Releasing the Full Potential of Cannabis through Biotechnology
Abstract
1. Introduction
2. The Advances of Cannabis Tissue Culture
3. The Genetic Improvement of Cannabis
4. Overcoming Limitations in Cannabis Gene Editing
4.1. Transformation Strategies
4.2. Target Tissue Selection
4.3. Gene Editing Strategies
4.4. Rapid Detection System for Editing Specificity
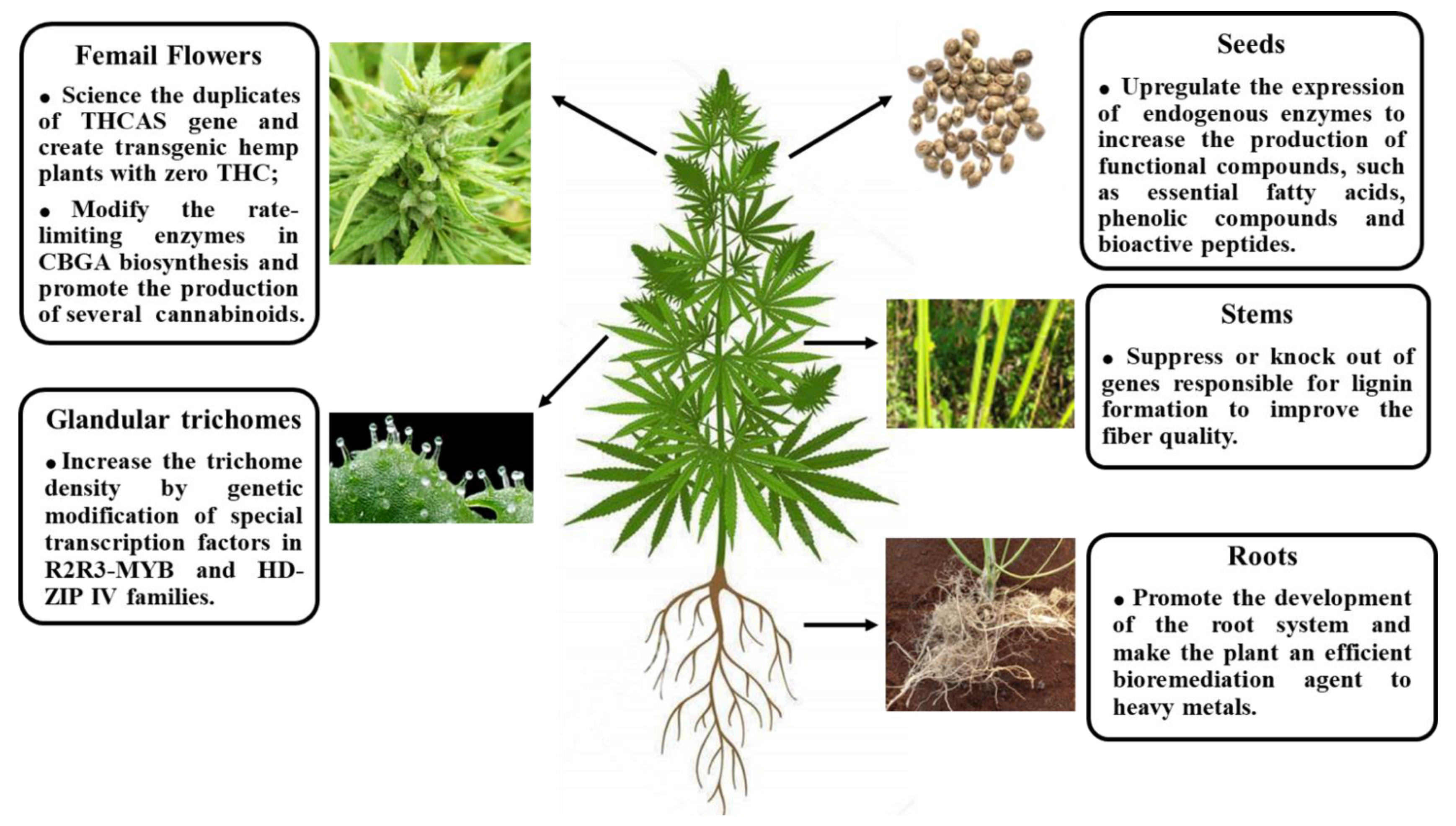
5. Releasing the Full Potential of Cannabis
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Deguchi, M.; Potlakayala, S.; Spuhler, Z.; George, H.; Sheri, V.; Agili, R.; Patel, A.; Rudrabhatla, S. Selection and validation of reference genes for normalization of qRT-PCR data to study the cannabinoid pathway genes in industrial hemp. PLoS ONE 2021, 16, e0260660. [Google Scholar]
- McKernan, K.J.; Helbert, Y.; Kane, L.T.; Ebling, H.; Zhang, L.; Liu, B.; Eaton, Z.; McLaughlin, S.; Kingan, S.; Baybayan, P.; et al. Sequence and annotation of 42 cannabis genomes reveals extensive copy number variation in cannabinoid synthesis and pathogen resistance genes. BioRxiv 2020. [Google Scholar] [CrossRef]
- Nelson, E.U.E. Consumption, not decriminalization: How Nigerian drug dealers/users account for cannabis harms. Int. J. Drug Policy 2022, 106, 103763. [Google Scholar] [CrossRef] [PubMed]
- Mi, R.; Taylor, A.G.; Smart, L.B.; Mattson, N.S. Developing Production Guidelines for Baby Leaf Hemp (Cannabis Sativa L.) as an Edible Salad Green: Cultivar, Sowing Density and Seed Size. Agriculture 2020, 10, 617. [Google Scholar] [CrossRef]
- Corrado, G.; Pannico, A.; Zarrelli, A.; Kyriacou, M.C.; De Pascale, S.; Rouphael, Y. Macro and trace element mineral composition of six hemp varieties grown as microgreens. J. Food Compos. Anal. 2022, 114, 104750. [Google Scholar] [CrossRef]
- Bewley-Taylor, D.; Jelsma, M. Regime change: Re-visiting the 1961 Single Convention on Narcotic Drugs. Int. J. Drug Policy 2012, 23, 72–81. [Google Scholar] [CrossRef]
- Olivas-Aguirre, M.; Torres-López, L.; Villatoro-Gómez, K.; Perez-Tapia, S.M.; Pottosin, I.; Dobrovinskaya, O. Cannabidiol on the Path from the Lab to the Cancer Patient: Opportunities and Challenges. Pharmaceuticals 2022, 15, 366. [Google Scholar] [CrossRef]
- Hesami, M.; Baiton, A.; Alizadeh, M.; Pepe, M.; Torkamaneh, D.; Jones, A.M.P. Advances and perspectives in tissue culture and genetic engineering of cannabis. Int. J. Mol. Sci. 2021, 22, 5671. [Google Scholar] [CrossRef]
- Linder, E.R.; Young, S.; Li, S.; Henriquez Inoa, S.; Suchoff, D.H. The Effect of Transplant Date and Plant Spacing on Biomass Production for Floral Hemp (Cannabis sativa L.). Agronomy 2022, 12, 1856. [Google Scholar] [CrossRef]
- Pattnaik, F.; Nanda, S.; Mohanty, S.; Dalai, A.K.; Kumar, V.; Ponnusamy, S.K.; Naik, S. Cannabis: Chemistry, extraction and therapeutic applications. Chemosphere 2022, 289, 133012. [Google Scholar] [CrossRef]
- Brand, A.; Tissier, A. Control of resource allocation between primary and specialized metabolism in glandular trichomes. Curr. Opin. Plant Biol. 2022, 66, 102172. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.J.H.; Hartono, Y.D.; Xue, B.; Go, M.K.; Fan, H.; Yew, W.S. Structure-Guided Engineering of Prenyltransferase NphB for High-Yield and Regioselective Cannabinoid Production. ACS Catal. 2022, 12, 4628–4639. [Google Scholar] [CrossRef]
- Spitzer-Rimon, B.; Duchin, S.; Bernstein, N.; Kamenetsky, R. Architecture and florogenesis in female Cannabis sativa plants. Front. Plant Sci. 2019, 10, 350. [Google Scholar] [CrossRef] [PubMed]
- Melzer, R.; McCabe, P.F.; Schilling, S. Evolution, genetics and biochemistry of plant cannabinoid synthesis: A challenge for biotechnology in the years ahead. Curr. Opin. Biotechnol. 2022, 75, 102684. [Google Scholar] [CrossRef] [PubMed]
- Ubeed, H.M.S.A.; Bhuyan, D.J.; Alsherbiny, M.A.; Basu, A.; Vuong, Q.V. A Comprehensive Review on the Techniques for Extraction of Bioactive Compounds from Medicinal Cannabis. Molecules 2022, 27, 604. [Google Scholar] [CrossRef]
- Onofri, C.; de Meijer, E.P.M.; Mandolino, G. Sequence heterogeneity of cannabidiolic- and tetrahydrocannabinolic acid-synthase in Cannabis sativa L. and its relationship with chemical phenotype. Phytochemistry 2015, 116, 57–68. [Google Scholar] [CrossRef]
- Zirpel, B.; Stehle, F.; Kayser, O. Production of D9-tetrahydrocannabinolic acid from cannabigerolic acid by whole cells of Pichia (Komagataella) pastoris expressing D9-tetrahydrocannabinolic acid synthase from Cannabis sativa L. Biotechnol. Lett. 2015, 37, 1869–1875. [Google Scholar] [CrossRef]
- Wenger, T.; Watanabe, K.; Sasaki, Y.; Kanazawa, K.; Shimizu, K.; Sirikantaramas, S.; Shoyama, Y.; Taura, F.; Morimoto, S.; Shoyama, Y. Overview of Cannabis including Kampo Medicine and Therapy for Treatment of Dementia: A Review. Front. Pharmacol. 2021, 12, 713228. [Google Scholar] [CrossRef]
- Rudolf, J.D.; Chang, C.Y. Terpene synthases in disguise: Enzymology, structure, and opportunities of non-canonical terpene synthases. Nat. Prod. Rep. 2020, 37, 425–463. [Google Scholar] [CrossRef]
- Valliere, A.; Korman, P.; Woodall, B.; Khitrov, A.; Taylor, E.; Baker, D.; Bowie, U. A cell-free platform for the prenylation of natural products and application to cannabinoid production. Nat. Commun. 2019, 10, 565. [Google Scholar] [CrossRef]
- Aguillón, A.R.; Leão, R.A.; Miranda, L.S.; de Souza, R.O. Cannabidiol Discovery and Synthesis—A Target—Oriented Analysis in Drug Production Processes. CHEM-EUR J. 2021, 27, 5577–5600. [Google Scholar] [CrossRef] [PubMed]
- Ren, G.; Zhang, X.; Li, Y.; Ridout, K.; Serrano-Serrano, M.L.; Yang, Y.; Liu, A.; Ravikanth, G.; Nawaz, M.A.; Mumtaz, A.S.; et al. Large-Scale Whole-Genome Resequencing Unravels the Domestication History of Cannabis sativa. Sci. Adv. 2021, 7, eabg2286. [Google Scholar] [CrossRef] [PubMed]
- Fulvio, F.; Paris, R.; Montanari, M.; Citti, C.; Cilento, V.; Bassolino, L.; Moschella, A.; Alberti, I.; Pecchioni, N.; Cannazza, G.; et al. Analysis of Sequence Variability and Transcriptional Profile of Cannabinoid synthase Genes in Cannabis sativa L. Chemotypes with a Focus on Cannabichromenic acid synthase. Plants 2021, 10, 1857. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, S.E.; Yates, A.S.; Porter, R.K. The Peripheral Cannabinoid Receptor Type 1 (CB1) as a Molecular Target for Modulating Body Weight in Man. Molecules 2021, 26, 6178. [Google Scholar] [CrossRef]
- Rohbeck, E.; Eckel, J.; Romacho, T. Cannabinoid receptors in metabolic regulation and diabetes. Physiology 2021, 36, 102–113. [Google Scholar] [CrossRef]
- Lim, K.J.; Lim, Y.P.; Hartono, Y.D.; Go, M.K.; Fan, H.; Yew, W.S. Biosynthesis of Nature-Inspired Unnatural Cannabinoids. Molecules 2021, 26, 2914. [Google Scholar] [CrossRef]
- Krishna Kumar, K.; Shalev-Benami, M.; Robertson, M.J.; Hu, H.; Banister, S.D.; Hollingsworth, S.A.; Latorraca, N.R.; Kato, H.E.; Hilger, D.; Maeda, S.; et al. Structure of a Signaling Cannabinoid Receptor 1-G Protein Complex. Cell 2019, 176, 448–458. [Google Scholar] [CrossRef]
- Franco, V.; Bialer, M.; Perucca, E. Cannabidiol in the treatment of epilepsy: Current evidence and perspectives for further research. Neuropharmacology 2021, 185, 108442. [Google Scholar] [CrossRef]
- Raïch, I.; Rivas-Santisteban, R.; Lillo, A.; Lillo, J.; Reyes-Resina, I.; Nadal, X.; Ferreiro-Vera, C.; Medina, V.F.; Majellarog, M.; Sotelog, E.; et al. Similarities and differences upon binding of naturally occurring Δ9-tetrahydrocannabinol-derivatives to cannabinoid CB1 and CB2 receptors. Pharmacol. Res. 2021, 174, 105970. [Google Scholar] [CrossRef]
- Laprairie, R.B.; Bagher, A.M.; Kelly, M.E.; Denovan-Wright, E.M. Cannabidiol is a negative allosteric modulator of the cannabinoid CB1 receptor. Br. J. Pharmacol. 2015, 172, 4790–4805. [Google Scholar] [CrossRef]
- Chung, H.; Fierro, A.; Pessoa-Mahana, C.D. Cannabidiol binding and negative allosteric modulation at the cannabinoid type 1 receptor in the presence of delta-9-tetrahydrocannabinol: An In Silico study. PLoS ONE 2019, 14, e0220025. [Google Scholar] [CrossRef] [PubMed]
- Ryu, B.R.; Kim, C.H.; Kwon, T.H.; Han, J.H.; Gim, G.J.; Jahirul Islam, M.; Azad, M.O.K.; Hafizur Rahman, M.; Soyel Rana, M.; Lim, J.D.; et al. Industrial Hemp Clone Selection Method under LED Smart Farm Condition Based on CBD Production per Cubic Meter. Agronomy 2022, 12, 1809. [Google Scholar] [CrossRef]
- Patel, A.; Agili-Shaban, R.; Patel, S.; Proano, R.; Riarh, F.; Hasan, E.; Potlakayala, S.; Rudrabhatla, S. Pharmacological Effects of Cannabinoids Extracted from Industrial hemp on Epilepsy. J. Pharm. Pharm. Sci. 2022, 6, 35–48. [Google Scholar] [CrossRef]
- Brown, J.D.; Winterstein, A.G. Potential adverse drug events and drug-drug interactions with medical and consumer cannabidiol (CBD) use. J. Clin. Med. 2019, 2019, 989. [Google Scholar] [CrossRef]
- Dolgin, E. Inner workings: Genomics blazes a trail to improved cannabis cultivation. Proc. Natl. Acad. Sci. USA 2019, 116, 8638–8640. [Google Scholar] [CrossRef]
- Luo, X.; Reiter, M.A.; d’Espaux, L.; Wong, J.; Denby, C.M.; Lechner, A.; Zhang, Y.; Grzybowski, A.T.; Harth, S.; Lin, W. Complete biosynthesis of cannabinoids and their unnatural analogues in yeast. Nature 2019, 567, 123. [Google Scholar] [CrossRef]
- Gülck, T.; Booth, J.K.; Carvalho, A.; Khakimov, B.; Crocoll, C.; Motawia, M.S.; Møller, B.L.; Bohlmann, J.; Gallage, N.J. Synthetic Biology of Cannabinoids and Cannabinoid Glucosides in Nicotiana benthamiana and Saccharomyces cerevisiae. J. Nat. Prod. 2020, 83, 2877–2893. [Google Scholar] [CrossRef]
- Jakočiūnas, T.; Jensen, M.K.; Keasling, J.D. CRISPR/Cas9 advances engineering of microbial cell factories. Metab. Eng. 2016, 34, 44–59. [Google Scholar] [CrossRef]
- Jakočiūnas, T.; Bonde, I.; Herrgård, M.; Harrison, S.J.; Kristensen, M.; Pedersen, L.E.; Jensen, M.K.; Keasling, J.D. Multiplex metabolic pathway engineering using CRISPR/Cas9 in Saccharomyces cerevisiae. Metab. Eng. 2015, 28, 213–222. [Google Scholar] [CrossRef]
- Dreger, M.; Szalata, M. The Effect of TIBA and NPA on Shoot Regeneration of Cannabis sativa L. Epicotyl Explants. Agronomy 2021, 12, 104. [Google Scholar] [CrossRef]
- Liu, W.; An, C.; Shu, X.; Meng, X.; Yao, Y.; Zhang, J.; Chen, F.; Xiang, H.; Yang, S.; Gao, X.; et al. A dual-plasmid CRISPR/cas system for mycotoxin elimination in polykaryotic industrial fungi. ACS Synth. Biol. 2020, 9, 2087–2095. [Google Scholar] [CrossRef] [PubMed]
- Blatt-Janmaat, K.; Qu, Y. The biochemistry of phytocannabinoids and metabolic engineering of their production in heterologous systems. Int. J. Mol. Sci. 2021, 22, 2454. [Google Scholar] [CrossRef] [PubMed]
- Lata, H.; Chandra, S.; Techen, N.Z.; Khan, I.A.; ElSohly, M.A. In vitro mass propagation of Cannabis sativa L.: A protocol refinement using novel aromatic cytokinin meta-topolin and the assessment of eco-physiological, biochemical and genetic fidelity of micropropagated plants. J. Appl. Res. Med. Aromat. Plants 2016, 3, 18–26. [Google Scholar] [CrossRef]
- Zhang, X.; Xu, G.; Cheng, C.; Lei, L.; Sun, J.; Xu, Y.; Deng, C.; Dai, Z.; Yang, Z.; Chen, X.; et al. Establishment of an Agrobacterium-mediated genetic transformation and CRISPR/Cas9-mediated targeted mutagenesis in Hemp (Cannabis Sativa L.). Plant Biotechnol. J. 2021, 19, 1979–1987. [Google Scholar] [CrossRef] [PubMed]
- Chaohua, C.; Gonggu, Z.; Lining, Z.; Chunsheng, G.; Qing, T.; Jianhua, C.; Xinbo, G.; Dingxiang, P.; Jianguang, S. A rapid shoot regeneration protocol from the cotyledons of hemp (Cannabis sativa L.). Ind. Crops Prod. 2016, 83, 61–65. [Google Scholar] [CrossRef]
- Kodym, A.; Leeb, C. Back to the roots: Protocol for the photoautotrophic micropropagation of medicinal Cannabis. Plant Cell Tissue Organ Cult. 2019, 138, 399–402. [Google Scholar] [CrossRef]
- Deguchi, M.; Kane, S.; Potlakayala, S.; George, H.; Proano, R.; Sheri, V.; Curtis, W.; Rudrabhatla, S. Metabolic engineering strategies of industrial hemp (Cannabis sativa L.): A brief review of the advances and challenges. Front. Plant Sci. 2020, 11, 580621. [Google Scholar] [CrossRef]
- Hesami, M.; Jones, A.M.P. Modeling and optimizing callus growth and development in Cannabis sativa using random forest and support vector machine in combination with a genetic algorithm. Appl. Microbiol. Biotechnol. 2021, 105, 5201–5212. [Google Scholar] [CrossRef]
- Holmes, J.E.; Lung, S.; Collyer, D.; Punja, Z.K. Variables Affecting Shoot Growth and Plantlet Recovery in Tissue Cultures of Drug-Type Cannabis sativa L. Front. Plant Sci. 2021, 12, 732344. [Google Scholar] [CrossRef] [PubMed]
- Page, S.R.G.; Monthony, A.S.; Jones, A.M.P. DKW basal salts improve micropropagation and callogenesis compared to MS basal salts in multiple commercial cultivars of Cannabis sativa. Botany 2021, 99, 269–279. [Google Scholar] [CrossRef]
- Shen, L.; Hua, Y.; Fu, Y.; Li, J.; Liu, Q.; Jiao, X.; Xin, G.; Wang, J.; Wang, X.; Yan, C.; et al. Rapid generation of genetic diversity by multiplex CRISPR/Cas9 genome editing in rice. Sci. China Life Sci. 2017, 60, 506–515. [Google Scholar] [CrossRef] [PubMed]
- Wahby, I.; Caba, J.M.; Ligero, F. Agrobacterium infection of hemp (Cannabis sativa L.): Establishment of hairy root cultures. J. Plant Interact. 2013, 8, 312–320. [Google Scholar] [CrossRef]
- Lowe, K.; La Rota, M.; Hoerster, G.; Hastings, C.; Wang, N.; Chamberlin, M.; Wu, E.; Jones, T.; Gordon-Kamm, W. Rapid genotype “independent” Zea mays L. (maize) transformation via direct somatic embryogenesis. Vitr. Cell. Dev. Biol. Plant 2018, 54, 240–252. [Google Scholar] [CrossRef] [PubMed]
- Matchett-Oates, L.; Cogan, N.O.; Spangenberg, G.C. Manipulation of cannabinoid biosynthesis via transient RNAi expression. Front. Plant Sci. 2021, 12, 773474. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.; Gao, X.; Jahan, M.A.; Adams, M.; Wu, N.; Kovinich, N. Nanoparticle-based genetic transformation of Cannabis sativa. J. Biotechnol. 2021, 326, 48–51. [Google Scholar] [CrossRef] [PubMed]
- Ozyigit, I.I.; Kurtoglu, K.Y. Particle bombardment technology and its applications in plants. Mol. Biol. Rep. 2020, 47, 9831–9847. [Google Scholar] [CrossRef]
- Sorokin, A.; Yadav, N.S.; Gaudet, D.; Kovalchuk, I. Transient expression of the β-glucuronidase gene in Cannabis sativa varieties. Plant Signal. Behav. 2020, 15, 1780037. [Google Scholar] [CrossRef]
- Kong, J.; Martin-Ortigosa, S.; Finer, J.; Orchard, N.; Gunadi, A.; Batts, L.A.; Thakare, D.; Rush, B.; Schmitz, O.; Stuiver, M.; et al. Overexpression of the transcription factor GROWTH-REGULATING FACTOR5 improves transformation of dicot and monocot species. Front. Plant Sci. 2020, 11, 1389. [Google Scholar] [CrossRef]
- Cody, J.P.; Graham, N.D.; Birchler, J.A. BiBAC Modification and Stable Transfer into Maize (Zea mays) Hi-II Immature Embryos via Agrobacterium-Mediated Transformation. Curr. Protoc. Plant Biol. 2017, 2, 350–369. [Google Scholar] [CrossRef]
- Nadakuduti, S.S.; Enciso-Rodríguez, F. Advances in genome editing with CRISPR systems and transformation technologies for plant DNA manipulation. Front. Plant Sci. 2021, 11, 2267. [Google Scholar] [CrossRef]
- Debernardi, J.M.; Tricoli, D.M.; Ercoli, M.F.; Hayta, S.; Ronald, P.; Palatnik, J.F.; Dubcovsky, J. A GRF–GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat. Biotechnol. 2020, 38, 1274–1279. [Google Scholar] [CrossRef]
- Beard, K.M.; Boling, A.W.H.; Bargmann, B.O.R. Protoplast Isolation, Transient Transformation, and Flow-Cytometric Analysis of Reporter-Gene Activation in Cannabis Sativa L. Ind. Crops Prod. 2021, 164, 113360. [Google Scholar] [CrossRef]
- Tsaliki, E.; Kalivas, A.; Jankauskiene, Z.; Irakli, M.; Cook, C.; Grigoriadis, I.; Panoras, I.; Vasilakoglou, I.; Dhima, K. Fibre and seed productivity of industrial hemp (Cannabis sativa L.) varieties under mediterranean conditions. Agronomy 2021, 11, 171. [Google Scholar] [CrossRef]
- Apicella, P.V.; Sands, L.B.; Ma, Y.; Berkowitz, G.A. Delineating genetic regulation of cannabinoid biosynthesis during female flower development in Cannabis sativa. Plant Direct 2022, 6, e412. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, C.; Huang, S.; Chang, L.; Li, J.; Tang, H.; Dey, S.; Biswas, A.; Du, D.; Li, D.; et al. Key Cannabis Salt-Responsive Genes and Pathways Revealed by Comparative Transcriptome and Physiological Analyses of Contrasting Varieties. Agronomy 2021, 11, 2338. [Google Scholar] [CrossRef]
- Haiden, S.R.; Apicella, P.V.; Ma, Y.; Berkowitz, G.A. Overexpression of CsMIXTA, a Transcription Factor from Cannabis sativa, Increases Glandular Trichome Density in Tobacco Leaves. Plants 2022, 11, 1519. [Google Scholar] [CrossRef]
- Tanney, C.A.S.; Backer, R.; Geitmann, A.; Smith, D.L. Cannabis Glandular Trichomes: A Cellular Metabolite Factory. Front. Plant Sci. 2021, 12, 721986. [Google Scholar] [CrossRef]
- Yin, M.; Pan, G.; Tao, J.; Doblin, M.S.; Zeng, W.; Pan, L.; Zhao, L.; Li, Z.; Jiang, H.; Chang, L.; et al. Identification of MYB genes reveals their potential functions in cadmium stress response and the regulation of cannabinoid biosynthesis in hemp. Ind. Crops Prod. 2022, 180, 114607. [Google Scholar] [CrossRef]
- Sayre, R.; Soto-Aguilar, M.; Zidenga, T.; Goncalves, E.C. Systems and Methods for Enhancing Trichome Formation and Density in Cannabis. U.S. Patent No. 10,724,048 B2, 28 July 2020. [Google Scholar]
- Jang, J.H.; Lee, O.R. Patatin-related phospholipase AtpPLAIIIα affects lignification of xylem in Arabidopsis and hybrid poplars. Plants 2020, 9, 451. [Google Scholar] [CrossRef]
- Simiyu, D.C.; Jang, J.H.; Lee, O.R. Understanding Cannabis sativa L.: Current Status of Propagation, Use, Legalization, and Haploid-Inducer-Mediated Genetic Engineering. Plants 2022, 11, 1236. [Google Scholar] [CrossRef]
- Shiels, D.; Prestwich, B.D.; Koo, O.; Kanchiswamy, C.N.; O’Halloran, R.; Badmi, R. Hemp Genome Editing-Challenges and Opportunities. Front. Genome Ed. 2022, 4, 823486. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, L.; Yu, S.; Chen, J.; Cheng, C.; Sun, J.; Xu, Y.; Deng, C.; Dai, Z.; Yang, Z.; Chen, X.; et al. Releasing the Full Potential of Cannabis through Biotechnology. Agronomy 2022, 12, 2439. https://doi.org/10.3390/agronomy12102439
Li L, Yu S, Chen J, Cheng C, Sun J, Xu Y, Deng C, Dai Z, Yang Z, Chen X, et al. Releasing the Full Potential of Cannabis through Biotechnology. Agronomy. 2022; 12(10):2439. https://doi.org/10.3390/agronomy12102439
Chicago/Turabian StyleLi, Li, Shuhao Yu, Jiquan Chen, Chaohua Cheng, Jian Sun, Ying Xu, Canhui Deng, Zhigang Dai, Zemao Yang, Xiaojun Chen, and et al. 2022. "Releasing the Full Potential of Cannabis through Biotechnology" Agronomy 12, no. 10: 2439. https://doi.org/10.3390/agronomy12102439
APA StyleLi, L., Yu, S., Chen, J., Cheng, C., Sun, J., Xu, Y., Deng, C., Dai, Z., Yang, Z., Chen, X., Tang, Q., Su, J., & Zhang, X. (2022). Releasing the Full Potential of Cannabis through Biotechnology. Agronomy, 12(10), 2439. https://doi.org/10.3390/agronomy12102439





