Diversity of Colletotrichum Species Associated with Olive Anthracnose and New Perspectives on Controlling the Disease in Portugal
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Sample Collection
2.2. Isolation of Colletotrichum spp.
2.3. Fungal DNA Extraction and Identification
2.4. Multivariate Data Analysis
3. Results
3.1. Isolation, Amplification, and Identification of Colletotrichum spp.
3.2. Colletotrichum spp. in Olive Cultivars, Sites, and Plant Organs
3.3. Multivariate Data Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Graniti, A.; Frisullo, S.; Pennisi, A.; Magnano di San Lio, G. Infections of Glomerella cingulata on olive in Italy. EPPO Bulletin 1993, 23, 457–465. [Google Scholar] [CrossRef]
- Talhinhas, P.; Sreenivasaprasad, S.; Neves-Martins, J.; Oliveira, H. Molecular and phenotypic analyses reveal the association of diverse Colletotrichum acutatum groups and a low level of C. gloeosporioides with olive anthracnose. Appl. Environ. Microbiol. 2005, 71, 2987–2998. [Google Scholar] [CrossRef] [PubMed]
- Moral, J.; Bouhmidi, K.; Trapero, A. Influence of fruit maturity, cultivar susceptibility, and inoculation method on infection of olive fruit by Colletotrichum acutatum. Plant disease 2008, 92, 1421–1426. [Google Scholar] [CrossRef]
- Cacciola, S.; Faedda, O.; Sinatra, R.; Agosteo, F.; Schena, G.; Frisullo, L.; di San, S.M.; Lio, G. Olive anthracnose. J. Plant Pathol. 2012, 94, 29–44. [Google Scholar]
- Mosca, S.; Li Destri Nicosia, M.; Cacciola, S.; Schena, L. Molecular Analysis of Colletotrichum Species in the Carposphere and Phyllosphere of Olive. PLoS ONE 2014, 9, e114031. [Google Scholar] [CrossRef] [PubMed]
- Talhinhas, P.; Gonçalves, E.; Sreenivasaprasad, S.; Oliveira, H. Virulence diversity of anthracnose pathogens (Colletotrichum acutatum and C. gloeosporioides species complexes) on eight olive cultivars commonly grown in Portugal. Eur. J. Plant Pathol. 2015, 142, 73–83. [Google Scholar] [CrossRef]
- Moral, J.; Xaviér, C.; Roca, L.; Romero, J.; Moreda, W.; Trapero, A. La Antracnosis del olivo y su efecto en la calidad del aceite. Grasas y Aceites 2014, 65, e028. [Google Scholar] [CrossRef]
- Jeffries, P.; Dodd, J.; Jeger, M.; Plumbley, R. The biology and control of Colletotrichum species on tropical fruit crops. Plant Pathol. 1990, 39, 343–366. [Google Scholar] [CrossRef]
- Moral, J.; Trapero, A. Assessing the susceptibility of olive cultivars to anthracnose caused by Colletotrichum acutatum. Plant Disease 2009, 93, 1028–1036. [Google Scholar] [CrossRef]
- Talhinhas, P.; Neves-Martins, J.; Oliveira, H.; Sreenivasaprasad, S. The distinctive population structure of Colletotrichum species associated with olive anthracnose in the Algarve region of Portugal reflects a host–pathogen diversity hot spot. FEMS Microbiol. Lett. 2009, 296, 31–38. [Google Scholar] [CrossRef]
- Gomes, S.; Martins-Lopes, P.; Lopes, L.; Guedes-Pinto, H. Assessing genetic diversity in Olea europaea L. using ISSR and SSR markers. Plant Mol. Biol. Rep. 2009, 123, 82–89. [Google Scholar] [CrossRef]
- Martelli, G. A first contribution to the knowledge of the biology of Gloeosporium olivarum. Phytopathol. Mediterr. 1960, 1, 101–112. [Google Scholar]
- Trapero, A.; Blanco-López, M. Enfermedades. In El cultivo de Olivo; Barranco, D., Fernández-Escobar, R., Rallo, L., Eds.; Junta de Andalucía/Mundi-Prensa: Madrid, Spain, 2008; pp. 57–614. [Google Scholar]
- Moral, J.; Oliveira, R.; Trapero, A. Elucidation of the Disease Cycle of Olive Anthracnose caused by Colletotrichum acutatum. Phytopathology 2009, 99, 548–556. [Google Scholar] [CrossRef]
- Sergeeva, V. Anthracnose in olives symptoms disease cycle and management. In Proceedings of the 4th Olivebioteq International Conference for olive products, Crete, Greece, 31 October–4 November 2011; pp. 269–274. [Google Scholar]
- Doyle, J.; Doyle, J. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Varanda, C.M.R.; Oliveira, M.; Materatski, P.; Landum, M.; Clara, M.I.E.; Félix, M.R. Fungal endophytic communities associated to phyllospheres of grapevine cultivars under different types of management. Fungal Biol. 2016, 1–12. [Google Scholar] [CrossRef] [PubMed]
- White, T.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- O’Donnell, K.; Cigelnik, E. Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Mol. Phylogenet. Evol. 1997, 7, 103–116. [Google Scholar] [CrossRef]
- Damm, U.; Cannon, P.; Woudenberg, J.; Crous, P. The Colletotrichum acutatum species complex. Stud. Mycol. 2012, 73, 37–113. [Google Scholar] [CrossRef]
- Guerber, J.; Liu, B.; Correll, J.; Johnston, P. Characterization of diversity in Colletotrichum acutatum sensu lato by sequence analysis of two gene introns, mtDNA and intron RFLPs, and mating compatibility. Mycologia 2003, 95, 872–895. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0. Mol. Biol. Evol. 2015, 33, 1870–1874. [Google Scholar] [CrossRef]
- Clarke, K.R.; Warwick, R.M. Changes in Marine Communities: An Approach to Statistical Analysis and Interpretation, 2nd ed.; Plymouth Marine Laboratory: Plymouth, UK, 2001. [Google Scholar]
- Anderson, M.J.; Gorley, R.N.; Clarke, K.R. PERMANOVA A+ for PRIMER: Guide to Software and Statistical Methods; PRIMER-E: Plymouth, UK, 2008. [Google Scholar]
- Clarke, K.; Green, R. Statistical design and analysis for a biological effects study. Mar. Ecol. Prog. Ser. 1988, 46, 213–226. [Google Scholar] [CrossRef]
- Agosteo, G.; Magnano di San Lio, G.; Cacciola, S.; Frisullo, S. Characterisation of the causal agent of olive anthracnose in Southern Italy. Acta Horticulturae 2002, 586, 713–716. [Google Scholar] [CrossRef]
- Chattaoui, M.; Raya, M.; Bouri, M.; Moral, J.; Perez-Rodriguez, M.; Trapero, A.; Msallem, M.; Rhouma, A. Characterization of a Colletotrichum population causing anthracnose disease on Olive in northern Tunisia. J. Appl. Microbiol. 2016, 120, 1364–1381. [Google Scholar] [CrossRef] [PubMed]
- Sutton, B. The genus Glomerella and its anamorph Colletotrichum. In Colletotrichum Biology, Pathology and Control; Bailey, J., Jeger, M., Eds.; CAB International: Wallingford, UK, 1992; pp. 1–26. [Google Scholar]
- Talhinhas, P.; Sreenivasaprasad, S.; Neves-Martins, J.; Oliveira, H. Genetic and morphological characterization of Colletotrichum acutatum causing anthracnose of lupins. Phytopathology 2002, 92, 986–996. [Google Scholar] [CrossRef] [PubMed]
- Schenaa, L.; Moscaa, S.; Cacciola, S.; Faeddab, R.; Sanzanic, S.; Agosteo, G.; Sergeevad, V.; Magnano di San Lio, G. Species of the Colletotrichum gloeosporioides and C. boninense complexes associated with olive anthracnose. Plant Pathol. 2014, 63, 437–446. [Google Scholar] [CrossRef]
- Faedda, R.; Agosteo, G.; Schena, L.; Mosca, S.; Frisullo, S.; Maganano Di San Lio, G.; Cacciola, S. Colletotrichum clavatum sp. nov. identified as the causal agent of olive anthracnose in Italy. Phytopathol. Mediterr. 2011, 50, 283–302. [Google Scholar]
- Cacciola, S.; Pane, A.; Agosteo, G.; Magnano di San Lio, G. Osservazioni sull’epidemiologia dell’anthracnosi dell’olivo in Calabria. Inf. Fitopatol. 1996, 6, 27–32. [Google Scholar]
- Margarita, L.; Porta-Puglia, A.; Quacuarelli, A. China—Colletotrichum acutatum on olive trees. FAO Plant Prot. Bull. 1988, 36, 185–186. [Google Scholar]
- Barranco, D. Variedades y patrones. In El Cultivo de Olivo; Barranco, D., Fernández-Escobar, R., Rallo, L., Eds.; Coedición Junta de Andalucía and Mundi-Prensa: Madrid, Spain, 2008; pp. 65–92. [Google Scholar]
- Benlloch, M. Observaciones sobre algunas enfermedades del olivo. Boletin de Patologia Vegetal y Entomologia Agricola 1942, 11, 1–12. [Google Scholar]
- de Andrés, F. Enfermedades y Plagas del Olivo; Riquelme y Vargas Ediciones S.L.: Jaén, Spain, 1991. [Google Scholar]
- Freeman, S.; Horowitz, S.; Sharon, A. Pathogenic and nonpathogenic lifestyles in Colletotrichum acutatum from strawberry and other plants. Phytopathology 2001, 91, 986–992. [Google Scholar] [CrossRef]
- Mateo-Sagasta, E. Estudios básicos sobre Gloeosporium olivarum Alm. (Deuteromiceto Melanconial). Bol. Patol. Veg. Entomol. Agric. 1968, 30, 31–135. [Google Scholar]
- Gomes, S.; Bacelar, E.; Martins-Lopes, P.; Carvalho, T.; Guedes-Pinto, H. Infection Process of Olive Fruits by Colletotrichum acutatum and the Protective Role of the Cuticle and Epidermis. J. Agric. Sci. 2012, 4, 101–110. [Google Scholar] [CrossRef]

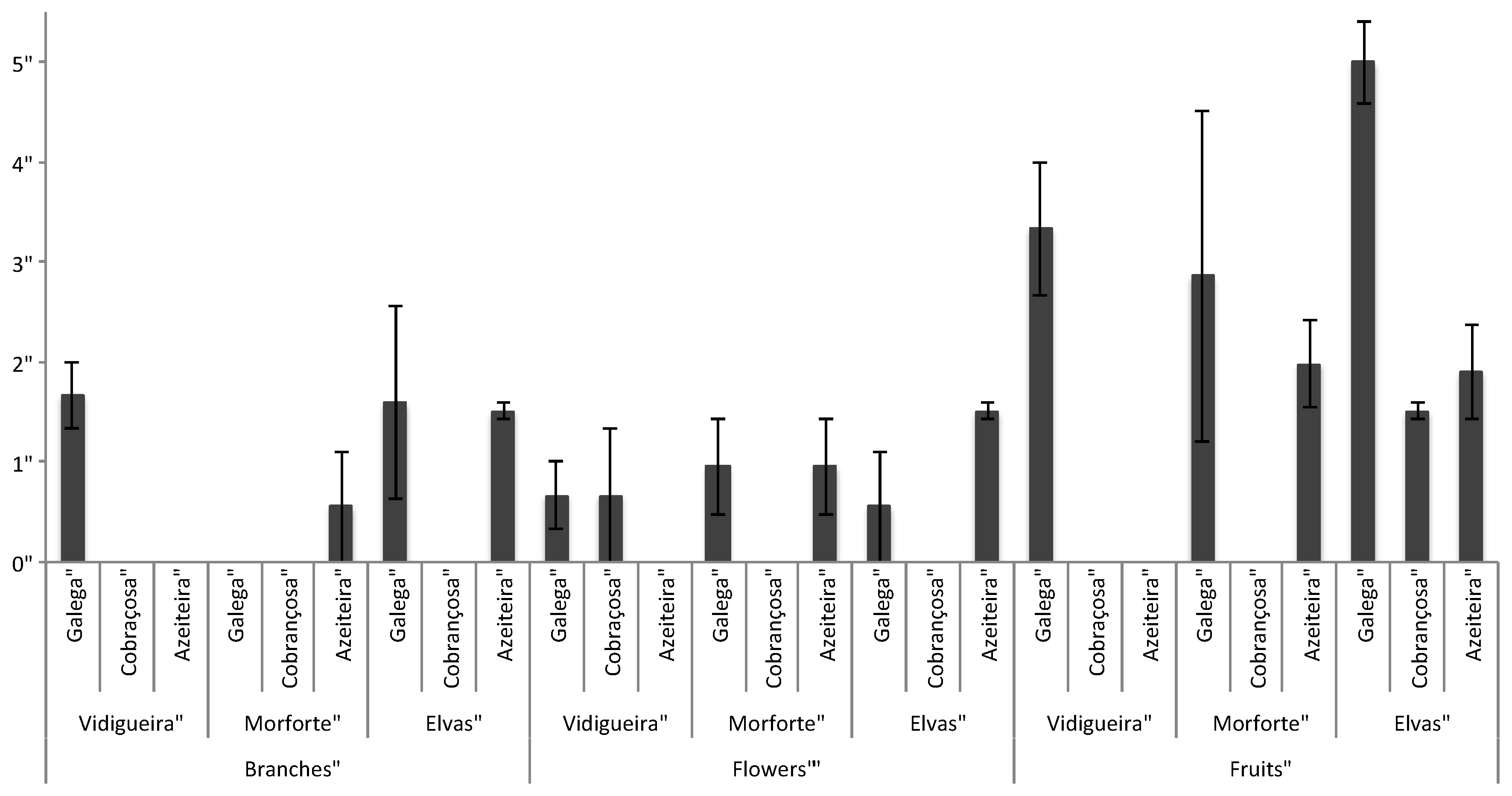
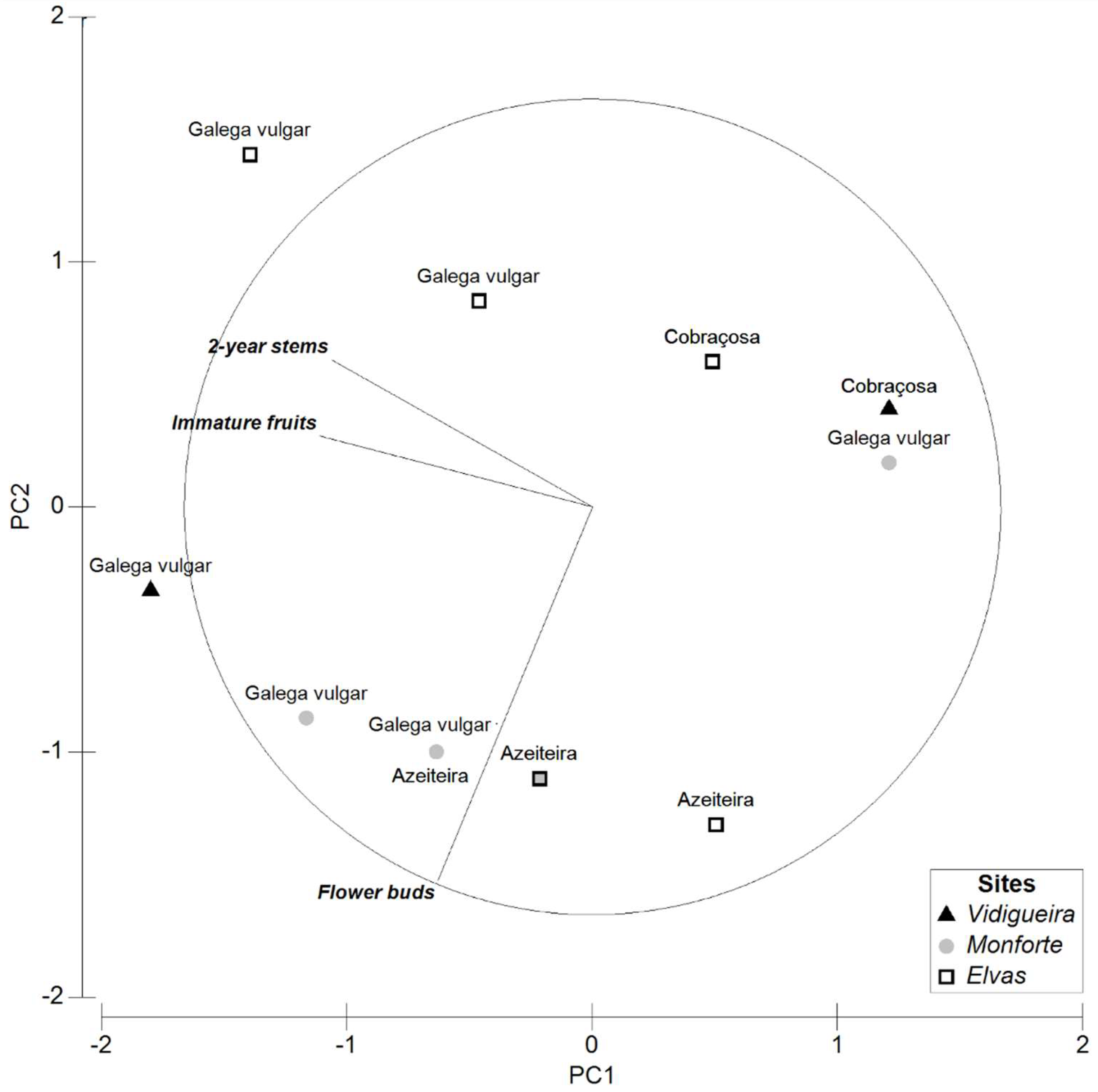
| Sites | Cultivars | 2-Year Stems | Flower Buds | Immature Fruits |
|---|---|---|---|---|
| Vidigueira | Galega vulgar | - | - | C. nymphaeae 1 and 3 |
| C. nymphaeae 1 and 3 | - | - | ||
| - | C. goditiae | C. goditiae | ||
| C. nymphaeae 1 and 2 | - | C. nymphaeae 1 and 2 | ||
| - | - | C. nymphaeae 1 | ||
| C. nymphaeae 1 | - | C. nymphaeae 1 | ||
| - | C. nymphaeae 1 | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 and 3 | ||
| - | - | C. nymphaeae 1 | ||
| C. nymphaeae 1 | - | - | ||
| C. nymphaeae 1 | - | - | ||
| - | - | C. nymphaeae 1 | ||
| Cobrançosa | - | C. goditiae | - | |
| - | C. nymphaeae 1 | - | ||
| Monforte | Galega vulgar | - | - | C. nymphaeae 1 |
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| - | C. nymphaeae 1 and 3 | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| - | C. nymphaeae 1 and 3 | - | ||
| - | - | C. nymphaeae 1 | ||
| Azeiteira | - | C. nymphaeae 1 | - | |
| - | - | C. nymphaeae 2 | ||
| - | C. nymphaeae 1 | - | ||
| - | - | C. nymphaeae 1 and 3 | ||
| Elvas | Galega vulgar | - | - | C. nymphaeae 1 and 3 |
| - | - | C. nymphaeae 1 | ||
| C. nymphaeae 1 | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 and 3 | ||
| - | - | C. nymphaeae 1 and 3 | ||
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 1 | ||
| C. nymphaeae 1 | - | - | ||
| - | - | C. nymphaeae 1 and 3 | ||
| C. nymphaeae 2 | C. nymphaeae 2 | C. nymphaeae 2 | ||
| - | - | C. nymphaeae 1 | ||
| Cobrançosa | - | - | C. nymphaeae 1 and 3 | |
| - | - | C. nymphaeae 1 | ||
| - | - | C. nymphaeae 2 | ||
| Azeiteira | - | C. nymphaeae 1 | - | |
| - | C. nymphaeae 2 | - | ||
| - | - | C. nymphaeae 1 and 3 | ||
| - | C. nymphaeae 1 | - | ||
| Total isolates (68) | 10 | 14 | 44 | |
| Site | Vidigueira 16.7% | ||||||||
| Cultivar | Galega Vulgar | Cobrançosa | Azeiteira | ||||||
| Infected trees | 43.3% | 6.7% | 0% | ||||||
| 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | |
| Infected organ | 10.0% | 10.0% | 24.6% | 0% | 10.0% | 0% | 0% | 0% | 0% |
| Site | Monforte 14.4% | ||||||||
| Cultivar | Galega Vulgar | Cobrançosa | Azeiteira | ||||||
| Infected trees | 30.0% | 0% | 13.3% | ||||||
| 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | |
| Infected organ | 0% | 14.0% | 48.5% | 0% | 0% | 0% | 0% | 15.0% | 28.0% |
| Site | Elvas 20.0% | ||||||||
| Cultivar | Galega Vulgar | Cobrançosa | Azeiteira | ||||||
| Infected trees | 36.7% | 10.0% | 13.3% | ||||||
| 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | 2-Year Stems | Flower Buds | Immature Fruits | |
| Infected organ | 16.0% | 16.0% | 30.4% | 0% | 0% | 16.0% | 0% | 16.0% | 30.0% |
| a. | Similarity % | ||
| 46.8 | 63.3 | 56.1 | |
| Plant organ | Vidigueira | Monforte | Elvas |
| 2-Year Stems | 24.57 | 0.0 | 0.9 |
| Flower Buds | 26.28 | 30.34 | 9.65 |
| Immature Fruits | 49.15 | 69.66 | 89.45 |
| Dissimilarity % | |||
| 44.0 | 52.5 | 38.7 | |
| Plant organ | Vidigueira vs. Monforte | Vidigueira vs. Elvas | Monforte vs. Elvas |
| 2-Year Stems | 27.43 | 26.67 | 29.05 |
| Flower Buds | 21.28 | 25.54 | 20.7 |
| Immature Fruits | 51.29 | 47.79 | 50.24 |
| b. | Similarity % | ||
| 68.7 | 47.4 | 73.0 | |
| Plant organ | Galega | Cobrançosa | Azeiteira |
| 2-Year Stems | 11.3 | 0.0 | 0.0 |
| Flower Buds | 9.34 | 0.0 | 38.25 |
| Immature Fruits | 79.36 | 100.0 | 61.75 |
| Dissimilarity % | |||
| 64.4 | 43.3 | 43.5 | |
| Plant organ | Galega vs. Cobrançosa | Galega vs. Azeiteira | Cobrançosa vs. Azeiteira |
| 2-Year Stems | 22.4 | 26.93 | 12.88 |
| Flower Buds | 18.42 | 18 | 51.55 |
| Immature Fruits | 59.18 | 55.07 | 35.57 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Materatski, P.; Varanda, C.; Carvalho, T.; Dias, A.B.; Campos, M.D.; Rei, F.; Félix, M.d.R. Diversity of Colletotrichum Species Associated with Olive Anthracnose and New Perspectives on Controlling the Disease in Portugal. Agronomy 2018, 8, 301. https://doi.org/10.3390/agronomy8120301
Materatski P, Varanda C, Carvalho T, Dias AB, Campos MD, Rei F, Félix MdR. Diversity of Colletotrichum Species Associated with Olive Anthracnose and New Perspectives on Controlling the Disease in Portugal. Agronomy. 2018; 8(12):301. https://doi.org/10.3390/agronomy8120301
Chicago/Turabian StyleMateratski, Patrick, Carla Varanda, Teresa Carvalho, António Bento Dias, M. Doroteia Campos, Fernando Rei, and Maria do Rosário Félix. 2018. "Diversity of Colletotrichum Species Associated with Olive Anthracnose and New Perspectives on Controlling the Disease in Portugal" Agronomy 8, no. 12: 301. https://doi.org/10.3390/agronomy8120301
APA StyleMateratski, P., Varanda, C., Carvalho, T., Dias, A. B., Campos, M. D., Rei, F., & Félix, M. d. R. (2018). Diversity of Colletotrichum Species Associated with Olive Anthracnose and New Perspectives on Controlling the Disease in Portugal. Agronomy, 8(12), 301. https://doi.org/10.3390/agronomy8120301







