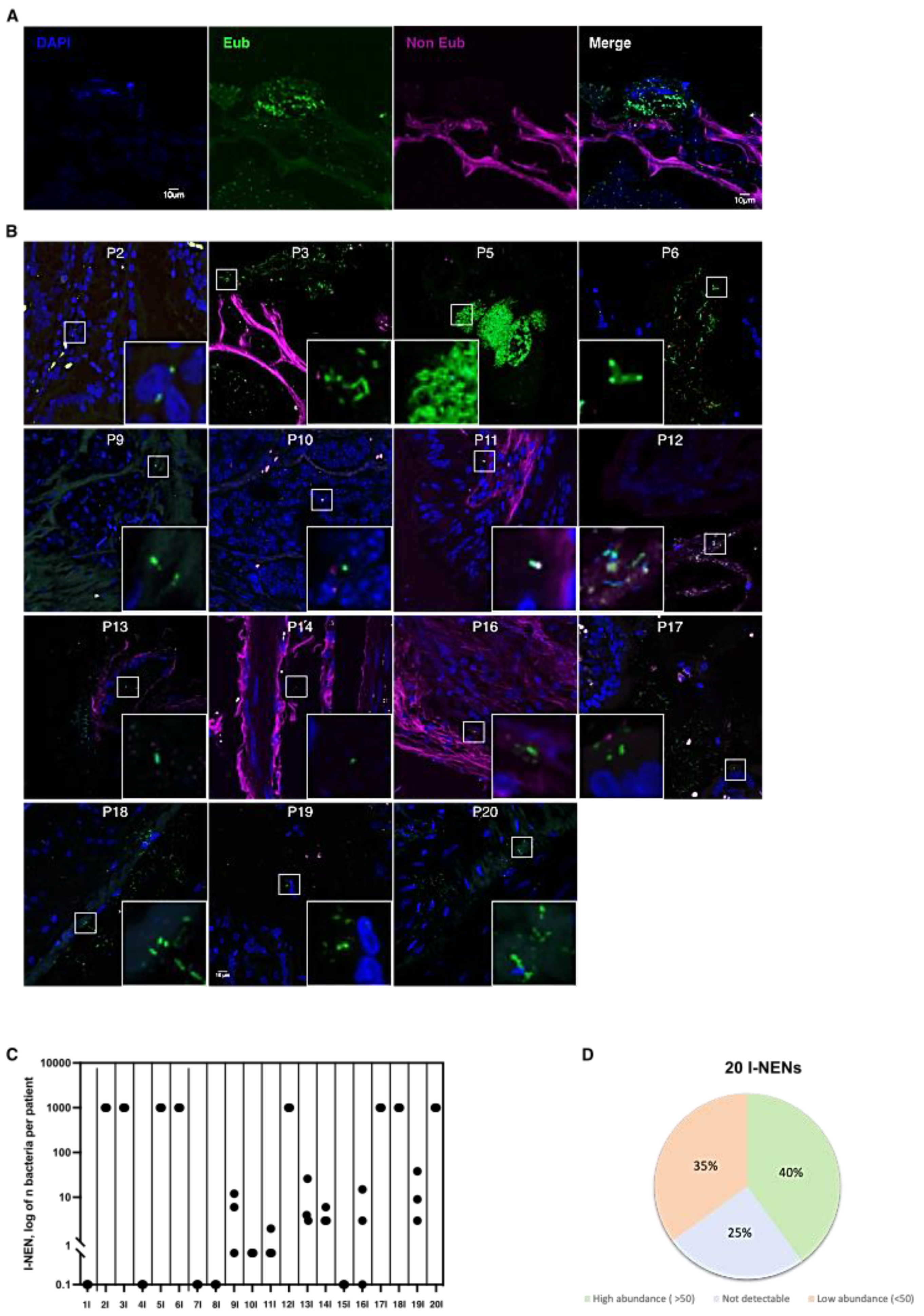
3.1. Bacteria Colonize I-NEN and pan-NEN Tissues
3.2. Bacteria Infiltrate pan-NEN Tissues
3.3. Clinical Variables
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kim, Y.I.; Yoo, C.; Oh, S.J.; Lee, S.J.; Kang, J.; Hwang, H.S.; Hong, S.M.; Ryoo, B.Y.; Ryu, J.S. Tumour-to-liver ratio determined by [68Ga]Ga-DOTA-TOC PET/CT as a prognostic factor of lanreotide efficacy for patients with well-differentiated gastroenteropancreatic-neuroendocrine tumours. EJNMMI Res. 2020, 10, 63. [Google Scholar] [CrossRef]
- Vitale, G.; Carra, S.; Ferraù, F.; Guadagno, E.; Faggiano, A.; Colao, A. Gastroenteropancreatic neuroendocrine neoplasms and inflammation: A complex cross-talk with relevant clinical implications. Crit. Rev. Oncol. Hematol. 2020, 146, 102840. [Google Scholar] [CrossRef] [PubMed]
- Hauso, O.; Gustafsson, B.I.; Kidd, M.; Waldum, H.L.; Drozdov, I.; Chan, A.K.; Modlin, I.M. Neuroendocrine tumor epidemiology: Contrasting Norway and North America. Cancer 2008, 113, 2655–2664. [Google Scholar] [CrossRef]
- Rindi, G.; Arnold, R.; Bosman, F.; Capella, C.; Kilmstra, D.; Kloppel, G. Nomenclature and classification of neuroendocrine neoplasms of the digestive system. In WHO Classification of Tumours of the Digestive System; Bosman, F.T., Carneiro, F., Hruban, R.H., Theise, N.D., Eds.; IARC Press: Lyon, France, 2010; pp. 13–14. [Google Scholar]
- Nagtegaal, I.D.; Odze, R.D.; Klimstra, D.; Paradis, V.; Rugge, M.; Schirmacher, P.; Washington, K.M.; Carneiro, F.; Cree, I.A. The 2019 WHO classification of tumours of the digestive system. Histopathology 2020, 76, 182–188. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pusceddu, S.; Vernieri, C.; Di Maio, M.; Marconcini, R.; Spada, F.; Massironi, S.; Ibrahim, T.; Brizzi, M.P.; Campana, D.; Faggiano, A.; et al. Metformin Use Is Associated with Longer Progression-Free Survival of Patients with Diabetes and Pancreatic Neuroendocrine Tumors Receiving Everolimus and/or Somatostatin Analogues. Gastroenterology 2018, 155, 479–489. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexandraki, K.I.; Karapanagioti, A.; Karoumpalis, I.; Boutzios, G.; Kaltsas, G.A. Advances and Current Concepts in the Medical Management of Gastroenteropancreatic Neuroendocrine Neoplasms. Biomed. Res. Int. 2017, 9856140. [Google Scholar] [CrossRef] [Green Version]
- Berardi, R.; Morgese, F.; Torniai, M.; Savini, A.; Partelli, S.; Rinaldi, S.; Caramanti, M.; Ferrini, C.; Falconi, M.; Cascinu, S. Medical treatment for gastro-entero-pancreatic neuroendocrine tumours. World J. Gastrointest. Oncol. 2016, 8, 389–401. [Google Scholar] [CrossRef]
- Ohtani, N.; Yoshimoto, S.; Hara, E. Obesity and cancer: A gut microbial connection. Cancer Res. 2014, 74, 1885–1889. [Google Scholar] [CrossRef] [Green Version]
- Tsuei, J.; Chau, T.; Mills, D.; Wan, Y.J. Bile acid dysregulation, gut dysbiosis, and gastrointestinal cancer. Exp. Biol. Med. 2014, 239, 1489–1504. [Google Scholar] [CrossRef] [Green Version]
- Amoroso, C.; Perillo, F.; Strati, F.; Fantini, M.C.; Caprioli, F.; Facciotti, F. The Role of Gut Microbiota Biomodulators on Mucosal Immunity and Intestinal Inflammation. Cells 2020, 9, 1234. [Google Scholar] [CrossRef]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zmora, N.; Suez, J.; Elinav, E. You are what you eat: Diet, health and the gut microbiota. Nat. Rev. Gastroenterol. Hepatol. 2019, 16, 35–56. [Google Scholar] [CrossRef] [PubMed]
- Cigrovski Berkovic, M.; Cacev, T.; Catela Ivkovic, T.; Zjacic-Rotkvic, V.; Kapitanovic, S. New insights into the role of chronic inflammation and cytokines in the etiopathogenesis of gastroenteropancreatic neuroendocrine tumors. Neuroendocrinology 2014, 99, 75–84. [Google Scholar] [CrossRef]
- Massironi, S.; Zilli, A.; Elvevi, A.; Invernizzi, P. The changing face of chronic autoimmune atrophic gastritis: An updated comprehensive perspective. Autoimmun. Rev. 2019, 18, 215–222. [Google Scholar] [CrossRef]
- Thomas, R.M.; Jobin, C. Microbiota in pancreatic health and disease: The next frontier in microbiome research. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 53–64. [Google Scholar] [CrossRef]
- Thomas, R.M.; Gharaibeh, R.Z.; Gauthier, J.; Beveridge, M.; Pope, J.L.; Guijarro, M.V.; Yu, Q.; He, Z.; Ohland, C.; Newsome, R.; et al. Intestinal microbiota enhances pancreatic carcinogenesis in preclinical models. Carcinogenesis 2018, 39, 1068–1078. [Google Scholar] [CrossRef]
- Pushalkar, S.; Hundeyin, M.; Daley, D.; Zambirinis, C.P.; Kurz, E.; Mishra, A.; Mohan, N.; Aykut, B.; Usyk, M.; Torres, L.E.; et al. The Pancreatic Cancer Microbiome Promotes Oncogenesis by Induction of Innate and Adaptive Immune Suppression. Cancer Discov. 2018, 8, 403–416. [Google Scholar] [CrossRef] [Green Version]
- Abdul Rahman, R.; Lamarca, A.; Hubner, R.A.; Valle, J.W.; McNamara, M.G. The Microbiome as a Potential Target for Therapeutic Manipulation in Pancreatic Cancer. Cancers 2021, 13, 3779. [Google Scholar] [CrossRef]
- Döerffel, Y.; Pavel, M.; Loening-Baucke, V.; Swidsinski, A. Common biostructure of the fecal flora in celiac disease, Crohn’s disease, and carcinoid tumors. Inflamm. Bowel Dis. 2008, 14, 1613–1614. [Google Scholar] [CrossRef]
- Ahuja, M.; Schwartz, D.M.; Tandon, M.; Son, A.; Zeng, M.; Swaim, W.; Eckhaus, M.; Hoffman, V.; Cui, Y.; Xiao, B.; et al. Orai1-Mediated Antimicrobial Secretion from Pancreatic Acini Shapes the Gut Microbiome and Regulates Gut Innate Immunity. Cell Metab. 2017, 25, 635–646. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kojima, T.; Yamaguchi, H.; Ito, T.; Kyuno, D.; Kono, T.; Konno, T.; Sawada, N. Tight junctions in human pancreatic duct epithelial cells. Tissue Barriers 2013, 1, e24894. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, S.; Fuhler, G.M.; Bn, N.; Jose, T.; Bruno, M.J.; Peppelenbosch, M.P.; Konstantinov, S.R. Pancreatic cyst fluid harbors a unique microbiome. Microbiome 2017, 5, 147. [Google Scholar] [CrossRef] [PubMed]
- Isenmann, R.; Beger, H.G. Bacterial infection of pancreatic necrosis: Role of bacterial translocation, impact of antibiotic treatment. Pancreatology 2001, 1, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Vitale, G.; Dicitore, A.; Barrea, L.; Sbardella, E.; Razzore, P.; Campione, S.; Faggiano, A.; Colao, A.; Albertelli, M.; Altieri, B.; et al. From microbiota toward gastro-enteropancreatic neuroendocrine neoplasms: Are we on the highway to hell? Rev. Endocr. Metab. Disord. 2020, 22, 511–525. [Google Scholar] [CrossRef]
- Yu, T.; Guo, F.; Yu, Y.; Sun, T.; Ma, D.; Han, J.; Qian, Y.; Kryczek, I.; Sun, D.; Nagarsheth, N.; et al. Fusobacterium nucleatum Promotes Chemoresistance to Colorectal Cancer by Modulating Autophagy. Cell 2017, 170, 548–563.e16. [Google Scholar] [CrossRef] [Green Version]
- Shi, Y.; Zheng, W.; Yang, K.; Harris, K.G.; Ni, K.; Xue, L.; Lin, W.; Chang, E.B.; Weichselbaum, R.R.; Fu, Y.X. Intratumoral accumulation of gut microbiota facilitates CD47-based immunotherapy via STING signaling. J. Exp. Med. 2020, 217, e20192282. [Google Scholar] [CrossRef]
- Geller, L.T.; Barzily-Rokni, M.; Danino, T.; Jonas, O.H.; Shental, N.; Nejman, D.; Gavert, N.; Zwang, Y.; Cooper, Z.A.; Shee, K.; et al. Potential role of intratumor bacteria in mediating tumor resistance to the chemotherapeutic drug gemcitabine. Science 2017, 357, 1156–1160. [Google Scholar] [CrossRef] [Green Version]
- Riquelme, E.; Zhang, Y.; Zhang, L.; Montiel, M.; Zoltan, M.; Dong, W.; Quesada, P.; Sahin, I.; Chandra, V.; San Lucas, A.; et al. Tumor Microbiome Diversity and Composition Influence Pancreatic Cancer Outcomes. Cell 2019, 178, 795–806. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 2018, 68, 7–30. [Google Scholar] [CrossRef]


| Characteristics | Pan-NEN (N=20) | I-NEN (N20) |
|---|---|---|
| Age in years, median (range) | 56 (34–76) | 62 (33–77) |
| Sex, M/F | 11/9 | 10/10 |
| Dimension mm, median (range) | 2.75 (0.8–6) | 1.6 (0.8–5) |
| Ki-67%, median (range) | 6.52 (0.1–45) | 3.4 (0.1–28) |
| Grading, N (%): | ||
| G1 | 13 (65) | 15 (75) |
| G2 | 5 (25) | 4 (20) |
| G3 | 2 (10) | 1 (5) |
| Treatment, N (%) | ||
| SSAs therapy | 9 (45) | 11 (55) |
| Chemotherapy | 3 (15) | 3 (15) |
| Target therapy | 3 (15) | 2 (10) |
| Radio ligand theraphy | 2 (10) | 2 (10) |
| Probes | Sequences | Vendor |
|---|---|---|
| EUB338-1 | 5′(A488)-GCTGCCTCCCGTAGGA | SIGMA |
| EUB338-2 | 5′(A488)-GCAGCCACCCGTAGGTG | SIGMA |
| EUB338-3 | 5′(A488)-GCTGCCACCCGTAGGTG | SIGMA |
| Non338 | 5′(Cyanine5)-CGACGGAGGGCATCCTCA | SIGMA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Massironi, S.; Facciotti, F.; Cavalcoli, F.; Amoroso, C.; Rausa, E.; Centonze, G.; Cribiù, F.M.; Invernizzi, P.; Milione, M. Intratumor Microbiome in Neuroendocrine Neoplasms: A New Partner of Tumor Microenvironment? A Pilot Study. Cells 2022, 11, 692. https://doi.org/10.3390/cells11040692
Massironi S, Facciotti F, Cavalcoli F, Amoroso C, Rausa E, Centonze G, Cribiù FM, Invernizzi P, Milione M. Intratumor Microbiome in Neuroendocrine Neoplasms: A New Partner of Tumor Microenvironment? A Pilot Study. Cells. 2022; 11(4):692. https://doi.org/10.3390/cells11040692
Chicago/Turabian StyleMassironi, Sara, Federica Facciotti, Federica Cavalcoli, Chiara Amoroso, Emanuele Rausa, Giovanni Centonze, Fulvia Milena Cribiù, Pietro Invernizzi, and Massimo Milione. 2022. "Intratumor Microbiome in Neuroendocrine Neoplasms: A New Partner of Tumor Microenvironment? A Pilot Study" Cells 11, no. 4: 692. https://doi.org/10.3390/cells11040692
APA StyleMassironi, S., Facciotti, F., Cavalcoli, F., Amoroso, C., Rausa, E., Centonze, G., Cribiù, F. M., Invernizzi, P., & Milione, M. (2022). Intratumor Microbiome in Neuroendocrine Neoplasms: A New Partner of Tumor Microenvironment? A Pilot Study. Cells, 11(4), 692. https://doi.org/10.3390/cells11040692








