Innovative Platform for the Advanced Online Monitoring of Three-Dimensional Cells and Tissue Cultures
Abstract
:1. Introduction
2. Materials and Methods
2.1. Rapid Prototyping
2.2. Cell Culture
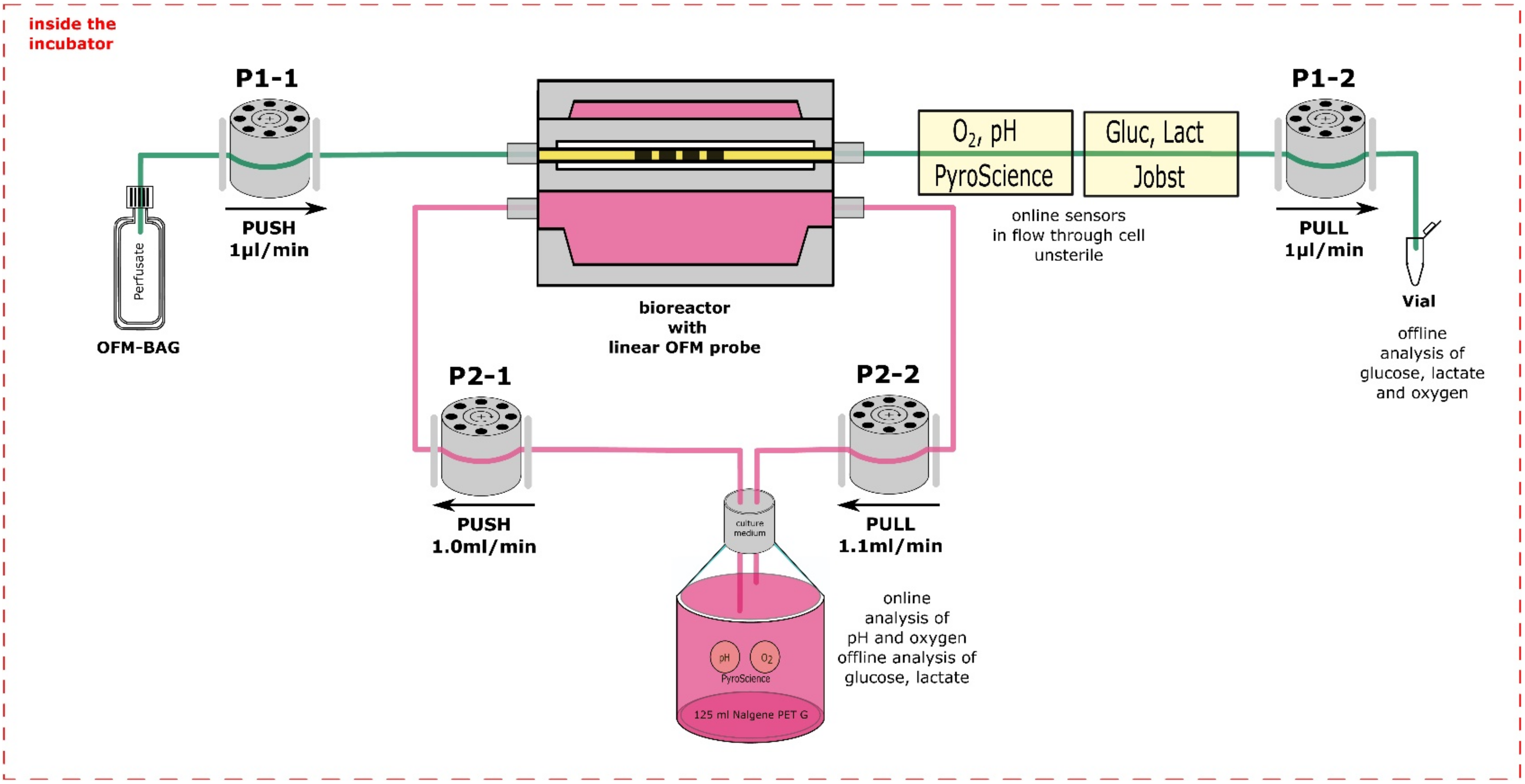
2.3. Perfusion Culture
2.4. Live/Dead Cell Staining
3. Results
3.1. Establishing a Modular Monitoring Platform
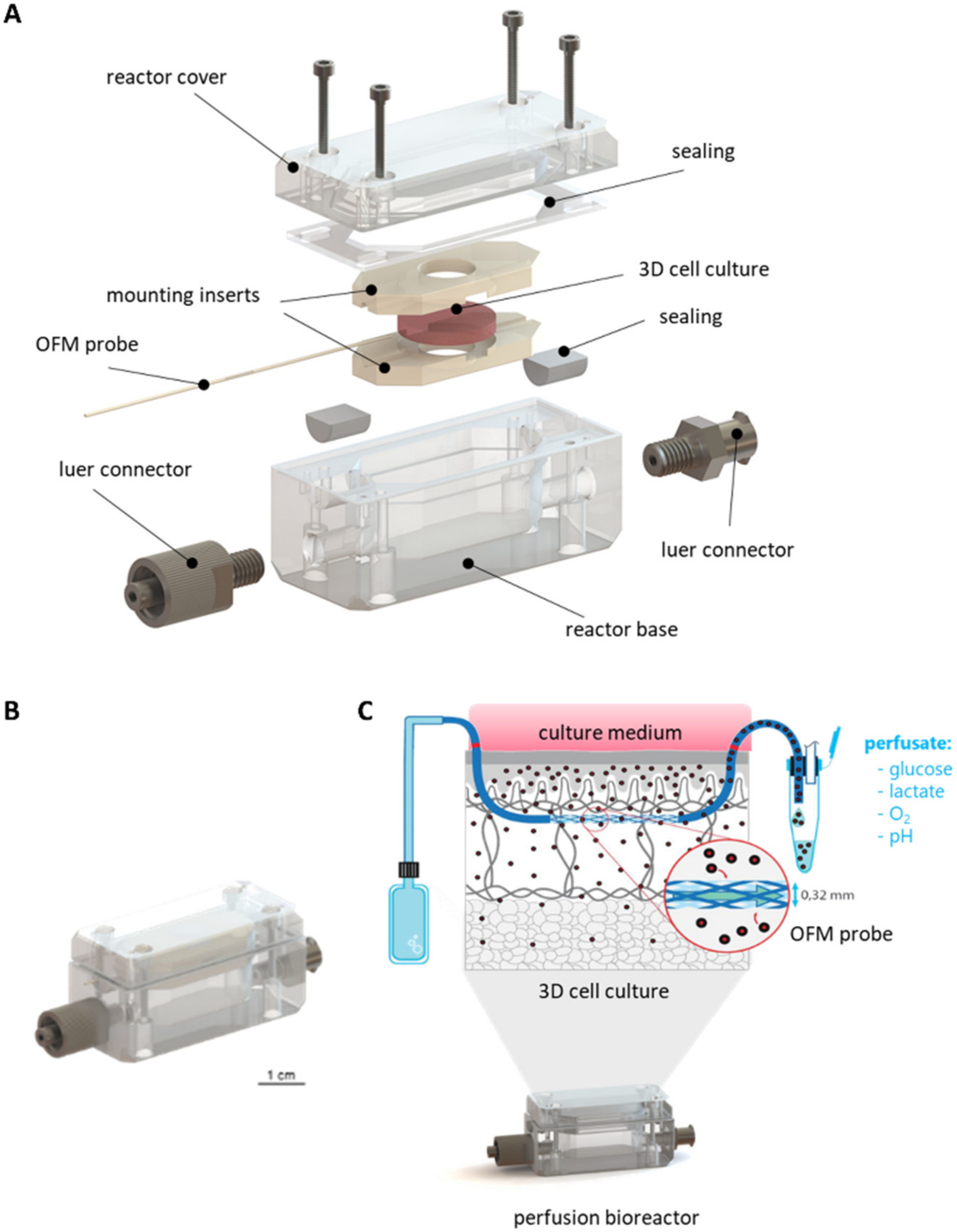
3.1.1. Concept and Design of the Perfusion Bioreactor
3.1.2. Mounting of 3D Cell and Tissue Cultures
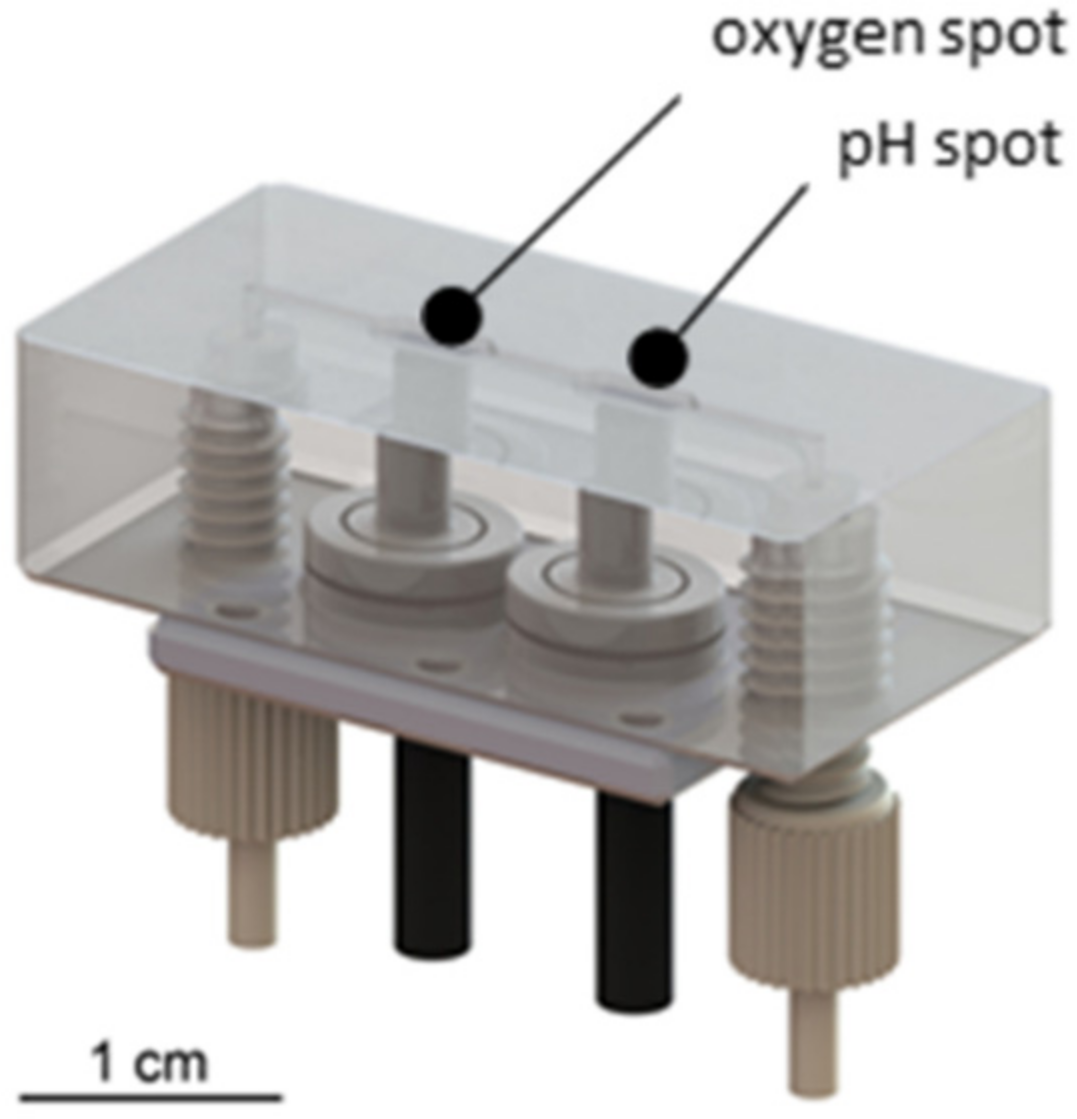
3.1.3. Development of a Sensor Platform for Online Monitoring
3.2. Assembly of the Monitoring Platform for 3D Cell Cultures
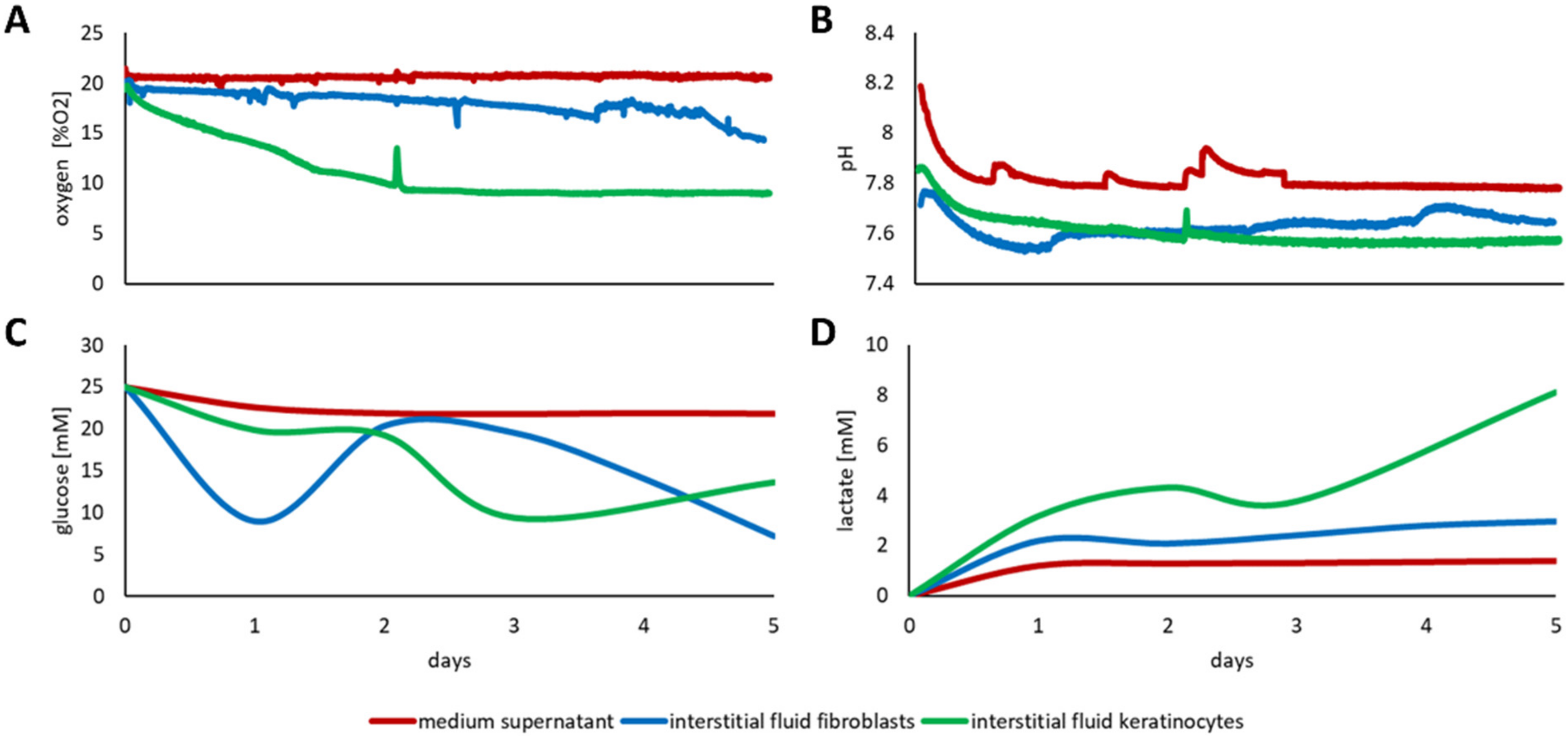
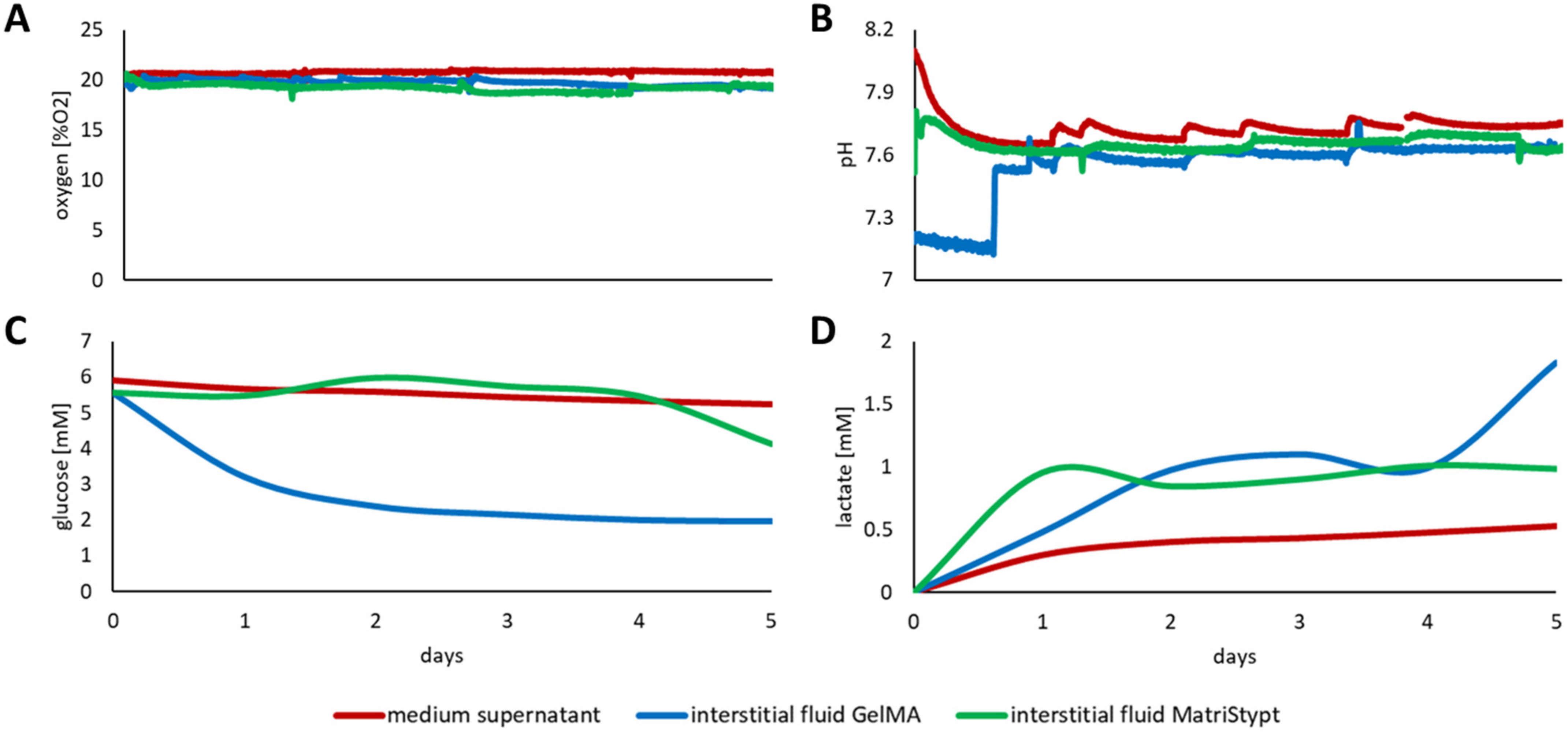
3.3. Discrepancy between Media Supernatant and Interstitial Fluid
3.3.1. Three-Dimensional Culture of Fibroblasts and Keratinocytes
3.3.2. Three-Dimensional Culture of Primary adMSCs
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Russell, W.M.S.; Burch, R.L. The Principles of Humane Experimental Technique; University College London: London, UK, 1959. [Google Scholar]
- Balls, M. It’s time to reconsider the principles of humane experimental technique. Altern. Lab. Anim. ATLA 2020, 48, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Ericsson, A.C.; Crim, M.J.; Franklin, C.L. A brief history of animal modeling. Mo. Med. 2013, 110, 201–205. [Google Scholar]
- Bédard, P.; Gauvin, S.; Ferland, K.; Caneparo, C.; Pellerin, È.; Chabaud, S.; Bolduc, S. Innovative human three-dimensional tissue-engineered models as an alternative to animal testing. Bioengineering 2020, 7, 115. [Google Scholar] [CrossRef] [PubMed]
- European Union. DIRECTIVE 2010/63/EU of the European parliament and of the council of 22 September 2010 on the protection of animals used for scientific purposes (text with EEA relevance). Off. J. Eur. Union 2010, 276, 33–79. [Google Scholar]
- European Union. Regulation (EC) No 1223/2009 of the European parliament and of the council of 30 November 2009 on cosmetic products. Off. J. Eur. Union 2009, 342, 59–209. [Google Scholar]
- Bell, C.C.; Dankers, A.C.A.; Lauschke, V.M.; Sison-Young, R.; Jenkins, R.; Rowe, C.; Goldring, C.E.; Park, K.; Regan, S.L.; Walker, T.; et al. Comparison of hepatic 2D sandwich cultures and 3d spheroids for long-term toxicity applications: A multicenter study. Toxicol. Sci. 2018, 162, 655–666. [Google Scholar] [CrossRef] [Green Version]
- Imamura, Y.; Mukohara, T.; Shimono, Y.; Funakoshi, Y.; Chayahara, N.; Toyoda, M.; Kiyota, N.; Takao, S.; Kono, S.; Nakatsura, T.; et al. Comparison of 2D- and 3D-culture models as drug-testing platforms in breast cancer. Oncol. Rep. 2015, 33, 1837–1843. [Google Scholar] [CrossRef] [Green Version]
- de Vries, R.B.M.; Leenaars, M.; Tra, J.; Huijbregtse, R.; Bongers, E.; Jansen, J.A.; Gordijn, B.; Ritskes-Hoitinga, M. The potential of tissue engineering for developing alternatives to animal experiments: A systematic review. J. Tissue Eng. Regen. Med. 2015, 9, 771–778. [Google Scholar] [CrossRef]
- Graf, B.W.; Boppart, S.A. Imaging and analysis of three-dimensional cell culture models. Methods Mol. Biol. 2010, 591, 211–227. [Google Scholar] [CrossRef] [Green Version]
- Folkman, J.; Hochberg, M. Self-regulation of growth in three dimensions. J. Exp. Med. 1973, 138, 745–753. [Google Scholar] [CrossRef] [PubMed]
- Edmondson, R.; Broglie, J.J.; Adcock, A.F.; Yang, L. Three-dimensional cell culture systems and their applications in drug discovery and cell-based biosensors. Assay Drug Dev. Technol. 2014, 12, 207–218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kreß, S.; Schaller-Ammann, R.; Feiel, J.; Priedl, J.; Kasper, C.; Egger, D. 3D printing of cell culture devices: Assessment and prevention of the cytotoxicity of photopolymers for stereolithography. Materials 2020, 13, 3011. [Google Scholar] [CrossRef] [PubMed]
- Formlabs Dental SG Safety Data Sheet. Available online: https://formlabs-media.formlabs.com/datasheets/Safety_Data_Sheet_EN-EU_-_Dental_SG.pdf (accessed on 15 November 2021).
- Formlabs Clear Resin Safety Data Sheet. Available online: https://formlabs-media.formlabs.com/datasheets/1801037-SDS-ENUS-0.pdf (accessed on 15 November 2021).
- Form Cure Time and Temperature Settings. Available online: https://support.formlabs.com/s/article/Form-Cure-Time-and-Temperature-Settings?language=en_US (accessed on 15 November 2021).
- Boukamp, P.; Petrussevska, R.T.; Breitkreutz, D.; Hornung, J.; Markham, A.; Fusenig, N.E. Normal keratinization in a spontaneously immortalized aneuploid human keratinocyte cell line. J. Cell Biol. 1988, 106, 761–771. [Google Scholar] [CrossRef] [Green Version]
- Meyer, M. Processing of collagen based biomaterials and the resulting materials properties. BioMed. Eng. Online 2019, 18, 24. [Google Scholar] [CrossRef] [Green Version]
- Pepelanova, I.; Kruppa, K.; Scheper, T.; Lavrentieva, A. Gelatin-methacryloyl (GelMA) hydrogels with defined degree of functionalization as a versatile toolkit for 3d cell culture and extrusion bioprinting. Bioengineering 2018, 5, 55. [Google Scholar] [CrossRef] [Green Version]
- Yue, K.; Trujillo-de Santiago, G.; Alvarez, M.M.; Tamayol, A.; Annabi, N.; Khademhosseini, A. Synthesis, properties, and biomedical applications of gelatin methacryloyl (GelMA) hydrogels. Biomaterials 2015, 73, 254–271. [Google Scholar] [CrossRef] [Green Version]
- Bodenlenz, M.; Aigner, B.; Dragatin, C.; Liebenberger, L.; Zahiragic, S.; Höfferer, C.; Birngruber, T.; Priedl, J.; Feichtner, F.; Schaupp, L.; et al. Clinical applicability of DOFM devices for dermal sampling. Ski. Res. Technol. 2013, 19, 474–483. [Google Scholar] [CrossRef]
- Bodenlenz, M.; Dragatin, C.; Hoefferer, C.; Birngruber, T.; Priedl, J.; Feichtner, F.; Schaller, R.; Aigner, B.; Korsatko, S.; Pieber, T.R.; et al. Certified DOFM sampling devices provide access to target tissue in pharmaceutical trials. Biomed. Eng./Biomed. Tech. 2013, 58, 000010151520134133. [Google Scholar] [CrossRef]
- Bodenlenz, M.; Tiffner, K.I.; Raml, R.; Augustin, T.; Dragatin, C.; Birngruber, T.; Schimek, D.; Schwagerle, G.; Pieber, T.R.; Raney, S.G.; et al. Open Flow Microperfusion as a dermal pharmacokinetic approach to evaluate topical bioequivalence. Clin. Pharmacokinet. 2017, 56, 91–98. [Google Scholar] [CrossRef] [Green Version]
- Höfferer, C.; Tutkur, D.; Fledelius, C.; Brand, C.L.; Alsted, T.J.; Damgaard, J.; Nishimura, E.; Jeppesen, C.B.; Mautner, S.I.; Pieber, T.R.; et al. Open flow microperfusion: Pharmacokinetics of human insulin and insulin detemir in the interstitial fluid of subcutaneous adipose tissue. Diabetes Obes. Metab. 2015, 17, 121–127. [Google Scholar] [CrossRef]
- Hummer, J.; Altendorfer-Kroath, T.; Birngruber, T. Cerebral open flow microperfusion to monitor drug transport across the blood-brain barrier. Curr. Protoc. Pharmacol. 2019, 85. [Google Scholar] [CrossRef] [PubMed]
- Wong, R.; Geyer, S.; Weninger, W.; Guimberteau, J.C.; Wong, J.K. The dynamic anatomy and patterning of skin. Exp. Dermatol. 2016, 25, 92–98. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Risueño, I.; Valencia, L.; Jorcano, J.L.; Velasco, D. Skin-on-a-chip models: General overview and future perspectives. APL Bioeng. 2021, 5, 030901. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.C.; Chang, Y.H.; Shyu, W.C.; Lin, S.Z. Human umbilical cord mesenchymal stem cells: A new era for stem cell therapy. Cell Transplant. 2015, 24, 339–347. [Google Scholar] [CrossRef]
- Cofano, F.; Boido, M.; Monticelli, M.; Zenga, F.; Ducati, A.; Vercelli, A.; Garbossa, D. Mesenchymal stem cells for spinal cord injury: Current options limitations, and future of cell therapy. Int. J. Mol. Sci. 2019, 20, 2698. [Google Scholar] [CrossRef] [Green Version]
- Squillaro, T.; Peluso, G.; Galderisi, U. Clinical trials with mesenchymal stem cells: An update. Cell Transplant. 2016, 25, 829–848. [Google Scholar] [CrossRef] [Green Version]
- Han, Y.; Li, X.; Zhang, Y.; Han, Y.; Chang, F.; Ding, J. Mesenchymal stem cells for regenerative medicine. Cells 2019, 8, 886. [Google Scholar] [CrossRef] [Green Version]
- Mazini, L.; Rochette, L.; Admou, B.; Amal, S.; Malka, G. Hopes and limits of adipose-derived stem cells (ADSCs) and mesenchymal stem cells (MSCs) in wound healing. Int. J. Mol. Sci. 2020, 21, 1306. [Google Scholar] [CrossRef] [Green Version]
- Komaki, M.; Numata, Y.; Morioka, C.; Honda, I.; Tooi, M.; Yokoyama, N.; Ayame, H.; Iwasaki, K.; Taki, A.; Oshima, N.; et al. Exosomes of human placenta-derived mesenchymal stem cells stimulate angiogenesis. Stem Cell Res. Ther. 2017, 8, 219. [Google Scholar] [CrossRef]
- Pers, Y.M.; Ruiz, M.; Noël, D.; Jorgensen, C. Mesenchymal stem cells for the management of inflammation in osteoarthritis: State of the art and perspectives. Osteoarthr. Cartil. 2015, 23, 2027–2035. [Google Scholar] [CrossRef] [Green Version]
- Singer, N.G.; Caplan, A.I. Mesenchymal stem cells: Mechanisms of inflammation. Annu. Rev. Pathol. Mech. Dis. 2011, 6, 457–478. [Google Scholar] [CrossRef] [Green Version]
- Harrell, C.R.; Jankovic, M.G.; Fellabaum, C.; Volarevic, A.; Djonov, V.; Arsenijevic, A.; Volarevic, V. Molecular mechanisms responsible for anti-inflammatory and immunosuppressive effects of mesenchymal stem cell-derived factors. In Advances in Experimental Medicine and Biology; Springer: New York, NY, USA, 2019; Volume 1084, pp. 187–206. [Google Scholar]
- Lavrentieva, A.; Kirsch, M.; Birnstein, L.; Pepelanova, I.; Handke, W.; Rach, J.; Seltsam, A.; Scheper, T. Gelatin-methacryloyl (GelMA) formulated with human platelet lysate supports mesenchymal stem cell proliferation and differentiation and enhances the hydrogel’s mechanical properties. Bioengineering 2019, 6, 76. [Google Scholar] [CrossRef] [Green Version]
- Xiao, W.; Shinohara, M.; Komori, K.; Sakai, Y.; Matsui, H.; Osada, T. The importance of physiological oxygen concentrations in the sandwich cultures of rat hepatocytes on gas-permeable membranes. Biotechnol. Prog. 2014, 30, 1401–1410. [Google Scholar] [CrossRef]
- Bellio, M.A.; Rodrigues, C.O.; Landin, A.M.; Hatzistergos, K.E.; Kuznetsov, J.; Florea, V.; Valasaki, K.; Khan, A.; Hare, J.M.; Schulman, I.H. Physiological and hypoxic oxygen concentration differentially regulates human C-Kit cardiac stem cell proliferation and migration. J. Physiol. Heart Circ. Physiol. 2016, 311, 1509–1519. [Google Scholar] [CrossRef] [Green Version]
- Ziółkowska-Suchanek, I. Mimicking tumor hypoxia in non-small cell lung cancer employing three-dimensional in vitro models. Cells 2021, 10, 141. [Google Scholar] [CrossRef]
- Liverani, C.; de Vita, A.; Minardi, S.; Kang, Y.; Mercatali, L.; Amadori, D.; Bongiovanni, A.; la Manna, F.; Ibrahim, T.; Tasciotti, E. A biomimetic 3D model of hypoxia-driven cancer progression. Sci. Rep. 2019, 9, 12263. [Google Scholar] [CrossRef] [Green Version]
- Heiden, M.G.V.; Cantley, L.C.; Thompson, C.B. Understanding the warburg effect: The metabolic requirements of cell proliferation. Science 2009, 324, 1029–1033. [Google Scholar] [CrossRef] [Green Version]
- Kruse, C.R.; Singh, M.; Targosinski, S.; Sinha, I.; Sørensen, J.A.; Eriksson, E.; Nuutila, K. The effect of PH on cell viability, cell migration, cell proliferation, wound closure, and wound reepithelialization: In vitro and in vivo study. Wound Repair Regen. 2017, 25, 260–269. [Google Scholar] [CrossRef]
- Gladden, L.B. Lactate metabolism: A new paradigm for the third millennium. J. Physiol. 2004, 558, 5–30. [Google Scholar] [CrossRef]
- Mulukutla, B.C.; Yongky, A.; Le, T.; Mashek, D.G.; Hu, W.S. Regulation of glucose metabolism—A perspective from cell bioprocessing. Trends Biotechnol. 2016, 34, 638–651. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, H. Reprogramming of glucose, fatty acid and amino acid metabolism for cancer progression. Cell. Mol. Life Sci. 2016, 73, 377–392. [Google Scholar] [CrossRef] [PubMed]
- Michl, J.; Park, K.C.; Swietach, P. Evidence-based guidelines for controlling PH in mammalian live-cell culture systems. Commun. Biol. 2019, 2, 144. [Google Scholar] [CrossRef] [PubMed]
- Al-Ani, A.; Toms, D.; Kondro, D.; Thundathil, J.; Yu, Y.; Ungrin, M. Oxygenation in cell culture: Critical parameters for reproducibility are routinely not reported. PLoS ONE 2018, 13, e0204269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, J.; Liu, Y.; Yin, J.; Li, Q.; Li, Y.; Gu, J.; Cai, W.; Yin, G. Oxygen-glucose-deprivation/reoxygenation-induced autophagic cell death depends on JNK-mediated phosphorylation of Bcl-2. Cell. Physiol. Biochem. 2016, 38, 1063–1074. [Google Scholar] [CrossRef] [PubMed]
- Nikolits, I.; Nebel, S.; Egger, D.; Kreß, S.; Kasper, C. Towards physiologic culture approaches to improve standard cultivation of mesenchymal stem cells. Cells 2021, 10, 886. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kreß, S.; Schaller-Ammann, R.; Feiel, J.; Wegener, J.; Priedl, J.; Dietrich, W.; Kasper, C.; Egger, D. Innovative Platform for the Advanced Online Monitoring of Three-Dimensional Cells and Tissue Cultures. Cells 2022, 11, 412. https://doi.org/10.3390/cells11030412
Kreß S, Schaller-Ammann R, Feiel J, Wegener J, Priedl J, Dietrich W, Kasper C, Egger D. Innovative Platform for the Advanced Online Monitoring of Three-Dimensional Cells and Tissue Cultures. Cells. 2022; 11(3):412. https://doi.org/10.3390/cells11030412
Chicago/Turabian StyleKreß, Sebastian, Roland Schaller-Ammann, Jürgen Feiel, Joachim Wegener, Joachim Priedl, Wolf Dietrich, Cornelia Kasper, and Dominik Egger. 2022. "Innovative Platform for the Advanced Online Monitoring of Three-Dimensional Cells and Tissue Cultures" Cells 11, no. 3: 412. https://doi.org/10.3390/cells11030412
APA StyleKreß, S., Schaller-Ammann, R., Feiel, J., Wegener, J., Priedl, J., Dietrich, W., Kasper, C., & Egger, D. (2022). Innovative Platform for the Advanced Online Monitoring of Three-Dimensional Cells and Tissue Cultures. Cells, 11(3), 412. https://doi.org/10.3390/cells11030412







