Genome-Wide Characterization and Expression Profiling of Squamosa Promoter Binding Protein-Like (SBP) Transcription Factors in Wheat (Triticum aestivum L.)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sequences Retrieval
2.2. Phylogenetic Analysis
2.3. Exon–Intron Structure and Motif Analysis
2.4. Chromosomal Location and Duplication Patterns of Wheat SBP Proteins
2.5. Cis-Acting Elements and miR156 Target Site Prediction
2.6. Multiple Conditional Transcriptome Analysis of TaSBP
2.7. Characterization of Wheat SBP Proteins
3. Results
3.1. SBP Sequences Search
3.2. Classification of the SBP Gene Family
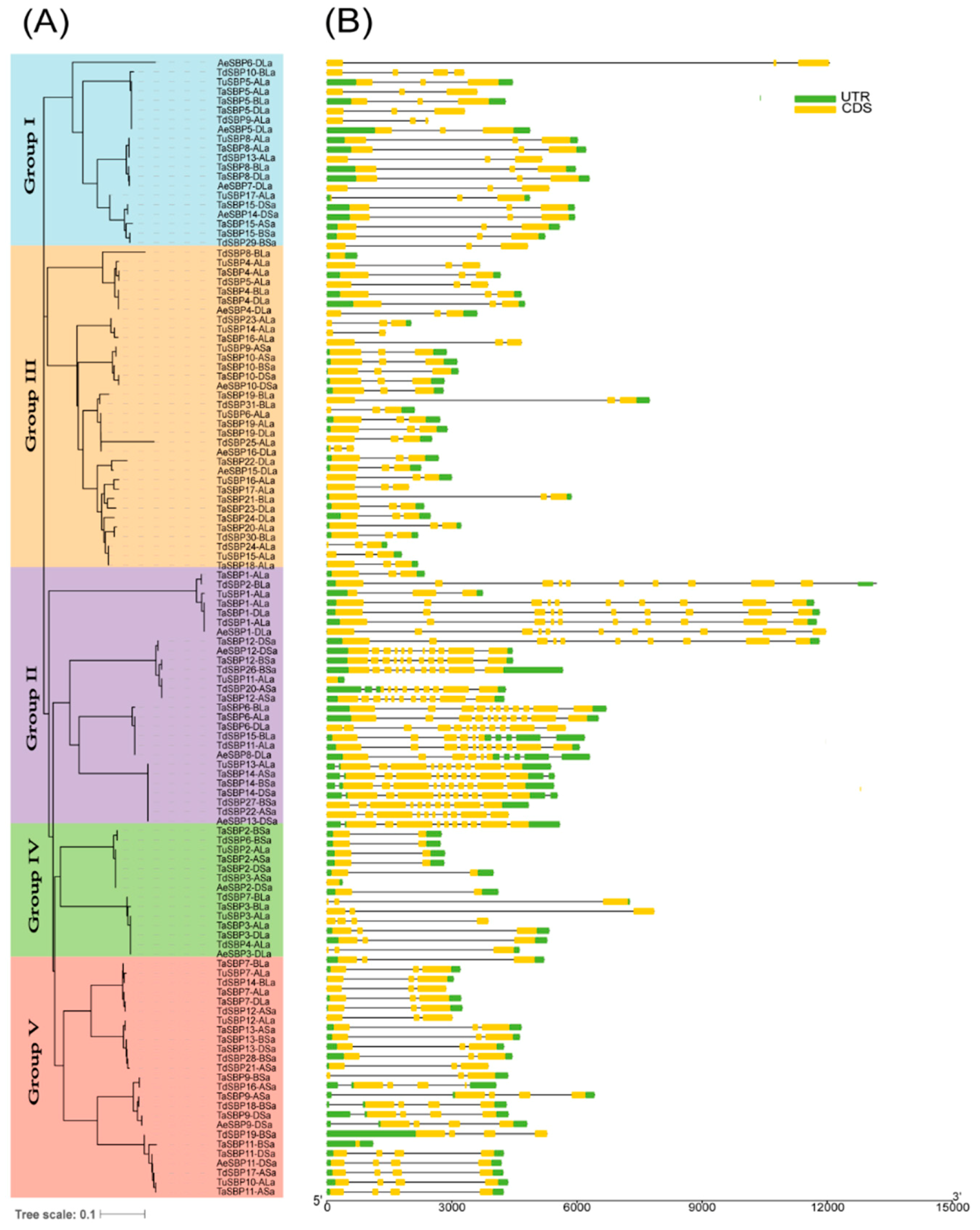
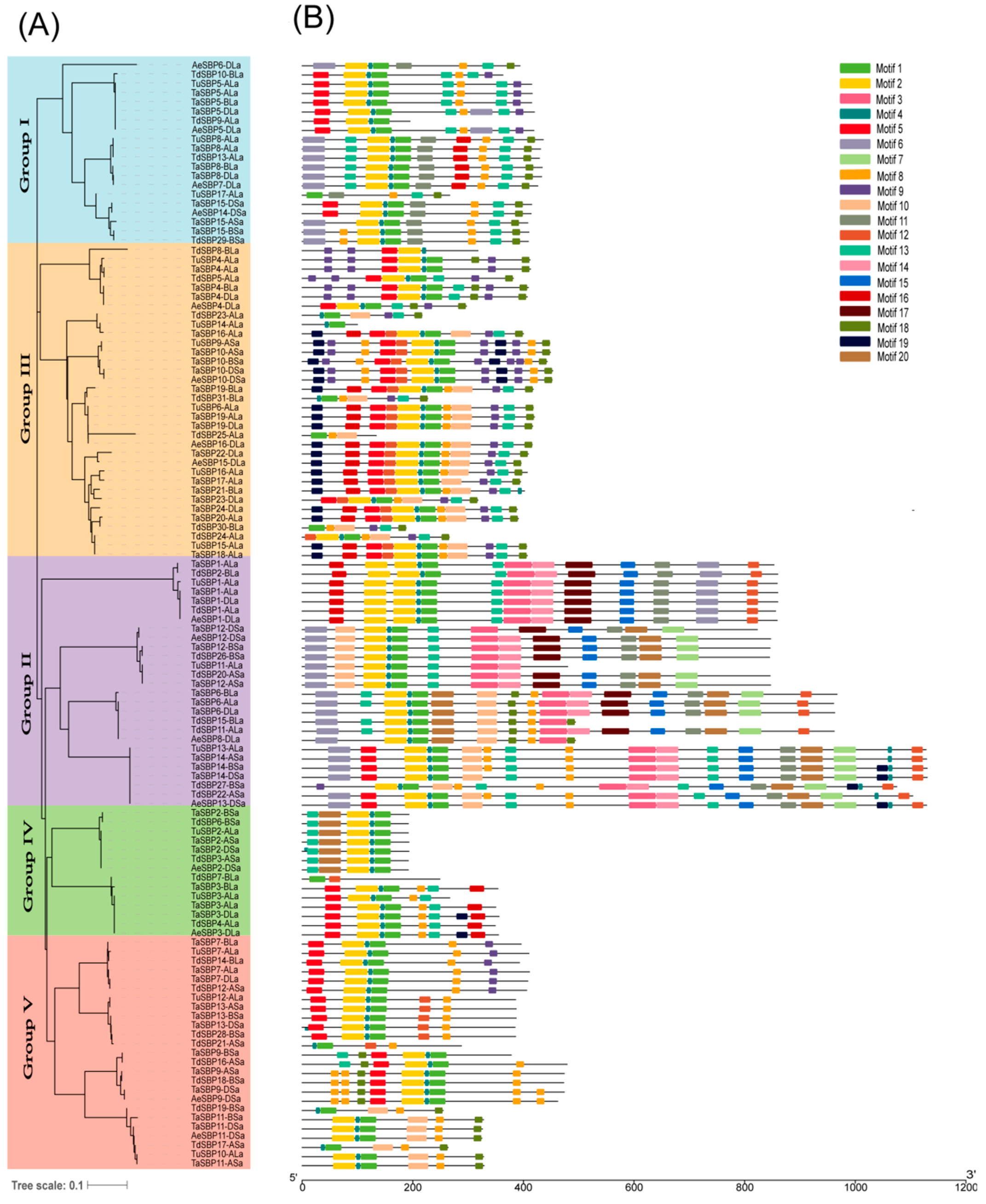
3.3. The Pattern of Gene Structure and Conserved Motifs
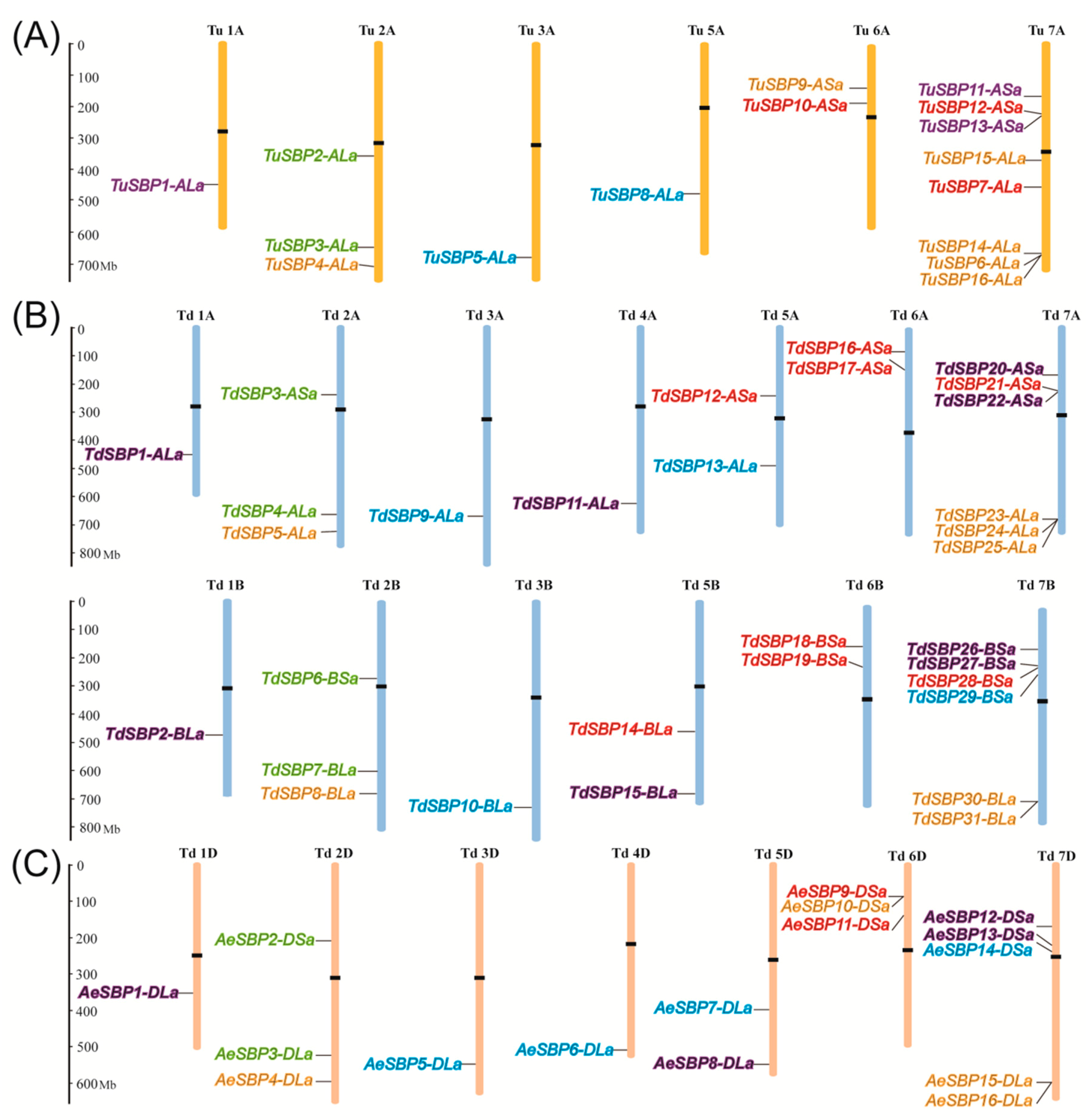
3.4. Chromosomal Distribution of SBP Genes
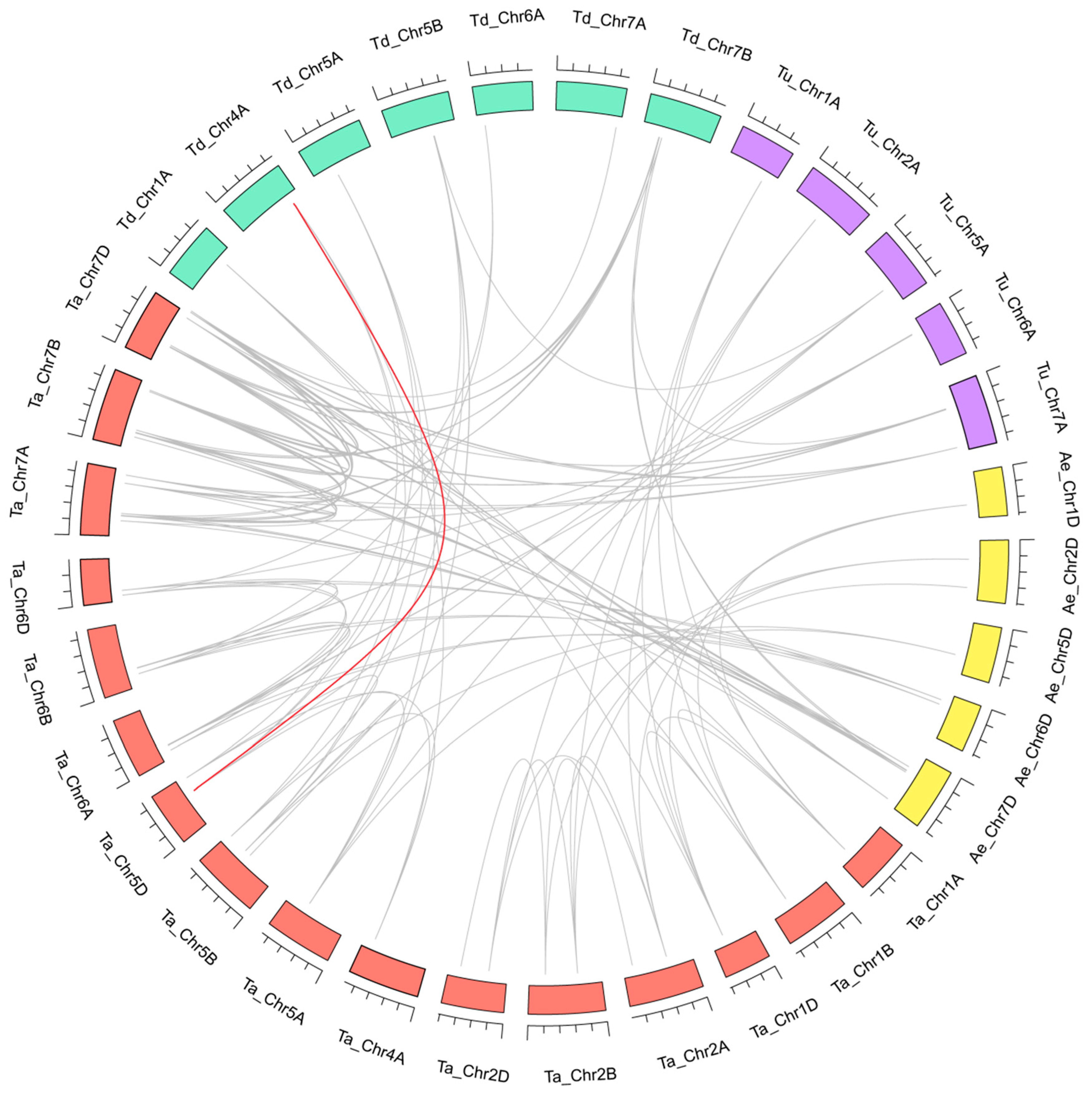
3.5. Homologous Gene Pairs and Synteny Analysis
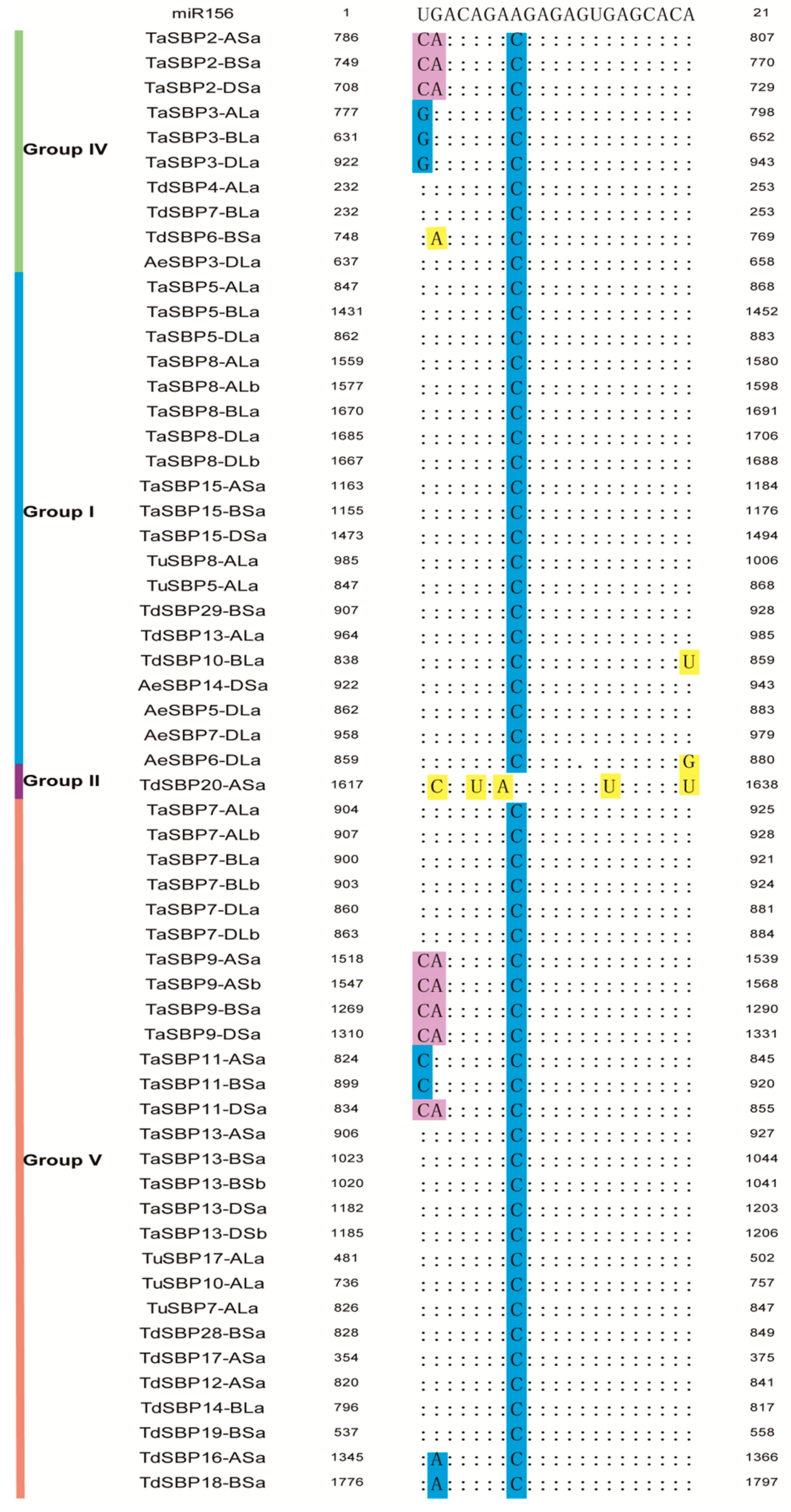
3.6. SBP Genes with MicroRNA156 Target Sites
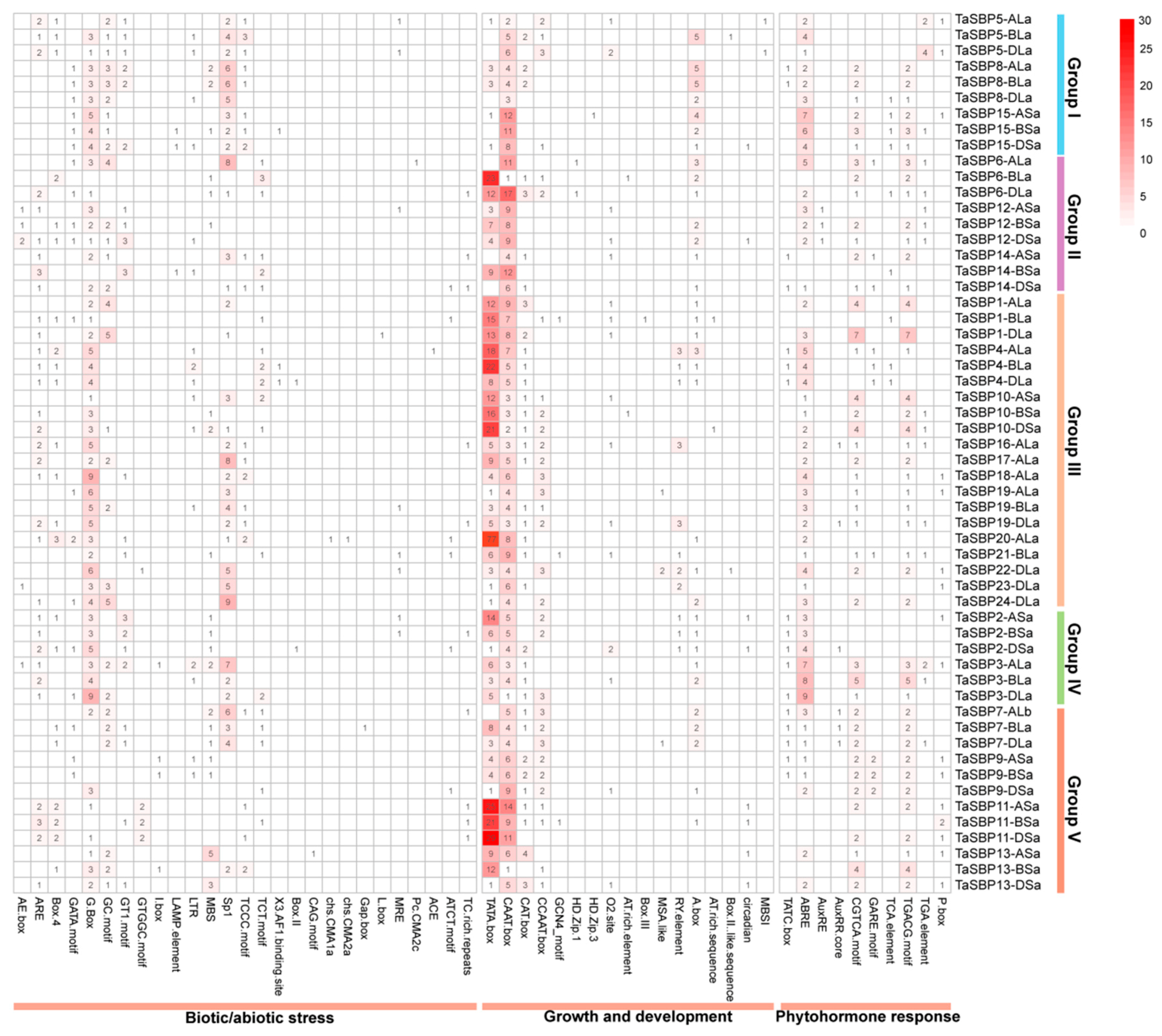
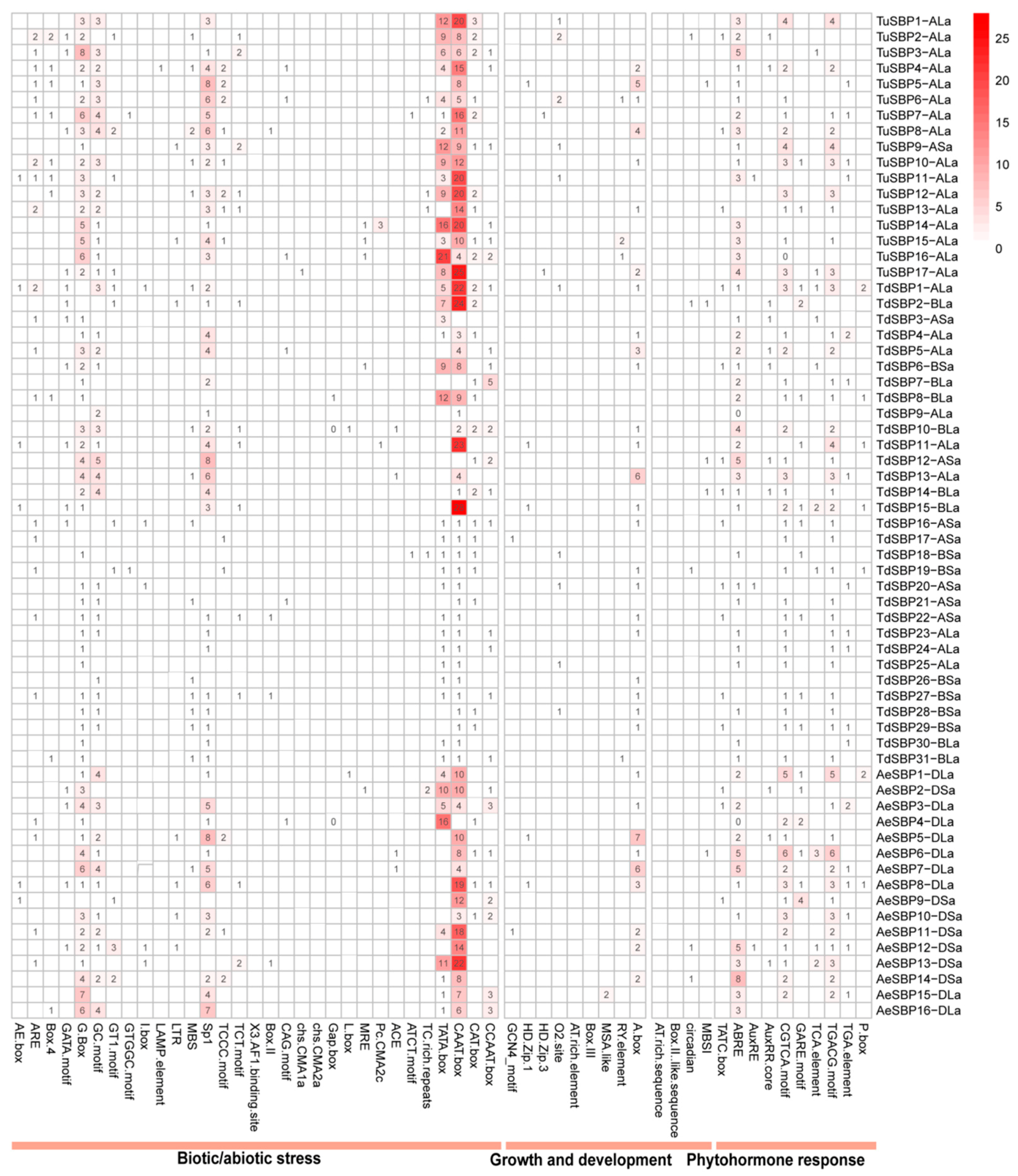
3.7. Identification of Cis-Acting Elements in the Promoter of SBP Genes
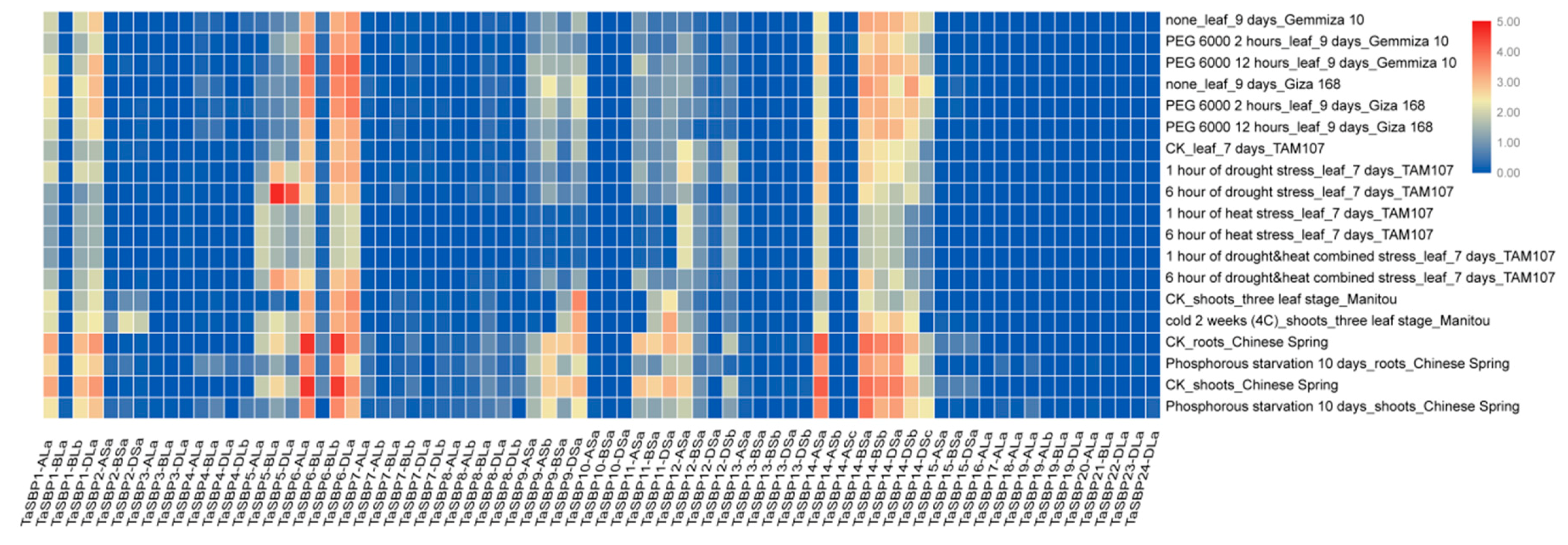
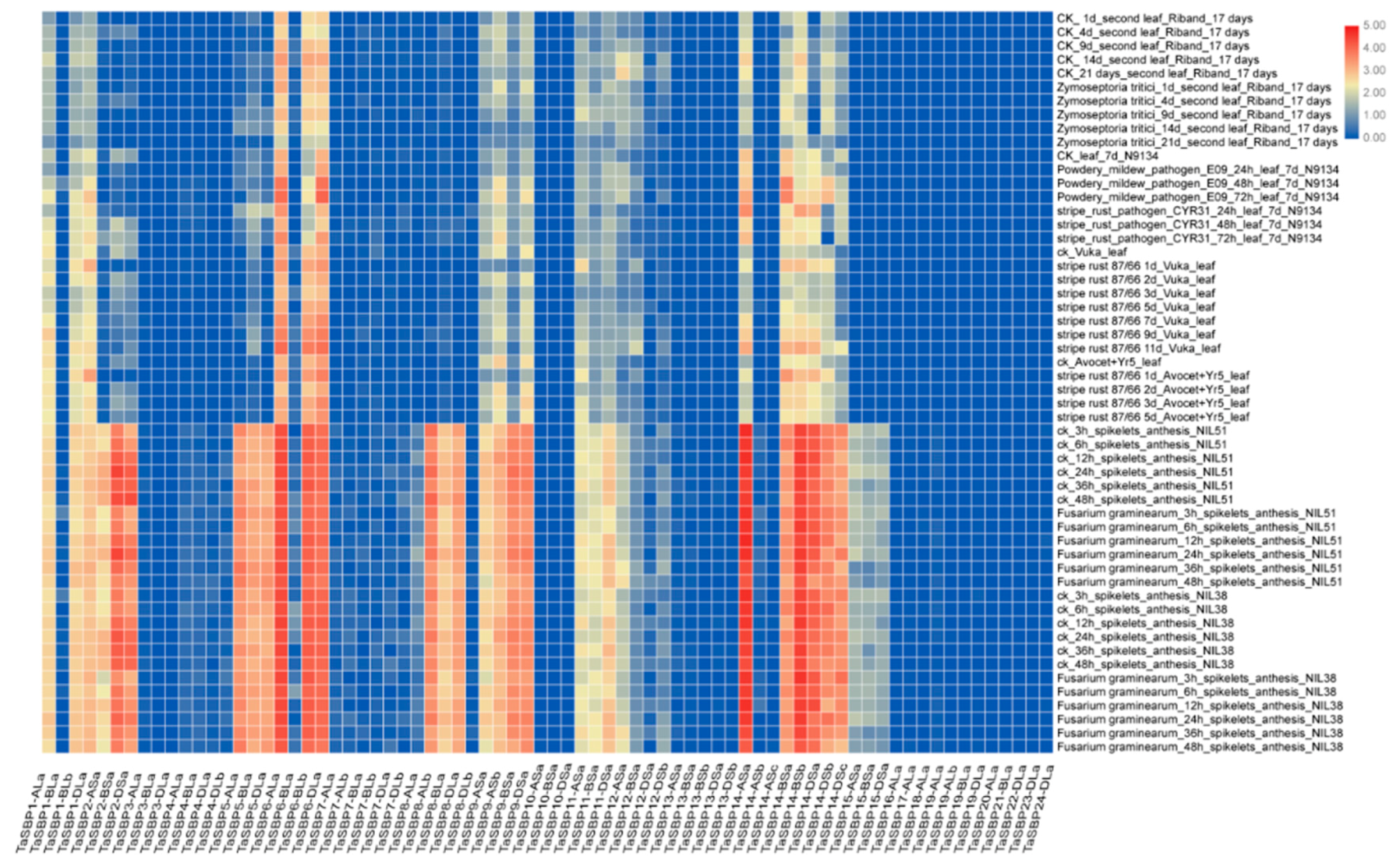
3.8. Expression Analysis of SBP Genes in Wheat
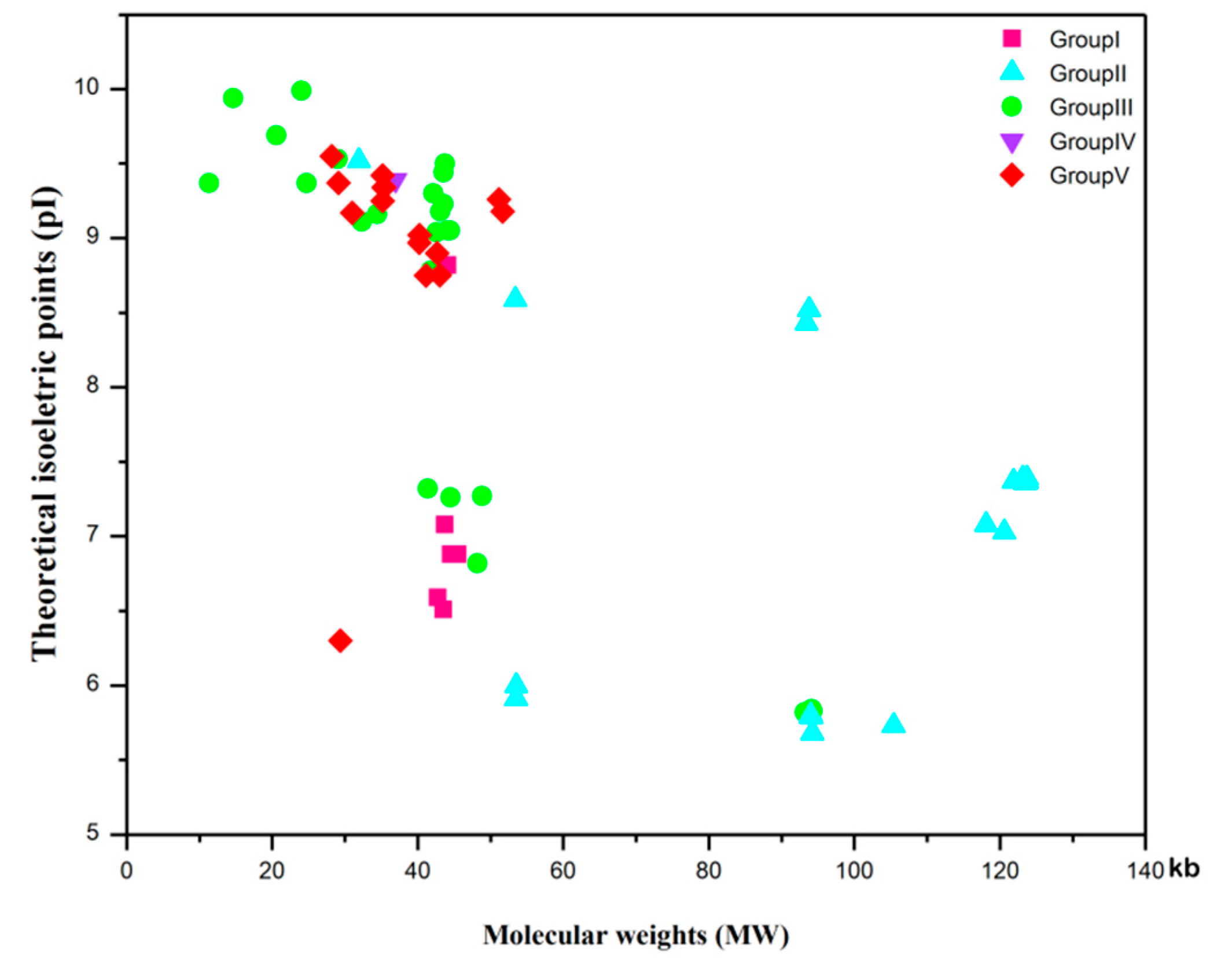
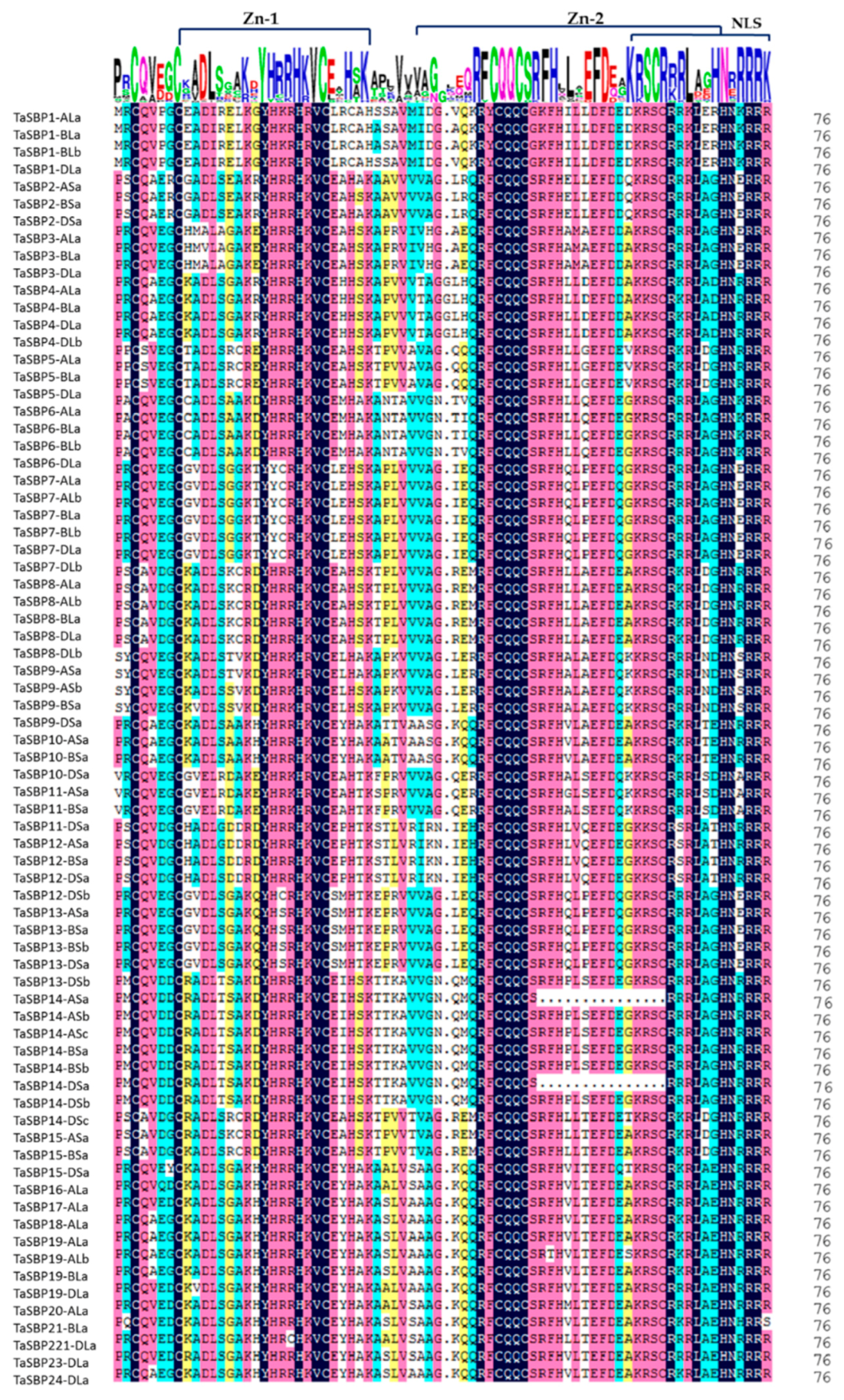
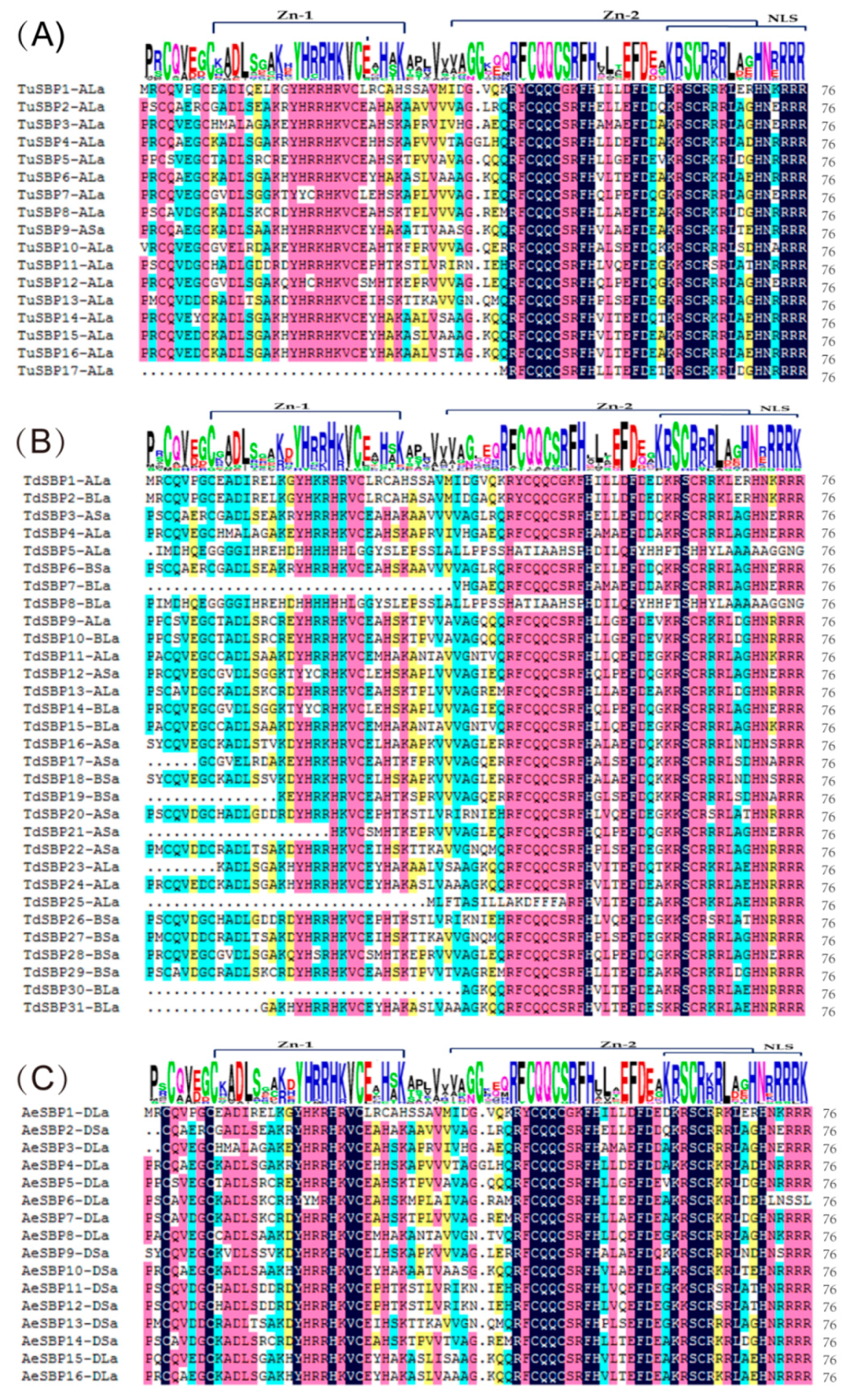
3.9. Protein Features and Conservative Domain Analyses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ilhan, E.; Büyük, I.; Inal, B. Transcriptome-Scale characterization of salt responsive bean TCP transcription factors. Gene 2018, 642, 64–73. [Google Scholar] [CrossRef] [PubMed]
- Shu, K.; Zhou, W.G.; Yang, W.Y. APETALA 2-domain-containing transcription factors: Focusing on abscisic acid and gibberellins antagonism. New Phytol. 2018, 217, 977–983. [Google Scholar] [CrossRef] [PubMed]
- Fu, F.F.; Xue, H.W. Coexpression Analysis Identifies Rice Starch Regulator1, a Rice AP2/EREBP Family Transcription Factor, as a Novel Rice Starch Biosynthesis Regulator. Plant Physiol. 2010, 154, 927–938. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Li, W.; Hu, X.; Guo, J.; Liu, A.; Zhang, B. A cotton MYB transcription factor, GbMYB5, is positively involved in plant adaptive response to drought stress. Plant Cell Physiol. 2015, 56, 917–929. [Google Scholar] [CrossRef] [PubMed]
- Heck, C.; Kuhn, H.; Heidt, S.; Walter, S.; Rieger, N.; Requena, N. Symbiotic Fungi Control Plant Root Cortex Development through the Novel GRAS Transcription Factor MIG1. Curr. Biol. 2016, 26, 2770–2778. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- An, J.P.; Qu, F.J.; Yao, J.F.; Wang, X.N.; You, C.X.; Wang, X.F.; Hao, Y.J. The bZIP transcription factor MdHY5 regulates anthocyanin accumulation and nitrate assimilation in apple. Hortic. Res. 2017, 4, 17023. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, K.; Kigawa, T.; Inoue, M.; Tateno, M.; Yamasaki, T.; Yabuki, T.; Aoki, M.; Seki, E.; Matsuda, T.; Nunokawa, E.; et al. A novel zinc-binding motif revealed by solution structures of DNA-binding domains of Arabidopsis SBP-family transcription factors. J. Mol. Biol. 2004, 337, 49–63. [Google Scholar] [CrossRef]
- Klein, J.; Saedler, H.; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene squamos. Mol. Gen. Genet. 1996, 250, 7–16. [Google Scholar] [CrossRef]
- Song, A.; Gao, T.W.; Wu, D.; Xin, J.J.; Chen, S.; Guan, Z.Y.; Wang, H.B.; Jin, L.L.; Chen, F.D. Transcriptome-wide identification and expression analysis of chrysanthemum SBP-like transcription factors. Plant Physiol. Biochem. 2016, 102, 10–16. [Google Scholar] [CrossRef]
- Birkenbihl, R.P.; Jach, G.; Saedler, H.; Huijser, P. Functional dissection of the plant-specific SBP-domain: Overlap of the DNA-binding and nuclear localization domains. J. Mol. Biol. 2005, 352, 585–596. [Google Scholar] [CrossRef]
- Feng, P.; Wang, Y.; Liu, H.L.; Wu, M.; Chu, W.Y.; Chen, D.M.; Yan, X. Genome-wide identification and expression analysis of SBP-like transcription factor genes in Moso Bamboo (Phyllostachys edulis). BMC Genom. 2017, 18, 486. [Google Scholar] [CrossRef]
- Harreman, M.T.; Kline, T.M.; Milford, H.G.; Harben, M.B.; Hodel, A.E.; Corbett, A.H. Regulation of nuclear import by phosphorylation adjacent to nuclear localization signals. J. Biol. Chem. 2004, 279, 20613–20621. [Google Scholar] [CrossRef] [PubMed]
- Huijser, P.; Klein, J.; Lönnig, W.E.; Meijer, H.; Saedler, H.; Sommer, H. Bracteomania, an inflorescence anomaly, is caused by the loss of function of the MADS-box gene squamosa in Antirrhinum majus. Embo J. 1992, 11, 1239–1249. [Google Scholar] [CrossRef]
- Cardon, G.; Höhmann, S.; Klein, J.; Nettesheim, K.; Saedler, H.; Huijser, P. Molecular characterisation of the Arabidopsis SBP-box genes. Gene. 1999, 237, 91. [Google Scholar] [CrossRef]
- Xie, K. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microrna156 in rice. Plant Physiol. 2006, 142, 280–293. [Google Scholar] [CrossRef] [PubMed]
- María, S.; Xing, S.; H?Hmann, S.; Berndtgen, R.; Huijser, P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta 2012, 235, 1171–1184. [Google Scholar] [CrossRef]
- Mao, H.D.; Yu, L.J.; Li, Z.J.; Yan, Y.; Han, R.; Liu, H.; Ma, M. Genome-wide analysis of the SPL family transcription factors and their responses to abiotic stresses in maize. Plant Gene 2016, 6, 1–12. [Google Scholar] [CrossRef] [Green Version]
- Hongmin, H.; Jun, L.; Min, G.; Singer, S.D.; Hao, W.; Linyong, M. Genomic Organization, Phylogenetic Comparison and Differential Expression of the SBP-Box Family Genes in Grape. PLoS ONE 2013, 8, e59358. [Google Scholar] [CrossRef]
- Zhang, X.; Dou, L.; Pang, C.; Song, M.; Wei, H.; Fan, S.; Wang, C.; Yu, S. Genomic organization, differential expression, and functional analysis of the SPL gene family in Gossypium hirsutum. Mol. Genet. Genom. MGG 2015, 290, 115–126. [Google Scholar] [CrossRef]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef]
- Jung, J.H.; Ju, Y.; Seo, P.J.; Lee, J.H.; Park, C.M. The SOC1-SPL module integrates photoperiod and gibberellic acid signals to control flowering time in Arabidopsis. Plant J. 2012, 69, 577–588. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Hu, Z.L.; Yang, Y.X.; Chen, X.Q.; Chen, G.P. Function annotation of an SBP-box gene in Arabidopsis based on analysis of co-expression networks and promoters. Int. J. Mol. Sci. 2009, 10, 116–132. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.J.; Lee, J.H.; Kim, W.; Jung, H.S.; Huijser, P.; Ahn, J.H. The microRNA156-SQUAMOSA PROMOTER BINDING PROTEIN-LIKE3 module regulates ambient temperature-responsive flowering via FLOWERING LOCUS T in Arabidopsis. Plant Physiol. 2012, 159, 461–478. [Google Scholar] [CrossRef] [PubMed]
- Yamasaki, K.; Hagiwara, H. Excess iron inhibits osteoblast metabolism. Toxicol. Lett. 2009, 191, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.M.; Liang, X.; Nekl, E.R.; Stiers, J.J. Arabidopsis AtSPL14, a plant-specific SBP-domain transcription factor, participates in plant development and sensitivity to fumonisin B1. Plant J. 2010, 41, 744–754. [Google Scholar] [CrossRef] [PubMed]
- Lan, T.; Zhang, S.; Liu, T.; Wang, B.; Guan, H.; Zhou, Y.; Duan, Y.; Wu, W. Fine mapping and candidate identification of SST, a gene controlling seedling salt tolerance in rice (Oryza sativa L.). Euphytica 2015, 205, 269–274. [Google Scholar] [CrossRef]
- Jung, J.H.; Lee, H.J.; Ryu, J.Y.; Park, C.M. SPL3/4/5 Integrate Developmental Aging and Photoperiodic Signals into the FT-FD Module in Arabidopsis Flowering. Mol. Plant. 2016, 9, 1647–1659. [Google Scholar] [CrossRef] [Green Version]
- Ma, Y.; Chen, X.L.; Zhang, H.X.; Tu, Y. Cloning and Expression Analysis of Transcription Factor CmSBP11 in melon. Genom. Appl. Biol. 2018, 37, 345–349. [Google Scholar] [CrossRef]
- Miura, K.; Ikeda, M.; Matsubara, A.; Song, X.J.; Ito, M.; Asano, K.; Matsuoka, M.; Kitano, H.; Ashikari, M. OsSPL14 promotes panicle branching and higher grain productivity in rice. Nat. Genet. 2010, 6, 545–549. [Google Scholar] [CrossRef]
- Kim, S.R.; Ramos, J.M.; Hizon, R.J.M.; Ashikari, M.; Virk, P.S.; Torres, E.A.; Nissila, E.; Jena, K.K. Introgression of a functional epigenetic OsSPL14WFP allele into elite indica rice genomes greatly improved panicle traits and grain yield. Sci. Rep. 2018, 8, 3833. [Google Scholar] [CrossRef]
- Yin, J.L.; Fang, Z.W.; Sun, C.; Zhang, P.; Zhang, X.; Lu, C.; Wang, S.P.; Ma, D.F.; Zhu, Y.X. Rapid identification of a stripe rust resistant gene in a space-induced wheat mutant using specific locus amplified fragment (SLAF) sequencing. Sci. Rep. 2018, 8, 3086. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Wei, J.; Song, N.; Wang, N.; Zhao, J.; Kang, Z. A novel wheat NAC transcription factor, TaNAC30, negatively regulates resistance of wheat to stripe rust. J. Integr. Plant Biol. 2018, 60, 432–443. [Google Scholar] [CrossRef] [PubMed]
- Schwessinger, B. Fundamental wheat stripe rust research in the 21st century. New Phytol. 2017, 213, 1625–1631. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Xu, K.; Liu, S.; Ya, J.; Su, F. Cloning, Tissue-specific Expression and Prokaryotic Expression Analysis of TaSPL17 Gene in Wheat(Triticum aestivum). Acta Agric. Boreal-Occident. Sin. 2015, 6, 1004–1389. [Google Scholar] [CrossRef]
- Zhang, B.; Xu, W.; Liu, X.; Mao, X.; Li, A.; Wang, J.; Chang, X.; Zhang, X.; Jing, R. Functional Conservation and Divergence among Homoeologs of TaSPL20 and TaSPL21, Two SBP-Box Genes Governing Yield-Related Traits in Hexaploid Wheat. Plant Physiol. 2017, 19, 1177–1191. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Cheng, X.; Liu, P.; Sun, J. miR156-Targeted SBP-Box Transcription Factors Interact with DWARF53 to Regulate TEOSINTE BRANCHED1 and BARREN STALK1 Expression in Bread Wheat. Plant Physiol. 2017, 174, 1931–1948. [Google Scholar] [CrossRef] [PubMed]
- Guo, A.Y.; Zhu, Q.H.; Gu, X.; Ge, S.; Yang, J.; Luo, J. Genome-wide identification and evolutionary analysis of the plant specific SBP-box transcription factor family. Gene 2008, 418, 1–8. [Google Scholar] [CrossRef]
- Abdullah, M.; Cao, Y.; Cheng, X.; Shakoor, A.; Su, X.; Gao, J.; Cai, Y. Genome-Wide Analysis Characterization and Evolution of SBP Genes in Fragaria vesca, Pyrus bretschneideri, Prunus persica and Prunus mume. Front. Genet. 2018, 9, 1–12. [Google Scholar] [CrossRef]
- Wang, B.N.; Geng, S.F.; Wang, D.; Feng, N.; Zhang, D.D.; Wu, L.; Hao, C.Y.; Zhang, X.Y.; Li, A.L.; Long, M. Characterization of Squamosa Promoter Binding Protein-LIKE Genes in Wheat. J. Plant Biol. 2015, 58, 220–229. [Google Scholar] [CrossRef]
- István, M.; Marie, K.; Hana, Š.; András, F.; András, C.; Mária, M.; Jan, V.; Márta, M.L.; Jaroslav, D. Flow cytometric chromosome sorting from diploid progenitors of bread wheat, T. urartu, Ae. speltoides and Ae. Tauschii. Theor. Appl. Genet. 2014, 127, 1091–1104. [Google Scholar] [CrossRef]
- Dvorák, J.; Terlizzi, P.D.; Zhang, H.B.; Resta, P. The evolution of polyploid wheat: Identification of the A genome donor species. Genome 1993, 36, 21–31. [Google Scholar] [CrossRef]
- Maestra, B.; Naranjo, T. Homoeologous relationships of Aegilops speltoides chromosomes to bread wheat. Theoret. Appl. Genet. 1998, 97, 181–186. [Google Scholar] [CrossRef]
- Avni, R.; Nave, M.; Barad, O.; Baruch, K.; Twardziok, S.O.; Gundlach, H.; Hale, I.; Mascher, M.; Spannagl, M.; Wiebe, K.; et al. Wild emmer genome architecture and diversity elucidate wheat evolution and domestication. Science 2017, 357, 93–97. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Mistry, J.; Schuster-Bockler, B.; Griffiths-Jones, S.; Hollich, V.; Lassmann, T.; Moxon, S.; Marshall, M.; Khanna, A.; Durbin, R.; et al. Pfam: Clans, web tools and services. Nucleic Acids Res. 2006, 34, 247–251. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Copley, R.R.; Schmidt, S.; Ciccarelli, F.D.; Doerks, T.; Schultz, J.; Ponting, C.P.; Bork, P. SMART 4.0: Towards genomic data integration. Nucleic Acids Res. 2004, 32, 142–144. [Google Scholar] [CrossRef] [PubMed]
- Quevillon, E.; Silventoinen, V.; Pillai, S.; Harte, N.; Mulder, N.; Apweiler, R.; Lopez, R. InterProScan: Protein domains identifier. Nucleic Acids Res. 2005, 33, 116–120. [Google Scholar] [CrossRef] [PubMed]
- Oliver, T.; Schmidt, B.; Nathan, D.; Clemens, R.; Maskell, D. Using reconfigurable hardware to accelerate multiple sequence alignment with clustalw. Bioinformatics 2005, 21, 3431–3432. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, 242–245. [Google Scholar] [CrossRef]
- Hu, B.; Jin, J.; Guo, A.Y.; Zhang, H.; Luo, J.; Gao, G. GSDS 2.0: An upgraded gene feature visualization server. Bioinformatics 2015, 31, 1296–1297. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Williams, N.; Misleh, C.; Li, W.W. MEME: Discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 2006, 34, 369–373. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.J.; Xia, R.; Chen, H.; He, Y.H. TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. Biorxiv 2018. [Google Scholar] [CrossRef]
- Wang, G.; Wang, T.; Jia, Z.H.; Xuan, J.P.; Pan, D.L.; Guo, Z.R.; Zhang, J.Y. Genome-Wide Bioinformatics Analysis of MAPK Gene Family in Kiwifruit (Actinidia Chinensis). Int. J. Mol. Sci. 2018, 19, 2510. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Cavalcanti, A.; Chen, F.C.; Bouman, P.; Li, W.H. Extent of Gene Duplication in the Genomes of Drosophila, Nematode, and Yeast. Mol. Biol. Evol. 2002, 19, 256–262. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, 49. [Google Scholar] [CrossRef] [PubMed]
- Gu, Z.; Gu, L.; Eils, R.; Schlesner, M.; Brors, B. circlize implements and enhances circular visualization in R. Genome Anal. 2014, 30, 2811–2812. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rozas, J. DNA sequence polymorphism analysis using DnaSP. Methods Mol. Biol. 2009, 537, 337–350. [Google Scholar] [CrossRef] [PubMed]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Dai, X.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server. Nucleic Acids Res. 2011, 39, 155–159. [Google Scholar] [CrossRef] [PubMed]
- Yin, J.L.; Liu, M.Y.; Ma, D.F.; Wu, J.W.; Lia, S.L.; Zhua, Y.X.; Han, B. Identification of circular RNAs and their targets during tomato fruit ripening. Postharvest Biol. Technol. 2018, 136, 90–98. [Google Scholar] [CrossRef]
- Trapnel, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Denise, M.; Timothy, W.H.; Han, S.J.; Michael, Y.G.; Arielle, G.Z.; Allyson, L.B.; Oliver, J.H.; Alexandra, M.O.; Mariam, Q.; Giorgio, T.; et al. Microbiota-dependent sequelae of acute infection compromise tissue-specific immunity. Cell 2015, 163, 354–366. [Google Scholar] [CrossRef]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein Identification and Analysis Tools in the ExPASy Server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [CrossRef] [PubMed]
- Chou, K.C.; Shen, H.B. Plant-mPLoc: A Top-Down Strategy to Augment the Power for Predicting Plant Protein Subcellular Localization. PLoS ONE 2010, 5, e11335. [Google Scholar] [CrossRef] [PubMed]
- Horton, P.; Park, K.J.; Obayashi, T.; Nakai, K. Protein subcellular localization prediction with WoLF PSORT. Asian Pac. Bioinform. Conf. 2006, 39–48. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; Heijne, G.V.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Ramírez-González, R.H.; Borrill, P.; Lang, D.; Harrington, S.A.; Brinton, J.; Venturini, L.; Davey, M.; Jacobs, J.; van Ex, F.; Pasha, A.; et al. The transcriptional landscape of polyploid wheat. Science 2018, 361, 662. [Google Scholar] [CrossRef]
- Feldman, M.; Levy, A.A. Allopolyploidy-a shaping force in the evolution of wheat genomes. Cytogenet Genome Res. 2005, 109, 250–258. [Google Scholar] [CrossRef]
- Griffiths-Jones, S.; Saini, H.K.; van Dongen, S.; Enright, A.J. miRBase: Tools for microRNA genomics. Nucleic Acids Res. 2008, 36, D154. [Google Scholar] [CrossRef]
- Zhu, Y.; Yang, L.; Liu, N.; Yang, J.; Zhou, X.; Xia, Y.; He, Y.; He, Y.; Gong, H.; Ma, D.; et al. Genome-wide identification, structure characterization, and expression pattern profiling of aquaporin gene family in cucumber. BMC Plant Biol. 2019, 19, 345. [Google Scholar] [CrossRef] [PubMed]
- Rouster, J.; Leah, R.; Mundy, J.; Cameron-Mills, V. Identification of a methyl jasmonate-responsive region in the promoter of a lipoxygenase 1 gene expressed in barley grain. Plant J. 2011, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Fujita, Y.; Fujita, M.; Satoh, R.; Maruyama, K.; Parvez, M.M.; Seki, M.; Hiratsu, K.; Ohme-Takagi, M.; Shinozaki, K.; Yamaguchi-Shinozaki, K. AREB1 is a transcription activator of novel ABRE-Dependent ABA signaling that enhances drought stress tolerance in Arabidopsis. Plant Cell 2005, 17, 3470. [Google Scholar] [CrossRef] [PubMed]
- Dai, F.G.; Hu, Z.L.; Chen, G.P.; Wang, B.Q.; Wang, Y. Progress in the plant specific SBP-box gene family. Chin. Bull. Life Sci. 2010, 22, 155–160. [Google Scholar] [CrossRef]
- Moreno, M.A.; Harper, L.C.; Krueger, R.W.; Dellaporta, S.L.; Freeling, M. Liguleless1 encodes a nuclear-localized protein required for induction of ligules and auricles during maize leaf organogenesis. Genes Dev. 1997, 11, 616–628. [Google Scholar] [CrossRef] [PubMed]
- Unte, U.S.; Sorensen, A.M.; Pesaresi, P.; Gandikota, M.; Leister, D.; Saedler, H.; Huijser, P. SPL8, an SBP-box gene that affects pollen sac development in Arabidopsis. Plant Cell 2003, 15, 1009–1019. [Google Scholar] [CrossRef] [PubMed]
- Ling, L.Z.; Zheng, S.D. Unraveling the Distribution and Evolution of miR156-targeted SPLs in Plants by Phylogenetic Analysis. Plant Divers. Resour. 2012, 34, 33–46. [Google Scholar] [CrossRef]
- Wong, D.C.J.; Gutierrez, R.L.; Gambetta, G.A.; Castellarin, S.D. Genome-wide analysis of cis-regulatory element structure and discovery of motif-driven gene co-expression networks in grapevine. DNA Res. 2017, 24, 311–326. [Google Scholar] [CrossRef] [Green Version]
- Riese, M.; Höhmann, S.; Saedler, H.; Thomas, M.; Huijser, P. Comparative analysis of the SBP-box gene families in P. patens and seed plants. Gene 2007, 401, 1–37. [Google Scholar] [CrossRef]
- Ma, J.; Yang, Y.; Luo, W.; Yang, C.; Ding, P.; Liu, Y.; Qiao, L.; Chang, Z.; Geng, H.; Wang, P.; et al. Genome-wide identification and analysis of the MADS-box gene family in bread wheat (Triticum aestivum L.). PLoS ONE 2017, 12, e0181443. [Google Scholar] [CrossRef]
- Yue, W.; Nie, X.; Cui, L.; Zhi, Y.; Zhang, T.; Du, X.; Song, W. Genome-wide sequence and expressional analysis of autophagy Gene family in bread wheat (Triticum aestivum L.). J. Plant Physiol. 2018, 229, 7–21. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Yue, H.; Feng, K.W.; Deng, P.C.; Song, W.N.; Nie, X.J. Genome-wide identification, phylogeny and expressional profiles of mitogen activated protein kinase kinase kinase (MAPKKK) gene family in bread wheat (Triticum aestivum L.). BMC Genom. 2016, 17, 668. [Google Scholar] [CrossRef] [PubMed]
- Feldman, M.; Liu, B.; Segal, G.; Abbo, S.; Levy, A.A.; Vega, J.M. Rapid elimination of low-copy DNA sequences in polyploidy wheat: A possible mechanism for differentiation of homoeologous chromosomes. Genetics 1997, 147, 1381–1387. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.C.; Belcram, H.; Gornicki, P.; Charles, M.; Just, J.; Huneau, C.; Magdelenat, G.; Couloux, A.; Samain, S.; Gill, B.S.; et al. Duplication and partitioning in evolution and function of homoeologous Q loci governing domestication characters in polyploid wheat. Proc. Natl. Acad. Sci. USA 2011, 108, 18737–18742. [Google Scholar] [CrossRef] [PubMed]
- Hao, J.; Yang, J.; Dong, J.; Fei, S.Z. Characterization of BdCBF, genes and genome-wide transcriptome profiling of BdCBF3 -dependent and -independent cold stress responses in Brachypodium Distachyon. Plant Sci. 2017, 262, 52. [Google Scholar] [CrossRef] [PubMed]
- Heidi, W.; Pierre, M.; Senthold, A.; Bruce, K.; Jeffrey, W.; Michael, O.; Gerard, W.W.; Giacomo, D.S.; Jordi, D.; Robert, G.; et al. Canopy temperature for simulation of heat stress in irrigated wheat in a semi-arid environment: A multi-model comparison. Field Crop. Res. 2015, 10, 6563. [Google Scholar] [CrossRef]
- Thomas, N.; Benedikt, W.; Sapna, S.; Christian, A.; Christoph, B.; Alexandra, P.; Matthias, P.; Gerald, S.; Barbara, S.; Marc, L.; et al. Joint Transcriptomic and Metabolomic Analyses Reveal Changes in the Primary Metabolism and Imbalances in the Subgenome Orchestration in the Bread Wheat Molecular Response to Fusarium graminearum. Genes Genomes Genet. 2015, 5, 2579–2592. [Google Scholar] [CrossRef]
- Lu, M.C.; Liu, Y.Q.; Chen, D.Y.; Xue, Y.X.; Mao, Y.B.; Chen, X.Y. Arabidopsis Transcription Factors SPL1 and SPL12 Confer Plant Thermotolerance at Reproductive Stage. Mol. Plant 2017, 10, 735–748. [Google Scholar]


| Gene Name | Accession Numbers | Chr | Location | Exo | CDS | Pro | Source of Information | Group |
|---|---|---|---|---|---|---|---|---|
| TaSBP1-ALa | TraesCS1A02G255300.1 | chr1A | 447623204–447635025 | 10 | 2580 | 859 | TaASPL9# | II |
| TaSBP1-BLa | TraesCS1B02G266100.1 | chr1B | 467540751–467553928 | 10 | 2559 | 852 | TaBSPL9# | II |
| TaSBP1-BLb | TraesCS1B02G266100.2 | chr1B | 467541883–467553928 | 10 | 861 | Identified in this study | II | |
| TaSBP1-DLa | TraesCS1D02G254700.1 | chr1D | 347199194–347210946 | 10 | 2580 | 859 | TaDSPL9# | II |
| TaSBP2-ASa | TraesCS2A02G232400.1 | chr2A | 276276336–276279155 | 2 | 579 | 192 | TaASPL13# | IV |
| TaSBP2-BSa | TraesCS2B02G250900.1 | chr2B | 260765427–260768180 | 2 | 579 | 192 | TaBSPL13# | IV |
| TaSBP2-DSa | TraesCS2D02G232800.1 | chr2D | 206632153–206636152 | 2 | 579 | 192 | Identified in this study | IV |
| TaSBP3-ALa | TraesCS2A02G413900.1 | chr2A | 670957294–670962629 | 3 | 1050 | 349 | TaASPL7# | IV |
| TaSBP3-BLa | TraesCS2B02G432700.1 | chr2B | 621960663–621968513 | 3 | 1062 | 353 | TaBSPL7# | IV |
| TaSBP3-DLa | TraesCS2D02G410700.1 | chr2D | 525512403–525517689 | 3 | 1068 | 355 | Identified in this study | IV |
| TaSBP4-ALa | TraesCS2A02G502300.1 | chr2A | 730613571–730617739 | 3 | 1239 | 412 | TaASPL8# | III |
| TaSBP4-BLa | TraesCS2B02G530400.1 | chr2B | 725575939–725580611 | 3 | 1227 | 408 | TaBSPL8# | III |
| TaSBP4-DLa | TraesCS2D02G502900.1 | chr2D | 596550011–596554762 | 3 | 1221 | 406 | TaDSPL8# | III |
| TaSBP4-DLb | TraesCS2D02G502900.2 | chr2D | 596550011–596554762 | 3 | 407 | Identified in this study | III | |
| TaSBP5-ALa | TraesCS3A02G432500.1 | chr3A | 673898993–673902601 | 3 | 1248 | 415 | Identified in this study | I |
| TaSBP5-BLa | TraesCS3B02G468400.1 | chr3B | 713320663–713324953 | 3 | 1245 | 414 | Identified in this study | I |
| TaSBP5-DLa | TraesCS3D02G425800.1 | chr3D | 538401064–538404375 | 3 | 1260 | 419 | Identified in this study | I |
| TaSBP6-ALa | TraesCS4A02G359500.1 | chr4A | 632857804–632864321 | 11 | 2883 | 960 | Identified in this study | II |
| TaSBP6-BLa | TraesCS5B02G512800.1 | chr5B | 677875145–677881852 | 11 | 2901 | 966 | TaBSPL6# | II |
| TaSBP6-BLb | TraesCS5B02G512800.2 | chr5B | 677875145–677881852 | 11 | 961 | Identified in this study | II | |
| TaSBP6-DLa | TraesCS5D02G513300.1 | chr5D | 537393493–537399223 | 12 | 2889 | 962 | TaDSPL6# | II |
| TaSBP7-ALa | TraesCS5A02G265900.1 | chr5A | 477584230–477587453 | 3 | 1233 | 410 | Identified in this study | V |
| TaSBP7-ALb | TraesCS5A02G265900.2 | chr5A | 477584230–477587453 | 3 | 411 | Identified in this study | V | |
| TaSBP7-BLa | TraesCS5B02G265600.1 | chr5B | 450105102–450108305 | 3 | 1188 | 395 | Identified in this study | V |
| TaSBP7-BLb | TraesCS5B02G265600.2 | chr5B | 450105198–450108305 | 3 | 396 | Identified in this study | V | |
| TaSBP7-DLa | TraesCS5D02G273900.1 | chr5D | 376936839–376940093 | 3 | 1224 | 407 | Identified in this study | V |
| TaSBP7-DLb | TraesCS5D02G273900.2 | chr5D | 376936978–376940093 | 3 | 408 | Identified in this study | V | |
| TaSBP8-ALa | TraesCS5A02G286700.1 | chr5A | 494569538–494575755 | 3 | 1293 | 430 | TaASPL6/16# | I |
| TaSBP8-ALb | TraesCS5A02G286700.2 | chr5A | 494569538–494575773 | 3 | 436 | Identified in this study | I | |
| TaSBP8-BLa | TraesCS5B02G286000.1 | chr5B | 471401006–471406978 | 3 | 1302 | 433 | TaBSPL16# | I |
| TaSBP8-DLa | TraesCS5D02G294400.1 | chr5D | 391372552–391378851 | 3 | 1299 | 432 | Identified in this study | I |
| TaSBP8-DLb | TraesCS5D02G294400.2 | chr5D | 391372552–391378851 | 3 | 426 | Identified in this study | I | |
| TaSBP9-ASa | TraesCS6A02G110100.1 | chr6A | 79267605–79271671 | 3 | 1134 | 377 | TaASPL3/14/18# | V |
| TaSBP9-ASb | TraesCS6A02G110100.2 | chr6A | 79267605–79271671 | 4 | 475 | Identified in this study | V | |
| TaSBP9-BSa | TraesCS6B02G138400.1 | chr6B | 135858620–135862931 | 4 | 1422 | 473 | TaBSPL3/18# | V |
| TaSBP9-DSa | TraesCS6D02G098500.1 | chr6D | 62113576–62118381 | 4 | 1422 | 473 | TaDSPL3/18# | V |
| TaSBP10-ASa | TraesCS6A02G152000.1 | chr6A | 136541404–136544531 | 3 | 1347 | 448 | Identified in this study | III |
| TaSBP10-BSa | TraesCS6B02G180300.1 | chr6B | 200508894–200512052 | 3 | 1329 | 442 | Identified in this study | III |
| TaSBP10-DSa | TraesCS6D02G142100.1 | chr6D | 111567310–111570133 | 3 | 1359 | 452 | Identified in this study | III |
| TaSBP11-ASa | TraesCS6A02G155300.1 | chr6A | 143965449–143969690 | 4 | 987 | 328 | Identified in this study | V |
| TaSBP11-BSa | TraesCS6B02G183400.1 | chr6B | 204923634–204927878 | 4 | 984 | 327 | Identified in this study | V |
| TaSBP11-DSa | TraesCS6D02G145200.1 | chr6D | 115545817–115550011 | 4 | 978 | 325 | Identified in this study | V |
| TaSBP12-ASa | TraesCS7A02G208000.1 | chr7A | 170629912–170634171 | 11 | 2541 | 846 | TaASPL1# | II |
| TaSBP12-BSa | TraesCS7B02G115200.1 | chr7B | 133742358–133748024 | 11 | 2538 | 845 | TaBSPL1# | II |
| TaSBP12-DSa | TraesCS7D02G210400.1 | chr7D | 168423642–168428099 | 11 | 2469 | 822 | TaDSPL1# | II |
| TaSBP12-DSb | TraesCS7D02G210400.2 | chr7D | 168423642–168428099 | 11 | 847 | Identified in this study | II | |
| TaSBP13-ASa | TraesCS7A02G246500.1 | chr7A | 225631628–225636261 | 3 | 1161 | 386 | Identified in this study | V |
| TaSBP13-BSa | TraesCS7B02G144900.1 | chr7B | 187777243–187781491 | 3 | 1161 | 386 | TaBSPL17# | V |
| TaSBP13-BSb | TraesCS7B02G144900.2 | chr7B | 187777243–187781786 | 3 | 385 | Identified in this study | V | |
| TaSBP13-DSa | TraesCS7D02G245200.1 | chr7D | 213786904–213791354 | 3 | 1155 | 384 | Identified in this study | V |
| TaSBP13-DSb | TraesCS7D02G245200.2 | chr7D | 213786904–213791354 | 3 | 385 | Identified in this study | V | |
| TaSBP14-ASa | TraesCS7A02G249100.2 | chr7A | 231544001–231549464 | 9 | 3387 | 898 | TaASPL15# | II |
| TaSBP14-ASb | TraesCS7A02G249100.3 | chr7A | 231544001–231549464 | 9 | 1113 | Identified in this study | II | |
| TaSBP14-ASc | TraesCS7A02G249100.4 | chr7A | 231544001–231549464 | 9 | 1123 | Identified in this study | II | |
| TaSBP14-BSa | TraesCS7B02G142200.1 | chr7B | 181033715–181039167 | 10 | 3390 | 1129 | TaBSPL15# | II |
| TaSBP14-BSb | TraesCS7B02G142200.2 | chr7B | 181033715–181039145 | 10 | 1124 | Identified in this study | II | |
| TaSBP14-DSa | TraesCS7D02G248000.1 | chr7D | 219291031–219296565 | 10 | 3390 | 1129 | TaDSPL15# | II |
| TaSBP14-DSb | TraesCS7D02G248000.2 | chr7D | 219291031–219296577 | 10 | 1114 | Identified in this study | II | |
| TaSBP14-DSc | TraesCS7D02G248000.3 | chr7D | 219291031–219296577 | 10 | 1124 | Identified in this study | II | |
| TaSBP15-ASa | TraesCS7A02G260500.1 | chr7A | 252715392–252720978 | 3 | 1224 | 407 | Identified in this study | I |
| TaSBP15-BSa | TraesCS7B02G158500.1 | chr7B | 214070642–214075881 | 3 | 1230 | 409 | Identified in this study | I |
| TaSBP15-DSa | TraesCS7D02G261500.1 | chr7D | 237410146–237416092 | 3 | 1245 | 414 | Identified in this study | I |
| TaSBP16-ALa | TraesCS7A02G494800.1 | chr7A | 684292548–684297219 | 3 | 1200 | 399 | Identified in this study | III |
| TaSBP17-ALa | TraesCS7A02G494900.1 | chr7A | 685090082–685095954 | 3 | 1185 | 394 | Identified in this study | III |
| TaSBP18-ALa | TraesCS7A02G495000.1 | chr7A | 685212558–685214906 | 3 | 1221 | 388 | TaASPL4/10# | III |
| TaSBP19-ALa | TraesCS7A02G495100.1 | chr7A | 685227875–685230770 | 3 | 1260 | 419 | Identified in this study | III |
| TaSBP19-ALb | TraesCS7A02G495100.2 | chr7A | 685227875–685230770 | 3 | 418 | Identified in this study | III | |
| TaSBP19-BLa | TraesCS7B02G402200.1 | chr7B | 668907244–668914987 | 3 | 1254 | 417 | Identified in this study | III |
| TaSBP19-DLa | TraesCS7D02G482500.1 | chr7D | 592857673–592860198 | 3 | 1251 | 317 | Identified in this study | III |
| TaSBP20-ALa | TraesCS7B02G402300.1 | chr7A | 668928550–668930740 | 3 | 1173 | 406 | Identified in this study | III |
| TaSBP21-BLa | TraesCS7B02G402400.1 | chr7B | 669139219–669141554 | 3 | 1206 | 401 | TaBSPL10# | III |
| TaSBP22-DLa | TraesCS7D02G482200.1 | chr7D | 592632237–592634499 | 3 | 1227 | 408 | TaDSPL5# | III |
| TaSBP23-DLa | TraesCS7D02G482300.1 | chr7D | 592677316–592679805 | 3 | 954 | 390 | Identified in this study | III |
| TaSBP24-DLa | TraesCS7D02G482400.1 | chr7D | 592816284–592819509 | 3 | 1167 | 394 | Identified in this study | III |
| Gene Name | Accession Numbers | Chr | Location | Exo | CDS | Pro | Source of Information | Group |
|---|---|---|---|---|---|---|---|---|
| TuSBP1-ALa | TuG1812G0100002957.01.T01 | chr1A | 447915501–447927189 | 10 | 2580 | 859 | Identified in this study | II |
| TuSBP2-ALa | TuG1812G0200002854.01.T01 | chr2A | 357384527–357387361 | 2 | 579 | 192 | Identified in this study | IV |
| TuSBP3-ALa | TuG1812G0200004639.01.T01 | chr2A | 647145178–647149050 | 4 | 801 | 267 | Identified in this study | IV |
| TuSBP4-ALa | TuG1812G0200005460.01.T01 | chr2A | 707789017–707792687 | 3 | 1239 | 412 | Identified in this study | III |
| TuSBP5-ALa | TuG1812G0300004828.01.T01 | chr3A | 676882033–676886494 | 3 | 1248 | 415 | Identified in this study | I |
| TuSBP6-ALa | TuG1812G0400000984.01.T01 | chr7A | 671389265–671391985 | 3 | 1257 | 418 | Identified in this study | III |
| TuSBP7-ALa | TuG1812G0500002978.01.T01 | chr7A | 455468876–455471920 | 3 | 1233 | 410 | Identified in this study | V |
| TuSBP8-ALa | TuG1812G0500003204.01.T01 | chr5A | 474168850–474174870 | 3 | 1311 | 436 | Identified in this study | I |
| TuSBP9-ASa | TuG1812G0600001583.01.T01 | chr6A | 130811290–130814169 | 3 | 1347 | 448 | Identified in this study | III |
| TuSBP10-ALa | TuG1812G0600001622.01.T01 | chr6A | 136757763–136762113 | 4 | 987 | 328 | Identified in this study | V |
| TuSBP11-ALa | TuG1812G0700002205.01.T01 | chr7A | 166227361–166231653 | 9 | 1785 | 480 | Identified in this study | II |
| TuSBP12-ALa | TuG1812G0700002579.01.T01 | chr7A | 221012709–221017378 | 3 | 1161 | 386 | Identified in this study | V |
| TuSBP13-ALa | TuG1812G0700002627.01.T01 | chr7A | 229641959–229647338 | 10 | 3387 | 1128 | Identified in this study | II |
| TuSBP14-ALa | TuG1812G0700005298.01.T01 | chr7A | 666539171–666540579 | 2 | 300 | 100 | Identified in this study | III |
| TuSBP15-ALa | TuG1812G0700005339.01.T01 | chr7A | 671400175–67140236 | 3 | 1221 | 406 | Identified in this study | III |
| TuSBP16-ALa | TuG1812G0700005341.01.T01 | chr7A | 671555473–671557441 | 3 | 1221 | 407 | Identified in this study | III |
| TuSBP17-ALa | TuG1812S0001265400.01.T01 | chrUn | 1790–6661 | 3 | 804 | 267 | Identified in this study | V |
| TdSBP1-ALa | TRIDC1AG038160.1 | chr1A | 449905748–449917720 | 10 | 2580 | 856 | Identified in this study | II |
| TdSBP2-BLa | TRIDC1BG043410.1 | chr1B | 472706171–472709916 | 3 | 1779 | 860 | Identified in this study | II |
| TdSBP3-ASa | TRIDC2AG030380.1 | chr2A | 238282797–238283178 | 1 | 382 | 192 | Identified in this study | IV |
| TdSBP4-ALa | TRIDC2AG059730.1 | chr2A | 663899405–663904029 | 3 | 784 | 349 | Identified in this study | IV |
| TdSBP5-ALa | TRIDC2AG070620.1 | chr2A | 723682186–723686050 | 3 | 1149 | 382 | Identified in this study | III |
| TdSBP6-BSa | TRIDC2BG034150.1 | chr2B | 269177236–269179967 | 2 | 1087 | 192 | Identified in this study | IV |
| TdSBP7-BLa | TRIDC2BG063080.1 | chr2B | 618377307–618384575 | 3 | 799 | 249 | Identified in this study | IV |
| TdSBP8-BLa | TRIDC2BG076330.1 | chr2B | 721397248–72139797 | 1 | 731 | 318 | Identified in this study | III |
| TdSBP9-ALa | TRIDC3AG061260.1 | chr3A | 669762928–669765360 | 3 | 585 | 195 | Identified in this study | I |
| TdSBP10-BLa | TRIDC3BG068940.1 | chr3B | 726994175–726997469 | 4 | 1092 | 363 | Identified in this study | I |
| TdSBP11-ALa | TRIDC4AG053710.1 | chr4A | 623528485–623534548 | 11 | 3298 | 962 | Identified in this study | II |
| TdSBP12-ASa | TRIDC5AG019810.1 | chr5A | 241751264–241754274 | 3 | 1218 | 406 | Identified in this study | V |
| TdSBP13-ALa | TRIDC5AG042970.1 | chr5A | 489480670–489485837 | 3 | 1280 | 429 | Identified in this study | I |
| TdSBP14-BLa | TRIDC5BG043980.1 | chr5B | 457191664–457194525 | 3 | 1182 | 393 | Identified in this study | V |
| TdSBP15-BLa | TRIDC5BG076500.1 | chr5B | 679130247–679136436 | 6 | 3342 | 494 | Identified in this study | II |
| TdSBP16-ASa | TRIDC6AG014380.1 | chr6A | 78107952–78114376 | 4 | 1842 | 479 | Identified in this study | V |
| TdSBP17-ASa | TRIDC6AG021720.1 | chr6A | 142480257–142483359 | 3 | 864 | 264 | Identified in this study | V |
| TdSBP18-BSa | TRIDC6BG019820.1 | chr6B | 138664345–138668705 | 4 | 2347 | 473 | Identified in this study | V |
| TdSBP19-BSa | TRIDC6BG027420.1 | chr6B | 210397416–210398530 | 1 | 1115 | 255 | Identified in this study | V |
| TdSBP20-ASa | TRIDC7AG026190.1 | chr7A | 168308982–168313393 | 4 | 3421 | 294 | Identified in this study | II |
| TdSBP21-ASa | TRIDC7AG031600.1 | chr7A | 223119140–223123491 | 3 | 1186 | 288 | Identified in this study | V |
| TdSBP22-ASa | TRIDC7AG032020.2 | chr7A | 229102299–229106659 | 10 | 3315 | 1104 | Identified in this study | II |
| TdSBP23-ALa | TRIDC7AG069100.1 | chr7A | 680628704–680630730 | 3 | 789 | 217 | Identified in this study | III |
| TdSBP24-ALa | TRIDC7AG069150.1 | chr7A | 681544947–681546745 | 3 | 999 | 266 | Identified in this study | III |
| TdSBP25-ALa | TRIDC7AG069170.1 | chr7A | 681560236–681560880 | 3 | 447 | 134 | Identified in this study | III |
| TdSBP26-BSa | TRIDC7BG016800.1 | chr7B | 138211281–138211703 | 1 | 423 | 845 | Identified in this study | II |
| TdSBP27-BSa | TRIDC7BG022390.1 | chr7B | 197125736–197130581 | 10 | 3875 | 1075 | Identified in this study | II |
| TdSBP28-BSa | TRIDC7BG022920.1 | chr7B | 203851538–203855411 | 3 | 1220 | 386 | Identified in this study | V |
| TdSBP29-BSa | TRIDC7BG025060.1 | chr7B | 230000670–230005489 | 3 | 1230 | 409 | Identified in this study | I |
| TdSBP30-BLa | TRIDC7BG063530.1 | chr7B | 677708807–677710254 | 3 | 708 | 188 | Identified in this study | III |
| TdSBP31-BLa | TRIDC7BG063560.1 | chr7B | 677720377–677722485 | 3 | 1005 | 227 | Identified in this study | III |
| AeSBP1-DLa | AET1Gv20619300.1 | chr1D | 352650035–352661850 | 10 | 2580 | 859 | Identified in this study | II |
| AeSBP2-DSa | AET2Gv20486300.1 | chr2D | 208545838–208549953 | 2 | 579 | 192 | Identified in this study | IV |
| AeSBP3-DLa | AET2Gv20915400.2 | chr2D | 524298824–524304038 | 3 | 1068 | 355 | Identified in this study | IV |
| AeSBP4-DLa | AET2Gv21102800.1 | chr2D | 595292761–595296371 | 3 | 894 | 297 | Identified in this study | III |
| AeSBP5-DLa | AET3Gv20960100.2 | chr3D | 547095656–547100533 | 3 | 1260 | 419 | Identified in this study | I |
| AeSBP6-DLa | AET4Gv20824900.1 | chr4D | 510058422–510070480 | 3 | 1185 | 394 | Identified in this study | I |
| AeSBP7-DLa | AET5Gv20667300.1 | chr5D | 398586010–398591342 | 3 | 1281 | 426 | Identified in this study | I |
| AeSBP8-DLa | AET5Gv21144000.1 | chr5D | 549430767–549437076 | 6 | 1485 | 494 | Identified in this study | II |
| AeSBP9-DSa | AET6Gv20284700.1 | chr6D | 86357505–86362787 | 4 | 1386 | 462 | Identified in this study | V |
| AeSBP10-DSa | AET6Gv20390500.2 | chr6D | 86362787–86362787 | 3 | 1359 | 452 | Identified in this study | III |
| AeSBP11-DSa | AET6Gv20396500.3 | chr6D | 139561288–139565520 | 4 | 978 | 325 | Identified in this study | V |
| AeSBP12-DSa | AET7Gv20522500.3 | chr7D | 169524973–169529438 | 11 | 2544 | 847 | Identified in this study | II |
| AeSBP13-DSa | AET7Gv20612200.1 | chr7D | 221003305–221008896 | 10 | 3390 | 1129 | Identified in this study | II |
| AeSBP14-DSa | AET7Gv20637900.1 | chr7D | 239178640–239184595 | 3 | 1245 | 414 | Identified in this study | I |
| AeSBP15-DLa | AET7Gv21205900.1 | chr7D | 598534268–598537267 | 3 | 1230 | 396 | Identified in this study | III |
| AeSBP16-DLa | AET7Gv21206500.1 | chr7D | 598574981–598577665 | 3 | 1251 | 416 | Identified in this study | III |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, J.; Ma, D.; Yin, J.; Yang, L.; He, Y.; Zhu, Z.; Tong, H.; Chen, L.; Zhu, G.; Liu, Y.; et al. Genome-Wide Characterization and Expression Profiling of Squamosa Promoter Binding Protein-Like (SBP) Transcription Factors in Wheat (Triticum aestivum L.). Agronomy 2019, 9, 527. https://doi.org/10.3390/agronomy9090527
Song J, Ma D, Yin J, Yang L, He Y, Zhu Z, Tong H, Chen L, Zhu G, Liu Y, et al. Genome-Wide Characterization and Expression Profiling of Squamosa Promoter Binding Protein-Like (SBP) Transcription Factors in Wheat (Triticum aestivum L.). Agronomy. 2019; 9(9):527. https://doi.org/10.3390/agronomy9090527
Chicago/Turabian StyleSong, Jinghan, Dongfang Ma, Junliang Yin, Lei Yang, Yiqin He, Zhanwang Zhu, Hanwen Tong, Lin Chen, Guang Zhu, Yike Liu, and et al. 2019. "Genome-Wide Characterization and Expression Profiling of Squamosa Promoter Binding Protein-Like (SBP) Transcription Factors in Wheat (Triticum aestivum L.)" Agronomy 9, no. 9: 527. https://doi.org/10.3390/agronomy9090527






