Correction: Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470
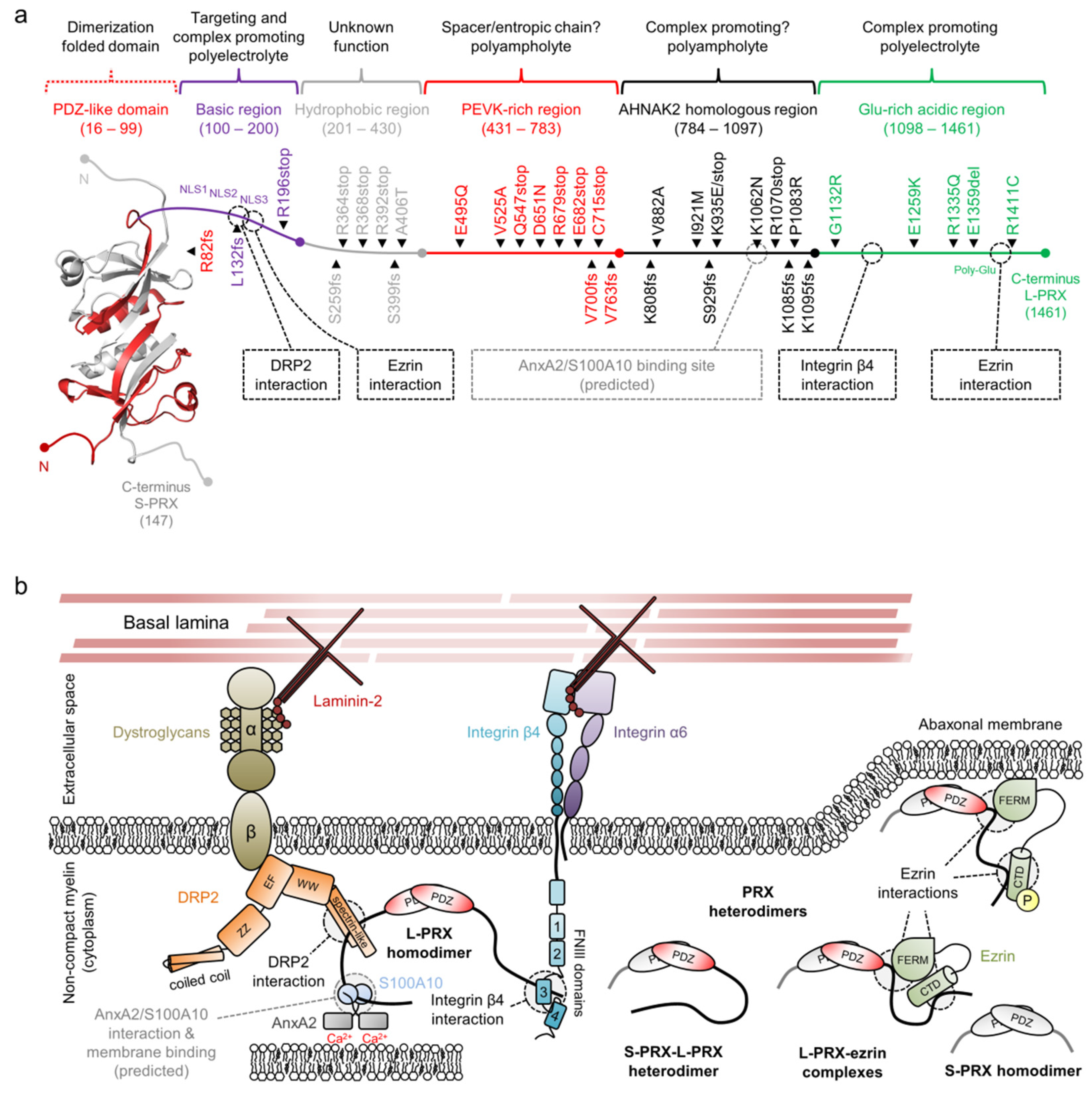
| Mutation 1 | Neuropathy | (Potential) Molecular Impact | Reference(s) |
|---|---|---|---|
| R82fs | DSS | Tail loss; loss of interactions | [258] |
| L132fs | CMT4F | Tail loss; loss of interactions | [259] |
| R196stop | CMT4F | [260] | |
| S259fs | CMT4F | Loss of hydrophobic, PEVK-rich, AHNAK2 homology and acidic regions; loss of interactions | [261] |
| R364stop | CMT4F | Loss of PEVK-rich, AHNAK2 homology and acidic regions; loss of interactions | [262] |
| R368stop | DSS | Loss of PEVK-rich, AHNAK2 homology and acidic regions; loss of interactions | [263] |
| R392stop | DSS | Loss of PEVK-rich, AHNAK2 homology and acidic regions; loss of interactions | [264] |
| S399fs | CMT4F | Loss of PEVK-rich, AHNAK2 homology and acidic regions; loss of interactions | [265] |
| A406T | DSS | [263] | |
| E495Q | DSS | [263] | |
| V525A | CMT4F | [260,266] | |
| Q547stop | CMT4F | Loss of PEVK-rich (partial), AHNAK2 homology and acidic regions; loss of interactions | [261] |
| D651N | CMT4F | [267] | |
| R679stop | DSS | Loss of PEVK-rich (partial), AHNAK2 homology and acidic regions; loss of interactions | [264] |
| E682stop | CMT4F | Loss of PEVK-rich (partial), AHNAK2 homology and acidic regions; loss of interactions | [261] |
| A700fs | CMT4F | [268] | |
| C715stop | DSS | Loss of PEVK-rich (partial), AHNAK2 homology and acidic regions; loss of interactions | [258] |
| V763fs | DSS | Loss of PEVK-rich (partial), AHNAK2 homology and acidic regions; loss of interactions | [263] |
| K808fs | CMT4F | Loss of AHNAK2 homology and acidic regions; loss of interactions | [261] |
| V882A | DSS | [263,269] | |
| I921M | DSS | [263] | |
| S929fs | DSS | Loss of AHNAK2 homology and acidic regions; loss of interactions | [263] |
| K935E | DSS | [263] | |
| K935stop | DSS | Loss of acidic domain; loss of integrin interaction | [263] |
| K1062N | CMT4F | (Loss of predicted AnxA2/S100A10 interaction?) | [257] |
| R1070stop | CMT4F | Loss of acidic domain; loss of integrin interaction | [208,259,267,270–272] |
| P1083R | DSS | [265] | |
| E1085fs | CMT4F | Loss of acidic domain; loss of integrin interaction | [273] |
| K1095fs | CMT4F | Loss of acidic domain; loss of integrin interaction | [274] |
| G1132R | DSS | [263] | |
| E1259K | DSS | [263] | |
| R1335Q 2 | CMT | [266] | |
| E1359del | DSS | [263] | |
| R1411C | DSS | [263] |
References
- Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, Y.; Shi, Y. Retraction: Spectrin-like domain 2 of DRP2 serves as a novel binding region for the NLS2 and 3 sub-domains of L-periaxin. RSC Adv. 2021, 11, 15160. [Google Scholar] [CrossRef]
- Yang, Y.; Liang, M.; Shi, Y. Retraction: Self-association of L-periaxin occurs via its acidic domain and NLS2/NLS3, and affects its trafficking in RSC96 cells. RSC Adv. 2021, 11, 15203. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Raasakka, A.; Kursula, P. Correction: Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470. Cells 2022, 11, 662. https://doi.org/10.3390/cells11040662
Raasakka A, Kursula P. Correction: Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470. Cells. 2022; 11(4):662. https://doi.org/10.3390/cells11040662
Chicago/Turabian StyleRaasakka, Arne, and Petri Kursula. 2022. "Correction: Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470" Cells 11, no. 4: 662. https://doi.org/10.3390/cells11040662
APA StyleRaasakka, A., & Kursula, P. (2022). Correction: Raasakka, A.; Kursula, P. Flexible Players within the Sheaths: The Intrinsically Disordered Proteins of Myelin in Health and Disease. Cells 2020, 9, 470. Cells, 11(4), 662. https://doi.org/10.3390/cells11040662






