Morphological Assessment of Cultivated and Wild Amaranth Species Diversity
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Greenhouse Planting and Field Transplanting
2.3. Phenotyping and Morphological Evaluations
2.4. Data Analysis of Phenotypic Data
3. Results
3.1. Morphological Variability in the SSE Collection
3.2. Morphological Variability in the USDA Collection
3.3. Cluster Analysis of Full Collection from Both USDA and SSE
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Suresh, S.; Chung, J.W.; Cho, G.T.; Sung, J.S.; Park, J.H.; Gwag, J.G.; Baek, H.J. Analysis of molecular genetic diversity and population structure in Amaranthus germplasm using SSR markers. Plant Biosyst. Int. J. Deal. Asp. Plant Biol. 2014, 148, 635–644. [Google Scholar]
- Schmid, K.J.; Stetter, M.G. Analysis of phylogenetic relationships and genome size evolution of the Amaranthus genus using GBS indicates the ancestors of an ancient crop. Mol. Phylogenet. Evol. 2017, 109. [Google Scholar] [CrossRef]
- Anjali, K.; Joshi, A.; Maloo, S.R.; Sharma, R. Assessment of the morphological and molecular diversity in Amaranthus spp. Afr. J. Agric. Res. 2013, 8, 2307–2311. [Google Scholar]
- Sauer, J.D. The grain amaranths and their relatives: A revised taxonomic and geographic survey. Ann. Mo. Bot. Gard. 1967, 54, 103–137. [Google Scholar] [CrossRef]
- He, Q.; Park, Y.J. Evaluation of Genetic Structure of Amaranth Accessions from the United States. Weed Turfgrass Sci. 2013, 2, 230–235. [Google Scholar] [CrossRef]
- Costea, M.; Weaver, S.E.; Tardif, F.J. The biology of Canadian weeds. 130. Amaranthus retroflexus L., A. powellii S. Watson and A. hybridus L. Can. J. Plant Sci. 2004, 84, 631–668. [Google Scholar] [CrossRef]
- Coons, M.P. The status of Amaranthus in Ecuador. Ph.D. Thesis, Indiana University, Bloomington, IN, USA, 1975. [Google Scholar]
- Omondi, E.O.; Debener, T.; Linde, M.; Abukutsa-Onyango, M.; Dinssa, F.F.; Winkelmann, T. Molecular Markers for Genetic Diversity Studies in African Leafy Vegetables. Adv. Biosci. Biotechnol. 2016, 7, 188. [Google Scholar] [CrossRef]
- Erum, S.; Ambreen, F.; Naeemullah, M.; Masood, S.; Qayyum, A.; Rabbani, M.A. Genetic divergence in Amaranthus collected from Pakistan. J. Anim. Plant Sci. 2012, 22, 653–658. [Google Scholar]
- Brenner, D.M.; Baltensperger, D.D.; Kulakow, P.A.; Lehmann, J.W.; Myers, R.L.; Slabbert, M.M.; Sleugh, B.B. Genetic resources and breeding of Amaranthus. Plant Breed. Rev. 2010, 19, 227–285. [Google Scholar]
- Espitia Rangel, E.; Mapes Sánchez, C.; Escobedo Lopez, D.; de la O Olán, M.; Rivas Valencia, P.; Martínez Trejo, G.; Cortés Espinoza, L.; Hernández Casillas, J.M. Conservacion y Uso de los Recursos Geneticos de Amaranto en México; CINVESTAF, D.F.: Mexico City, Mexico, 2010. [Google Scholar]
- Gamel, T.H.; Linssen, J.P.; Mesallam, A.S.; Damir, A.A.; Shekib, L.A. Seed treatments affect functional and antinutritional properties of amaranth flours. J. Sci. Food Agric. 2006, 86, 1095–1102. [Google Scholar] [CrossRef]
- Espitia, E. Amaranth germplasm development and agronomic studies in Mexico. Food Rev. Int. 1992, 8, 71–86. [Google Scholar] [CrossRef]
- Bressani, R.; Gonzales, J.M.; Zuniga, J.; Breuner, M.; Elias, L.G. Yield, selected chemical composition and nutritive value of 14 selections of amaranth grain representing four species. J. Sci. Food Agric. 1987, 38, 347–356. [Google Scholar] [CrossRef]
- Dodok, L.; Modhir, A.A.; Buchtova, V.; Halasova, G.; Polaček, I. Importance and utilization of amaranth in food industry. Part 2. Composition of amino acids and fatty acids. Food/Nahrung 1997, 41, 108–110. [Google Scholar] [CrossRef]
- Das, S. Amaranthus: A Promising Crop of Future; Springer Science & Business Media: Singapore, 2016; ISBN 978-981-10-1469-7. [Google Scholar]
- Gonor, K.V.; Pogozheva, A.V.; Derbeneva, S.A.; Mal’tsev, G.; Trushina, E.N.; Mustafina, O.K. The influence of a diet with including amaranth oil on antioxidant and immune status in patients with ischemic heart disease and hyperlipoproteidemia. Vopr. Pitan. 2005, 75, 30–33. [Google Scholar]
- Brown, A.H.D. Core collections: A practical approach to genetic resources management. Genome 1989, 31, 818–824. [Google Scholar] [CrossRef]
- Fatinah, A.A.; Arumingtyas, E.L.; Mastuti, R. Morphological and genetic variation of Amaranthus spinosus L.: An adaptation evidence of climate differences and gene interaction. Int. J. Biosci. 2013, 3, 205–212. [Google Scholar]
- Khaing, A.A.; Moe, K.T.; Chung, J.W.; Baek, H.J.; Park, Y.J. Genetic diversity and population structure of the selected core set in Amaranthus using SSR markers. Plant Breed. 2013, 132, 165–173. [Google Scholar] [CrossRef]
- Sammour, R.H.; Radwan, S.A.; Mira, M. Genetic diversity in genus Amaranthus: From morphology to genomic DNA. Res. Rev. Biosci. 2012, 6, 351–360. [Google Scholar]
- Mallory, M.A.; Hall, R.V.; McNabb, A.R.; Pratt, D.B.; Jellen, E.N.; Maughan, P.J. Development and characterization of microsatellite markers for the grain amaranths. Crop Sci. 2008, 48, 1098–1106. [Google Scholar] [CrossRef]
- Mandal, N.; Das, P.K. Intra-and interspecific genetic diversity in grain Amaranthus using random amplified polymorphic DNA markers. Plant Tissue Cult. 2002, 12, 49–56. [Google Scholar]
- Maughan, P.J.; Smith, S.M.; Fairbanks, D.J.; Jellen, E.N. Development, characterization, and linkage mapping of single nucleotide polymorphisms in the grain amaranths (Amaranthus sp.). Plant Genome 2011, 4, 92–101. [Google Scholar] [CrossRef]
- Maughan, P.J.; Yourstone, S.M.; Jellen, E.N.; Udall, J.A. SNP discovery via genomic reduction, barcoding, and 454-pyrosequencing in amaranth. Plant Genome 2009, 2, 260–270. [Google Scholar] [CrossRef]
- Oo, W.H.; Park, Y.J. Analysis of the Genetic Diversity and Population Structure of Amaranth Accessions from South America Using 14 SSR Markers. Korean J. Crop Sci. 2013, 58, 336–346. [Google Scholar] [CrossRef]
- Popa, G.; Cornea, C.P.; Ciuca, M.; Babeanu, N.; Popa, O.; Marin, D. Studies on genetic diversity in Amaranthus species using the RAPD markers. Tom 2010, 17, 280–285. [Google Scholar]
- Ray, T.; Roy, S.C. Genetic diversity of Amaranthus species from the Indo-Gangetic Plains revealed by RAPD analysis leading to the development of ecotype-specific SCAR marker. J. Hered. 2009, 100, 338–347. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.; Pandey, S.; Kumar, J. A Comparative Study of Inter Simple Sequence Repeat (ISSR), Random Amplified Polymorphic DNA (RAPD) and Simple Sequence Repeat (SSR) Loci in Assessing Genetic Diversity in Amaranthus. Indian J. Genet. Plant Breed. 2013, 73, 411–418. [Google Scholar] [CrossRef]
- Xu, F.; Sun, M. Comparative analysis of phylogenetic relationships of grain amaranths and their wild relatives (Amaranthus; Amaranthaceae) using internal transcribed spacer, amplified fragment length polymorphism, and double-primer fluorescent intersimple sequence repeat markers. Mol. Phylogenet. Evol. 2001, 21, 372–387. [Google Scholar] [PubMed]
- Pratt, D.B.; Clark, L.G. Amaranthus rudis and A. tuberculatus, One Species or Two? J. Torrey Bot. Soc. 2001, 128, 282–296. [Google Scholar] [CrossRef]
- Shukla, S.; Bhargava, A.; Chatterjee, A.; Pandey, A.C.; Mishra, B.K. Diversity in phenotypic and nutritional traits in vegetable amaranth (Amaranthus tricolor), a nutritionally underutilized crop. J. Sci. Food Agric. 2010, 90, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Snezana, D.M.; Marija, K.; Danijela, R.; Milena, S.; Lidija, S. Assessment of genetic relatedness of the two Amaranthus retroflexus populations by protein and random amplified polymorphic DNA (RAPD) markers. Afr. J. Biotechnol. 2012, 11, 7331–7337. [Google Scholar]
- Tony-Odigie, A.E.; Adekoya, K.O.; Makinde, S.C.O.; Oboh, B.O.; Ogunkanmi, L.A.; Fowora, M.A. Assessment of Genetic Interspecies Relationships among Five Selected Amaranthus Species Using Phenotypic and RAPD Markers. Int. J. Bot. 2012, 8, 145. [Google Scholar]
- Transue, D.K.; Fairbanks, D.J.; Robison, L.R.; Andersen, W.R. Species identification by RAPD analysis of grain amaranth genetic resources. Crop Sci. 1994, 34, 1385–1389. [Google Scholar] [CrossRef]
- Wassom, J.J.; Tranel, P.J. Amplified fragment length polymorphism-based genetic relationships among weedy Amaranthus species. J. Hered. 2005, 96, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Wetzel, D.K.; Horak, M.J.; Skinner, D.Z. Use of PCR-based molecular markers to identify weedy Amaranthus species. Weed Sci. 1999, 47, 518–523. [Google Scholar]
- Zheleznov, A.V.; Solonenko, L.P.; Zheleznova, N.B. Seed proteins of the wild and the cultivated Amaranthus species. Euphytica 1997, 97, 177–182. [Google Scholar] [CrossRef]
- Yudina, R.S.; Zheleznova, N.B.; Zakharova, O.V.; Zheleznov, A.V.; Shumny, V.K. Isozyme analysis in a genetic collection of amaranths (Amaranthus L.). Russ. J. Genet. 2005, 41, 1395–1400. [Google Scholar] [CrossRef]
- Wu, X.; Blair, M.W. Genotyping by Sequencing (GBS) Polymorphism Diversity in Grain Amaranths and Relatives. Front. Plant Sci. 2017, 8, 1960. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Sun, M.; Yue, S.; Sun, H.; Cai, Y.; Huang, R.; Brenner, D.; Corke, H. Field evaluation of an Amaranthus genetic resource collection in China. Genet. Resour. Crop Evol. 2000, 47, 43–53. [Google Scholar] [CrossRef]
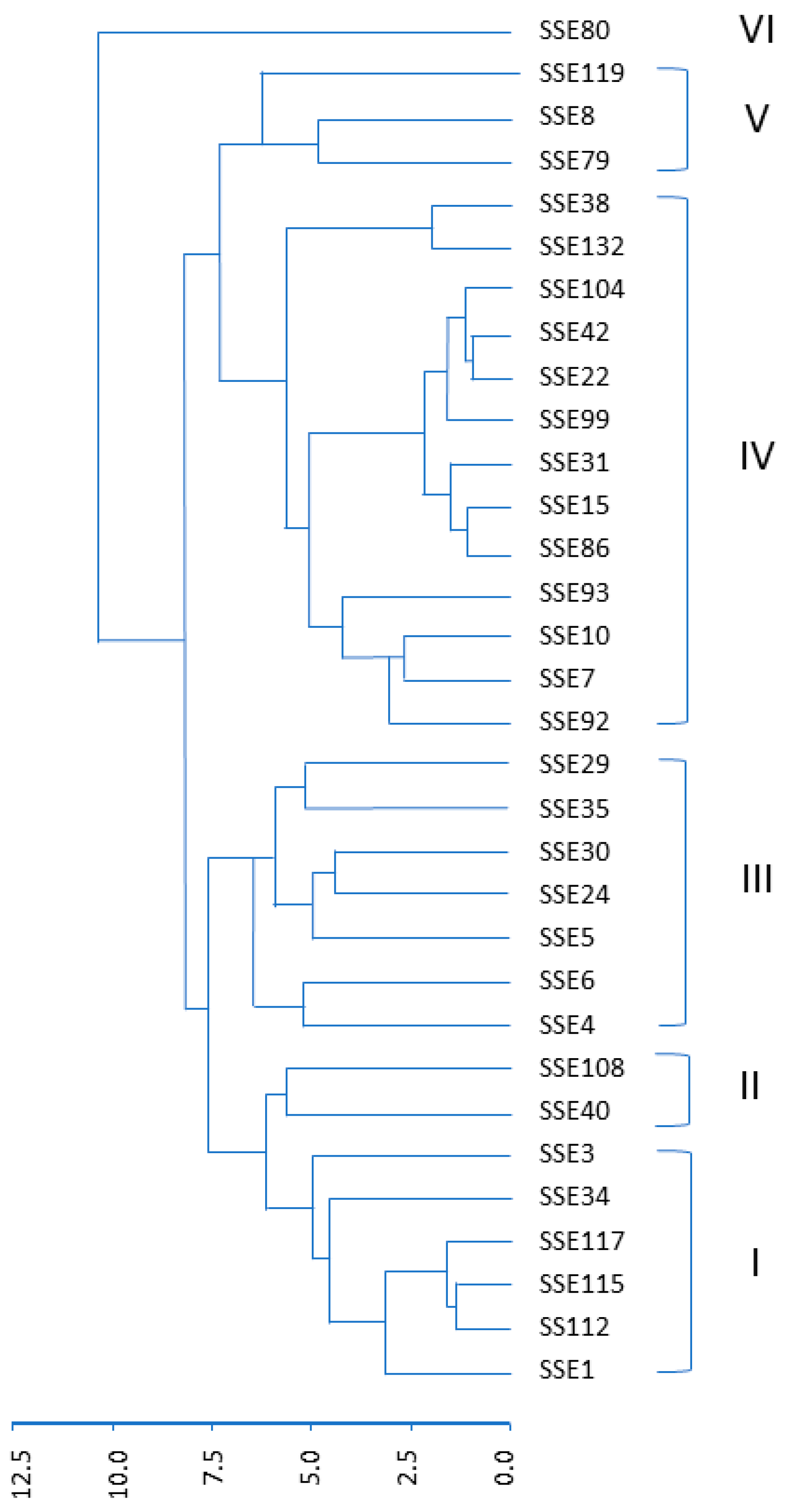
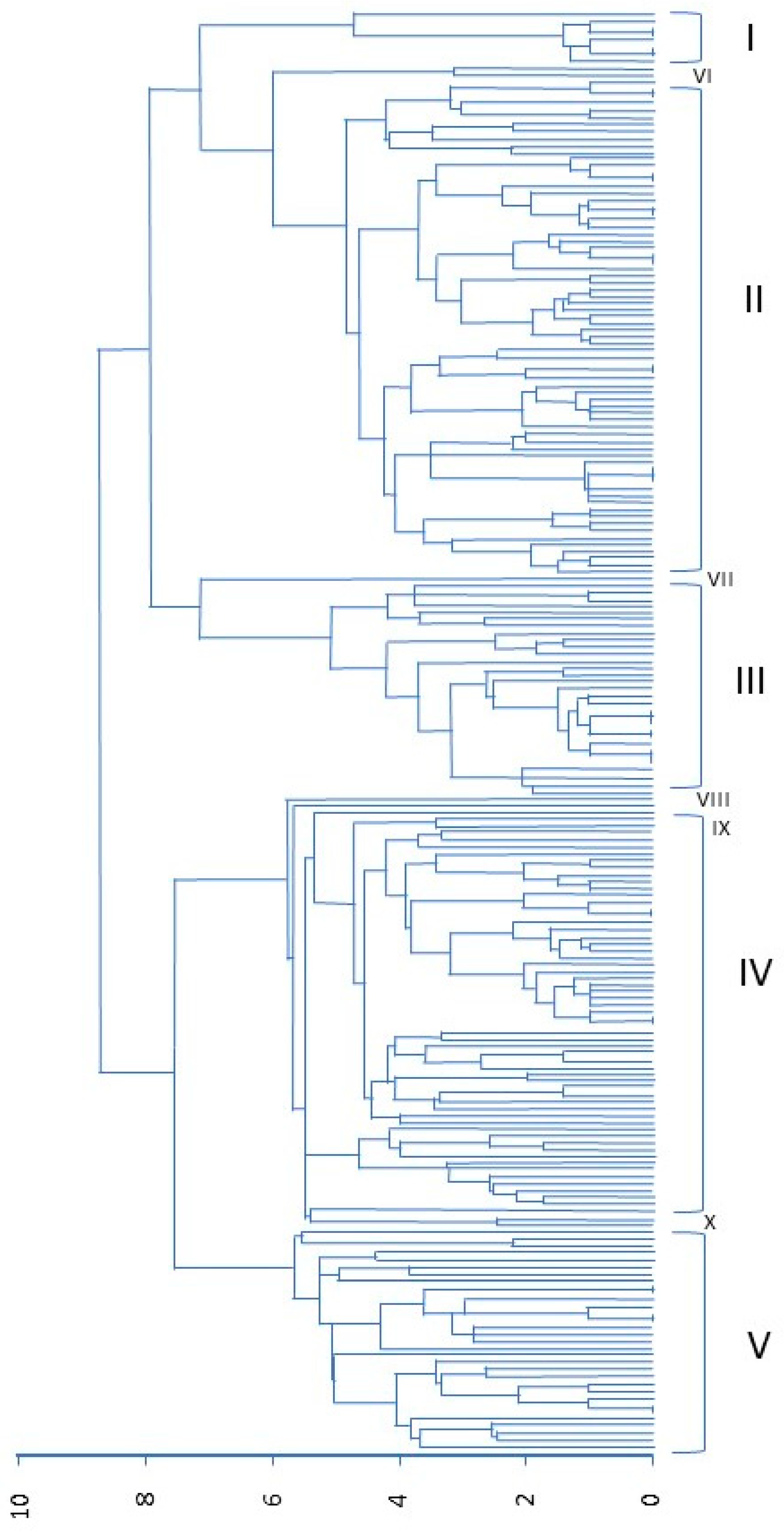
| Morphological Trait | Cluster MS | Error MS F Value | p > F |
|---|---|---|---|
| BP | 66.10 ** | 3.54 | <0.0001 |
| BS | 95.62 ** | 2.54 | <0.0001 |
| PP | 6.18 | 1.37 | 0.0041 |
| BI | 7.21 | 3.48 | 0.1008 |
| FC | 18.02 ** | 1.58 | <0.0001 |
| IS | 0.67 | 0.26 | 0.0539 |
| ID | 1.04 | 0.97 | 0.3981 |
| TIA | 0.49 | 0.4 | 0.3356 |
| SC | 11.35 | 2.83 | 0.0076 |
| Cluster | 1 | 2 | 3 | 4 | 5 | 6 |
|---|---|---|---|---|---|---|
| 1 | 9.521 | 11.47 | 8.485 | 7.553 | 10.41 | |
| 2 | 9.521 | 9.869 | 11.68 | 6.209 | 11.14 | |
| 3 | 11.47 | 9.869 | 5.566 | 8.762 | 13.33 | |
| 4 | 8.485 | 11.68 | 5.566 | 9.008 | 12.3 | |
| 5 | 7.553 | 6.209 | 8.762 | 9.008 | 9.936 | |
| 6 | 10.41 | 11.14 | 13.33 | 12.3 | 9.936 |
| Morphological Trait | Clusters | Error | p > F |
|---|---|---|---|
| BP | 436.21 ** | 0.98 | <0.0001 |
| BS | 297.44 ** | 2.47 | <0.0001 |
| PP | 5.52 ** | 0.88 | <0.0001 |
| BI | 7.41 | 4.27 | 0.0835 |
| FC | 17.029 ** | 3.21 | <0.0001 |
| IS | 0.188 | 0.23 | 0.6261 |
| ID | 3.39 | 1.38 | 0.0113 |
| TIA | 0.35 | 0.147 | 0.0138 |
| SC | 48.79 ** | 3.87 | <0.0001 |
| Cluster | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 9.631 | 12.7 | 5.28 | 13.19 | 6.902 | 9.26 | 9.302 | 6.598 | 10.42 | |
| 2 | 9.631 | 9.803 | 9.536 | 11.45 | 9.176 | 5.888 | 13 | 10.39 | 13.46 | |
| 3 | 12.7 | 9.803 | 13.04 | 5.222 | 7.766 | 10.32 | 8.176 | 13.7 | 9.306 | |
| 4 | 5.28 | 9.536 | 13.04 | 12.14 | 7.448 | 7.997 | 9.976 | 4.892 | 9.004 | |
| 5 | 13.19 | 11.45 | 5.222 | 12.14 | 9.221 | 9.269 | 8.262 | 12.78 | 7.748 | |
| 6 | 6.902 | 9.176 | 7.766 | 7.448 | 9.221 | 9.882 | 5.168 | 8.788 | 6.257 | |
| 7 | 9.26 | 5.888 | 10.32 | 7.997 | 9.269 | 9.882 | 12.24 | 9.075 | 11.95 | |
| 8 | 9.302 | 13 | 8.176 | 9.976 | 8.262 | 5.168 | 12.24 | 10.38 | 4.094 | |
| 9 | 6.598 | 10.39 | 13.7 | 4.892 | 12.78 | 8.788 | 9.075 | 10.38 | 10.28 | |
| 10 | 10.42 | 13.46 | 9.306 | 9.004 | 7.748 | 6.257 | 11.95 | 4.094 | 10.28 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thapa, R.; Blair, M.W. Morphological Assessment of Cultivated and Wild Amaranth Species Diversity. Agronomy 2018, 8, 272. https://doi.org/10.3390/agronomy8110272
Thapa R, Blair MW. Morphological Assessment of Cultivated and Wild Amaranth Species Diversity. Agronomy. 2018; 8(11):272. https://doi.org/10.3390/agronomy8110272
Chicago/Turabian StyleThapa, Ranjita, and Matthew W. Blair. 2018. "Morphological Assessment of Cultivated and Wild Amaranth Species Diversity" Agronomy 8, no. 11: 272. https://doi.org/10.3390/agronomy8110272
APA StyleThapa, R., & Blair, M. W. (2018). Morphological Assessment of Cultivated and Wild Amaranth Species Diversity. Agronomy, 8(11), 272. https://doi.org/10.3390/agronomy8110272





