Abstract
Bioreducible, cationic linear poly(amino ether)s (PAEs) were designed as promising gene vectors. These polymers were synthesized by the reaction of a disulfide-functional monomer, N,N′-dimethylcystamine (DMC), and several different diglycidyl ethers. The resulting PAEs displayed a substantial buffer capacity (up to 64%) in the endosomal acidification region of pH 7.4–5.1. The PAEs condense plasmid DNA into 80–200 nm sized polyplexes, and have surface charges ranging from +20 to +40 mV. The polyplexes readily release DNA upon exposure to reducing conditions (2.5 mM DTT) due to the cleavage of the disulfide groups that is present in the main chain of the polymers, as was demonstrated by agarose gel electrophoresis. Upon exposing COS-7 cells to polyplexes that were prepared at polymer/DNA w/w ratios below 48, cell viabilities between 80–100% were observed, even under serum-free conditions. These polyplexes show comparable or higher transfection efficiencies (up to 38%) compared to 25 kDa branched polyethylenimine (PEI) polyplexes (12% under serum-free conditions). Moreover, the PAE-based polyplexes yield transfection efficiencies as high as 32% in serum-containing medium, which makes these polymers interesting for gene delivery applications.
1. Introduction
Non-viral gene vectors are extensively sought after to deliver DNA as therapeutic agents to treat or cure genetic disorders [1,2,3,4,5,6,7]. The revolutionary advancement of CRISPR/Cas gene editing techniques has especially inspired renewed interest in the development of safe gene delivery agents [8,9,10]. Moreover, numerous non-viral vectors are able to deliver proteins which is a crucial step in the CRISPR/Cas process [11,12]. Several properties are required to render vectors attractive for gene delivery, such as high transfection efficiency, minimal or non-toxicity, and ease of manufacturing [13,14,15]. Cationic polymers are interesting candidates that are designed to meet these conditions, using modern developments in polymer chemistry [16,17,18,19] as well as nanotechnology [20,21,22]. These polymers spontaneously form polyplexes with DNA tightly packed inside by means of electrostatic interactions [23,24,25,26]. These interactions are aided by the release of sodium ions upon condensation through multivalent, polycationic polymers, leading to a net entropy increase [27,28,29]. After entering the cells through endocytosis, polyplexes must escape from the endosomal-lysosomal pathway and unpack the DNA in the cell nucleus in order to initiate gene expression [30,31,32,33]. The cationic nature of the polymers, which is often due to the incorporation of amine moieties, promotes endosomal escape by means of the proton sponge effect [34].
Many approaches were proposed to facilitate DNA unpacking, such as the use of acid-labile cationic polymers [35,36], ester-based cationic polymers [37,38], and biologically triggered cationic polymers [39,40,41,42]. Among these approaches, the latter approach is arguably the most interesting since it takes advantage of intracellular enzymes or related biomolecules [43,44]. Glutathione, an enzyme-related antioxidant that dictates the reductive potential of cells, is more concentrated inside the cells compared to the extracellular environment, which thereby increases the intracellular reductive potential [45,46,47]. Through the introduction of disulfide-linkages in cationic polymers, this higher reductive potential has been utilized as a trigger to cleave the disulfide bonds and unpack DNA from their polyplexes after the successful endosomal escape of the polyplexes into the cytosol [48,49,50].
The introduction of disulfide moieties was successfully employed in polyethylenimine (PEI) [51,52,53,54,55], poly(L-lysine) (PLL) [56,57], peptides [58,59], and poly(amido amine) dendrimers [60], resulting in increased gene transfection and improved biocompatibility. Green and coworkers reported cationic poly(amino ester)s with disulfides along the backbone for efficient gene delivery [61]. Our group, as well as other groups, reported on disulfide-linked linear and branched poly(amido amine)s that display excellent transfection efficiency and biocompatibility [62,63,64,65]. However, the clinical application of these polymers has so far been restrained due to their impaired transfection in the presence of serum [22,66]. The introduction of hydrophilic poly(ethylene glycol) (PEG) onto these vectors was found to increase the serum tolerance of these polymers to a limited extent, however they still failed to meet the requirements for clinical application [66,67]. Non-degradable poly(amino ether)s (PAEs), which are cationic polymers that bear ether groups that are similar to those of PEG, were shown to be effective gene delivery agents, however their serum tolerance was not investigated [68].
Here, we report a modular synthetic approach to generate disulfide-linked PAEs through the polyaddition of N,N′-dimethylcystamine (DMC) as a bifunctional diamine monomer, and through a series of diglycidyl ethers as epoxy monomers. The resulting bioreducible PAEs with structural varieties in the main polymer chain, will be evaluated for cell viabilities and transfection efficiencies.
2. Results and Discussion
2.1. Polymer Preparation and Characterization
Cystamine is an interesting building block for biologically applied polymers, since its disulfide moiety renders the resulting polymers redox responsive. However, in combination with diglycidyl ethers, it may form branched or crosslinked structures due to the twofold reactivity of each of the primary amines [69,70]. The resulting structures suffer from poor water solubility, which impedes their application as gene vectors. In order to form linear polymers that are derived from cystamine disulfide, N,N′-dimethylcystamine disulfide (DMC), a cystamine analogue in which both amine groups are monomethylated was employed as the bifunctional amine monomer. The reaction of this amine with a series of diglycidyl ethers with structural variations yields water-soluble bioreducible linear poly(amino ether)s (PAEs) that are designed to be well applicable for efficient gene delivery (Scheme 1).
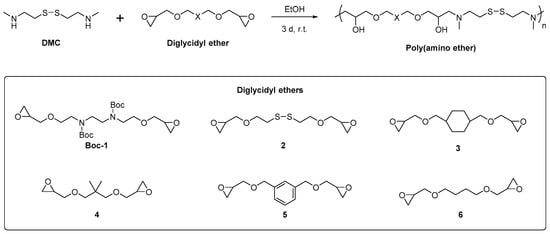
Scheme 1.
The modular synthesis of bioreducible poly(amino ether)s through the amine-epoxide reaction of N,N′-dimethylcystamine (DMC) with different diglycidyl ethers.
The set of diglycidyl ethers (DEs) consists of six different molecules, as depicted in Scheme 1. These are classified into three monomer groups: (i) high charge density (1); (ii) additional bioreducible linker (2); and (iii) varying hydrophobicity (3–6). All three factors are rationalized to play a role in the transfection performance of the cationic polymers. From these DEs, linear cationic PAEs were prepared by stirring the disulfide-containing diamine DMC with the corresponding diglycidyl ethers in the presence of excess triethylamine. The mixtures typically became homogenous after a couple of hours and subsequently became viscous, indicating the formation of polymers. No occurrence of gelation during the polymerization was observed, which indicated the formation of linear polymers. After removal of impurities and small oligomers through dialysis, the polymers were obtained in appreciable yields (ca. 14–45%). All of the polymer structures were validated by 1H NMR spectroscopy (see Figures S1–S6 in the supplementary information). No residual epoxide signals were observed in all cases. Molecular weights of the polymers were determined through size exclusion chromatography (SEC) and ranged from 2.5 kDa to 4.2 kDa, with polydispersities between 1.5–2.0 (summarized in Table 1).

Table 1.
Characteristics of linear bioreducible poly(amino ether)s prepared through the reaction of N,N′-dimethylcystamine (DMC) with different diglycidyl ethers (1–6).
The obtained PAEs contain tertiary amines in their main chain that are prone to protonation in acidic milieu. Protonation not only grants the necessary positive charge to condense DNA into polyplexes, it also provides a buffer capacity to facilitate the endosomal escape of polyplexes through the proton sponge effect [34]. To evaluate the capacity of these polymers to accept protons in the endosomal acidification range (pH 7.4–5.1), titration experiments were performed. The titration curves for all PAEs are displayed in Figure 1, including the titration of reference component 25 kDa branched PEI, a commonly employed transfection agent, and blank NaCl solution (0.15 M). Plateaus along the titration curves in the pH 5.1–7.4 region are observed for all PAEs, which implies that there are pronounced buffer capacities. These buffer capacities range from 34% up to 64%, and are superior over PEI (18%). High buffer capacity of polymers is an excellent property to enhance endosomal escape which is necessary for the efficient gene delivery into the cytosol.
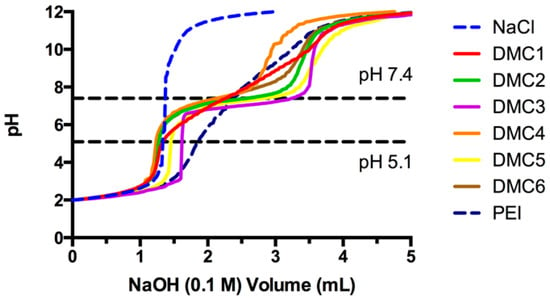
Figure 1.
Titration curves of poly(amino ether)s by NaOH (0.1 M) from pH 2–12. PEI (branched, 25 kDa) and NaCl (150 mM) are included as references.
2.2. Polyplex Formation and Characterization
For efficient transfection, naked DNA requires protection by vectors, which can be realized by forming polyplexes through electrostatic interaction between DNA and cationic polymers [71]. Here, polyplexes were prepared by combining polymer solutions of DMC1-6 and DNA solutions at polymer/DNA mass ratios of 6, 12, 24, and 48. The resulting size and surface charge of the obtained polyplexes at these ratios are displayed in Figure 2. The polyplexes that were formed at polymer/DNA ratios above 6 exhibit hydrodynamic sizes from 80 to 200 nm and zeta-potentials from +20 to +40 mV. With increasing polymer/DNA mass ratios, the polyplexes become smaller in dimension and carry increased positive charge, corresponding to more tightly packed polyplexes and an increased number of cationic PAEs incorporated in the formation of these polyplexes [23,26,72]. Polyplexes formed at a polymer/DNA mass ratio of 6 display a relatively large size (over 400 nm) and an average surface charge of +15 mV, which indicates a rather loose structure. Polyplexes prepared from DMC1 and DMC2 have a relatively small size compared to other polymers at the same polymer/DNA ratios. This phenomenon is related to the higher cationic density of polymer DMC1, which bears additional amine groups, resulting in increased electrostatic interaction with DNA. The additional disulfide content of polymer DMC2 makes this polymer more flexible along the main chain, since the disulfide bond has a low rotational barrier, which also leads to more compact polyplexes. The other four polymers DMC3, DMC4, DMC5, and DMC6 form polyplexes with similar values in size and zeta-potential at identical polymer/DNA mass ratios, and no clear relation can be drawn between the polymer structure and the characteristics of the polyplexes here.
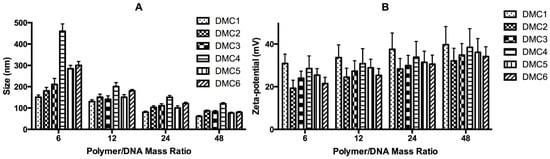
Figure 2.
Characteristics of polyplexes prepared from bioreducible poly(amino ether)s and plasmid pCMV-GFP under various polymer/DNA mass ratios. (A) Hydrodynamic sizes versus mass ratios; (B) Zeta-potentials versus mass ratios.
The stability of the polyplexes under normal and reductive conditions was validated through agarose gel electrophoresis experiments, and results are shown in Figure 3. When untreated polyplexes (w/w = 48) that were dissolved in a HEPES buffer were subjected to electrophoresis, DNA bands did not migrate along the electric field, indicating that DNA was fully retained by the cationic PAEs. However, it is expected that due to the relatively high intracellular concentration of glutathione (GSH, 2.5–10 mM), the disulfides in the polymer chain can be cleaved resulting in the disassembly of the polyplexes and the release of the DNA content. This biological reaction was mimicked by the incubation of the polyplexes with dithiothreitol (DTT, 2.5 mM) as a reducing agent for 30 min under ambient conditions. These polyplexes displayed clear bands migrating along the electric force direction, similarly to free DNA, illustrating the release of DNA from the degraded polyplexes.
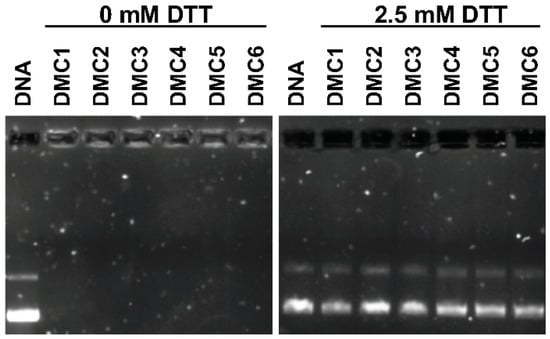
Figure 3.
Agarose gel electrophoresis of polyplexes (w/w = 48) treated without DTT (left) and with 2.5 mM DTT (right).
2.3. Cytotoxicity
For biomedical applications, gene vectors require minimal toxicity and excellent biocompatibility towards cells and tissues [6,73]. It is therefore important to evaluate the cytotoxicity of the polyplexes before exploring their transfection performance. MTT assays were used to study the cytotoxicity profiles of the polyplexes towards COS-7 cells under serum-free (0%) and serum-present (10%) conditions. As shown in Figure 4, the viability of COS-7 cells under serum-free conditions is above 80% when treated with polyplexes prepared at polymer/DNA ratios of 6, 12, and 24 for all polymers, except for DMC1 which has the highest cationic density in the series due to the presence of extra secondary amino groups in the chain. In contrast, branched PEI (25 kDa) is considerably more toxic at the same polymer/DNA ratios. Under 10% serum conditions, cells show an even higher viability, and significant toxicity is only observed at the polymer/DNA mass ratio 48, indicating the promising potential of these PAEs for gene delivery under serum conditions.
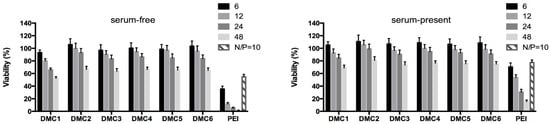
Figure 4.
Cell viabilities of COS-7 cells exposed to polyplexes at different polymer/DNA mass ratios for 1h and incubated for 2 days, as determined via MTT assays under 0% serum conditions (left) and 10% serum conditions (right). Branched PEI (25 kDa) is included as a reference.
2.4. Transfection Efficiency
After the evaluation of the polyplex toxicity, the transfection performance of the polyplexes at polymer/DNA ratios 6 and 12 was investigated on COS-7 cells using green fluorescent protein (GFP) as a reporter gene. Three different serum conditions of 0%, 10%, and 25% were evaluated. In all experiments, PEI polyplexes at the same polymer/DNA ratios were included as a reference. Transfection efficiency was quantified by flow cytometry and was reported as the percentage of GFP-expressing cells of the total population (results are shown in Figure 5). Under serum-free conditions, the transfection efficiencies of the polyplexes prepared from PAEs at a polymer/DNA ratio of 6 were similar or enhanced (12–25%) compared to PEI (12.1%). The transfection efficiency increased at polymer/DNA ratio 12, reaching the highest efficiency of 36% for polyplexes based on DMC2. Relatively low transfection efficiencies were observed for DMC1, DMC4, and DMC6. This is most likely related to their lower buffer capacities as compared to polymers DMC2, DMC3, and DMC5. At polymer/DNA ratio 6, the transfection efficiencies of DMC2, DMC3, and DMC5 were 25 ± 5.9%, 20 ± 2.2%, and 21 ± 2.8%, respectively, which increased to 36 ± 5.1%, 27 ± 4.0%, and 29 ± 3.4%, respectively at a polymer/DNA ratio of 12.
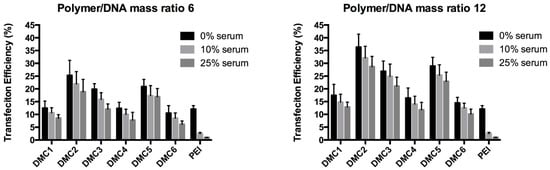
Figure 5.
Transfection efficiencies of polyplexes prepared from bioreducible PAEs DMC1-6 and plasmid DNA at polymer/DNA mass ratios of 6 (left) and 12 (right). PEI is included as a reference at its optimal ratio of N/P = 10 in both graphs. COS-7 cells were exposed to polyplexes for 1 h and were incubated using various serum concentrations for 2 days. Transfection efficiency was quantified using FACS.
As illustrated in Figure 5, the PAEs remained efficient in transfecting COS-7 cells, with efficiencies ranging from 10–32% at 10% serum and 10–28% at 25% serum, which is considerably better than that of PEI (2.5% at 10% serum and 1.0% at 25%). This effect of decreased transfection efficiency under serum-present conditions is consistently reported in literature, for example employing poly(amino ester)s [74], poly(amido amine)s [62], and several methacrylate-based cationic polymers [75]. The retained transfection of PAEs in the presence of serum is related to the hydroxyl groups that are present on the polymers causing a protein repelling effect, similar to polyethylene glycol [76]. A similar effect was reported for hydroxyl-functionalized methacrylates, where the addition of hydroxyls resulted in polyplex shielding against the serum, which was attributed to the formation of hydroxyl-DNA hydrogen bonding [77]. The transfection efficiency of DMC2-based polyplexes at a polymer/DNA mass ratio of 12 in the presence of 25% serum is 32%, and this is even further enhanced to 50% for polyplexes with a polymer/DNA mass ratio of 24 with a cell viability of up to 85% (data not shown). These results underline the high potential of these disulfide-based PAEs for non-viral gene delivery.
3. Materials and Methods
Epichlorohydrin (≥98%, Sigma-Aldrich, St. Louis, MO, USA), 2-hydroxyethyl disulfide (technical grade, Sigma-Aldrich), tetrabutylammonium bromide (TBAB, ≥98%, Sigma-Aldrich), sodium hydroxide (NaOH, pellets, Sigma-Aldrich), 1,3-benzenedimethanol (98%, Sigma-Aldrich), N,N′-dimethylethylene diamine (>97.0%, TCI Europe N.V., Zwijndrecht, Belgium), 1,4-butanediol diglycidyl ether (≥95%, Sigma-Aldrich), 1,4-cyclohexanedimethanol diglycidyl ether (3, Sigma-Aldrich), neopentyl glycol diglycidyl ether (4, Sigma-Aldrich), branched polyethylenimine (PEI, Mw 25 kDa, Sigma-Aldrich), dithiothreitol (DTT, Sigma-Aldrich), triethylamine (TEA, ≥99%, Sigma-Aldrich), borane dimethyl sulfide complex (BMS, Sigma-Aldrich), N,N′-bis(2-hydroxylethyl) ethylenediamine (>95.0%, TCI Europe N.V., Zwijndrecht, Belgium), di-tert-butyl dicarbonate (>95.0%, TCI Europe N.V.), and thiazolidine (98%, ACROS Organics) were used as received. The water used in these experiments was treated through a Milli-Q Gradient System (Millipore, Bedford, MA, USA). Plasmid DNA and plasmids pCMV-GFP and pCMV-ΔGFP, were purchased from a Plasmid Factory (Bielefeld, Germany).
1H NMR and 13C NMR were recorded on a Bruker NMR spectrometer (400 MHz) in deuterated solvents. The molecular weights were determined by size exclusion chromatography (SEC, Waters Alliance 2695) relative to PEG standards, using a Mixed-M column (PL-aquagel-OH 8 micron, 300 × 7.5 mm) with N,N-dimethylformamide containing LiCl (50 mM) as a mobile phase.
3.1. The Synthesis of N,N′-Dimethylcystamine (DMC)
The monomer was synthesized in its HCl salt form following our previous report [78]. Then, into a solution of thiazolidine (5.0 g, 56 mmol) in anhydrous tetrahydrofuran (THF, 20 mL) under a nitrogen atmosphere, borane dimethyl sulfide complex (BMS, 10.6 g, 140 mmol) in anhydrous THF (20 mL) was slowly added under stirring, and the mixture was subsequently refluxed overnight. After cooling down to the ambient temperature, methanol (20 mL) was added by syringe to neutralize any residual BMS, and the mixture was refluxed for an additional 3 h. After cooling down to the ambient temperature, the solution was purged with HCl gas, and the mixture was concentrated under reduced pressure. The resulting liquid was consecutively dissolved in methanol (5.0 mL) and was precipitated twice in diethyl ether (50 mL) to obtain a white oily precipitate. The precipitate was dissolved in water (7.5 mL) and was titrated with saturated I2/KI solution until a persistent yellow color was obtained. After stirring for an additional 2 h, the yellow solution turned colorless upon raising the pH to 10. The solution was extracted with chloroform (4 × 30 mL). Afterwards, the organic fractions were combined, dried over MgSO4, and were concentrated under reduced pressure to yield a clear liquid. Subsequently, the liquid was dissolved in isopropanol (20 mL) and was purged with HCl gas, yielding a turbid yellow mixture and a white precipitate. The precipitate was filtered off, washed with cold diethyl ether, and was finally recrystallized from isopropanol/methanol to obtain the HCl salt as a white solid. (2.12 g, 30.0% yield). 1H NMR (D2O, 400 MHz): δH 2.76 (s, 6H, CH2NHCH3), 3.05 (t, 4H, SCH2CH2), 3.44 (t, 4H, SCH2CH2). 13C NMR (D2O, 400 MHz): δC 31.92, 32.83, 47.02.
3.2. The Synthesis of N,N′-Bis(2-hydroxyethyl)-N,N′-bis(tert-butoxycarbonyl) ethylenediamine
Di-tert-butyl dicarbonate (17.6 g, 40 mmol) in dichloromethane (200 mL) was added dropwise under stirring at 0 °C to a mixture of N,N′-bis(2-hydroxylethyl) ethylenediamine (6.00 g, 40.0 mmol) and triethylamine (14.0 mL, 10.2 g, 100 mmol) in dichloromethane (100 mL). The mixture was stirred at the ambient temperature overnight. The solvent was removed under reduced pressure. The resulting solid was washed with ethyl acetate and diethyl ether and was subsequently dried under vacuum to yield a white solid (11.2 g, 80.4% yield). 1H NMR (CDCl3, 400 MHz): δH 1.45 (s, 18H), 3.25–3.60 (m, 8H), 3.71–3.82 (m, 4H).
3.3. The Synthesis of Diglycidyl Ethers
Diglycidyl ether monomers were synthesized via the reaction of epichlorohydrin with the corresponding diols. A typical example of the Boc-protected form of 1 (Boc-1) was: into a mixture of N,N′-bis(2-hydroxyethyl)-N,N′-bis(tert-butoxycarbonyl) ethylenediamine (1.75 g, 5.00 mmol), sodium hydroxide (0.800 g, 20.0 mmol), and tetrabutylammonium bromide (12.37 mg, 34.8 µmol), epichlorohydrin (1.45 mL, 18.4 mmol) was added gradually. The mixture was heated to 40 °C and was stirred for 3 h. The resulting mixture was subjected to column chromatography to obtain the diglycidyl ether of Boc-1 (1.95 g, 84.7% yield). 1H NMR (CDCl3, 400 MHz): δH 1.46 (s, 18H), 2.57 (s, 2H), 2.76 (t, 2H), 3.10 (m, 2H), 3.36 (m, 10H), 3.60 (m, 4H), 3.72 (m, 2H). 13C NMR (CDCl3, 400 MHz): δ 28.43, 44.12, 46.13, 47.34, 50.75, 69.89, 71.76, 79.76, 155.41.
Diglycidyl ether of 5 (1.12 g, 89.5 % yield). 1H NMR (CDCl3, 400 MHz): δH 2.62 (m, 2H, epoxide ring OCHHCHO), 2.81 (m, 2H, epoxide ring OCHHCHO), 3.20 (m, 2H, epoxide ring OCHHCHO), 3.45 (m, 2H, CHCHHO), 3.78 (m, 2H, CHCHHO), 4.59 (m, 4H, OCH2-phenyl ring), 7.27–7.36 (m, 4H, phenyl ring). 13C NMR (CDCl3, 400 MHz): δC 44.36, 50.93, 71.01, 73.27, 127.13, 128.64, 138.22.
3.4. The Synthesis of Poly(amino ether)s
Poly(amino ether)s (PAEs) were synthesized by the ring open polymerization of bifunctional epoxide monomers with equimolar amounts of primary amines. A typical procedure is given for the synthesis of DMC1:
Into a glass vial, diglycidyl ether Boc-1 (0.461 g, 1.00 mmol), DMC (0.254 g, 1.00 mmol), triethylamine (0.400 mL, 2.87 mmol), and ethanol (0.500 mL) were charged. After 3 d stirring under ambient temperature, the resulting viscous mixture was diluted with methanol (50.0 mL) and purged with HCl gas for 30 min in order to remove the Boc protecting groups. The mixture was concentrated, and the residue was dissolved in water (15.0 mL), acidified with 4 M HCl solution to pH 4, and was purified via dialysis (MWCO 1 kDa) against water overnight. Polymer DMC1 was obtained after lyophilization as an amorphous solid. The other PAE polymers DMC2-6 were synthesized analogously, however these polymers do not require the Boc removal step.
Polymer DMC1 (off-white solid, 0.162 g, 31.6%): 1H NMR (D2O, 400 MHz): δH 2.98 (s, 6H, NCH3), 3.10–3.16 (m, 4H, CH2NHCH2CH2O), 3.26 (m, 4H, NCH2CH2S), 3.32–3.38 (b, 8H, CH2NHCH2CH2O + OCH2CHOHCH2NCH2CH2S), 3.35–3.64 (m, 8H, CH2OCH2), 3.79 (t, 4H, OCH2CHOHCH2NCH2CH2S), 4.30 (bs, 2H, OCH2CHOHCH2NCH2CH2S).
Polymer DMC2 (white solid, 0.132 g, 33.2%): 1H NMR (D2O, 400 MHz): δH 2.95 (s, 6H, NCH3), 2.96 (m, 4H, OCH2CH2S), 3.09–3.14 (m, 4H, NCH2CH2S), 3.30 (m, 4H, CH2OCH2CHOHCH2NCH2CH2S), 3.55–3.65 (m, 8H, CH2OCH2CHOHCH2NCH2CH2S), 3.80–3.89 (m, 4H, OCH2CH2S), 4.25 (bs, 2H, CH2OCH2CHOHCH2NCH2CH2S).
Polymer of DMC3 (white solid, 0.086 g, 16.9% yield): 1H NMR (D2O, 400 MHz): δH 0.90–1.01 and 1.75–1.79 (m, 8H, 4 × CH2 of cyclohexane ring), 1.29 and 1.49 (m, 2H, CH of cyclohexane ring), 2.98 (s, 6H, NCH3), 3.11–3.18 (m, 4H, NCH2CH2S), 3.20–3.31 (m, 4H, CH2OCH2CHOHCH2NCH2CH2S), 3.37–3.47 (m, 4H, CH2OCH2CHOHCH2NCH2CH2S), 3.51–3.67 (m, 8H, CH2OCH2CHOHCH2NCH2CH2S), 4.25 (bs, 2H, CH2OCH2CHOHCH2NCH2CH2S).
Polymer DMC4 (amorphous solid, 0.064 g, 13.9% yield): 1H NMR (D2O, 400 MHz): δH 0.91 (s, 6H, C(CH2CH3)2), 2.92 (s, 6H, NCH3), 3.08–3.12 (m, 4H, NCH2CH2S), 3.26–3.32 (m, 8H, C(CH2CH3)2 + CH2OCH2CHOHCH2NCH2CH2S), 3.66–3.75 (m, 8H, CH2OCH2CHOHCH2NCH2CH2S), 4.25 (bs, 2H, CH2OCH2CHOHCH2NCH2CH2S).
Polymer of DMC5 (white solid, 0.220g, 43.7%): 1H NMR (D2O, 400 MHz): δH 2.96 (s, 6H, NCH3), 3.09–3.12 (m, 4H, NCH2CH2S), 3.24–3.30 (m, 4H, phenyl ring-CH2OCH2CHOHCH2NCH2CH2S), 3.57–3.64 (m, 8H, phenyl ring-CH2OCH2CHOHCH2NCH2CH2S), 4.26 (bs, 2H, phenyl ring-CH2OCH2CHOHCH2NCH2CH2S), 4.62–4.65 (m, 4H, phenyl ring-CH2OCH2CHOHCH2NCH2CH2S), 7.40–7.45 (m, 4H, phenyl ring).
Polymer of DMC6 (amorphous solid, 0.183 g, 40.1% yield): 1H NMR (D2O, 400 MHz): δH 1.63 (bs, 4H, OCH2CH2CH2CH2O), 2.96 (s, 6H, NCH3), 3.04–3.13 (m, 4H, NCH2CH2S), 3.23–3.30 (m, 4H, CH2OCH2CHOHCH2NCH2CH2S), 3.52–3.62 (m, 12H, CH2OCH2CHOHCH2NCH2CH2S), 4.25 (bs, 2H, CH2OCH2CHOHCH2N).
3.5. Buffer Capacity Titration
Buffering capacities of the polymers were determined via acid-base titration. The amount of PAE polymer equal to 5 mmol protonable amine groups was dissolved in NaCl aqueous solution (150 mM, 10 mL). The pH of the polymer solution was acidified to ≤2.0 and the solution was titrated with NaOH solution (0.1 M) using an automatic titrator (Metrohm 702 SM Tirino). As a reference, b-PEI (25 kDa) and NaCl aqueous solution were titrated following the same method. The percentage of amine groups protonated from pH 5.1 to 7.4 was defined as the buffer capacity that can be calculated from the following equation [79]:
wherein, ΔVNaOH denotes the NaOH volume that is required to bring the pH value of the polymer solution from 5.1 to 7.4, and N mole (5 mmol) denotes the total moles of protonable amine groups in the PAE polymer.
3.6. Polyplex Preparation
Polyplexes were prepared by adding polymer solution into the DNA solution at designated polymer/DNA mass ratios. All polymer and DNA solutions were prepared in the HEPES buffer (20 mM, pH 7.4). The procedure of polyplexes prepared at the mass ratio of 48 is as follows:
In an Eppendorf tube (1.5 mL), DNA solution (75 µg/mL, 200 µL) and polymer solution (900 µg/mL, 800 µL) were successively added. The mixture was subjected to vortexing for 5 s and was incubated at room temperature for 30 min to obtain the polyplex suspensions. The size and surface charge of the polyplexes were measured on a Zetasizer Nano ZS (Malvern Instruments, Malvern, UK) at 25 °C. The data present the means of the three measurements with the corresponding standard deviations as error bars.
3.7. Agarose Gel Electrophoresis
Polyplexes were prepared at a polymer/DNA mass ratio of 48, as described in the previous protocol. Polyplex solutions (90 µL) were diluted with either DTT HEPES solution (25 mM, 10 µL) or HEPES buffer (10 µL), and were incubated under ambient conditions for 30 min. The resulting dispersions (20 µL) were mixed with 6× loading buffer containing bromophenol (Ferments, 5.0 µL) and were aliquoted (10 µL) to load on agarose gel (0.8% w/v) containing 1× SYBR® Safe DNA Gel Stain (Invitrogen™, Carlsbad, CA, USA). The gel was developed at 90 V for 60 min in a TBE (Tris-borate-EDTA, 1×) running buffer and was consecutively imaged via FluorChem (Proteinsimple, Westburg, Leusden, The Netherlands) under UV excitation.
3.8. Cytotoxicity
The cytotoxicity of polyplexes towards COS-7 cells was evaluated through MTT assays. In 96 well plates, ca. 104 cells were seeded per well and were allowed to grow overnight in complete medium (100 µL, DMEM medium supplemented with 10% FBS) at 37 °C under 5% CO2 conditions to reach 60–80% confluency. The medium was replaced with fresh medium (100 µL, w/o 10% FBS). After 30 min of incubation, the cells were loaded with either polyplex solutions at various polymer/DNA mass ratios (100 µL, 0.25 µg DNA per well) or HEPES buffer (20 mM, pH 7.4, supplemented with 5.0 wt % glucose) and were incubated for an additional 1 h. All cells were refreshed with complete warm culture medium (100 µL) and were cultured for another 48 h. Half of the cells that were untreated with polyplexes were killed by incubating in 100× triton (4%, 10 µL) for 15 min, while the residual half were taken as a control to set the viability to 100%. All of the cells were washed with DPBS (100 µL) and were successively incubated in MTT solution (0.5 mg/mL, 100 µL) at 37 °C under 5% CO2 condition for 4 h. After refreshing MTT with DMSO (100 µL), the resultant formazone crystals were quantified through a plate reader (Tecan Infinite M200) at wavelength 540 nm with reference 680 nm. Measurements were performed in triplicate.
3.9. Transfection Efficiency
The plasmids of pCMV-GFP and pCMV-ΔGFP were respectively used as positive and negative controls. In 48 well plates, the cells were seeded at a density of 1.6 × 104 per well (1.6 × 104 cells/cm2) and were cultured in complete medium (DMEM with 10% FBS, 200 µL) at 37 °C under humidified atmosphere with 5% CO2 to reach confluencies of 60–80%. The cells were refreshed with warm medium (200 µL) with designated serum concentration and were maintained in new medium for 30 min until they were finally treated with polyplex solutions (200 µL, 0.5 µg DNA per well). After 60 min of incubation at 37 °C, the polyplexes were replaced with fresh warm complete medium (200 µL), and the cells were allowed to grow for another 48 h. After replacing the old medium with trypsin solution (0.25%, 200 µL), the cells were spun down (600 g, 5 min, R.T.), resuspended in HBSS buffer (200 µL), and were measured by the FACSCalibur (Becton-Dickinson, Breda, The Netherlands) at an excitation wavelength of 488 nm and emission at 530 nm. FACS Cellquest Software was applied to process the data. A 25 kDa branched PEI/DNA formulation, prepared at its recommended ratio of N/P = 10, was used as a reference.
4. Conclusions
A modular approach towards the incorporation of bioreducibility into linear poly(amino ether)s was developed through the reaction of N,N′-dimethylcystamine with various diglycidyl ethers. The obtained PAEs have good to excellent buffer capacities, ranging from 40 to 67%. Upon mixing PAEs with DNA at appropriate mass ratios, polyplexes were formed with hydrodynamic diameters ranging from 80–200 nm and zeta-potentials of up to +40 mV. The polyplexes displayed no significant toxicity towards COS-7 cells at polymer/DNA ratios up to 24 with transfection of COS-7 cells with notable efficiencies of up to 36%. In addition, the presence of serum did not markedly interfere with the transfection efficiency, even at serum concentrations of 10% and 25%. This effect is attributed to the presence of the protein repelling hydroxyl moieties within the PAEs. All transfection efficiencies and biocompatibility profiles of PAE-based polyplexes outpace the common polymeric vector PEI (branched, 25 kDa). Overall, these results present a novel modular approach towards a new generation of easily accessible cationic polymers for gene and drug delivery.
Supplementary Materials
The following supplementary materials are available online at http://www.mdpi.com/2073-4360/10/6/687/s1, Figures S1–S6: 1H NMR spectra of polymers DMC1-6.
Author Contributions
Conceptualization, G.S., J.F.J.E. and J.M.J.P.; Methodology, G.S., M.R.E.; Investigation, M.R.E., G.S.; Resources, J.F.J.E., J.M.J.P.; Data Curation, G.S, J.M.J.P.; Writing-Original Draft Preparation, M.R.E., G.S., J.M.J.P.; Writing-Review & Editing, M.R.E., J.M.J.P.
Funding
This research was funded by the provinces of Overijssel and Gelderland and the project consortium of the Center for Medical Imaging—North East Netherlands (CMI-NEN), as well as by NanoNextNL, a micro and nanotechnology consortium of the Government of the Netherlands, and 130 partners.
Acknowledgments
Karin Roelofs is acknowledged for her assistance in the cell culture and FACS measurements, and Marc Ankoné for his valuable advice regarding the synthesis of DMC.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Mingozzi, F.; High, K.A. Therapeutic in Vivo Gene Transfer for Genetic Disease Using Aav: Progress and Challenges. Nat. Rev. Genet. 2011, 12, 341–355. [Google Scholar] [CrossRef] [PubMed]
- Kay, M.A. State-of-the-Art Gene-Based Therapies: The Road Ahead. Nat. Rev. Genet. 2011, 12, 316–328. [Google Scholar] [CrossRef] [PubMed]
- Seow, Y.; Wood, M.J. Biological Gene Delivery Vehicles: Beyond Viral Vectors. Mol. Ther. 2009, 17, 767–777. [Google Scholar] [CrossRef] [PubMed]
- Jones, C.H.; Chen, C.K.; Ravikrishnan, A.; Rane, S.; Pfeifer, B.A. Overcoming Nonviral Gene Delivery Barriers: Perspective and Future. Mol. Pharm. 2013, 10, 4082–4098. [Google Scholar] [CrossRef] [PubMed]
- Draghici, B.; Ilies, M.A. Synthetic Nucleic Acid Delivery Systems: Present and Perspectives. J. Med. Chem. 2015, 58, 4091–4130. [Google Scholar] [CrossRef] [PubMed]
- Davis, M.E. Non-Viral Gene Delivery Systems. Curr. Opin. Biotechnol. 2002, 13, 128–131. [Google Scholar] [CrossRef]
- Rezaee, M.; Oskuee, R.K.; Nassirli, H.; Malaekeh-Nikouei, B. Progress in the Development of Lipopolyplexes as Efficient Non-Viral Gene Delivery Systems. J. Control. Release 2016, 236, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.B.; Zhang, S.; Kos, P.; Xiong, H.; Zhou, K.; Perelman, S.S.; Zhu, H.; Siegwart, D.J. Non-Viral Crispr/Cas Gene Editing in Vitro and in Vivo Enabled by Synthetic Nanoparticle Co-Delivery of Cas9 Mrna and Sgrna. Angew. Chem. Int. Ed. Engl. 2017, 56, 1059–1063. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.X.; Li, M.; Lee, C.M.; Chakraborty, S.; Kim, H.W.; Bao, G.; Leong, K.W. Crispr/Cas9-Based Genome Editing for Disease Modeling and Therapy: Challenges and Opportunities for Nonviral Delivery. Chem. Rev. 2017, 117, 9874–9906. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Song, C.Q.; Dorkin, J.R.; Zhu, L.J.; Li, Y.; Wu, Q.; Park, A.; Yang, J.; Suresh, S.; Bizhanova, A.; et al. Therapeutic Genome Editing by Combined Viral and Non-Viral Delivery of Crispr System Components in Vivo. Nat. Biotechnol. 2016, 34, 328–333. [Google Scholar] [CrossRef] [PubMed]
- El-Sherbiny, I.; Khalil, I.; Ali, I.; Yacoub, M. Updates on Smart Polymeric Carrier Systems for Protein Delivery. Drug Dev. Ind. Pharm. 2017, 43, 1567–1583. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.Y.; Yuk, S.H. Polymeric Protein Delivery Systems. Prog. Polym. Sci. 2007, 32, 669–697. [Google Scholar] [CrossRef]
- Kataoka, K.; Harashima, H. Gene Delivery Systems: Viral Vs. Non-Viral Vectors. Adv. Drug Deliv. Rev. 2001, 52, 151. [Google Scholar] [CrossRef]
- Miyata, K.; Nishiyama, N.; Kataoka, K. Rational Design of Smart Supramolecular Assemblies for Gene Delivery: Chemical Challenges in the Creation of Artificial Viruses. Chem. Soc. Rev. 2012, 41, 2562–2574. [Google Scholar] [CrossRef] [PubMed]
- Yin, H.; Kanasty, R.L.; Eltoukhy, A.A.; Vegas, A.J.; Dorkin, J.R.; Anderson, D.G. Non-Viral Vectors for Gene-Based Therapy. Nat. Rev. Genet. 2014, 15, 541–555. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.; Narain, R. Progress of Raft Based Polymers in Gene Delivery. Prog. Polym. Sci. 2013, 38, 767–790. [Google Scholar] [CrossRef]
- Pack, D.W.; Hoffman, A.S.; Pun, S.; Stayton, P.S. Design and Development of Polymers for Gene Delivery. Nat. Rev. Drug Discov. 2005, 4, 581–593. [Google Scholar] [CrossRef] [PubMed]
- Chu, D.S.; Schellinger, J.G.; Shi, J.; Convertine, A.J.; Stayton, P.S.; Pun, S.H. Application of Living Free Radical Polymerization for Nucleic Acid Delivery. Acc. Chem. Res. 2012, 45, 1089–1099. [Google Scholar] [CrossRef] [PubMed]
- Tong, R.; Tang, L.; Ma, L.; Tu, C.; Baumgartner, R.; Cheng, J. Smart Chemistry in Polymeric Nanomedicine. Chem. Soc. Rev. 2014, 43, 6982–7012. [Google Scholar] [CrossRef] [PubMed]
- Park, K. Facing the Truth About Nanotechnology in Drug Delivery. ACS Nano 2013, 7, 7442–7447. [Google Scholar] [CrossRef] [PubMed]
- Farokhzad, O.C.; Langer, R. Impact of Nanotechnology on Drug Delivery. ACS Nano 2009, 3, 16–20. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, V.P.; Jain, R.K. Strategies for Advancing Cancer Nanomedicine. Nat. Mater. 2013, 12, 958–962. [Google Scholar] [CrossRef] [PubMed]
- Bloomfield, V.A. DNA Condensation by Multivalent Cations. Biopolymers 1997, 44, 269–282. [Google Scholar] [CrossRef]
- Perevyazko, I.Y.; Bauer, M.; Pavlov, G.M.; Hoeppener, S.; Schubert, S.; Fischer, D.; Schubert, U.S. Polyelectrolyte Complexes of DNA and Linear PEI: Formation, Composition and Properties. Langmuir 2012, 28, 16167–16176. [Google Scholar] [CrossRef] [PubMed]
- Meneksedag-Erol, D.; Tang, T.; Uludag, H. Molecular Modeling of Polynucleotide Complexes. Biomaterials 2014, 35, 7068–7076. [Google Scholar] [CrossRef] [PubMed]
- Schaaf, P.; Schlenoff, J.B. Saloplastics: Processing Compact Polyelectrolyte Complexes. Adv. Mater. 2015, 27, 2420–2432. [Google Scholar] [CrossRef] [PubMed]
- Agostinelli, E.; Vianello, F.; Magliulo, G.; Thomas, T.; Thomas, T.J. Nanoparticle Strategies for Cancer Therapeutics: Nucleic Acids, Polyamines, Bovine Serum Amine Oxidase and Iron Oxide Nanoparticles (Review). Int. J. Oncol. 2015, 46, 5–16. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.J.; Tajmir-Riahi, H.A.; Thomas, T. Polyamine-DNA Interactions and Development of Gene Delivery Vehicles. Amino Acids 2016, 48, 2423–2431. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.J.; Thomas, T. Collapse of DNA in Packaging and Cellular Transport. Int. J. Biol. MacroMol. 2018, 109, 36–48. [Google Scholar] [CrossRef] [PubMed]
- Midoux, P.; Breuzard, G.; Gomez, J.P.; Pichon, C. Polymer-Based Gene Delivery: A Current Review on the Uptake and Intracellular Trafficking of Polyplexes. Curr. Gene Ther. 2008, 8, 335–352. [Google Scholar] [CrossRef] [PubMed]
- Boussif, O.; Lezoualch, F.; Zanta, M.A.; Mergny, M.D.; Scherman, D.; Demeneix, B.; Behr, J.P. A Versatile Vector for Gene and Oligonucleotide Transfer into Cells in Culture and in-Vivo-Polyethylenimine. Proc. Natl. Acad. Sci. USA 1995, 92, 7297–7301. [Google Scholar] [CrossRef] [PubMed]
- Behr, J.-P. The Proton Sponge: A Trick to Enter Cells the Viruses Did Not Exploit. CHIMIA Int. J. Chem. 1997, 51, 34–36. [Google Scholar]
- Liang, W.; Lam, J.K.W. Endosomal escape pathways for non-viral nucleic acid delivery systems. In Molecular Regulation of Endocytosis; InTech: Rijeka, Croatia, 2012. [Google Scholar]
- Sonawane, N.D.; Szoka, F.C., Jr.; Verkman, A.S. Chloride Accumulation and Swelling in Endosomes Enhances DNA Transfer by Polyamine-DNA Polyplexes. J. Biol. Chem. 2003, 278, 44826–44831. [Google Scholar] [CrossRef] [PubMed]
- Knorr, V.; Russ, V.; Allmendinger, L.; Ogris, M.; Wagner, E. Acetal Linked Oligoethylenimines for Use as Ph-Sensitive Gene Carriers. Bioconjug. Chem. 2008, 19, 1625–1634. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Zheng, M.; Meng, F.; Zhong, Z. Non-Viral Gene Transfection in Vitro Using Endosomal Ph-Sensitive Reversibly Hydrophobilized Polyethylenimine. Biomaterials 2011, 32, 9109–9119. [Google Scholar] [CrossRef] [PubMed]
- Lynn, D.M.; Anderson, D.G.; Putnam, D.; Langer, R. Accelerated Discovery of Synthetic Transfection Vectors: Parallel Synthesis and Screening of a Degradable Polymer Library. J. Am. Chem. Soc. 2001, 123, 8155–8156. [Google Scholar] [CrossRef] [PubMed]
- Lynn, D.M.; Langer, R. Degradable Poly(Beta-Amino Esters): Synthesis, Characterization, and Self-Assembly with Plasmid DNA. J. Am. Chem. Soc. 2000, 122, 10761–10768. [Google Scholar] [CrossRef]
- Law, B.; Tung, C.H. Proteolysis: A Biological Process Adapted in Drug Delivery, Therapy, and Imaging. Bioconj. Chem. 2009, 20, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Kate, P.; Torchilin, V.P. Matrix Metalloprotease 2-Responsive Multifunctional Liposomal Nanocarrier for Enhanced Tumor Targeting. ACS Nano 2012, 6, 3491–3498. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Wang, T.; Perche, F.; Taigind, A.; Torchilin, V.P. Enhanced Anticancer Activity of Nanopreparation Containing an Mmp2-Sensitive Peg-Drug Conjugate and Cell-Penetrating Moiety. Proc. Natl. Acad. Sci. USA 2013, 110, 17047–17052. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Perche, F.; Wang, T.; Torchilin, V.P. Matrix Metalloproteinase 2-Sensitive Multifunctional Polymeric Micelles for Tumor-Specific Co-Delivery of siRNA and Hydrophobic Drugs. Biomaterials 2014, 35, 4213–4222. [Google Scholar] [CrossRef] [PubMed]
- Mura, S.; Nicolas, J.; Couvreur, P. Stimuli-Responsive Nanocarriers for Drug Delivery. Nat. Mater. 2013, 12, 991–1003. [Google Scholar] [CrossRef] [PubMed]
- Stuart, M.A.; Huck, W.T.; Genzer, J.; Muller, M.; Ober, C.; Stamm, M.; Sukhorukov, G.B.; Szleifer, I.; Tsukruk, V.V.; Urban, M.; et al. Emerging Applications of Stimuli-Responsive Polymer Materials. Nat. Mater. 2010, 9, 101–113. [Google Scholar] [CrossRef] [PubMed]
- Bauhuber, S.; Hozsa, C.; Breunig, M.; Gopferich, A. Delivery of Nucleic Acids Via Disulfide-Based Carrier Systems. Adv. Mater. 2009, 21, 3286–3306. [Google Scholar] [CrossRef] [PubMed]
- Cheng, R.; Feng, F.; Meng, F.; Deng, C.; Feijen, J.; Zhong, Z. Glutathione-Responsive Nano-Vehicles as a Promising Platform for Targeted Intracellular Drug and Gene Delivery. J. Control. Release 2011, 152, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.H.; Yang, Z.; Lim, C.W.; Lee, Y.H.; Dongbang, S.; Kang, C.; Kim, J.S. Disulfide-Cleavage-Triggered Chemosensors and Their Biological Applications. Chem. Rev. 2013, 113, 5071–5109. [Google Scholar] [CrossRef] [PubMed]
- Oupicky, D.; Li, J. Bioreducible Polycations in Nucleic Acid Delivery: Past, Present, and Future Trends. MacroMol. Biosci. 2014, 14, 908–922. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; Engbersen, J.F. The Role of the Disulfide Group in Disulfide-Based Polymeric Gene Carriers. Expert Opin. Drug Deliv. 2009, 6, 421–439. [Google Scholar] [CrossRef] [PubMed]
- Elzes, M.R.; Akeroyd, N.; Engbersen, J.F.; Paulusse, J.M. Disulfide-Functional Poly(Amido Amine)S with Tunable Degradability for Gene Delivery. J. Control. Release 2016, 244, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Gosselin, M.A.; Guo, W.J.; Lee, R.J. Efficient Gene Transfer Using Reversibly Cross-Linked Low Molecular Weight Polyethylenimine. Bioconj. Chem. 2001, 12, 989–994. [Google Scholar] [CrossRef]
- Wang, Y.; Chen, P.; Shen, J. The Development and Characterization of a Glutathione-Sensitive Cross-Linked Polyethylenimine Gene Vector. Biomaterials 2006, 27, 5292–5298. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, R.C.; Etrych, T.; Briggs, S.S.; Preece, J.A.; Ulbrich, K.; Seymour, L.W. Polymer-Coated Polyethylenimine/DNA Complexes Designed for Triggered Activation by Intracellular Reduction. J. Gene Med. 2004, 6, 337–344. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Mo, H.; Koo, H.; Park, J.Y.; Cho, M.Y.; Jin, G.W.; Park, J.S. Visualization of the Degradation of a Disulfide Polymer, Linear Poly(Ethylenimine Sulfide), for Gene Delivery. Bioconj. Chem. 2007, 18, 13–18. [Google Scholar] [CrossRef] [PubMed]
- Peng, Q.; Zhong, Z.; Zhuo, R. Disulfide Cross-Linked Polyethylenimines (PEI) Prepared Via Thiolation of Low Molecular Weight PEI as Highly Efficient Gene Vectors. Bioconj. Chem. 2008, 19, 499–506. [Google Scholar] [CrossRef] [PubMed]
- Kakizawa, Y.; Harada, A.; Kataoka, K. Environment-Sensitive Stabilization of Core-Shell Structured Polyion Complex Micelle by Reversible Cross-Linking of the Core through Disulfide Bond. J. Am. Chem. Soc. 1999, 121, 11247–11248. [Google Scholar] [CrossRef]
- Miyata, K.; Kakizawa, Y.; Nishiyama, N.; Harada, A.; Yamasaki, Y.; Koyama, H.; Kataoka, K. Block Catiomer Polyplexes with Regulated Densities of Charge and Disulfide Cross-Linking Directed to Enhance Gene Expression. J. Am. Chem. Soc. 2004, 126, 2355–2361. [Google Scholar] [CrossRef] [PubMed]
- McKenzie, D.L.; Smiley, E.; Kwok, K.Y.; Rice, K.G. Low Molecular Weight Disulfide Cross-Linking Peptides as Nonviral Gene Delivery Carriers. Bioconj. Chem. 2000, 11, 901–909. [Google Scholar] [CrossRef]
- McKenzie, D.L.; Kwok, K.Y.; Rice, K.G. A Potent New Class of Reductively Activated Peptide Gene Delivery Agents. J. Biol. Chem. 2000, 275, 9970–9977. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, H.; Yang, W.; Cheng, Y. Disulfide Cross-Linked Low Generation Dendrimers with High Gene Transfection Efficacy, Low Cytotoxicity, and Low Cost. J. Am. Chem. Soc. 2012, 134, 17680–17687. [Google Scholar] [CrossRef] [PubMed]
- Kozielski, K.L.; Tzeng, S.Y.; Green, J.J. A Bioreducible Linear Poly(Beta-Amino Ester) for siRNA Delivery. Chem. Commun. 2013, 49, 5319–5321. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.; Zhong, Z.; Lok, M.C.; Jiang, X.; Hennink, W.E.; Feijen, J.; Engbersen, J.F. Novel Bioreducible Poly(Amido Amine)S for Highly Efficient Gene Delivery. Bioconj. Chem. 2007, 18, 138–145. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.; Yan, J.; You, Y. Synthesis of Bioreducible and Acid Labile Poly(Amido Amine)S Via Michael-Addition Reactions and Their Application in Gene Delivery. J. Control. Release 2011, 152 (Suppl. 1), e179–e181. [Google Scholar] [CrossRef] [PubMed]
- Ou, M.; Xu, R.; Kim, S.H.; Bull, D.A.; Kim, S.W. A Family of Bioreducible Poly(Disulfide Amine)S for Gene Delivery. Biomaterials 2009, 30, 5804–5814. [Google Scholar] [CrossRef] [PubMed]
- Ferruti, P. Poly(Amidoamine)S: Past, Present, and Perspectives. J. Polym. Sci. Polym. Chem. 2013, 51, 2319–2353. [Google Scholar] [CrossRef]
- Hubbell, J.A.; Langer, R. Translating Materials Design to the Clinic. Nat. Mater. 2013, 12, 963–966. [Google Scholar] [CrossRef] [PubMed]
- Hatakeyama, H.; Akita, H.; Harashima, H. A Multifunctional Envelope Type Nano Device (Mend) for Gene Delivery to Tumours Based on the Epr Effect: A Strategy for Overcoming the Peg Dilemma. Adv. Drug Deliv. Rev. 2011, 63, 152–160. [Google Scholar] [CrossRef] [PubMed]
- Vu, L.; Ramos, J.; Potta, T.; Rege, K. Generation of a Focused Poly(Amino Ether) Library: Polymer-Mediated Transgene Delivery and Gold-Nanorod Based Theranostic Systems. Theranostics 2012, 2, 1160–1173. [Google Scholar] [CrossRef] [PubMed]
- Ferruti, P.; Marchisio, M.A.; Duncan, R. Poly(Amido-Amine)S: Biomedical Applications. MacroMol. Rapid Commun. 2002, 23, 332–355. [Google Scholar] [CrossRef]
- Emilitri, E.; Ferruti, P.; Annunziata, R.; Ranucci, E.; Rossi, M.; Falciola, L.; Mussini, P.; Chiellini, F.; Bartoli, C. Novel Amphoteric Cystine-Based Poly(Amidoamine)S Responsive to Redox Stimuli. Macromolecules 2007, 40, 4785–4793. [Google Scholar] [CrossRef]
- Nguyen, D.N.; Green, J.J.; Chan, J.M.; Longer, R.; Anderson, D.G. Polymeric Materials for Gene Delivery and DNA Vaccination. Adv. Mater. 2009, 21, 847–867. [Google Scholar] [CrossRef] [PubMed]
- Niebel, Y.; Buschmann, M.D.; Lavertu, M.; De Crescenzo, G. Combined Analysis of Polycation/Odn Polyplexes by Analytical Ultracentrifugation and Dynamic Light Scattering Reveals Their Size, Refractive Index Increment, Stoichiometry, Porosity, and Molecular Weight. Biomacromolecules 2014, 15, 940–947. [Google Scholar] [CrossRef] [PubMed]
- Mintzer, M.A.; Simanek, E.E. Nonviral Vectors for Gene Delivery. Chem. Rev. 2009, 109, 259–302. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Su, R.C.; Yi, W.J.; Zhao, Z.G. Biodegradable Poly(Amino Ester) with Aromatic Backbone as Efficient Nonviral Gene Delivery Vectors. Molecules 2017, 22, 566. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Jung, S.; Si, G.; Cheng, R.; Meng, F.; Zhu, X.; Park, T.G.; Zhong, Z. Cationic Methacrylate Copolymers Containing Primary and Tertiary Amino Side Groups: Controlled Synthesis Via Raft Polymerization, DNA Condensation, and in Vitro Gene Transfection. J. Polym. Sci. Part A Polym. Chem. 2010, 48, 2869–2877. [Google Scholar] [CrossRef]
- Luo, X.H.; Huang, F.W.; Qin, S.Y.; Wang, H.F.; Feng, J.; Zhang, X.Z.; Zhuo, R.X. A Strategy to Improve Serum-Tolerant Transfection Activity of Polycation Vectors by Surface Hydroxylation. Biomaterials 2011, 32, 9925–9939. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Li, F.; Yuan, Z.F.; Zhuo, R.X. Influence of Hydroxyl Groups on the Biological Properties of Cationic Polymethacrylates as Gene Vectors. Acta Biomater. 2010, 6, 2658–2665. [Google Scholar] [CrossRef] [PubMed]
- Piest, M.; Lin, C.; Mateos-Timoneda, M.A.; Lok, M.C.; Hennink, W.E.; Feijen, J.; Engbersen, J.F. Novel Poly(Amido Amine)S with Bioreducible Disulfide Linkages in Their Diamino-Units: Structure Effects and in Vitro Gene Transfer Properties. J. Control. Release 2008, 130, 38–45. [Google Scholar] [CrossRef] [PubMed]
- Zhong, Z.; Feijen, J.; Lok, M.C.; Hennink, W.E.; Christensen, L.V.; Yockman, J.W.; Kim, Y.H.; Kim, S.W. Low Molecular Weight Linear Polyethylenimine-B-Poly(Ethylene Glycol)-B-Polyethylenimine Triblock Copolymers: Synthesis, Characterization, and in Vitro Gene Transfer Properties. Biomacromolecules 2005, 6, 3440–3448. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).