Polymer-Based Microfluidic Devices for Pharmacy, Biology and Tissue Engineering
Abstract
:Abbreviations
| AC | Alternating current |
| ATP | Adenosine tri-phosphate |
| CNC | Computer numerical control |
| COC | Cyclic olefin copolymer |
| DC | Direct current |
| DEP | Dielectrophoresis |
| DNA | Deoxyribonucleic acid |
| ECM | Extracellular matrix coating |
| E. coli | Escherichia coli |
| EDEP | Electrodeless dielectrophoresis |
| EOF | Electro-osmotic flow |
| hMSCs | Human primary mesenchymal stem cells |
| HTC | High throughput screening |
| IC | Integrated circuit |
| MACS | Magnetically actuated cell sorter |
| μTAS | Micro total analysis system |
| PC | Polycarbonate |
| PCR | Polymerase chain reaction |
| PDMS | Polydimethylsiloxane |
| pH | Measure of the acidity or alkalinity of an aqueous solution |
| PMMA | Polymethylmethacrylate |
| PMT | Photomultiplier tube |
| PS | Polystyrene |
| Re | Reynolds number |
| RNA | Ribonucleic acid |
| SPRI | Solid-phase reversible immobilization |
| Tg | Glass transition temperature |
| UV | Ultraviolet |
| UV-Vis | Ultraviolet-Visible |
| UVLIGA | Ultraviolet lithography, electroforming and molding |
| VSCEL | Vertical cavity surface emitting lasers |
1. Introduction
2. Microfluidic Systems, Flow Dynamics on the Small Scale, and Manufacturing
2.1. Characterization of the Fluid Flow
2.2. The Fluid Transport
2.2.1. Pressure-Driven Flow
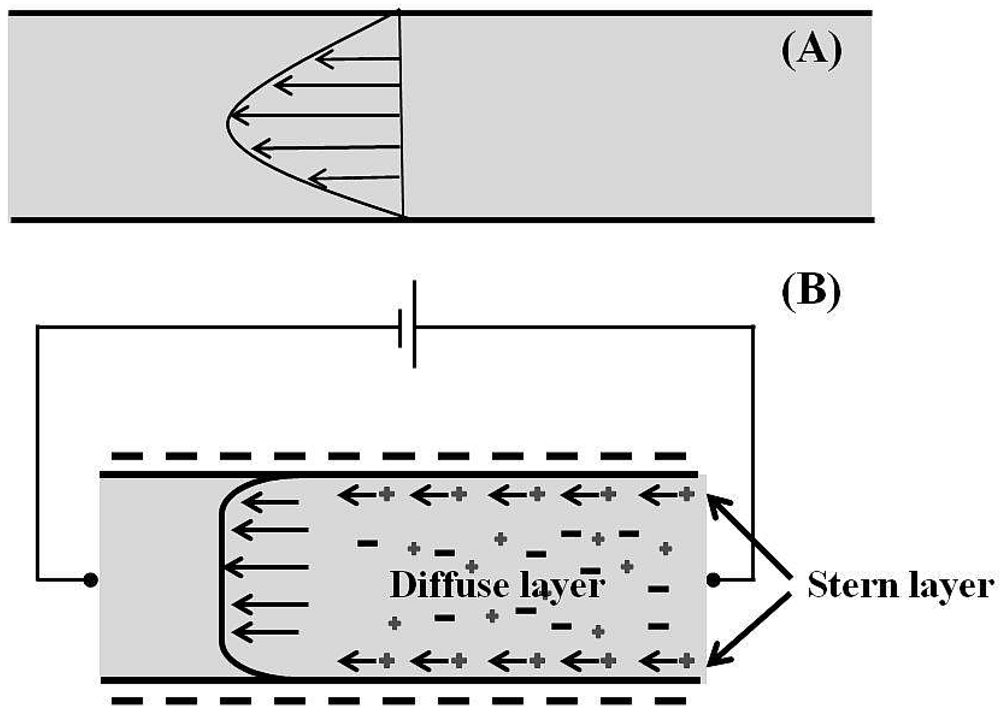
2.2.2. Electro-Kinetic Flow
2.2.2.1. Electro-Osmotic Flow (EOF)
2.2.2.2. Electrophoresis Flow
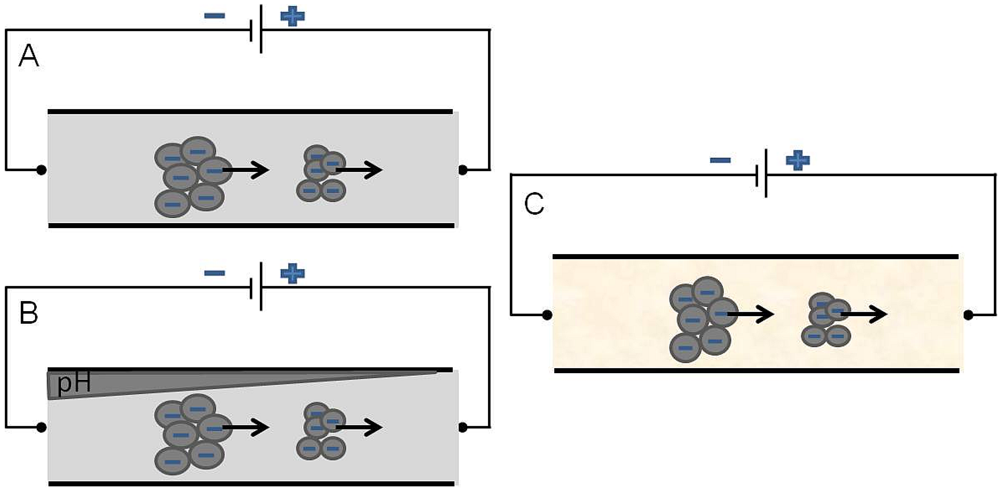
3. Fabrication of Microfluidic Devices
3.1. Integrated Micromachined Devices
3.2. Direct Fabrication Techniques
3.2.1. Photolithography
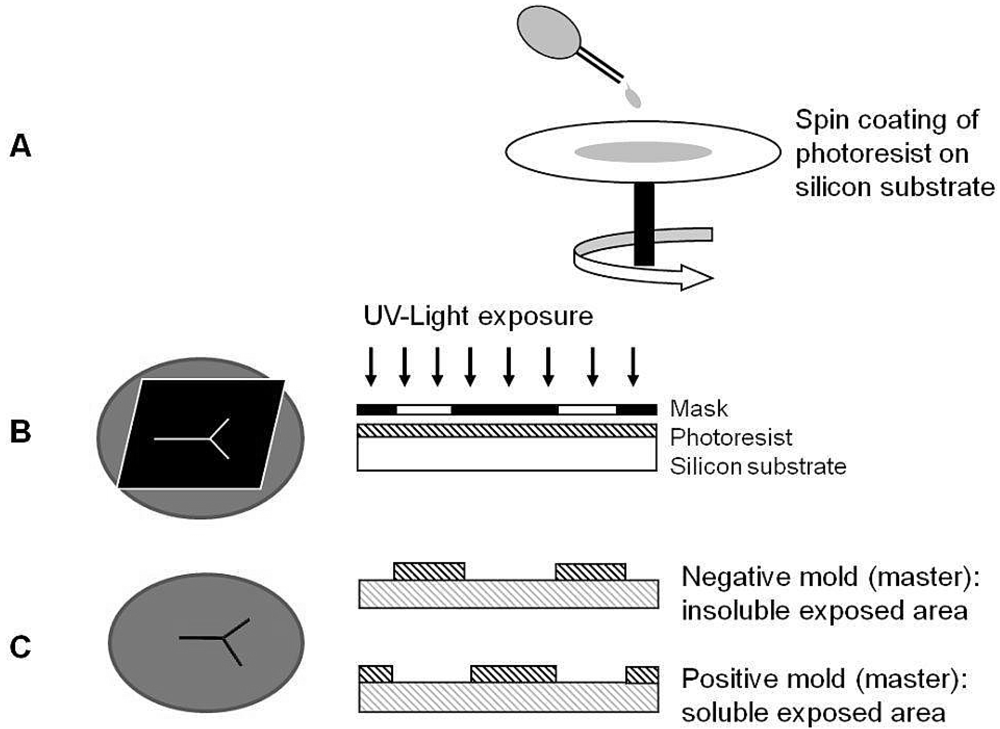
3.2.2. The Conventional Surface Micromachining
3.2.3. Laser Ablation Micromachining
3.3. Prototyping Techniques
3.3.1. The Hot Embossing Technique
3.3.2. Microinjection Molding
3.3.3. Computer Numerical Control (CNC) Micromachining
3.3.4. Soft Lithography
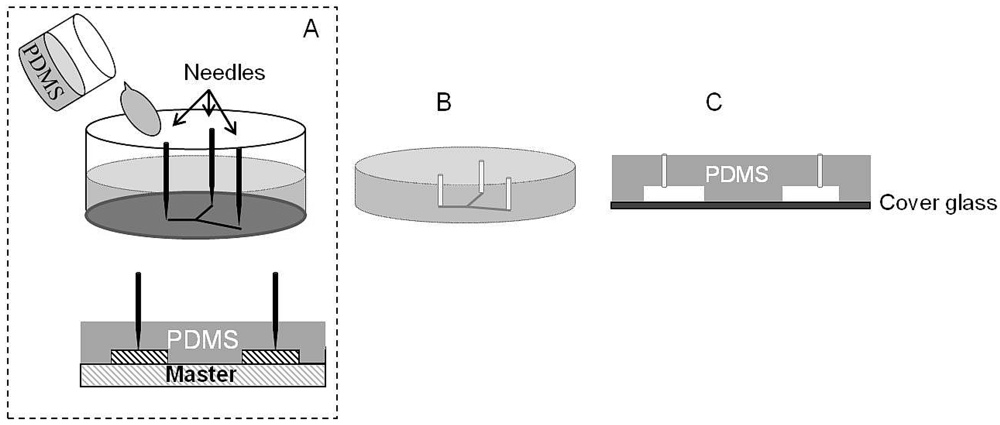
3.4. Bonding: Complementary Step for Fabrication of Microfluidic Devices
4. Microfluidic Applications in Pharmaceutical, Biological and Biomedical Engineering
4.1. Manipulation of Biological Cells on-Chip
4.1.1. Magnetic Manipulation

4.1.2. Mechanical Manipulation
4.1.3. Electric Manipulation
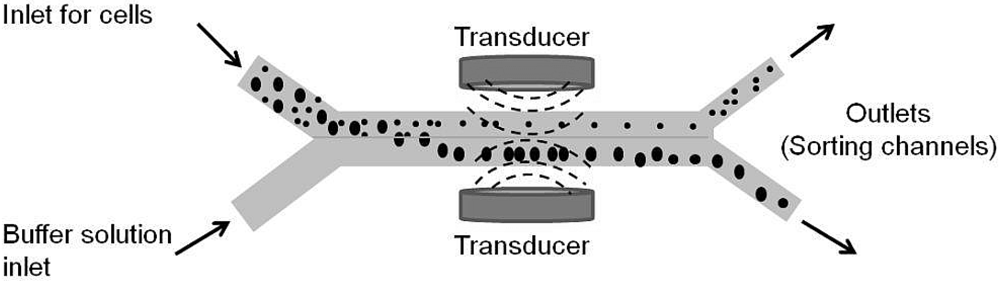
4.1.4. Acoustic Manipulation
4.1.5. Optical Manipulation
4.1.6. Flow Cytometry on Microfluidic Chip
4.1.7. Cell Analysis
4.1.8. Cell Culturing
4.1.9. Cell-Based High Throughput Screening (HTC)
4.1.10. Tissue Engineering Models on Microfluidic Chips
4.2. Biocompatibility and Cell Viability within Microfluidic Systems
5. Discussion
6. Conclusions
References
- Gravesen, P.; Branebjerg, J.; Jensen, O.S. Microfluidics: A review. J. Micromech. Microeng. 1993, 3, 168–182. [Google Scholar] [CrossRef]
- Terry, S.C.; Jerman, J.H.; Angell, J.B. A gas chromatographic air analyzer fabricated on a silicon wafer. IEEE Trans. Electron Devices 1979, 26, 1880–1886. [Google Scholar]
- Petersen, K.E. Fabrication of an integrated, planar silicon ink-jet structure. IEEE Trans. Electron Devices 1979, 26, 1918–1920. [Google Scholar] [CrossRef]
- Manz, A.; Graber, N.; Widmer, H.M. Miniaturized total chemical analysis systems: A novel concept for chemical sensing. Sens. Actuators B Chem. 1990, 1, 244–248. [Google Scholar] [CrossRef]
- Beebe, D.J.; Mensing, G.A.; Walker, G.M. Physics and applications of microfluidics in biology. Annu. Rev. Biomed. Eng. 2002, 4, 261–286. [Google Scholar] [CrossRef]
- Koch, M.; Evans, A.; Brunnschweiler, A. Microfluidic Technology and Applications; Research Studies Press: Baldock, UK, 2000. [Google Scholar]
- Becker, H.; Locascio, L.E. Polymer microfluidic devices. Talanta 2002, 56, 267–287. [Google Scholar] [CrossRef]
- Stroock, A.D.; Whitesides, G.M. Components for integrated poly(dimethylsiloxane) microfluidic systems. Electrophoresis 2002, 23, 3461–3473. [Google Scholar] [CrossRef]
- Anderson, J.R.; Chiu, D.T.; Wu, H.; Schueller, O.J.A.; Whitesides, G.M. Fabrication of microfluidic systems in poly(dimethylsiloxane). Electrophoresis 2000, 21, 27–40. [Google Scholar] [CrossRef]
- Gast, F.U.; Fiehn, H. Profile. The development of integrated microfluidic systems at GeSiM. Lab Chip 2003, 3, 6–10. [Google Scholar]
- Reyes, D.R.; Iossifidis, D.; Auroux, P.A.; Manz, A. Micro total analysis systems. 1. Introduction, theory, and technology. Anal. Chem. 2002, 74, 2623–2636. [Google Scholar]
- Auroux, P.A.; Iossifidis, D.; Reyes, D.R.; Manz, A. Micro total analysis systems. 2. Analytical standard operations and applications. Anal. Chem. 2002, 74, 2637–2652. [Google Scholar] [CrossRef]
- Pol, F.; Branebjerg, J. Micro Liquid-Handling Devices: A Review; MST: Berlin, Germany, 1990. [Google Scholar]
- Jakeway, S.C.; de Mello, A.J.; Russell, E.L. Miniaturized total analysis systems for biological analysis. Fresenius J. Anal. Chem. 2000, 366, 525–539. [Google Scholar] [CrossRef]
- Chován, T.; Guttman, A. Microfabricated devices in biotechnology and biochemical processing. Trends Biotechnol. 2002, 20, 116–122. [Google Scholar] [CrossRef]
- Tüdős, A.J.; Besselink, G.A.J.; Schasfoort, R.B.M. Trends in miniaturized total analysis systems for point-of-care testing in clinical chemistry. Lab Chip 2001, 1, 83–95. [Google Scholar] [CrossRef]
- Vo-Dinh, T.; Cullum, B. Biosensors and biochips: Advances in biological and medical diagnostics. Fresenius J. Anal. Chem. 2000, 366, 540–551. [Google Scholar] [CrossRef]
- Verpoorte, E. Microfluidic chips for clinical and forensic analysis. Electrophoresis 2002, 23, 677–712. [Google Scholar] [CrossRef]
- Huang, Y.; Mather, E.L.; Bell, J.L.; Madou, M. MEMS-based sample preparation for molecular diagnostics. Anal. Bioanal. Chem. 2002, 372, 49–65. [Google Scholar] [CrossRef]
- Erickson, D.; Li, D.; Krull, U.J. Modeling of DNA hybridization kinetics for spatially resolved biochips. Anal. Biochem. 2003, 317, 186–200. [Google Scholar] [CrossRef]
- Ho, C.M.; Tai, Y.C. Micro-Electro-Mechanical-Systems (MEMS) and fluid flows. Annu. Rev. Fluid Mech. 1998, 30, 579–612. [Google Scholar] [CrossRef]
- Currie, I.G. Fundamental Mechanics of Fluids; CRC Press: Boca Raton, FL, USA, 2002. [Google Scholar]
- Kim, C.; Lee, K.; Kim, J.H.; Shin, K.S.; Lee, K.J.; Kim, T.S.; Kang, J.Y. A serial dilution microfluidic device using a ladder network generating logarithmic or linear concentrations. Lab Chip 2008, 8, 473–479. [Google Scholar] [CrossRef]
- Cardiovascular Physiology Concepts Homepage. Available online: http://www.cvphysiology.com/Hemodynamics/H006.htm (accessed on 28 February 2012).
- Avila, K.; Moxey, D.; de Lozar, A.; Avila, M.; Barkley, D.; Hof, B. The onset of turbulence in pipe flow. Science 2011, 333, 192–196. [Google Scholar]
- Kovarik, M.L.; Lai, H.H.; Xiong, J.C.; Allbritton, N.L. Sample transport and electrokinetic injection in a microchip device for chemical cytometry. Electrophoresis 2011, 32, 1–8. [Google Scholar] [CrossRef]
- Yang, R.; Feeback, D.L.; Wang, W. Microfabrication and test of a three-dimensional polymer hydro-focusing unit for flow cytometry applications. Sens. Actuators A Phys. 2005, 118, 259–267. [Google Scholar] [CrossRef]
- Weigl, B.H.; Yager, P. Microfluidic diffusion-based separation and detection. Science 1999, 283, 346–347. [Google Scholar] [CrossRef]
- Burns, M.A.; Johnson, B.N.; Brahmasandra, S.N.; Handique, K.; Webster, J.R.; Krishnan, M.; Sammarco, T.S.; Man, P.M.; Jones, D.; Heldsinger, D. An integrated nanoliter DNA analysis device. Science 1998, 282, 484–487. [Google Scholar]
- Ocvirk, G.; Munroe, M.; Tang, T.; Oleschuk, R.; Westra, K.; Harrison, D.J. Electrokinetic control of fluid flow in native poly(dimethylsiloxane) capillary electrophoresis devices. Electrophoresis 2000, 21, 107–115. [Google Scholar] [CrossRef]
- Thompson, P.A.; Troian, S.M. A general boundary condition for liquid flow at solid surfaces. Nature 1997, 389, 360–362. [Google Scholar]
- Tallarek, U.; Rapp, E.; Scheenen, T.; Bayer, E.; van As, H. Electroosmotic and pressure-driven flow in open and packed capillaries: Velocity distributions and fluid dispersion. Anal. Chem. 2000, 72, 2292–2301. [Google Scholar]
- Clemmens, J.; Hess, H.; Doot, R.; Matzke, C.M.; Bachand, G.D.; Vogel, V. Motor-protein “roundabouts”: Microtubules moving on kinesin-coated tracks through engineered networks. Lab Chip 2004, 4, 83–86. [Google Scholar] [CrossRef]
- Laser, D.; Santiago, J. A review of micropumps. J. Micromech. Microeng. 2004, 14, R35–R64. [Google Scholar] [CrossRef]
- Celectricon Simply Stimulating Homepage. Available online: http://www.cellectricon.se (accessed on 28 February 2012).
- Beebe, D.J.; Moore, J.S.; Yu, Q.; Liu, R.H.; Kraft, M.L.; Jo, B.H.; Devadoss, C. Microfluidic tectonics: A comprehensive construction platform for microfluidic systems. Proc. Natl. Acad. Sci. USA 2000, 97, 13488–13493. [Google Scholar]
- Yu, C.; Parikh, A.N.; Groves, J.T. Direct patterning of membrane-derivatized colloids using in situ UV-ozone photolithography. Adv. Mater. 2005, 17, 1477–1480. [Google Scholar] [CrossRef]
- Yang, M.; Li, C.W.; Yang, J. Cell docking and on-chip monitoring of cellular reactions with a controlled concentration gradient on a microfluidic device. Anal. Chem. 2002, 74, 3991–4001. [Google Scholar] [CrossRef]
- Clarson, S.J.; Semlyen, J.A.; Clarson, S.J. Siloxane Polymers; Prentice Hall: Englewood Cliffs, NJ, USA, 1993. [Google Scholar]
- Kim, P.; Jeong, H.E.; Khademhosseini, A.; Suh, K.Y. Fabrication of non-biofouling polyethylene glycol micro-and nanochannels by ultraviolet-assisted irreversible sealing. Lab Chip 2006, 6, 1432–1437. [Google Scholar] [CrossRef]
- Kaneta, T.; Ogura, T.; Yamato, S.; Imasaka, T. Band broadening of DNA fragments isolated by polyacrylamide gel electrophoresis in capillary electrophoresis. J. Sep. Sci. 2012, 35, 431–435. [Google Scholar] [CrossRef]
- Li, P.C.H.; Harrison, D.J. Transport, manipulation, and reaction of biological cells on-chip using electrokinetic effect. Anal. Chem. 1997, 69, 1564–1568. [Google Scholar]
- Waters, L.C.; Jacobson, S.C.; Kroutchinina, N.; Khandurina, J.; Foote, R.S.; Ramsey, J.M. Microchip device for cell lysis, multiplex PCR amplification, and electrophoretic sizing. Anal. Chem. 1998, 70, 158–162. [Google Scholar]
- Ichiki, T.; Ujiie, T.; Shinbashi, S.; Okuda, T.; Horiike, Y. Immunoelectrophoresis of red blood cells performed on microcapillary chips. Electrophoresis 2002, 23, 2029–2034. [Google Scholar]
- Lee, H.; Sun, E.; Ham, D.; Weissleder, R. Chip-NMR biosensor for detection and molecular analysis of cells. Nat. Med. 2008, 14, 869–874. [Google Scholar] [CrossRef]
- Qu, B.Y.; Wu, Z.Y.; Fang, F.; Bai, Z.M.; Yang, D.Z.; Xu, S.K. A glass microfluidic chip for continuous blood cell sorting by a magnetic gradient without labeling. Anal. Bioanal. Chem. 2008, 392, 1317–1324. [Google Scholar] [CrossRef]
- Omasu, F.; Nakano, Y.; Ichiki, T. Measurement of the electrophoretic mobility of sheep erythrocytes using microcapillary chips. Electrophoresis 2005, 26, 1163–1167. [Google Scholar]
- Pilarski, P.M.; Adamia, S.; Backhouse, C.J. An adaptable microvalving system for on-chip polymerase chain reactions. J. Immunol. Methods 2005, 305, 48–58. [Google Scholar] [CrossRef]
- Unger, M.A.; Chou, H.P.; Thorsen, T.; Scherer, A.; Quake, S.R. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 2000, 288, 113–116. [Google Scholar]
- Harrison, D.J.; Fluri, K.; Seiler, K.; Fan, Z.; Effenhauser, C.S.; Manz, A. Micromachining a miniaturized capillary electrophoresis-based chemical analysis system on a chip. Science 1993, 261, 895–897. [Google Scholar]
- Raley, N.F.; Davidson, J.C.; Balch, J.W. Examination of glass-silicon and glass-glass bonding techniques for microfluidic systems. Proc. SPIE 2639, 40–45. [Google Scholar]
- Kopp, M.U.; de Mello, A.J.; Manz, A. Chemical amplification: Continuous-flow PCR on a chip. Science 1998, 280, 1046–1048. [Google Scholar]
- Stjernström, M.; Roeraade, J. Method for fabrication of microfluidic systems in glass. J. Micromech. Microeng. 1998, 8, 33–38. [Google Scholar] [CrossRef]
- Bings, N.H.; Wang, C.; Skinner, C.D.; Colyer, C.L.; Thibault, P.; Harrison, D.J. Microfluidic devices connected to fused-silica capillaries with minimal dead volume. Anal. Chem. 1999, 71, 3292–3296. [Google Scholar]
- Ruano, J.M.; Benoit, V.; Aitchison, J.S.; Cooper, J.M. Flame hydrolysis deposition of glass on silicon for the integration of optical and microfluidic devices. Anal. Chem. 2000, 72, 1093–1097. [Google Scholar]
- Grover, W.H.; Skelley, A.M.; Liu, C.N.; Lagally, E.T.; Mathies, R.A. Monolithic membrane valves and diaphragm pumps for practical large-scale integration into glass microfluidic devices. Sens. Actuators B Chem. 2003, 89, 315–323. [Google Scholar] [CrossRef]
- Jia, Z.J.; Fang, Q.; Fang, Z.L. Bonding of glass microfluidic chips at room temperatures. Anal. Chem. 2004, 76, 5597–5602. [Google Scholar] [CrossRef]
- Cheng, Y.; Sugioka, K.; Midorikawa, K. Microfluidic laser embedded in glass by three-dimensional femtosecond laser microprocessing. Opt. Lett. 2004, 29, 2007–2009. [Google Scholar] [CrossRef]
- Bu, M.; Melvin, T.; Ensell, G.J.; Wilkinson, J.S.; Evans, A.G.R. A new masking technology for deep glass etching and its microfluidic application. Sens. Actuators A Phys. 2004, 115, 476–482. [Google Scholar] [CrossRef]
- Allen, P.B.; Chiu, D.T. Calcium-assisted glass-to-glass bonding for fabrication of glass microfluidic devices. Anal. Chem. 2008, 80, 7153–7157. [Google Scholar] [CrossRef]
- Vulto, P.; Huesgen, T.; Albrecht, B.; Urban, G. A full-wafer fabrication process for glass microfluidic chips with integrated electroplated electrodes by direct bonding of dry film resist. J. Micromech. Microeng. 2009, 19, 077001:1–077001:5. [Google Scholar]
- Giordano, B.; Ferrance, J.; Swedberg, S.; Hühmer, A.; Landers, J. Polymerase chain reaction in polymeric microchips: DNA amplification in less than 240 seconds. Anal. Biochem. 2001, 291, 124–132. [Google Scholar]
- Rossier, J.; Reymond, F.; Michel, P.E. Polymer microfluidic chips for electrochemical and biochemical analyses. Electrophoresis 2002, 23, 858–867. [Google Scholar] [CrossRef]
- Cao, H.; Tegenfeldt, J.O.; Austin, R.H.; Chou, S.Y. Gradient nanostructures for interfacing microfluidics and nanofluidics. Appl. Phys. Lett. 2002, 81, 3058–3060. [Google Scholar]
- Mappes, T.; Achenbach, S.; Mohr, J. X-ray lithography for devices with high aspect ratio polymer submicron structures. Microelectron. Eng. 2007, 84, 1235–1239. [Google Scholar] [CrossRef]
- Becker, H.; Heim, U. Hot embossing as a method for the fabrication of polymer high aspect ratio structures. Sens. Actuators A Phys. 2000, 83, 130–135. [Google Scholar] [CrossRef]
- Qi, S.; Liu, X.; Ford, S.; Barrows, J.; Thomas, G.; Kelly, K.; McCandless, A.; Lian, K.; Goettert, J.; Soper, S.A. Microfluidic devices fabricated in poly(methyl methacrylate) using hot-embossing with integrated sampling capillary and fiber optics for fluorescence detection. Lab Chip 2002, 2, 88–95. [Google Scholar] [CrossRef]
- Chien, R.D. Micromolding of biochip devices designed with microchannels. Sens. Actuators A Phys. 2006, 128, 238–247. [Google Scholar] [CrossRef]
- Attia, U.M.; Marson, S.; Alcock, J.R. Micro-injection moulding of polymer microfluidic devices. Microfluid. Nanofluidics 2009, 7, 1–28. [Google Scholar] [CrossRef]
- Maillefer, D.; Gamper, S.; Frehner, B.; Balmer, P.; van Lintel, H.; Renaud, P. A high-performance silicon micropump for disposable drug delivery systems. In Proceedings of 14th IEEE International Conference on Micro Electro Mechanical SystemsMEMS 2001, Interlaken, Switzerland, 21-25 January 2001; pp. 413–417.
- Pemble, C.M.; Towe, B.C. A miniature shape memory alloy pinch valve. Sens. Actuators A Phys. 1999, 77, 145–148. [Google Scholar] [CrossRef]
- Weibel, D.B.; Siegel, A.C.; Lee, A.; George, A.H.; Whitesides, G.M. Pumping fluids in microfluidic systems using the elastic deformation of poly(dimethylsiloxane). Lab Chip 2007, 7, 1832–1836. [Google Scholar] [CrossRef]
- Weibel, D.B.; Kruithof, M.; Potenta, S.; Sia, S.K.; Lee, A.; Whitesides, G.M. Torque-actuated valves for microfluidics. Anal. Chem. 2005, 77, 4726–4733. [Google Scholar]
- Sundararajan, N.; Kim, D.; Berlin, A.A. Microfluidic operations using deformable polymer membranes fabricated by single layer soft lithography. Lab Chip 2005, 5, 350–354. [Google Scholar] [CrossRef]
- Studer, V.; Hang, G.; Pandolfi, A.; Ortiz, M.; Anderson, W.F.; Quake, S.R. Scaling properties of a low-actuation pressure microfluidic valve. J. Appl. Phys. 2004, 95, 393–398. [Google Scholar]
- Hosokawa, K.; Maeda, R. A pneumatically-actuated three-way microvalve fabricated with polydimethylsiloxane using the membrane transfer technique. J. Micromech. Microeng. 2000, 10, 415–420. [Google Scholar] [CrossRef]
- Yang, B.; Lin, Q. A latchable microvalve using phase change of paraffin wax. Sens. Actuators A Phys. 2007, 134, 194–200. [Google Scholar] [CrossRef]
- Yoo, J.C.; Choi, Y.; Kang, C.; Kim, Y.S. A novel polydimethylsiloxane microfluidic system including thermopneumatic-actuated micropump and paraffin-actuated microvalve. Sens. Actuators A Phys. 2007, 139, 216–220. [Google Scholar] [CrossRef]
- Irimia, D.; Toner, M. Cell handling using microstructured membranes. Lab Chip 2006, 6, 345–352. [Google Scholar] [CrossRef]
- Van der Wijngaart, W.; Chugh, D.; Man, E.; Melin, J.; Stemme, G. A low-temperature thermopneumatic actuation principle for gas bubble microvalves. J. Microelectromech. Syst. 2007, 16, 765–774. [Google Scholar] [CrossRef]
- Lee, D.E.; Soper, S.; Wang, W. Design and fabrication of an electrochemically actuated microvalve. Microsyst. Technol. 2008, 14, 1751–1756. [Google Scholar] [CrossRef]
- Kaigala, G.V.; Hoang, V.N.; Backhouse, C.J. Electrically controlled microvalves to integrate microchip polymerase chain reaction and capillary electrophoresis. Lab Chip 2008, 8, 1071–1078. [Google Scholar] [CrossRef]
- Jacobson, S.C.; Ermakov, S.V.; Ramsey, J.M. Minimizing the number of voltage sources and fluid reservoirs for electrokinetic valving in microfluidic devices. Anal. Chem. 1999, 71, 3273–3276. [Google Scholar] [CrossRef]
- Schasfoort, R.B.M.; Schlautmann, S.; Hendrikse, J.; van den Berg, A. Field-effect flow control for microfabricated fluidic networks. Science 1999, 286, 942–945. [Google Scholar]
- Liu, R.H.; Bonanno, J.; Yang, J.; Lenigk, R.; Grodzinski, P. Single-use, thermally actuated paraffin valves for microfluidic applications. Sens. Actuators B Chem. 2004, 98, 328–336. [Google Scholar] [CrossRef]
- Gui, L.; Liu, J. Ice valve for a mini/micro flow channel. J. Micromech. Microeng. 2004, 14, 242–246. [Google Scholar] [CrossRef]
- Yu, Q.; Bauer, J.M.; Moore, J.S.; Beebe, D.J. Responsive biomimetic hydrogel valve for microfluidics. Appl. Phys. Lett. 2001, 78, 2589–2591. [Google Scholar]
- Liu, C.; Park, J.Y.; Xu, Y.; Lee, S.H. Arrayed pH-responsive microvalves controlled by multiphase laminar flow. J. Micromech. Microeng. 2007, 17, 1985–1991. [Google Scholar] [CrossRef]
- Cho, H.; Kim, H.Y.; Kang, J.Y.; Kim, T.S. How the capillary burst microvalve works. J. Colloid Interface Sci. 2007, 306, 379–385. [Google Scholar] [CrossRef]
- Chen, J.M.; Huang, P.C.; Lin, M.G. Analysis and experiment of capillary valves for microfluidics on a rotating disk. Microfluidics Nanofluidics 2008, 4, 427–437. [Google Scholar] [CrossRef]
- Riegger, L.; Mielnik, M.; Gulliksen, A.; Mark, D.; Steigert, J.; Lutz, S.; Clad, M.; Zengerle, R.; Koltay, P.; Hoffmann, J. Dye-based coatings for hydrophobic valves and their application to polymer labs-on-a-chip. J. Micromech. Microeng. 2010, 20, 045021:1–045021:6. [Google Scholar]
- Allain, M.; Berthier, J.; Basrour, S.; Pouteau, P. Electrically actuated sacrificial membranes for valving in microsystems. J. Micromech. Microeng. 2010, 20, 035006:1–035006:7. [Google Scholar]
- Moreno, J.M.; Quero, J.M. A Novel single-use SU-8 microvalve for pressure-driven microfluidic applications. J. Micromech. Microeng. 2010, 20, 015005:1–015005:11. [Google Scholar]
- Daniel, K.; Duc, H.L.H.; Cima, M.; Langer, R. Controlled release microchips. Chronopharmaceutics 2009, 187–215. [Google Scholar]
- Juncker, D.; Schmid, H.; Drechsler, U.; Wolf, H.; Wolf, M.; Michel, B.; de Rooij, N.; Delamarche, E. Autonomous microfluidic capillary system. Anal. Chem. 2002, 74, 6139–6144. [Google Scholar]
- Walker, G.M.; Beebe, D.J. A passive pumping method for microfluidic devices. Lab Chip 2002, 2, 131–134. [Google Scholar] [CrossRef]
- Schönfeld, F.; Hessel, V.; Hofmann, C. An optimised split-and-recombine micro-mixer with uniform “chaotic” mixing. Lab Chip 2004, 4, 65–69. [Google Scholar] [CrossRef]
- Jacobson, S.C.; Ramsey, J.M. Integrated microdevice for DNA restriction fragment analysis. Anal. Chem. 1996, 68, 720–723. [Google Scholar] [CrossRef]
- Yuen, P.K.; Kricka, L.J.; Fortina, P.; Panaro, N.J.; Sakazume, T.; Wilding, P. Microchip module for blood sample preparation and nucleic acid amplification reactions. Genome Res. 2001, 11, 405–412. [Google Scholar] [CrossRef]
- Kozlowski, F.; Lindmair, N.; Scheiter, T.; Hierold, C.; Lang, W. A novel method to avoid sticking of surface-micromachined structures. Sens. Actuators A Phys. 1996, 54, 659–662. [Google Scholar] [CrossRef]
- Steiner, P.; Lang, W. Micromachining applications of porous silicon. Thin Solid Films 1995, 255, 52–58. [Google Scholar] [CrossRef]
- De Boer, M.J.; Tjerkstra, R.W.; Berenschot, J.; Jansen, H.V.; Burger, G.; Gardeniers, J.; Elwenspoek, M.; van den Berg, A. Micromachining of buried micro channels in silicon. J. Microelectromech. Syst. 2000, 9, 94–103. [Google Scholar] [CrossRef]
- Schwarz, A.; Rossier, J.; Bianchi, F.; Reymond, F.; Ferrigno, R.; Girault, H. Micro-TAS on polymer substrates micromachined by laser photoablation. In Proceedings of the µTAS’98 Workshop, Banff, AB, Canada, 13-16 October 1998; pp. 241–244.
- Becker, H.; Klotzbuecher, T. Polymer nanowell plates with variable slope angles. In Proceedings of the 3rd International Conference on Microreaction Technology, Frankfurt, Germany, 19-21 April 1999.
- Pethig, R.; Burt, J.; Parton, A.; Rizvi, N.; Talary, M.; Tame, J. Development of biofactory-on-a-chip technology using excimer laser micromachining. J. Micromech. Microeng. 1998, 8, 57–63. [Google Scholar] [CrossRef]
- Roberts, M.A.; Rossier, J.S.; Bercier, P.; Girault, H. UV Laser machined polymer substrates for the development of microdiagnostic systems. Anal. Chem. 1997, 69, 2035–2042. [Google Scholar] [CrossRef]
- Rötting, O.; Röpke, W.; Becker, H.; Gärtner, C. Polymer microfabrication technologies. Microsyst. Technol. 2002, 8, 32–36. [Google Scholar] [CrossRef]
- Becker, H.; Dietz, W. Microfluidic devices for μ-TAS applications fabricated by polymer hot embossing. Proc. SPIE 3515, 177–182. [Google Scholar]
- Shao, P.; van Kan, A.; Wang, L.; Ansari, K.; Bettiol, A.; Watt, F. Fabrication of enclosed nanochannels in poly(methylmethacrylate) using proton beam writing and thermal bonding. Appl. Phys. Lett. 2006, 88, 093515:1–093515:4. [Google Scholar]
- Schift, H.; Heyderman, L.; Padeste, C.; Gobrecht, J. Chemical nano-patterning using hot embossing lithography. Microelectron. Eng. 2002, 61, 423–428. [Google Scholar] [CrossRef]
- Heckele, M.; Schomburg, W. Review on micro molding of thermoplastic polymers. J. Micromech. Microeng. 2004, 14, R1–R14. [Google Scholar] [CrossRef]
- Worgull, M.; Heckele, M.; Schomburg, W. Large-scale hot embossing. Microsyst. Technol. 2005, 12, 110–115. [Google Scholar] [CrossRef]
- Grund, T.; Heckele, M.; Kohl, M. Batch fabrication methods for polymer-based active microsystems using hot embossing and transfer bonding strategies. In Proceedings of Multi-Material Micro Manufacture, Cardiff, UK, 9-11 September 2008.
- Park, D.S.W.; Hupert, M.; Witek, M.; You, B.; Datta, P.; Guy, J.; Lee, J.B.; Soper, S.; Nikitopoulos, D.; Murphy, M. A titer plate-based polymer microfluidic platform for high throughput nucleic acid purification. Biomed.Microdevices 2008, 10, 21–33. [Google Scholar] [CrossRef]
- Park, D.S.W.; Chen, P.C.; You, B.H.; Kim, N.; Park, T.; Lee, T.Y.; Datta, P.; Desta, Y.; Soper, S.A.; Nikitopoulos, D.E. Titer plate formatted continuous flow thermal reactors for high throughput applications: Fabrication and testing. J. Micromech. Microeng. 2010, 20, 055003:1–055003:11. [Google Scholar]
- Hardt, D.E.; Hale, M. Development of a Low-Cost, Rapid-Cycle Hot Embossing System for Microscale Parts; Massachusetts Institute of Technology: Cambridge, MA, USA, 2009. Available online: http://hdl.handle.net/1721.1/50565 (accessed on 25 June 2012).
- Liu, S.J.; Huang, C.C.; Liao, C.T. Continuous infrared-assisted double-sided roll-to-roll embossing of flexible polymer substrates. Polym. Eng. Sci. 2012, 52, 1395–1401. [Google Scholar] [CrossRef]
- Su, Y.C.; Shah, J.; Lin, L. Implementation and analysis of polymeric microstructure replication by micro injection molding. J. Micromech. Microeng. 2004, 14, 415–422. [Google Scholar] [CrossRef]
- Yao, D.; Kim, B. Development of rapid heating and cooling systems for injection molding applications. Polym. Eng. Sci. 2002, 42, 2471–2481. [Google Scholar] [CrossRef]
- Lee, B.K.; Kwon, T.H. A Novel monolithic fabrication method for a plastic microfluidic chip with liquid interconnecting ports. J. Micromech. Microeng. 2010, 20, 105004:1–105004:11. [Google Scholar]
- Mecomber, J.S.; Stalcup, A.M.; Hurd, D.; Halsall, H.B.; Heineman, W.R.; Seliskar, C.J.; Wehmeyer, K.R.; Limbach, P.A. Analytical performance of polymer-based microfluidic devices fabricated by computer numerical controlled machining. Anal. Chem. 2006, 78, 936–941. [Google Scholar]
- Thiel, A.; Scheffold, A.; Radbruch, A. Immunomagnetic cell sorting—pushing the limits. Immunotechnology 1998, 4, 89–96. [Google Scholar] [CrossRef]
- Luo, X.; Cheng, K.; Webb, D.; Wardle, F. Design of ultraprecision machine tools with applications to manufacture of miniature and micro components. J. Mater. Process. Technol. 2005, 167, 515–528. [Google Scholar] [CrossRef]
- Okazaki, K. Micromachine tool to machine micro-parts. In Proceedings of the ASPE Annual Meeting, Scottsdale, AZ, USA, 22-27 October 2000; pp. 197–204.
- Miao, J.; Chen, G.; Lai, X.; Li, H.; Li, C. Review of dynamic issues in micro-end-milling. Int. J. Adv. Manuf. Technol. 2007, 31, 897–904. [Google Scholar] [CrossRef]
- Hupert, M.L.; Guy, W.J.; Llopis, S.D.; Shadpour, H.; Rani, S.; Nikitopoulos, D.E.; Soper, S.A. Evaluation of micromilled metal mold masters for the replication of microchip electrophoresis devices. Microfluid. Nanofluidics 2007, 3, 1–11. [Google Scholar]
- Wilding, P.; Kricka, L.J.; Cheng, J.; Hvichia, G.; Shoffner, M.A.; Fortina, P. Integrated cell isolation and polymerase chain reaction analysis using silicon microfilter chambers. Anal. Biochem. 1998, 257, 95–100. [Google Scholar]
- Park, T.; Zimmerman, T.J.; Park, D.; Lowrey, B.; Murphy, M.C. Thermoplastic fusion bonding of polymer-based micro devices using a pressure cooker. In Proceedings of the ASME IMECE, Lake Buena Vista, FL, USA, 13-19 November 2009; pp. 295–300.
- Sniadecki, N.J.; Lamb, C.M.; Liu, Y.; Chen, C.S.; Reich, D.H. Magnetic microposts for mechanical stimulation of biological cells: Fabrication, characterization, and analysis. Rev. Sci. Instrum. 2008, 79, 044302:1–044302:8. [Google Scholar]
- Häfeli, U. Scientific and Clinical Applications of Magnetic Carriers; Plenum Press: New York, NY, USA, 1997. [Google Scholar]
- Berger, M.; Castelino, J.; Huang, R.; Shah, M.; Austin, R.H. Design of a microfabricated magnetic cell separator. Electrophoresis 2001, 22, 3883–3892. [Google Scholar] [CrossRef]
- Lee, H.; Purdon, A.; Westervelt, R. Manipulation of biological cells using a microelectromagnet matrix. Appl. Phys. Lett. 2004, 85, 1063–1065. [Google Scholar]
- Miltenyi, S.; Müller, W.; Weichel, W.; Radbruch, A. High gradient magnetic cell separation with MACS. Cytometry 1990, 11, 231–238. [Google Scholar] [CrossRef]
- Toner, M.; Irimia, D. Blood-on-a-chip. Annu. Rev. Biomed. Eng. 2005, 7, 77–103. [Google Scholar] [CrossRef]
- Furdui, V.I.; Harrison, D.J. Immunomagnetic T cell capture from blood for PCR analysis using microfluidic systems. Lab Chip 2004, 4, 614–618. [Google Scholar] [CrossRef]
- Grodzinski, P.; Yang, J.; Liu, R.; Ward, M. A modular microfluidic system for cell pre-concentration and genetic sample preparation. Biomed. Microdevices 2003, 5, 303–310. [Google Scholar] [CrossRef]
- Inglis, D.W.; Riehn, R.; Austin, R.; Sturm, J. Continuous microfluidic immunomagnetic cell separation. Appl. Phys. Lett. 2004, 85, 5093–5095. [Google Scholar]
- Miltenyibiotec Homepage. Available online: http://www.miltenyibiotec.com (accessed on 28 February 2012).
- Zborowski, M.; Sun, L.; Moore, L.R.; Stephen Williams, P.; Chalmers, J.J. Continuous Cell separation using novel magnetic quadrupole flow sorter. J. Magn. Magn. Mater. 1999, 194, 224–230. [Google Scholar] [CrossRef]
- Murayama, A.; Furuta, T.; Hyomi, K.; Souma, I.; Oka, Y.; Dagnelund, D.; Buyanova, I.; Chen, W. Dynamics of exciton-spin injection, transfer, and relaxation in self-assembled quantum dots of cdse coupled with a diluted magnetic semiconductor layer of Zn0.80Mn0.20Se. Phys. Rev. B 2007, 75, 195308–195317. [Google Scholar]
- Han, K.H.; Frazier, A.B. Continuous magnetophoretic separation of blood cells in microdevice format. J. Appl. Phys. 2004, 96, 5797–5802. [Google Scholar]
- Zborowski, M.; Fuh, C.B.; Green, R.; Sun, L.; Chalmers, J.J. Analytical magnetapheresis of ferritin-labeled lymphocytes. Anal. Chem. 1995, 67, 3702–3712. [Google Scholar]
- Zborowski, M.; Ostera, G.R.; Moore, L.R.; Milliron, S.; Chalmers, J.J.; Schechter, A.N. Red blood cell magnetophoresis. Biophys. J. 2003, 84, 2638–2645. [Google Scholar] [CrossRef]
- Han, K.H.; Frazier, A.B. Paramagnetic capture mode magnetophoretic microseparator for high efficiency blood cell separations. Lab Chip 2006, 6, 265–273. [Google Scholar] [CrossRef]
- Pamme, N. Magnetism and microfluidics. Lab Chip 2006, 6, 24–38. [Google Scholar] [CrossRef]
- Pankhurst, Q.A.; Connolly, J.; Jones, S.; Dobson, J. Applications of magnetic nanoparticles in biomedicine. J. Phys. D 2003, 36, R167–R181. [Google Scholar] [CrossRef]
- Zhu, L.; Zhang, Q.; Feng, H.; Ang, S.; Chau, F.S.; Liu, W.T. Filter-based microfluidic device as a platform for immunofluorescent assay of microbial cells. Lab Chip 2004, 4, 337–341. [Google Scholar] [CrossRef]
- Mohamed, H.; McCurdy, L.D.; Szarowski, D.H.; Duva, S.; Turner, J.N.; Caggana, M. Development of a rare cell fractionation device: Application for cancer detection. IEEE Trans. NanoBiosci. 2004, 3, 251–256. [Google Scholar] [CrossRef]
- Moorthy, J.; Beebe, D.J. In situ fabricated porous filters for microsystems. Lab Chip 2003, 3, 62–66. [Google Scholar] [CrossRef]
- Huang, L.R.; Cox, E.C.; Austin, R.H.; Sturm, J.C. Continuous particle separation through deterministic lateral displacement. Science 2004, 304, 987–990. [Google Scholar]
- Khademhosseini, A.; Yeh, J.; Jon, S.; Eng, G.; Suh, K.Y.; Burdick, J.A.; Langer, R. Molded polyethylene glycol microstructures for capturing cells within microfluidic channels. Lab Chip 2004, 4, 425–430. [Google Scholar] [CrossRef]
- Tani, H.; Maehana, K.; Kamidate, T. Chip-based bioassay using bacterial sensor strains immobilized in three-dimensional microfluidic network. Anal. Chem. 2004, 76, 6693–6697. [Google Scholar] [CrossRef]
- Revzin, A.; Tompkins, R.G.; Toner, M. Surface engineering with poly(ethylene glycol) photolithography to create high-density cell arrays on glass. Langmuir 2003, 19, 9855–9862. [Google Scholar] [CrossRef]
- Chronis, N.; Lee, L.P. Electrothermally activated SU-8 microgripper for single cell manipulation in solution. J. Microelectromech. Syst. 2005, 14, 857–863. [Google Scholar] [CrossRef]
- Yang, J.; Li, C.W.; Yang, M. Hydrodynamic simulation of cell docking in microfluidic channels with different dam structures. Lab Chip 2004, 4, 53–59. [Google Scholar] [CrossRef]
- Li, C.W.; Cheung, C.N.; Yang, J.; Tzang, C.H.; Yang, M. PDMS-based microfluidic device with multi-height structures fabricated by single-step photolithography using printed circuit board as masters. Analyst 2003, 128, 1137–1142. [Google Scholar] [CrossRef]
- Lahann, J.; Balcells, M.; Lu, H.; Rodon, T.; Jensen, K.F.; Langer, R. Reactive polymer coatings: A first step toward surface engineering of microfluidic devices. Anal. Chem. 2003, 75, 2117–2122. [Google Scholar]
- Kirby, B.J.; Wheeler, A.R.; Zare, R.N.; Fruetel, J.A.; Shepodd, T.J. Programmable modification of cell adhesion and zeta potential in silica microchips. Lab Chip 2003, 3, 5–10. [Google Scholar] [CrossRef]
- Cox, J.D.; Curry, M.S.; Skirboll, S.K.; Gourley, P.L.; Sasaki, D.Y. Surface passivation of a microfluidic device to glial cell adhesion: A comparison of hydrophobic and hydrophilic SAM coatings. Biomaterials 2002, 23, 929–935. [Google Scholar] [CrossRef]
- Murthy, S.K.; Sin, A.; Tompkins, R.G.; Toner, M. Effect of flow and surface conditions on human lymphocyte isolation using microfluidic chambers. Langmuir 2004, 20, 11649–11655. [Google Scholar]
- Revzin, A.; Sekine, K.; Sin, A.; Tompkins, R.G.; Toner, M. Development of a microfabricated cytometry platform for characterization and sorting of individual leukocytes. Lab Chip 2005, 5, 30–37. [Google Scholar] [CrossRef]
- Chang, W.C.; Lee, L.P.; Liepmann, D. Biomimetic technique for adhesion-based collection and separation of cells in a microfluidic channel. Lab Chip 2005, 5, 64–73. [Google Scholar] [CrossRef]
- Chen, T.; Small, D.A.; McDermott, M.K.; Bentley, W.E.; Payne, G.F. Enzymatic methods for in situ cell entrapment and cell release. Biomacromolecules 2003, 4, 1558–1563. [Google Scholar] [CrossRef]
- Carlson, R.H.; Gabel, C.V.; Chan, S.S.; Austin, R.H.; Brody, J.P.; Winkelman, J.W. Self-sorting of white blood cells in a lattice. Phys. Rev. Lett. 1997, 79, 2149–2152. [Google Scholar]
- Bakajin, O.; Carlson, R.; Chou, C.; Chan, S.; Gabel, C.; Knight, J.; Cox, T.; Austin, R. Sizing, fractionation and mixing of biological objects via microfabricated devices. In Proceedings of Micro Total Analysis Systems, Banff, AB, Canada, 13-16 October 1998; pp. 193–198.
- Andersson, H.; van der Wijngaart, W.; Enoksson, P.; Stemme, G. Micromachined flow-through filter-chamber for chemical reactions on beads. Sens. Actuators B Chem. 2000, 67, 203–208. [Google Scholar] [CrossRef]
- Gossett, D.R.; Weaver, W.M.; Mach, A.J.; Hur, S.C.; Tse, H.T.K.; Lee, W.; Amini, H.; Di Carlo, D. Label-free cell separation and sorting in microfluidic systems. Anal. Bioanal. Chem. 2010, 397, 3249–3267. [Google Scholar] [CrossRef]
- Ji, H.M.; Samper, V.; Chen, Y.; Heng, C.K.; Lim, T.M.; Yobas, L. Silicon-based microfilters for whole blood cell separation. Biomed. Microdevices 2008, 10, 251–257. [Google Scholar] [CrossRef]
- Rusu, C.; van Oever, R.; de Boer, M.J.; Jansen, H.V.; Berenschot, J.; Bennink, M.L.; Kanger, J.S.; de Grooth, B.G.; Elwenspoek, M.; Greve, J. Direct integration of micromachined pipettes in a flow channel for single DNA molecule study by optical tweezers. J. Microelectromech. Syst. 2001, 10, 238–246. [Google Scholar] [CrossRef]
- Dennison, C. Principles of electrophoresis. In A Guide to Protein Isolation; Springer: Berlin, Germany, 2003; pp. 115–149. [Google Scholar]
- Manz, A.; Effenhauser, C.S.; Burggraf, N.; Harrison, D.J.; Seiler, K.; Fluri, K. Electroosmotic pumping and electrophoretic separations for miniaturized chemical analysis systems. J. Micromech. Microeng. 1994, 4, 257–265. [Google Scholar] [CrossRef]
- Ramsey, J.M.; Jacobson, S.C.; Knapp, M.R. Microfabricated chemical measurement systems. Nat. Med. 1995, 1, 1093–1095. [Google Scholar] [CrossRef]
- Fu, A.Y.; Spence, C.; Scherer, A.; Arnold, F.H.; Quake, S.R. A microfabricated fluorescence-activated cell sorter. Nat. Biotechnol. 1999, 17, 1109–1111. [Google Scholar] [CrossRef]
- Umehara, S.; Wakamoto, Y.; Inoue, I.; Yasuda, K. On-chip single-cell microcultivation assay for monitoring environmental effects on isolated cells. Biochem. Biophys. Res. Commun. 2003, 305, 534–540. [Google Scholar] [CrossRef]
- Enger, J.; Goksör, M.; Ramser, K.; Hagberg, P.; Hanstorp, D. Optical tweezers applied to a microfluidic system. Lab Chip 2004, 4, 196–200. [Google Scholar] [CrossRef]
- Chou, C.F.; Tegenfeldt, J.O.; Bakajin, O.; Chan, S.S.; Cox, E.C.; Darnton, N.; Duke, T.; Austin, R.H. Electrodeless dielectrophoresis of single-and double-stranded DNA. Biophys. J. 2002, 83, 2170–2179. [Google Scholar] [CrossRef]
- Fiedler, S.; Shirley, S.G.; Schnelle, T.; Fuhr, G. Dielectrophoretic sorting of particles and cells in a microsystem. Anal. Chem. 1998, 70, 1909–1915. [Google Scholar]
- Markx, G.H.; Huang, Y.; Zhou, X.F.; Pethig, R. Dielectrophoretic characterization and separation of micro-organisms. Microbiology 1994, 140, 585–591. [Google Scholar] [CrossRef]
- Cheng, J.; Sheldon, E.L.; Wu, L.; Uribe, A.; Gerrue, L.O.; Carrino, J.; Heller, M.J.; O’Connell, J.P. Preparation and hybridization analysis of DNA/RNA from E. Coli on microfabricated bioelectronic chips. Nat. Biotechnol. 1998, 16, 541–546. [Google Scholar]
- Becker, F.F.; Wang, X.B.; Huang, Y.; Pethig, R.; Vykoukal, J.; Gascoyne, P. Separation of human breast cancer cells from blood by differential dielectric affinity. Proc. Natl. Acad. Sci. USA 1995, 92, 860–864. [Google Scholar]
- Wang, X.B.; Yang, J.; Huang, Y.; Vykoukal, J.; Becker, F.F.; Gascoyne, P.R.C. Cell separation by dielectrophoretic field-flow-fractionation. Anal. Chem. 2000, 72, 832–839. [Google Scholar]
- Xu, J.; Wu, L.; Wang, X. Functional biochips for cell and molecular manipulation. Micro Total Anal. Syst. 2001, 313–314. [Google Scholar]
- Huang, Y.; Joo, S.; Duhon, M.; Heller, M.; Wallace, B.; Xu, X. Dielectrophoretic cell separation and gene expression profiling on microelectronic chip arrays. Anal. Chem. 2002, 74, 3362–3371. [Google Scholar] [CrossRef]
- Voldman, J.; Gray, M.L.; Toner, M.; Schmidt, M.A. A microfabrication-based dynamic array cytometer. Anal. Chem. 2002, 74, 3984–3990. [Google Scholar] [CrossRef]
- Huang, Y.; Ewalt, K.L.; Tirado, M.; Haigis, R.; Forster, A.; Ackley, D.; Heller, M.J.; O’Connel, J.P.; Krihak, M. Electric manipulation of bioparticles and macromolecules on microfabricated electrodes. Anal. Chem. 2001, 73, 1549–1559. [Google Scholar]
- Dürr, M.; Kentsch, J.; Müller, T.; Schnelle, T.; Stelzle, M. Microdevices for manipulation and accumulation of micro-and nanoparticles by dielectrophoresis. Electrophoresis 2003, 24, 722–731. [Google Scholar] [CrossRef]
- Chou, C.F.; Morgan, M.; Zenhausern, F.; Prinz, C.; Austin, R.H. Electrodeless dielectrophoretic trapping and separation of cells. Micro Total Anal. Syst. 2002, 1, 25–27. [Google Scholar]
- Arai, F.; Ichikawa, A.; Ogawa, M.; Fukuda, T.; Horio, K.; Itoigawa, K. High-speed separation system of randomly suspended single living cells by laser trap and dielectrophoresis. Electrophoresis 2001, 22, 283–288. [Google Scholar] [CrossRef]
- Cui, L.; Holmes, D.; Morgan, H. The dielectrophoretic levitation and separation of latex beads in microchips. Electrophoresis 2001, 22, 3893–3901. [Google Scholar]
- Li, H.; Bashir, R. Dielectrophoretic separation and manipulation of live and heat-treated cells of listeria on microfabricated devices with interdigitated electrodes. Sens. Actuators B Chem. 2002, 86, 215–221. [Google Scholar]
- Li, Y.; Kaler, K.V.I.S. Dielectrophoretic fluidic cell fractionation system. Anal. Chim. Acta 2004, 507, 151–161. [Google Scholar] [CrossRef]
- Seger, U.; Gawad, S.; Johann, R.; Bertsch, A.; Renaud, P. Cell immersion and cell dipping in microfluidic devices. Lab Chip 2004, 4, 148–151. [Google Scholar] [CrossRef]
- Hultström, J.; Manneberg, O.; Dopf, K.; Hertz, H.; Brismar, H.; Wiklund, M. Proliferation and viability of adherent cells manipulated by standing-wave ultrasound in a microfluidic chip. Ultrasound Med. Biol. 2007, 33, 145–151. [Google Scholar] [CrossRef]
- Bazou, D.; Kuznetsova, L.A.; Coakley, W.T. Physical enviroment of 2-D animal cell aggregates formed in a short pathlength ultrasound standing wave trap. Ultrasound Med. Biol. 2005, 31, 423–430. [Google Scholar] [CrossRef]
- Evander, M.; Johansson, L.; Lilliehorn, T.; Piskur, J.; Lindvall, M.; Johansson, S.; Almqvist, M.; Laurell, T.; Nilsson, J. Noninvasive acoustic cell trapping in a microfluidic perfusion system for online bioassays. Anal. Chem. 2007, 79, 2984–2991. [Google Scholar]
- Liu, J.; Kuznetsova, L.A.; Edwards, G.O.; Xu, J.; Ma, M.; Purcell, W.M.; Jackson, S.K.; Coakley, W.T. Functional three-dimensional HepG2 aggregate cultures generated from an ultrasound trap: Comparison with HepG2 spheroids. J. Cell. Biochem. 2007, 102, 1180–1189. [Google Scholar] [CrossRef]
- Bazou, D.; Blain, E.J.; Coakley, W.T. NCAM and PSA-NCAM dependent membrane spreading and F-Actin reorganization in suspended adhering neural cells. Mol. Membr. Biol. 2008, 25, 102–114. [Google Scholar] [CrossRef]
- Edwards, G.O.; Bazou, D.; Kuznetsova, L.A.; Coakley, W.T. Cell adhesion dynamics and actin cytoskeleton reorganization in HepG2 cell aggregates. Cell Commun. Adhes. 2007, 14, 9–20. [Google Scholar] [CrossRef]
- Gherardini, L.; Cousins, C.M.; Hawkes, J.J.; Spengler, J.; Radel, S.; Lawler, H.; Devcic-Kuhar, B.; Gröschl, M.; Coakley, W.T.; McLoughlin, A.J. A new immobilisation method to arrange particles in a gel matrix by ultrasound standing waves. Ultrasound Med. Biol. 2005, 31, 261–272. [Google Scholar] [CrossRef]
- Bazou, D.; Coakley, W.T.; Hayes, A.; Jackson, S.K. Long-term viability and proliferation of alginate-encapsulated 3-D HepG2 aggregates formed in an ultrasound trap. Toxicol. in vitro 2008, 22, 1321–1331. [Google Scholar] [CrossRef]
- Ruedas-Rama, M.J.; Domínguez-Vidal, A.; Radel, S.; Lendl, B. Ultrasonic trapping of microparticles in suspension and reaction monitoring using raman microspectroscopy. Anal. Chem. 2007, 79, 7853–7857. [Google Scholar]
- Ashkin, A.; Dziedzic, J.; Yamane, T. Optical trapping and manipulation of single cells using infrared laser beams. Nature 1987, 330, 769–771. [Google Scholar]
- Arai, F.; Ng, C.; Maruyama, H.; Ichikawa, A.; El-Shimy, H.; Fukuda, T. On chip single-cell separation and immobilization using optical tweezers and thermosensitive hydrogel. Lab Chip 2005, 5, 1399–1403. [Google Scholar] [CrossRef]
- Kovac, J.; Voldman, J. Intuitive, image-based cell sorting using optofluidic cell sorting. Anal. Chem. 2007, 79, 9321–9330. [Google Scholar] [CrossRef]
- Arai, Y.; Yasuda, R.; Akashi, K.; Harada, Y.; Miyata, H.; Kinosita, K.; Itoh, H. Tying a molecular knot with optical tweezers. Nature 1999, 399, 446–448. [Google Scholar]
- Smith, S.B.; Cui, Y.; Bustamante, C. Overstretching B-DNA: The elastic response of individual double-stranded and single-stranded DNA molecules. Science 1996, 271, 795–799. [Google Scholar]
- Rodrigo, P.; Eriksen, R.; Daria, V.; Glueckstad, J. Interactive light-driven and parallel manipulation of inhomogeneous particles. Opt. Express 2002, 10, 1550–1556. [Google Scholar]
- Flynn, R.A.; Birkbeck, A.L.; Gross, M.; Ozkan, M.; Shao, B.; Wang, M.M.; Esener, S.C. Parallel transport of biological cells using individually addressable VCSEL arrays as optical tweezers. Sens. Actuators B Chem. 2002, 87, 239–243. [Google Scholar] [CrossRef]
- Birkbeck, A.L.; Flynn, R.A.; Ozkan, M.; Song, D.; Gross, M.; Esener, S.C. VCSEL arrays as micromanipulators in chip-based biosystems. Biomed. Microdevices 2003, 5, 47–54. [Google Scholar] [CrossRef]
- Ozkan, M.; Wang, M.; Ozkan, C.; Flynn, R.; Esener, S. Optical manipulation of objects and biological cells in microfluidic devices. Biomed. Microdevices 2003, 5, 61–67. [Google Scholar] [CrossRef]
- MacDonald, M.; Spalding, G.; Dholakia, K. Microfluidic sorting in an optical lattice. Nature 2003, 426, 421–424. [Google Scholar]
- Ramser, K.; Wenseleers, W.; Dewilde, S.; van Doorslaer, S.; Moens, L. The combination of resonance raman spectroscopy, optical tweezers and microfluidic systems applied to the study of various heme-containing single cells. Spectroscopy 2008, 22, 287–295. [Google Scholar]
- Alrifaiy, A.; Ramser, K. How to integrate a micropipette into a closed microfluidic system: Absorption spectra of an optically trapped erythrocyte. Biomed. Opt. Express 2011, 2, 2299–2306. [Google Scholar] [CrossRef]
- Moldavan, A. Photo-electric technique for the counting of microscopical cells. Science 1934, 80, 188–189. [Google Scholar]
- Crosland-Taylor, P. A device for counting small particles suspended in a fluid through a tube. Nature 1953, 171, 37–38. [Google Scholar] [CrossRef]
- Coulter, W.H. High speed automatic blood cell counter and cell size analyzer. Proc. Natl. Electron. Conf. 1956, 12, 1034–1042. [Google Scholar]
- Fulwyler, M.J. Electronic separation of biological cells by volume. Science 1965, 150, 910–911. [Google Scholar]
- Kamentsky, L.A.; Melamed, M.R.; Derman, H. Spectrophotometer: New instrument for ultrarapid cell analysis. Science 1965, 150, 630–631. [Google Scholar]
- Kamentsky, L.A.; Melamed, M.R. Spectrophotometric cell sorter. Science 1967, 156, 1364. [Google Scholar]
- Melamed, M.; Mullaney, P.; Shapiro, H. An historical review of the development of Ow cytometry and sorters. Flow Cytometry Sorting 1979, 1–10. [Google Scholar]
- Shapiro, H.M.; Leif, R.C. Practical Flow Cytometry; Wiley-Liss: Wilmington, DE, USA, 2003. [Google Scholar]
- Lindmo, T.; Peters, D.; Sweet, R. Flow Sorters for Biological Cells. Flow Cytometry and Sorting, 2nd ed; Wiley-Liss: New York, NY, USA, 1990; pp. 145–169. [Google Scholar]
- Herzenberg, L.A.; Herzenberg, L.A. Genetics, FACS, immunology, and redox: A tale of two lives intertwined. Annu. Rev. Immunol. 2004, 22, 1–31. [Google Scholar] [CrossRef]
- Ormerod, M.G. Flow Cytometry–A Basic Introduction; De Novo software: Los Angeles, CA, USA, 2008. Available online: http://flowbook.denovosoftware.com/ (accessed on 28 February 2012).
- El-Naggar, A.K. Concurrent flow cytometric analysis of DNA and RNA. Methods Mol. Biol. 2004, 263, 371–384. [Google Scholar]
- Rabinovitch, P.; June, C. Intracellular ionized calcium, magnesium, membrane potential, and pH. In Flow Cytometry: A Practical Approach; Oxford University Press: New York, NY, USA, 2000; pp. 203–234. [Google Scholar]
- Darzynkiewicz, Z.; Juan, G.; Li, X.; Gorczyca, W.; Murakami, T.; Traganos, F. Cytometry in cell necrobiology: Analysis of apoptosis and accidental cell death (Necrosis). Cytometry 1997, 27, 1–20. [Google Scholar] [CrossRef]
- Pruitt, S.C.; Mielnicki, L.M.; Stewart, C.C. Analysis of fluorescent protein expressing cells by flow cytometry. Methods Mol. Biol. 2004, 263, 239–258. [Google Scholar]
- Edwards, A.D.; Manickasingham, S.P.; Spörri, R.; Diebold, S.S.; Schulz, O.; Sher, A.; Kaisho, T.; Akira, S.; Reis, E.S.C. Microbial recognition via toll-like receptor-dependent and-independent pathways determines the cytokine response of murine dendritic cell subsets to CD40 triggering. J. Immunol. 2002, 169, 3652–3660. [Google Scholar]
- Suzuki, A.; Zheng, Y.; Fukao, K.; Nakauchi, H.; Taniguchi, H. Liver repopulation by c-Met-Positive stem/progenitor cells isolated from the developing rat liver. Hepatogastroenterology 2004, 51, 423–426. [Google Scholar]
- Bomberger, C.; Singh-Jairam, M.; Rodey, G.; Guerriero, A.; Yeager, A.M.; Fleming, W.H.; Holland, H.K.; Waller, E.K. Lymphoid reconstitution after autologous PBSC transplantation with FACS-sorted CD34 hematopoietic progenitors. Blood 1998, 91, 2588–2600. [Google Scholar]
- Cran, D.; Johnson, L. The predetermination of embryonic sex using flow cytometrically separated X and Y spermatozoa. Hum. Reprod. Update 1996, 2, 355–363. [Google Scholar] [CrossRef]
- Gift, E.A.; Park, H.J.; Paradis, G.A.; Demain, A.L.; Weaver, J.C. FACS-based isolation of slowly growing cells: Double encapsulation of yeast in gel microdrops. Nat. Biotechnol. 1996, 14, 884–887. [Google Scholar] [CrossRef]
- Deere, D.; Shen, J.; Vesey, G.; Bell, P.; Bissinger, P.; Veal, D. Flow cytometry and cell sorting for yeast viability assessment and cell selection. Yeast 1998, 14, 147–160. [Google Scholar] [CrossRef]
- Wallner, G.; Fuchs, B.; Spring, S.; Beisker, W.; Amann, R. Flow sorting of microorganisms for molecular analysis. Appl. Environ. Microbiol. 1997, 63, 4223–4231. [Google Scholar]
- Sekar, R.; Fuchs, B.M.; Amann, R.; Pernthaler, J. Flow sorting of marine bacterioplankton after fluorescence in situ hybridization. Appl. Environ. Microbiol. 2004, 70, 6210–6219. [Google Scholar] [CrossRef]
- Jochem, F.J. Short-term physiologic effects of mechanical flow sorting and the becton-dickinson cell concentrator in cultures of the marine phytoflagellata emiliania huxleyi and micromonas pusilla. Cytom. A 2005, 65, 77–83. [Google Scholar] [CrossRef]
- Guasch, R.; Guerri, C.; O’Connor, J. Flow cytometric analysis of concanavalin A binding to isolated golgi fractions from rat liver. Exp. Cell Res. 1993, 207, 136–141. [Google Scholar] [CrossRef]
- Dolezel, J.; Kubalakova, M.; Cihalikova, J.; Suchankova, P.; Simkova, H. Chromosome analysis and sorting using flow cytometry. Methods Mol. Biol. 2011, 701, 221–238. [Google Scholar] [CrossRef]
- Ferguson-Smith, M.; Yang, F.; Rens, W.; O’Brien, P. the impact of chromosome sorting and painting on the comparative analysis of primate genomes. Cytogenet. Genome Res. 2005, 108, 112–121. [Google Scholar] [CrossRef]
- Doležel, J.; Kubalakova, M.; Bartoš, J.; Macas, J. Flow cytogenetics and plant genome mapping. Chromosome Res. 2004, 12, 77–91. [Google Scholar] [CrossRef]
- Avni, R.D.; Miron, A.; Mok, S.; Randrianarison, V. Generation of chromo-some paints: approach for increasing specificity and intensity of signals. Biotechniques 2003, 34, 530–536. [Google Scholar]
- Szaniszlo, P.; Wang, N.; Sinha, M.; Reece, L.M.; van Hook, J.W.; Luxon, B.A.; Leary, J.F. Getting the right cells to the array: Gene expression microarray analysis of cell mixtures and sorted cells. Cytom. A 2004, 59, 191–202. [Google Scholar]
- Battye, F.L.; Light, A.; Tarlinton, D.M. Single cell sorting and cloning. J. Immunol. Methods 2000, 243, 25–32. [Google Scholar] [CrossRef]
- Williams, C.; Davies, D.; Williamson, R. Segregation of ΔF508 and normal CFTR alleles in human sperm. Hum. Mol. Genet. 1993, 2, 445–448. [Google Scholar] [CrossRef]
- Telleman, P.; Larsen, U.; Philip, J.; Blankenstein, G.; Wolff, A. Cell sorting in microfluidic systems. In Proceedings of Micro Total Analysis Systems, Banff, AB, Canada, 13-16 October 1998; pp. 39–44.
- Blankenstein, G.; Darling Larsen, U. Modular concept of a laboratory on a chip for chemical and biochemical analysis. Biosens. Bioelectron. 1998, 13, 427–438. [Google Scholar] [CrossRef]
- McClain, M.A.; Culbertson, C.T.; Jacobson, S.C.; Ramsey, J.M. Flow cytometry of escherichia coli on microfluidic devices. Anal. Chem. 2001, 73, 5334–5338. [Google Scholar]
- Müller, T.; Gradl, G.; Howitz, S.; Shirley, S.; Schnelle, T.; Fuhr, G. A 3-D Microelectrode system for handling and caging single cells and particles. Biosens. Bioelectron. 1999, 14, 247–256. [Google Scholar] [CrossRef]
- Fu, A.Y.; Chou, H.P.; Spence, C.; Arnold, F.H.; Stephen, R. An integrated microfabricated cell sorter. Anal. Chem. 2002, 74, 2451–2457. [Google Scholar]
- Gradl, G.; Müller, T.; Pfennig, A.; Shirley, S.; Schnelle, T.; Führ, G. New micro device for single cell analysis, cell sorting and cloning on a chip: The cytocon instrument. Micro Total Anal. 2000, 443–446. [Google Scholar]
- Muller, T.; Schnelle, T.; Gradl, T.; Pfennig, A.; Fuhr, G. Live cells in cell processors. Bioworld 2002, 2, 12–13. [Google Scholar]
- Gawad, S.; Metz, S.; Laurent, S.; Renaud, P. Impedance spectroscopy cell analysis in microchannels. Micro Total Anal. Syst. 2002, 253–255. [Google Scholar]
- Nieuwenhuis, J.; Vellekoop, M.J. FEM study of coulter counter with water-based adaptable aperture. Micro Total Anal. Syst. 2002, 67–69. [Google Scholar]
- Hayenga, J.; Bardell, R.; Morris, C.; Graham, P.; Kesler, C.; Lancaster, A.; Padmanabhan, A.; Cabuz, C.; Schulte, T.; Weigl, B.; et al. Enabling technologies for a personal flow cytometer. Part II. Integrated analysis cartridges. In Proceedings of the Mocro Total Analysis Systems 2002 Symposium, Nara, Japan, 3-7 November 2002.
- Kruger, J.; Porta, P.; Morrison, A.; Singh, K.; O’Neill, A.; O’Brien, P. Micro-optical laser induced fluorescence detection on a miniaturized flow cytometry device. In Proceedings of the Mocro Total Analysis Systems 2002 Symposium, Nara, Japan, 3-7 November 2002.
- Glasgow, I.; Zeringue, H.; Beebe, D.; Choi, S.; Lyman, J.; Wheeler, M. Individual embryo transport and retention on a chip. In Proceedings of the Mocro Total Analysis Systems 2000 Symposium,, Ensched, The Netherlans, 14-18 May 2000.
- Kricka, L.; Nozaki, O.; Heyner, S.; Garside, W.; Wilding, P. Applications of a microfabricated device for evaluating sperm function. Clin. Chem. 1993, 39, 1944–1947. [Google Scholar]
- Tracey, M.; Johnston, I.; Greenaway, R.; Davis, J.; Sutton, N.; Doetzel, W. Microfluidics based microcytometer: Interfacing microfluidics with macrofluidics. In Proceedings of the Mocro Total Analysis Systems 1998 Symposium, Banff, Canada, 13-16 October 1998.
- Takayama, S.; Ostuni, E.; Qian, X.; McDonald, J.C.; Jiang, X.; LeDuc, P.; Wu, M.H.; Ingber, D.E.; Whitesides, G.M. Topographical micropatterning of poly(dimethylsiloxane) using laminar flows of liquids in capillaries. Adv. Mater. 2001, 13, 570–574. [Google Scholar] [CrossRef]
- Turner, A.; Dowell, N.; Turner, S.; Kam, L.; Isaacson, M.; Turner, J.; Craighead, H.; Shain, W. Attachment of astroglial cells to microfabricated pillar arrays of different geometries. J. Biomed. Mater. Res. 2000, 51, 430–441. [Google Scholar] [CrossRef]
- Lu, H.; Koo, L.; Griffith, L.; Jensen, K. Development of microfluidic shear assays for quantitative analysis of cell adhesion. Micro Total Anal. Syst. 2002, 784–786. [Google Scholar]
- Monaghan, P.; Manz, A.; Nichols, W. Microbiology on a chip. In Proceedings of the Mocro Total Analysis Systems 2000 Symposium, Ensched, The Netherlans, 14-18 May 2000.
- Tamaki, E.; Sato, K.; Tokeshi, M.; Sato, K.; Aihara, M.; Kitamori, T. Single-cell analysis by a scanning thermal lens microscope with a microchip: Direct monitoring of cytochrome C distribution during apoptosis process. Anal. Chem. 2002, 74, 1560–1564. [Google Scholar]
- DeBusschere, B.D.; Kovacs, G.T.A. Portable cell-based biosensor system using integrated CMOS cell-cartridges. Biosens. Bioelectron. 2001, 16, 543–556. [Google Scholar] [CrossRef]
- Hediger, S.; Fontannaz, J.; Sayah, A.; Hunziker, W.; Gijs, M.A.M. Biosystem for the culture and characterization of epithelial cell tissues. Sens. Actuators B Chem. 2000, 63, 63–73. [Google Scholar] [CrossRef]
- Hediger, S.; Sayah, A.; Horisberger, J.D.; Gijs, M.A.M. Modular microsystem for epithelial cell culture and electrical characterisation. Biosens. Bioelectron. 2001, 16, 689–694. [Google Scholar] [CrossRef]
- Heuschkel, M.O.; Guerin, L.; Buisson, B.; Bertrand, D.; Renaud, P. Buried microchannels in photopolymer for delivering of solutions to neurons in a network. Sens. Actuators B Chem. 1998, 48, 356–361. [Google Scholar] [CrossRef]
- Xu, J.; Wang, X.; Ensign, B.; Li, M.; Wu, L.; Guia, A.; Xu, J. Ion-channel assay technologies: Quo vadis? Drug Discov. Today 2001, 6, 1278–1287. [Google Scholar] [CrossRef]
- Schmidt, C.; Mayer, M.; Vogel, H. A chip-based biosensor for the functional analysis of single ion channels. Angew. Chem. 2000, 112, 3267–3270. [Google Scholar]
- Lehnert, T.; Gijs, M.A.M.; Netzer, R.; Bischoff, U. Realization of hollow SiO2 micronozzles for electrical measurements on living cells. Appl. Phys. Lett. 2002, 81, 5063–5065. [Google Scholar]
- Han, A.; Moss, E.; Rabbitt, R.; Frazier, B. A multi-purpose micro system for electrophysiological analyses of single cells. In Proceedings of the Mocro Total Analysis Systems 2002 Symposium, Nara, Japan, 3-7 November 2002.
- Farinas, J.; Chow, A.W.; Wada, H.G. A microfluidic device for measuring cellular membrane potential. Anal. Biochem. 2001, 295, 138–142. [Google Scholar]
- Walker, G.M.; Zeringue, H.C.; Beebe, D.J. Microenvironment design considerations for cellular scale studies. Lab Chip 2004, 4, 91–97. [Google Scholar] [CrossRef]
- Mahoney, M.J.; Chen, R.R.; Tan, J.; Mark, S.W. The influence of microchannels on neurite growth and architecture. Biomaterials 2005, 26, 771–778. [Google Scholar]
- Walker, G.; Ozers, M.; Beebe, D. Insect cell culture in microfluidic channels. Biomed. Microdevices 2002, 4, 161–166. [Google Scholar] [CrossRef]
- Li, N.; Tourovskaia, A.; Folch, A. Biology on a chip: Microfabrication for studying the behavior of cultured cells. Crit. Rev. Biomed. Eng. 2003, 31, 423–488. [Google Scholar] [CrossRef]
- Raty, S.; Walters, E.M.; Davis, J.; Zeringue, H.; Beebe, D.J.; Rodriguez-Zas, S.L.; Wheeler, M.B. Embryonic development in the mouse is enhanced via microchannel culture. Lab Chip 2004, 4, 186–190. [Google Scholar] [CrossRef]
- Leclerc, E.; Sakai, Y.; Fujii, T. Perfusion culture of fetal human hepatocytes in microfluidic environments. Biochem. Eng. J. 2004, 20, 143–148. [Google Scholar] [CrossRef]
- Groisman, A.; Lobo, C.; Cho, H.J.; Campbell, J.K.; Dufour, Y.S.; Stevens, A.M.; Levchenko, A. A microfluidic chemostat for experiments with bacterial and yeast cells. Nat. Methods 2005, 2, 685–689. [Google Scholar]
- Balagaddé, F.K.; You, L.; Hansen, C.L.; Arnold, F.H.; Quake, S.R. Long-term monitoring of bacteria undergoing programmed population control in a microchemostat. Science 2005, 309, 137–140. [Google Scholar]
- Thompson, D.M.; King, K.R.; Wieder, K.J.; Toner, M.; Yarmush, M.L.; Jayaraman, A. Dynamic gene expression profiling using a microfabricated living cell array. Anal. Chem. 2004, 76, 4098–4103. [Google Scholar]
- Gómez-Sjöberg, R.; Leyrat, A.A.; Pirone, D.M.; Chen, C.S.; Stephen, R. Versatile, fully automated, microfluidic cell culture system. Anal. Chem. 2007, 79, 8557–8563. [Google Scholar]
- Toh, Y.C.; Zhang, C.; Zhang, J.; Khong, Y.M.; Chang, S.; Samper, V.D.; van Noort, D.; Hutmacher, D.W.; Yu, H. A novel 3D mammalian cell perfusion-culture system in microfluidic channels. Lab Chip 2007, 7, 302–309. [Google Scholar]
- Lee, P.; Lin, R.; Moon, J.; Lee, L.P. Microfluidic alignment of collagen fibers for in vitro cell culture. Biomed. Microdevices 2006, 8, 35–41. [Google Scholar] [CrossRef]
- Hung, P.J.; Lee, P.J.; Sabounchi, P.; Aghdam, N.; Lin, R.; Lee, L.P. A novel high aspect ratio microfluidic design to provide a stable and uniform microenvironment for cell growth in a high throughput mammalian cell culture array. Lab Chip 2005, 5, 44–48. [Google Scholar] [CrossRef]
- Taylor, A.M.; Rhee, S.W.; Tu, C.H.; Cribbs, D.H.; Cotman, C.W.; Jeon, N.L. Microfluidic multicompartment device for neuroscience research. Langmuir 2003, 19, 1551–1556. [Google Scholar]
- Thiébaud, P.; Lauer, L.; Knoll, W.; Offenhäusser, A. PDMS device for patterned application of microfluids to neuronal cells arranged by microcontact printing. Biosens. Bioelectron. 2002, 17, 87–93. [Google Scholar] [CrossRef]
- Rhee, S.W.; Taylor, A.M.; Tu, C.H.; Cribbs, D.H.; Cotman, C.W.; Jeon, N.L. Patterned cell culture inside microfluidic devices. Lab Chip 2005, 5, 102–107. [Google Scholar] [CrossRef]
- Zhu, X.; Chu, L.Y.; Chueh, B.; Shen, M.; Hazarika, B.; Phadke, N.; Takayama, S. Arrays of horizontally-oriented mini-reservoirs generate steady microfluidic flows for continuous perfusion cell culture and gradient generation. Analyst 2004, 129, 1026–1031. [Google Scholar] [CrossRef]
- Maharbiz, M.M.; Holtz, W.J.; Sharifzadeh, S.; Keasling, J.D.; Howe, R.T. A Microfabricated electrochemical oxygen generator for high-density cell culture arrays. J. Microelectromech. Syst. 2003, 12, 590–599. [Google Scholar] [CrossRef]
- Tourovskaia, A.; Figueroa-Masot, X.; Folch, A. Differentiation-on-a-chip: A microfluidic platform for long-term cell culture studies. Lab Chip 2005, 5, 14–19. [Google Scholar] [CrossRef]
- Prokop, A.; Prokop, Z.; Schaffer, D.; Kozlov, E.; Wikswo, J.; Cliffel, D.; Baudenbacher, F. NanoLiterBioReactor: Long-term mammalian cell culture at nanofabricated scale. Biomed. Microdevices 2004, 6, 325–339. [Google Scholar] [CrossRef]
- Kojima, K.; Moriguchi, H.; Hattori, A.; Kaneko, T.; Yasuda, K. Two-dimensional network formation of cardiac myocytes in agar microculture chip with 1480 nm infrared laser photo-thermal etching. Lab Chip 2003, 3, 292–296. [Google Scholar] [CrossRef]
- Moriguchi, H.; Wakamoto, Y.; Sugio, Y.; Takahashi, K.; Inoue, I.; Yasuda, K. An agar-microchamber cell-cultivation system: Flexible change of microchamber shapes during cultivation by photo-thermal etching. Lab Chip 2002, 2, 125–132. [Google Scholar] [CrossRef]
- Tan, W.; Desai, T.A. Layer-by-layer microfluidics for biomimetic three-dimensional structures. Biomaterials 2004, 25, 1355–1364. [Google Scholar] [CrossRef]
- Leclerc, E.; Furukawa, K.; Miyata, F.; Sakai, Y.; Ushida, T.; Fujii, T. Fabrication of microstructures in photosensitive biodegradable polymers for tissue engineering applications. Biomaterials 2004, 25, 4683–4690. [Google Scholar] [CrossRef]
- Sakai, Y.; Leclerc, E.; Fujii, T. Microfluidic cell-culture devices. In Lab-on-Chips Cellomics; Springer: Amsterdam, The Netherlands, 2004; pp. 299–318. [Google Scholar]
- Martin, K.; Henkel, T.; Baier, V.; Grodrian, A.; Schön, T.; Roth, M.; Köhler, J.M.; Metze, J. Generation of larger numbers of separated microbial populations by cultivation in segmented-flow microdevices. Lab Chip 2003, 3, 202–207. [Google Scholar] [CrossRef]
- Pearce, T.M.; Wilson, J.A.; Oakes, S.G.; Chiu, S.Y.; Williams, J.C. Integrated microelectrode array and microfluidics for temperature clamp of sensory neurons in culture. Lab Chip 2005, 5, 97–101. [Google Scholar] [CrossRef]
- Cluzel, P.; Surette, M.; Leibler, S. An ultrasensitive bacterial motor revealed by monitoring signaling proteins in single cells. Science 2000, 287, 1652–1655. [Google Scholar]
- Yamaguchi, Y.; Arakawa, T.; Takeda, N.; Edagawa, Y.; Shoji, S. Development of a poly-dimethylsiloxane microfluidic device for single cell isolation and incubation. Sens. Actuators B Chem. 2009, 136, 555–561. [Google Scholar] [CrossRef]
- Wheeler, A.R.; Throndset, W.R.; Whelan, R.J.; Leach, A.M.; Zare, R.N.; Liao, Y.H.; Farrell, K.; Manger, I.D.; Daridon, A. Microfluidic device for single-cell analysis. Anal. Chem. 2003, 75, 3581–3586. [Google Scholar]
- Di Carlo, D.; Wu, L.Y.; Lee, L.P. Dynamic single cell culture array. Lab Chip 2006, 6, 1445–1449. [Google Scholar] [CrossRef]
- Lee, P.J.; Hung, P.J.; Shaw, R.; Jan, L.; Lee, L.P. Microfluidic application-specific integrated device for monitoring direct cell-cell communication via gap junctions between individual cell pairs. Appl. Phys. Lett. 2005, 86, 223902:1–223902:3. [Google Scholar]
- Kang, L.; Chung, B.G.; Langer, R.; Khademhosseini, A. Microfluidics for drug discovery and development: From target selection to product lifecycle management. Drug Discov. Today 2008, 13, 1–13. [Google Scholar] [CrossRef]
- Zheng, B.; Tice, J.D.; Ismagilov, R.F. Formation of arrayed droplets by soft lithography and two-phase fluid flow, and application in protein crystallization. Adv. Mater. 2004, 16, 1365–1368. [Google Scholar] [CrossRef]
- Zheng, W.; Spencer, R.H.; Kiss, L. High throughput assay technologies for ion channel drug discovery. Assay Drug Dev. Technol. 2004, 2, 543–552. [Google Scholar] [CrossRef]
- Dittrich, P.S.; Manz, A. Lab-on-a-chip: Microfluidics in drug discovery. Nat. Rev. Drug Discov. 2006, 5, 210–218. [Google Scholar] [CrossRef]
- Maerkl, S.J. Integration column: Microfluidic high-throughput screening. Integr. Biol. 2009, 1, 19–29. [Google Scholar] [CrossRef]
- Lii, J.; Hsu, W.J.; Parsa, H.; Das, A.; Rouse, R.; Sia, S.K. Real-time microfluidic system for studying mammalian cells in 3D microenvironments. Anal. Chem. 2008, 80, 3640–3647. [Google Scholar]
- Pregibon, D.C.; Toner, M.; Doyle, P.S. Multifunctional encoded particles for high-throughput biomolecule analysis. Science 2007, 315, 1393–1396. [Google Scholar]
- Hwang, D.K.; Oakey, J.; Toner, M.; Arthur, J.A.; Anseth, K.S.; Lee, S.; Zeiger, A.; van Vliet, K.J.; Doyle, P.S. Stop-flow lithography for the production of shape-evolving degradable microgel particles. J. Am. Chem. Soc. 2009, 131, 4499–4504. [Google Scholar]
- Brouzes, E.; Medkova, M.; Savenelli, N.; Marran, D.; Twardowski, M.; Hutchison, J.B.; Rothberg, J.M.; Link, D.R.; Perrimon, N.; Samuels, M.L. Droplet microfluidic technology for single-cell high-throughput screening. Proc. Natl. Acad. Sci. USA 2009, 106, 14195–14200. [Google Scholar]
- Chen, A.A.; Underhill, G.H.; Bhatia, S.N. Multiplexed, high-throughput analysis of 3D microtissue suspensions. Integr. Biol. 2010, 2, 517–527. [Google Scholar] [CrossRef]
- Choi, N.W.; Cabodi, M.; Held, B.; Gleghorn, J.P.; Bonassar, L.J.; Stroock, A.D. Microfluidic scaffolds for tissue engineering. Nat. Mater. 2007, 6, 908–915. [Google Scholar]
- Kumachev, A.; Greener, J.; Tumarkin, E.; Eiser, E.; Zandstra, P.W.; Kumacheva, E. High-throughput generation of hydrogel microbeads with varying elasticity for cell encapsulation. Biomaterials 2011, 32, 1477–1483. [Google Scholar]
- Trivedi, V.; Ereifej, E.S.; Doshi, A.; Sehgal, P.; VandeVord, P.J.; Basu, A.S. Microfluidic encapsulation of cells in alginate capsules for high throughput screening. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2009, 7037–7040. [Google Scholar]
- Choi, C.H.; Jung, J.H.; Rhee, Y.W.; Kim, D.P.; Shim, S.E.; Lee, C.S. Generation of monodisperse alginate microbeads and in situ encapsulation of cell in microfluidic device. Biomed. Microdevices 2007, 9, 855–862. [Google Scholar] [CrossRef]
- Chen, W.; Yang, Y.; Rinadi, C.; Zhou, D.; Shen, A.Q. Formation of supramolecular hydrogel microspheres via microfluidics. Lab Chip 2009, 9, 2947–2951. [Google Scholar] [CrossRef]
- Zeng, S.; Liu, X.; Xie, H.; Lin, B. Basic technologies for droplet microfluidics. Top. Curr. Chem. 2011, 304, 69–90. [Google Scholar] [CrossRef]
- Song, H.; Chen, D.L.; Ismagilov, R.F. Reactions in droplets in microfluidic channels. Angew. Chem. Int. Ed. 2006, 45, 7336–7356. [Google Scholar]
- Tan, Y.C.; Hettiarachchi, K.; Siu, M.; Pan, Y.R.; Lee, A.P. Controlled microfluidic encapsulation of cells, proteins, and microbeads in lipid vesicles. J. Am. Chem. Soc. 2006, 128, 5656–5658. [Google Scholar]
- Shah, R.K.; Shum, H.C.; Rowat, A.C.; Lee, D.; Agresti, J.J.; Utada, A.S.; Chu, L.Y.; Kim, J.W.; Fernandez-Nieves, A.; Martinez, C.J. Designer emulsions using microfluidics. Mater. Today 2008, 11, 18–27. [Google Scholar]
- Atencia, J.; Beebe, D.J. Controlled microfluidic interfaces. Nature 2005, 437, 648–655. [Google Scholar] [CrossRef]
- Gong, Z.; Penmetsa, S.; Zheng, Z.; Lvov, Y.; Que, L. Encapsulation of microparticles and biomolecules based on layer-by-layer nanoassembly techniques with microfluidic droplet devices. In Proceedings of Solid-State Sensors, Actuators and Microsystems Conference, TRANSDUCERS 2009, Denver, CO, USA, 21-25 June 2009; pp. 1043–1046.
- Gong, Z.; Zhao, H.; Zhang, T.; Nie, F.; Pathak, P.; Cui, K.; Wang, Z.; Wong, S.; Que, L. Drug effects analysis on cells using a high throughput microfluidic chip. Biomed. Microdevices 2011, 13, 215–219. [Google Scholar] [CrossRef]
- Häussinger, D. The role of cellular hydration in the regulation of cell function. Biochem. J. 1996, 313, 697. [Google Scholar]
- Hua, S.Z.; Pennell, T. A microfluidic chip for real-time studies of the volume of single cells. Lab Chip 2009, 9, 251–256. [Google Scholar] [CrossRef]
- El-Ali, J.; Sorger, P.K.; Jensen, K.F. Cells on chips. Nature 2006, 442, 403–411. [Google Scholar]
- Warrick, J.; Meyvantsson, I.; Ju, J.; Beebe, D.J. High-throughput microfluidics: Improved sample treatment and washing over standard wells. Lab Chip 2007, 7, 316–321. [Google Scholar] [CrossRef]
- Chen, D.L.; Ismagilov, R.F. microfluidic cartridges preloaded with nanoliter plugs of reagents: An alternative to 96-well plates for screening. Curr. Opin. Chem. Biol. 2006, 10, 226–231. [Google Scholar] [CrossRef]
- Clausell-Tormos, J.; Lieber, D.; Baret, J.C.; El-Harrak, A.; Miller, O.J.; Frenz, L.; Blouwolff, J.; Humphry, K.J.; Köster, S.; Duan, H. Droplet-based microfluidic platforms for the encapsulation and screening of mammalian cells and multicellular organisms. Chem. Biol. 2008, 15, 427–437. [Google Scholar] [CrossRef]
- Huebner, A.; Bratton, D.; Whyte, G.; Yang, M.; Abell, C.; Hollfelder, F. Static microdroplet arrays: A microfluidic device for droplet trapping, incubation and release for enzymatic and cell-based assays. Lab Chip 2009, 9, 692–698. [Google Scholar] [CrossRef]
- Huh, D.; Matthews, B.D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H.Y.; Ingber, D.E. Reconstituting organ-level lung functions on a chip. Science 2010, 328, 1662–1668. [Google Scholar]
- Discher, D.E.; Mooney, D.J.; Zandstra, P.W. Growth factors, matrices, and forces combine and control stem cells. Science 2009, 324, 1673–1677. [Google Scholar]
- Liu, J.; Gao, D.; Li, H.F.; Lin, J.M. Controlled photopolymerization of hydrogel microstructures inside microchannels for bioassays. Lab Chip 2009, 9, 1301–1305. [Google Scholar] [CrossRef]
- Cheung, Y.K.; Gillette, B.M.; Zhong, M.; Ramcharan, S.; Sia, S.K. Direct patterning of composite biocompatible microstructures using microfluidics. Lab Chip 2007, 7, 574–579. [Google Scholar] [CrossRef]
- Qi, H.; Du, Y.; Wang, L.; Kaji, H.; Bae, H.; Khademhosseini, A. Patterned differentiation of individual embryoid bodies in spatially organized 3D hybrid microgels. Adv. Mater. 2010, 22, 5276–5281. [Google Scholar]
- Bruzewicz, D.A.; McGuigan, A.P.; Whitesides, G.M. Fabrication of a modular tissue construct in a microfluidic chip. Lab Chip 2008, 8, 663–671. [Google Scholar] [CrossRef]
- Pautot, S.; Wyart, C.; Isacoff, E.Y. Colloid-guided assembly of oriented 3D neuronal networks. Nat. Methods 2008, 5, 735–740. [Google Scholar]
- Chung, S.; Sudo, R.; Mack, P.J.; Wan, C.R.; Vickerman, V.; Kamm, R.D. Cell migration into scaffolds under co-culture conditions in a microfluidic platform. Lab Chip 2009, 9, 269–275. [Google Scholar] [CrossRef]
- Huang, C.P.; Lu, J.; Seon, H.; Lee, A.P.; Flanagan, L.A.; Kim, H.Y.; Putnam, A.J.; Jeon, N.L. Engineering microscale cellular niches for three-dimensional multicellular co-cultures. Lab Chip 2009, 9, 1740–1748. [Google Scholar]
- Carrion, B.; Huang, C.P.; Ghajar, C.M.; Kachgal, S.; Kniazeva, E.; Jeon, N.L.; Putnam, A.J. Recreating the perivascular niche ex vivo using a microfluidic approach. Biotechnol. Bioeng. 2010, 107, 1020–1028. [Google Scholar] [CrossRef]
- Sudo, R.; Chung, S.; Zervantonakis, I.K.; Vickerman, V.; Toshimitsu, Y.; Griffith, L.G.; Kamm, R.D. Transport-mediated angiogenesis in 3D epithelial coculture. FASEB J. 2009, 23, 2155–2164. [Google Scholar] [CrossRef]
- Kim, L.; Toh, Y.C.; Voldman, J.; Yu, H. A practical guide to microfluidic perfusion culture of adherent mammalian cells. Lab Chip 2007, 7, 681–694. [Google Scholar] [CrossRef]
- Wolbers, F.; ter Braak, P.; le Gac, S.; Luttge, R.; Andersson, H.; Vermes, I.; van den Berg, A. Viability study of HL60 cells in contact with commonly used microchip materials. Electrophoresis 2006, 27, 5073–5080. [Google Scholar] [CrossRef]
- Davidsson, R.; Boketoft, Å.; Bristulf, J.; Kotarsky, K.; Olde, B.; Owman, C.; Bengtsson, M.; Laurell, T.; Emnéus, J. Developments toward a microfluidic system for long-term monitoring of dynamic cellular events in immobilized human cells. Anal. Chem. 2004, 76, 4715–4720. [Google Scholar]
- Blau, A.W.; Ziegler, C.M. Prototype of a novel autonomous perfusion chamber for long-term culturing and in situ investigation of various cell types. J. Biochem. Biophys. Methods 2001, 50, 15–27. [Google Scholar] [CrossRef]
- Lee, J.N.; Jiang, X.; Ryan, D.; Whitesides, G.M. Compatibility of mammalian cells on surfaces of poly(dimethylsiloxane). Langmuir 2004, 20, 11684–11691. [Google Scholar]
- Futai, N.; Gu, W.; Song, J.W.; Takayama, S. Handheld recirculation system and customized media for microfluidic cell culture. Lab Chip 2006, 6, 149–154. [Google Scholar] [CrossRef]
- Komen, J.; Wolbers, F.; Franke, H.R.; Andersson, H.; Vermes, I.; van den Berg, A. Viability analysis and apoptosis induction of breast cancer cells in a microfluidic device: Effect of cytostatic drugs. Biomed. Microdevices 2008, 10, 727–737. [Google Scholar] [CrossRef]
- Yu, H.; Meyvantsson, I.; Shkel, I.A.; Beebe, D.J. Diffusion dependent cell behavior in microenvironments. Lab Chip 2005, 5, 1089–1095. [Google Scholar] [CrossRef]
- Li, Y.S.J.; Haga, J.H.; Chien, S. Molecular basis of the effects of shear stress on vascular endothelial cells. J. Biomech. 2005, 38, 1949–1971. [Google Scholar] [CrossRef]
- Healy, Z.R.; Lee, N.H.; Gao, X.; Goldring, M.B.; Talalay, P.; Kensler, T.W.; Konstantopoulos, K. Divergent responses of chondrocytes and endothelial cells to shear stress: Cross-talk among COX-2, the phase 2 response, and apoptosis. Proc. Natl. Acad. Sci. USA 2005, 102, 14010–14015. [Google Scholar]
- Wang, M.M.; Tu, E.; Raymond, D.E.; Yang, J.M.; Zhang, H.; Hagen, N.; Dees, B.; Mercer, E.M.; Forster, A.H.; Kariv, I. Microfluidic sorting of mammalian cells by optical force switching. Nat. Biotechnol. 2004, 23, 83–87. [Google Scholar]
- Ramser, K.; Hanstorp, D. Optical manipulation for single-cell studies. J. Biophotonics 2010, 3, 187–206. [Google Scholar] [CrossRef]
- Li, H.; Friend, J.R.; Yeo, L.Y. A scaffold cell seeding method driven by surface acoustic waves. Biomaterials 2007, 28, 4098–4104. [Google Scholar]
- Gray, B.L.; Lieu, D.K.; Collins, S.D.; Smith, R.L.; Barakat, A.I. Microchannel platform for the study of endothelial cell shape and function. Biomed. Microdevices 2002, 4, 9–16. [Google Scholar] [CrossRef]
- Frame, M.D.; Sarelius, I.H. Flow-induced cytoskeletal changes in endothelial cells growing on curved surfaces. Microcirculation 2000, 7, 419–427. [Google Scholar]
- Cinamon, G.; Alon, R. A real time in vitro assay for studying leukocyte transendothelial migration under physiological flow conditions. J. Immunol. Methods 2003, 273, 53–62. [Google Scholar] [CrossRef]
- Frame, M.D.S.; Chapman, G.B.; Makino, Y.; Sarelius, I.H. Shear stress gradient over endothelial cells in a curved microchannel system. Biorheology 1998, 35, 245–262. [Google Scholar] [CrossRef]
- Schaff, U.Y.; Xing, M.M.Q.; Lin, K.K.; Pan, N.; Jeon, N.L.; Simon, S.I. Vascular mimetics based on microfluidics for imaging the leukocyte-endothelial inflammatory response. Lab Chip 2007, 7, 448–456. [Google Scholar] [CrossRef]
- Tanaka, Y.; Kikukawa, Y.; Sato, K.; Sugii, Y.; Kitamori, T. Culture and leukocyte adhesion assay of human arterial endothelial cells in a glass microchip. Anal. Sci. 2007, 23, 261–266. [Google Scholar] [CrossRef]
- Young, E.W.K.; Wheeler, A.R.; Simmons, C.A. Matrix-dependent adhesion of vascular and valvular endothelial cells in microfluidic channels. Lab Chip 2007, 7, 1759–1766. [Google Scholar] [CrossRef]
- Ku, C.J.; D’Amico, O.T.; Spence, D.M. Interactions between multiple cell types in parallel microfluidic channels: Monitoring platelet adhesion to an endothelium in the presence of an anti-adhesion drug. Anal. Chem. 2008, 80, 7543–7548. [Google Scholar] [CrossRef]
- Liu, K.; Pitchimani, R.; Dang, D.; Bayer, K.; Harrington, T.; Pappas, D. Cell culture chip using low-shear mass transport. Langmuir 2008, 24, 5955–5960. [Google Scholar]
- Shamloo, A.; Ma, N.; Poo, M.; Sohn, L.L.; Heilshorn, S.C. Endothelial cell polarization and chemotaxis in a microfluidic device. Lab Chip 2008, 8, 1292–1299. [Google Scholar] [CrossRef]
- Borenstein, J.T.; Terai, H.; King, K.R.; Weinberg, E.; Kaazempur-Mofrad, M.; Vacanti, J. Microfabrication technology for vascularized tissue engineering. Biomed. Microdevices 2002, 4, 167–175. [Google Scholar] [CrossRef]
- Song, J.W.; Gu, W.; Futai, N.; Warner, K.A.; Jacques, E.; Takayama, S. Computer-controlled microcirculatory support system for endothelial cell culture and shearing. Anal. Chem. 2005, 77, 3993–3999. [Google Scholar]
- Huh, D.; Fujioka, H.; Tung, Y.C.; Futai, N.; Paine, R.; Grotberg, J.B.; Takayama, S. Acoustically detectable cellular-level lung injury induced by fluid mechanical stresses in microfluidic airway systems. Proc. Natl. Acad. Sci. USA 2007, 104, 18886–18891. [Google Scholar]
- Antia, M.; Herricks, T.; Rathod, P.K. Microfluidic approaches to malaria pathogenesis. Cell. Microbiol. 2008, 10, 1968–1974. [Google Scholar] [CrossRef]
- Shelby, J.P.; White, J.; Ganesan, K.; Rathod, P.K.; Chiu, D.T. A microfluidic model for single-cell capillary obstruction by plasmodium falciparum-infected erythrocytes. Proc. Natl. Acad. Sci. USA 2003, 100, 14618. [Google Scholar]
- Becker, H.; Gärtner, C. Polymer microfabrication methods for microfluidic analytical applications. Electrophoresis 2000, 21, 12–26. [Google Scholar] [CrossRef]
- Paulus, A.; Williams, S.J.; Sassi, A.P.; Kao, P.H.; Tan, H.; Hooper, H.H. Integrated capillary electrophoresis using glass and plastic chips for multiplexed DNA analysis. Proc. SPIE 1998, 351594–103. [Google Scholar]
- Duffy, D.C.; McDonald, J.C.; Schueller, O.J.A.; Whitesides, G.M. Rapid prototyping of microfluidic systems in poly(dimethylsiloxane). Anal. Chem. 1998, 70, 4974–4984. [Google Scholar]
- Chen, Y.H.; Chen, S.H. Analysis of DNA fragments by microchip electrophoresis fabricated on poly(methyl methacrylate) substrates using a wire-imprinting method. Electrophoresis 2000, 21, 165–170. [Google Scholar] [CrossRef]
- Shoffner, M.A.; Cheng, J.; Hvichia, G.E.; Kricka, L.J.; Wilding, P. Chip PCR. I. Surface passivation of microfabricated silicon-glass chips for PCR. Nucleic Acids Res. 1996, 24, 375–379. [Google Scholar] [CrossRef]
- Bélanger, M.C.; Marois, Y. Hemocompatibility, biocompatibility, inflammatory and in vivo studies of primary reference materials low-density polyethylene and polydimethylsiloxane: A review. J. Biomed. Mater. Res. 2001, 58, 467–477. [Google Scholar] [CrossRef]
- Charati, S.; Stern, S. Diffusion of gases in silicone polymers: Molecular dynamics simulations. Macromolecules 1998, 31, 5529–5535. [Google Scholar] [CrossRef]
- Piruska, A.; Nikcevic, I.; Lee, S.H.; Ahn, C.; Heineman, W.R.; Limbach, P.A.; Seliskar, C.J. The autofluorescence of plastic materials and chips measured under laser irradiation. Lab chip 2005, 5, 1348–1354. [Google Scholar] [CrossRef]
- Toepke, M.W.; Beebe, D.J. PDMS absorption of small molecules and consequences in microfluidic applications. Lab Chip 2006, 6, 1484–1486. [Google Scholar]
- Prieto, A.; Basauri, O.; Rodil, R.; Usobiaga, A.; Fernández, L.; Etxebarria, N.; Zuloaga, O. Stir-bar sorptive extraction: A view on method optimisation, novel applications, limitations and potential solution. J. Chromatogr. A 1217, 2642–2666. [Google Scholar]
- Paguirigan, A.L.; Beebe, D.J. From the cellular perspective: Exploring differences in the cellular baseline in macroscale and microfluidic cultures. Integr. Biol. 2009, 1, 182–195. [Google Scholar] [CrossRef]
- Van Midwoud, P.M.; Groothuis, G.M.M.; Merema, M.T.; Verpoorte, E. Microfluidic biochip for the perifusion of precision-cut rat liver slices for metabolism and toxicology studies. Biotechnol. Bioeng. 2010, 105, 184–194. [Google Scholar] [CrossRef]
- Amstein, C.F.; Hartman, P.A. Adaptation of plastic surfaces for tissue culture by glow discharge. J. Clin. Microbiol. 1975, 2, 46–54. [Google Scholar]
- Makamba, H.; Kim, J.H.; Lim, K.; Park, N.; Hahn, J.H. Surface modification of poly(dimethylsiloxane) microchannels. Electrophoresis 2003, 24, 3607–3619. [Google Scholar] [CrossRef]
- Paguirigan, A.; Beebe, D. Gelatin based microfluidic devices for cell culture. Lab Chip 2006, 6, 407–413. [Google Scholar] [CrossRef]
- Zhang, X.; Haswell, S.J. Materials matter in microfluidic devices. MRS Bull. 2006, 31, 95–99. [Google Scholar] [CrossRef]
- Voskerician, G.; Shive, M.S.; Shawgo, R.S.; Recum, H.; Anderson, J.M.; Cima, M.J.; Langer, R. Biocompatibility and biofouling of MEMS drug delivery devices. Biomaterials 2003, 24, 1959–1967. [Google Scholar] [CrossRef]
- Van Midwoud, P.M. An alternative approach based on microfluidics to study drug metabolism and toxicity using liver and intestinal tissue. Ph.D. Dissertation, Faculty of Mathematics and Natural Sciences, University of Groningen, Groningen, The Netherlands, 2010. [Google Scholar]
- Berthier, E.; Young, E.W.K.; Beebe, D. Engineers are from PDMS-land, biologists are from polystyrenia. Lab Chip 2012, 12, 1224–1237. [Google Scholar]
- Lucas, J. Advances in plant disease and pest management. J. Agric. Sci. 2011, 149, 91–114. [Google Scholar] [CrossRef]
- Andersson, H.; van den Berg, A. Microfabrication and microfluidics for tissue engineering: State of the art and future opportunities. Lab Chip 2004, 4, 98–103. [Google Scholar] [CrossRef]
- Lutolf, M.P.; Gilbert, P.M.; Blau, H.M. Designing materials to direct stem-cell fate. Nature 2009, 462, 433–441. [Google Scholar]
- Quantalife Homepage. Available online: http://www.quantalife.com (accessed on 28 February 2012).
- Scientific Application Resource Center. Available online: http://www.cellectricon.com/sites/download/download_dynaflow.php (accessed on 28 February 2012).
- Whitesides, G.M.; Stroock, A.D. Flexible methods for microfluidics. Phys. Today 2001, 54, 42–48. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Alrifaiy, A.; Lindahl, O.A.; Ramser, K. Polymer-Based Microfluidic Devices for Pharmacy, Biology and Tissue Engineering. Polymers 2012, 4, 1349-1398. https://doi.org/10.3390/polym4031349
Alrifaiy A, Lindahl OA, Ramser K. Polymer-Based Microfluidic Devices for Pharmacy, Biology and Tissue Engineering. Polymers. 2012; 4(3):1349-1398. https://doi.org/10.3390/polym4031349
Chicago/Turabian StyleAlrifaiy, Ahmed, Olof A. Lindahl, and Kerstin Ramser. 2012. "Polymer-Based Microfluidic Devices for Pharmacy, Biology and Tissue Engineering" Polymers 4, no. 3: 1349-1398. https://doi.org/10.3390/polym4031349
APA StyleAlrifaiy, A., Lindahl, O. A., & Ramser, K. (2012). Polymer-Based Microfluidic Devices for Pharmacy, Biology and Tissue Engineering. Polymers, 4(3), 1349-1398. https://doi.org/10.3390/polym4031349



