Not Cleaving the His-tag of Thal Results in More Tightly Packed and Better-Diffracting Crystals
Abstract
1. Introduction
2. Materials and Methods
2.1. Plasmids and Molecular Cloning
2.2. Expression and Purification of Recombinant Thal-RebH5
2.3. Crystallization and Data Collection
2.4. Data Processing, Structure Determination and Refinement
2.5. Accession Numbers
3. Results
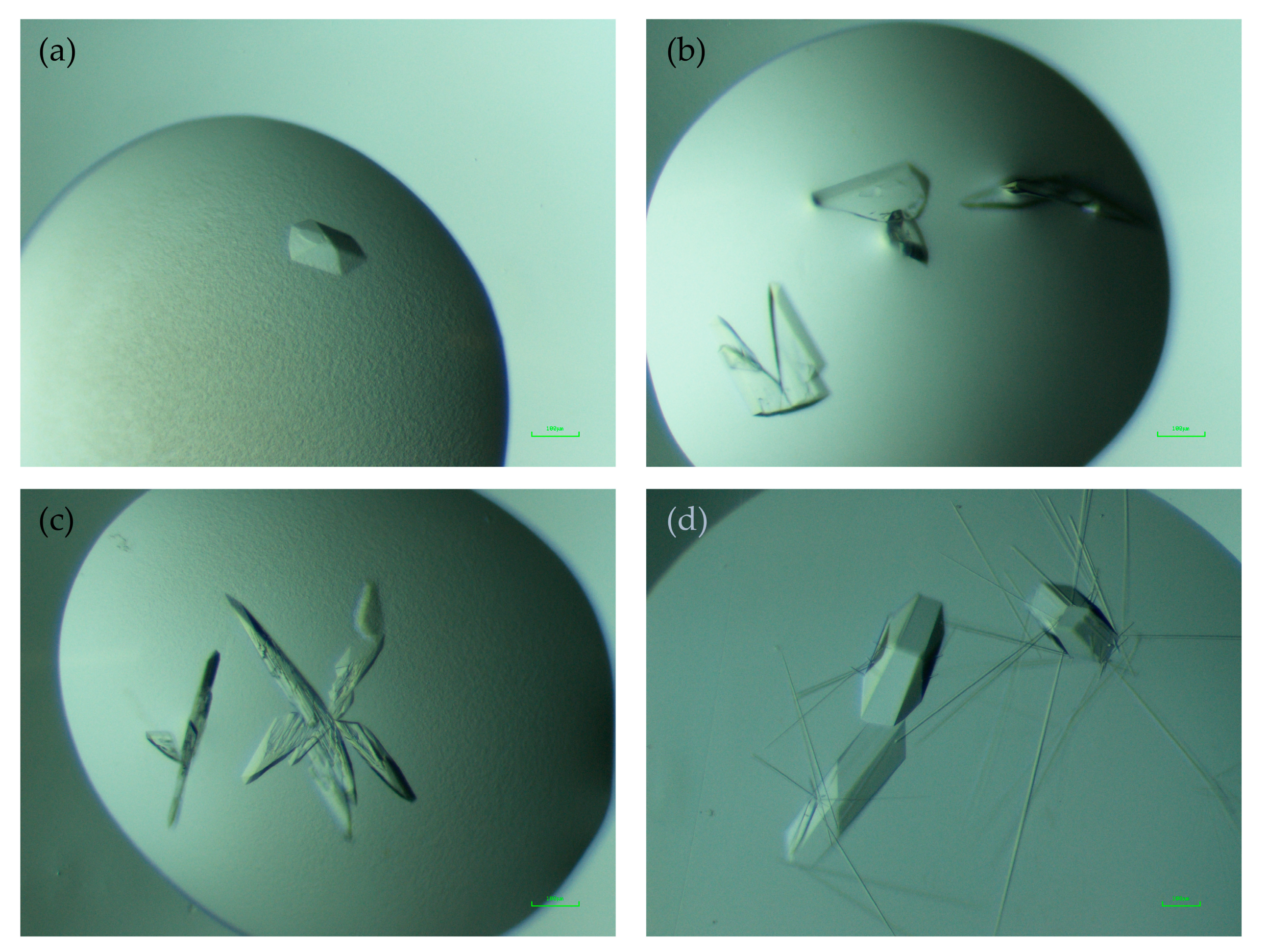
3.1. Crystallization Details and Comparison to Thal-RebH5
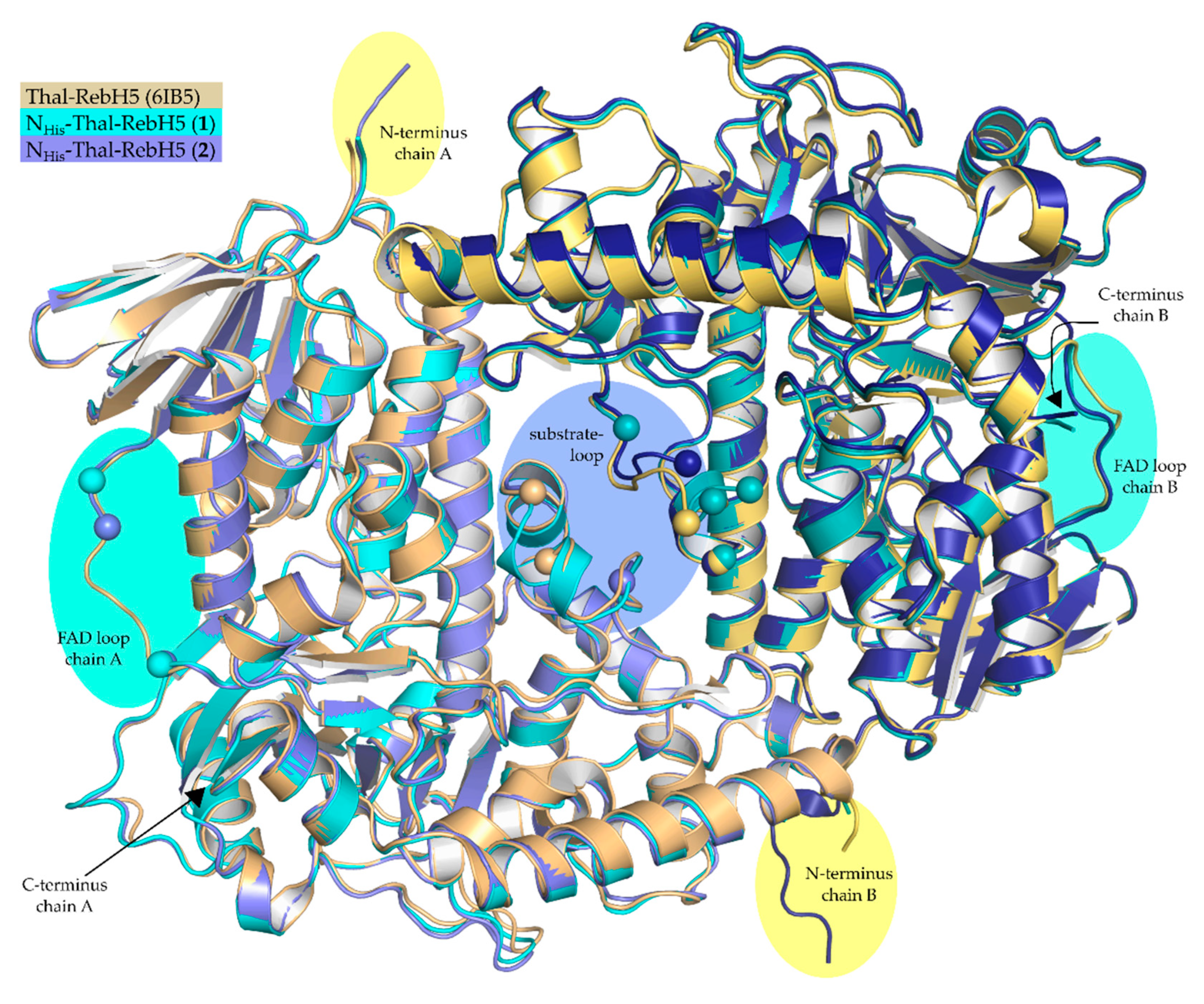
3.2. Comparison of the Previously Described P64 and the New P21 Crystal Forms
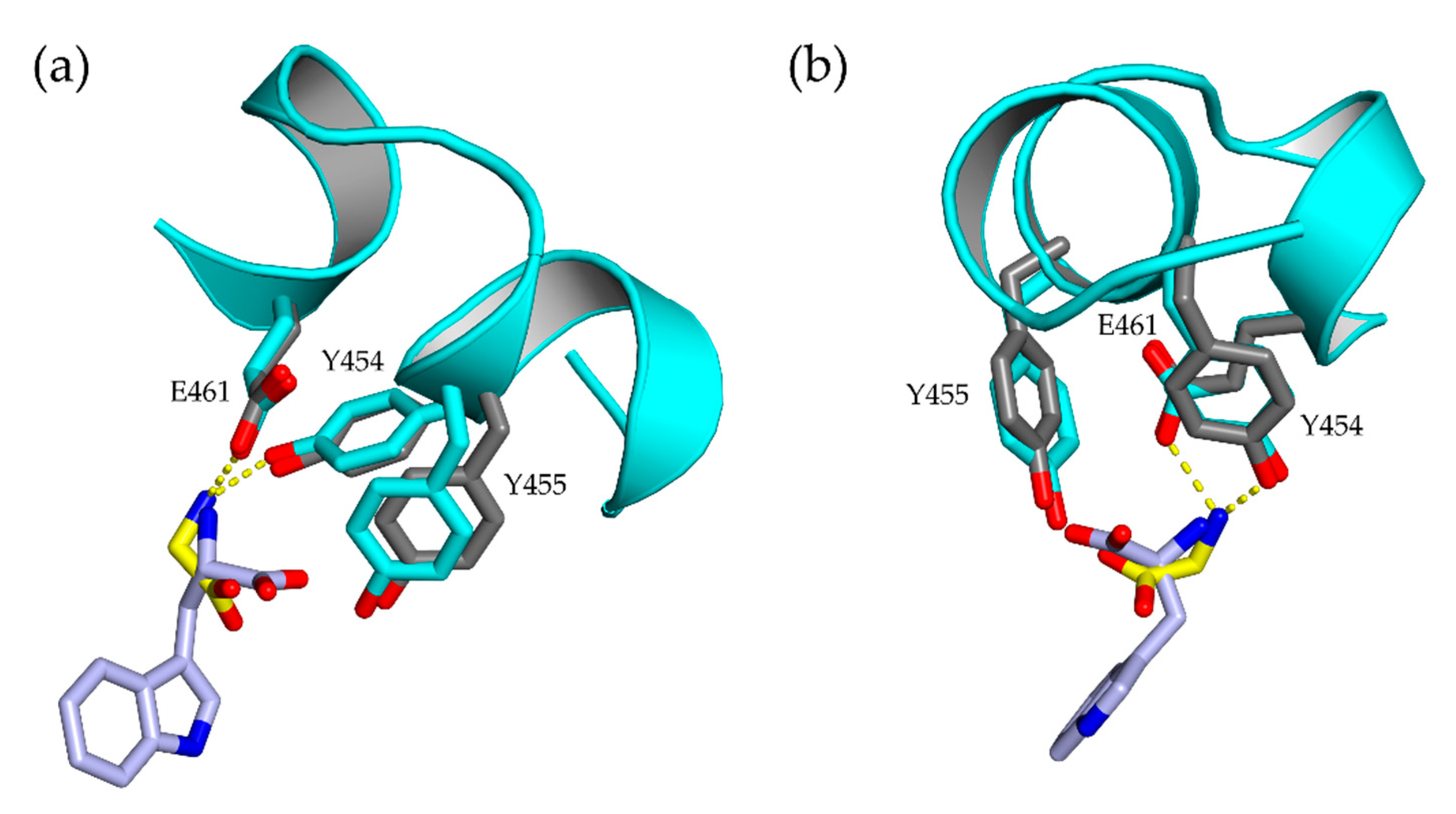
3.3. Crystal Contacts and Possible Impact of the His-tag
4. Discussion
4.1. Differing Functional Regions
4.2. Impact of VM on Diffraction
4.3. Impact of His-tag on Crystallization
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Podzelinska, K.; Latimer, R.; Bhattacharya, A.; Vining, L.C.; Zechel, D.L.; Jia, Z. Chloramphenicol Biosynthesis: The Structure of CmlS, a Flavin-Dependent Halogenase Showing a Covalent Flavin-Aspartate Bond. J. Mol. Biol. 2010, 397, 316–331. [Google Scholar] [CrossRef]
- Williams, D.H.; Bardsley, B. The vancomycin group of antibiotics and the fight against resistant bacteria. Angew. Chem. Int. Ed. 1999, 38, 1172–1193. [Google Scholar] [CrossRef]
- Bischoff, D.; Pelzer, S.; Bister, B.; Nicholson, G.J.; Stockert, S.; Schirle, M.; Wohlleben, W.; Jung, G.; Süssmuth, R.D. The biosynthesis of vancomycin-type glycopeptide antibiotics-The order of the cyclization steps. Angew. Chem. Int. Ed. 2001, 40, 4688–4691. [Google Scholar] [CrossRef]
- Weichold, V.; Milbredt, D.; van Pée, K.-H. Specific Enzymatic Halogenation-From the Discovery of Halogenated Enzymes to Their Applications In Vitro and In Vivo. Angew. Chem. Int. Ed. 2016, 55, 6374–6389. [Google Scholar] [CrossRef] [PubMed]
- Blasiak, L.C.; Drennan, C.L. Structural perspective on enzymatic halogenation. Acc. Chem. Res. 2009, 42, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Pereira, E.R.; Belin, L.; Sancelme, M.; Prudhomme, M.; Ollier, M.; Rapp, M.; Sevère, D.; Riou, J.F.; Fabbro, D.; Meyer, T. Structure-activity relationships in a series of substituted indolocarbazoles: Topoisomerase I and protein kinase C inhibition and antitumoral and antimicrobial properties. J. Med. Chem. 1996, 39, 4471–4477. [Google Scholar] [CrossRef] [PubMed]
- Frese, M.; Guzowska, P.H.; Voß, H.; Sewald, N. Regioselective Enzymatic Halogenation of Substituted Tryptophan Derivatives using the FAD-Dependent Halogenase RebH. ChemCatChem 2014, 6, 1270–1276. [Google Scholar] [CrossRef]
- Schnepel, C.; Sewald, N. Enzymatic Halogenation: A Timely Strategy for Regioselective C−H Activation. Chem. A Eur. J. 2017, 23, 12064–12086. [Google Scholar] [CrossRef]
- Zhu, X.; De Laurentis, W.; Leang, K.; Herrmann, J.; Ihlefeld, K.; van Pée, K.H.; Naismith, J.H. Structural Insights into Regioselectivity in the Enzymatic Chlorination of Tryptophan. J. Mol. Biol. 2009, 391, 74–85. [Google Scholar] [CrossRef]
- Moritzer, A.C.; Minges, H.; Prior, T.; Frese, M.; Sewald, N.; Niemann, H.H. Structure-based switch of regioselectivity in the flavin-dependent tryptophan 6-halogenase Thal. J. Biol. Chem. 2019, 294, 2529–2542. [Google Scholar] [CrossRef]
- Bitto, E.; Huang, Y.; Bingman, C.A.; Singh, S.; Thorson, J.S.; Phillips, G.N. The structure of Flavin-dependent tryptophan 7-halogenase RebH. Proteins Struct. Funct. Bioinform. 2007, 70, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Yeh, E.; Blasiak, L.C.; Koglin, A.; Drennan, C.L.; Walsh, C.T. Chlorination by a long-lived intermediate in the mechanism of flavin-dependent halogenases. Biochemistry 2007, 46, 1284–1292. [Google Scholar] [CrossRef] [PubMed]
- Dong, C.; Flecks, S.; Unversucht, S.; Haupt, C.; Van Pée, K.H.; Naismith, J.H. Structural biology: Tryptophan 7-halogenase (PrnA) structure suggests a mechanism for regioselective chlorination. Science 2005, 309, 2216–2219. [Google Scholar] [CrossRef] [PubMed]
- Moritzer, A.; Niemann, H.H. Binding of FAD and tryptophan to the tryptophan 6-halogenase Thal is negatively coupled. Protein Sci. 2019, 28, 2112–2118. [Google Scholar] [CrossRef] [PubMed]
- Neubauer, P.R.; Widmann, C.; Wibberg, D.; Schröder, L.; Frese, M.; Kottke, T.; Kalinowski, J.; Niemann, H.H.; Sewald, N. A flavin-dependent halogenase from metagenomic analysis prefers bromination over chlorination. PLoS ONE 2018, 13. [Google Scholar] [CrossRef]
- Shepherd, S.A.; Menon, B.R.K.; Fisk, H.; Struck, A.-W.; Levy, C.; Leys, D.; Micklefield, J. A Structure-Guided Switch in the Regioselectivity of a Tryptophan Halogenase. ChemBioChem 2016, 17, 821–824. [Google Scholar] [CrossRef]
- Luhavaya, H.; Sigrist, R.; Chekan, J.R.; McKinnie, S.M.K.; Moore, B.S. Biosynthesis of L-4-Chlorokynurenine, an Antidepressant Prodrug and a Non-Proteinogenic Amino Acid Found in Lipopeptide Antibiotics. Angew. Chem. Int. Ed. 2019, 58, 8394–8399. [Google Scholar] [CrossRef]
- Widmann, C.; Ismail, M.; Sewald, N.; Niemann, H.H. Structure of apo flavin-dependent halogenase Xcc4156 hints at a reason for cofactor-soaking difficulties. Acta Crystallogr. Sect. D Struct. Biol. 2020, 76, 687–697. [Google Scholar] [CrossRef]
- Milbredt, D.; Patallo, E.P.; van Pée, K.-H. A Tryptophan 6-Halogenase and an Amidotransferase Are Involved in Thienodolin Biosynthesis. ChemBioChem 2014, 15, 1011–1020. [Google Scholar] [CrossRef]
- Seibold, C.; Schnerr, H.; Rumpf, J.; Kunzendorf, A.; Hatscher, C.; Wage, T.; Ernyei, A.J.; Dong, C.; Naismith, J.H.; Van Pée, K.-H. A flavin-dependent tryptophan 6-halogenase and its use in modification of pyrrolnitrin biosynthesis. Biocatal. Biotransform. 2006, 24, 401–408. [Google Scholar] [CrossRef]
- Schnepel, C.; Minges, H.; Frese, M.; Sewald, N. A High-Throughput Fluorescence Assay to Determine the Activity of Tryptophan Halogenases. Angew. Chem. Int. Ed. 2016, 55, 14159–14163. [Google Scholar] [CrossRef] [PubMed]
- Cianci, M.; Bourenkov, G.; Pompidor, G.; Karpics, I.; Kallio, J.; Bento, I.; Roessle, M.; Cipriani, F.; Fiedler, S.; Schneider, T.R. P13, the EMBL macromolecular crystallography beamline at the low-emittance PETRA III ring for high- and low-energy phasing with variable beam focusing. J. Synchrotron Rad. 2017, 24, 323–332. [Google Scholar] [CrossRef] [PubMed]
- Kabsch, W. XDS. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Evans, P.R.; Murshudov, G.N. How good are my data and what is the resolution? Acta Crystallogr. Sect. D Biol. Crystallogr. 2013, 69, 1204–1214. [Google Scholar] [CrossRef]
- Winn, M.D.; Ballard, C.C.; Cowtan, K.D.; Dodson, E.J.; Emsley, P.; Evans, P.R.; Keegan, R.M.; Krissinel, E.B.; Leslie, A.G.W.; McCoy, A.; et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. Sect. D Biol. Crystallogr. 2011, 67, 235–242. [Google Scholar] [CrossRef]
- McCoy, A.J.; Grosse-Kunstleve, R.W.; Adams, P.D.; Winn, M.D.; Storoni, L.C.; Read, R.J. Phaser crystallographic software. J. Appl. Crystallogr. 2007, 40, 658–674. [Google Scholar] [CrossRef]
- Terwilliger, T.C.; Grosse-Kunstleve, R.W.; Afonine, P.V.; Moriarty, N.W.; Zwart, P.H.; Hung, L.W.; Read, R.J.; Adams, P.D. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. Sect. D Biol. Crystallogr. 2007, 64, 61–69. [Google Scholar] [CrossRef]
- Adams, P.D.; Afonine, P.V.; Bunkóczi, G.; Chen, V.B.; Davis, I.W.; Echols, N.; Headd, J.J.; Hung, L.W.; Kapral, G.J.; Grosse-Kunstleve, R.W.; et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 213–221. [Google Scholar] [CrossRef]
- Emsley, P.; Lohkamp, B.; Scott, W.G.; Cowtan, K. Features and development of Coot. Acta Crystallogr. Sect. D Biol. Crystallogr. 2010, 66, 486–501. [Google Scholar] [CrossRef]
- Kovalevskiy, O.; Nicholls, R.A.; Long, F.; Carlon, A.; Murshudov, G.N. Overview of refinement procedures within REFMAC 5: Utilizing data from different sources. Acta Crystallogr. Sect. D Struct. Biol. 2018, 74, 215–227. [Google Scholar] [CrossRef]
- Berman, H.; Henrick, K.; Nakamura, H. Announcing the worldwide Protein Data Bank. Nat. Struct. Biol. 2003, 10, 980. [Google Scholar] [CrossRef] [PubMed]
- Holm, L.; Laakso, L.M. Dali server update. Nucleic Acids Res. 2016, 44, W351–W355. [Google Scholar] [CrossRef] [PubMed]
- Gorrec, F. The MORPHEUS protein crystallization screen. J. Appl. Crystallogr. 2009, 42, 1035–1042. [Google Scholar] [CrossRef] [PubMed]
- Krissinel, E.; Henrick, K. Inference of Macromolecular Assemblies from Crystalline State. J. Mol. Biol. 2007, 372, 774–797. [Google Scholar] [CrossRef]
- Matthews, B.W. Solvent content of protein crystals. J. Mol. Biol. 1968, 33, 491–497. [Google Scholar] [CrossRef]
- Kantardjieff, K.A.; Rupp, B. Matthews coefficient probabilities: Improved estimates for unit cell contents of proteins, DNA, and protein-nucleic acid complex crystals. Protein Sci. 2003, 12, 1865–1871. [Google Scholar] [CrossRef]
- Zhao, D.; Huang, Z. Effect of His-tag on Expression, Purification, and Structure of Zinc Finger Protein, ZNF191(243-368). Bioinorg. Chem. Appl. 2016, 2016. [Google Scholar] [CrossRef]
- Munadziroh, E.; Ulfa, E.U.; Labiqah, A.; Asmarani, O.; Puspaningsih, N.N.T. Effect of Poly-Histidine Tag Position toward Inhibition Activity of Secretory Leukocyte Protease Inhibitor as Candidate for Material Wound Healing. Avicenna J. Med. Biotechnol. 2020, 12, 32–36. [Google Scholar]
- Smits, S.H.J.; Mueller, A.; Grieshaber, M.K.; Schmitt, L. Coenzyme- and His-tag-induced crystallization of octopine dehydrogenase. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2008, 64, 836–839. [Google Scholar] [CrossRef]
- Plavša, J.J.; Řezáčová, P.; Kugler, M.; Pachl, P.; Brynda, J.; Voburka, Z.; Celic, A.S.S.; Petri, E.T.; Škerlová, J. In situ proteolysis of an N-terminal His tag with thrombin improves the diffraction quality of human aldo-keto reductase 1C3 crystals. Acta Crystallogr. Sect. F Struct. Biol. Commun. 2018, 74, 300–306. [Google Scholar] [CrossRef]
- Aslantas, Y.; Surmeli, N.B. Effects of N-Terminal and C-Terminal Polyhistidine Tag on the Stability and Function of the Thermophilic P450 CYP119. Bioinorg. Chem. Appl. 2019, 2019, 8080697. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Su, S.; Liu, F.; Xu, T.; Wang, X.; Huang, Y.; Sun, X.; Ge, X.; Chen, T.; Liu, H.; et al. Removal of the tag from His-tagged ILYd4, a human CD59 inhibitor, significantly improves its physical properties and its activity. Curr. Pharm. Des. 2012, 18, 4187–4196. [Google Scholar] [CrossRef] [PubMed]
- Tajika, Y.; Sakai, N.; Tamura, T.; Yao, M.; Watanabe, N.; Tanaka, I. Crystal structure of hypothetical protein PH0828 fromPyrococcus horikoshii. Proteins Struct. Funct. Bioinform. 2004, 57, 862–865. [Google Scholar] [CrossRef] [PubMed]
- Bucher, M.H.; Evdokimov, A.G.; Waugh, D.S. Differential effects of short affinity tags on the crystallization of Pyrococcus furiosus maltodextrin-binding protein. Acta Crystallogr. Sect. D Biol. Crystallogr. 2002, 58, 392–397. [Google Scholar] [CrossRef] [PubMed]
- Carson, M.; Johnson, D.H.; McDonald, H.; Brouillette, C.; DeLucas, L.J. His-tag impact on structure. Acta Crystallogr. Sect. D Biol. Crystallogr. 2007, 63, 295–301. [Google Scholar] [CrossRef] [PubMed]
| NHis-Thal-RebH5 (1) | NHis-Thal-RebH5 (2) | Untagged Thal-RebH5 [10] | |
|---|---|---|---|
| PDB ID | 7AQU | 7AQV | 6IB5 |
| Space group | P21 | P21 | P64 |
| Unit-cell constants | |||
| a, b, c (Å) | 54.7, 118.6, 87.1 | 56.0, 119.1, 87.3 | 139.3, 139.3, 144.3 |
| α, β, γ (°) | 90.0, 104.0, 90.0 | 90.0, 104.8, 90.0 | 90, 90, 120 |
| Data collection statistics | |||
| Wavelength (Å) | 0.9763 | 0.9763 | 0.9763 |
| Resolution range (Å) | 50.0–1.63 (1.67–1.63) | 50.0–1.84 (1.89–1.84) | 50.0–2.12 (2.18–2.12) |
| No. of reflections (measured/unique) | 907,692 (56,713)/129,082 (8424) | 657,026 (49,313)/95,688 (7071) | 1,607,569 (124,056)/89845 (6649) |
| Completeness (%) | 96.6 (85.3) | 99.8 (99.9) | 100 (100) |
| Rmeas (%) | 7.9 (114.6) | 7.7 (118.2) | 7.6 (185.7) |
| Redundancy | 7.03 (6.73) | 6.87 (6.97) | 17.9 (18.7) |
| Mean I/σ(I) | 15.33 (1.97) | 14.44 (1.97) | 20.74 (2.05) |
| CC ½ (%) | 99.9 (74.8) | 99.9 (80.1) | 100 (76.3) |
| Wilson B factor(Å2) | 31.4 | 38.6 | 59.9 |
| Refinement and model statistics | |||
| Resolution range | 48.58–1.63 (1.67–1.63) | 49.28–1.84 (1.86–1.84) | 48.11–2.12 (2.14–2.12) |
| No. of reflections (work/test) | 129,042 (7970)/6474 (438) | 95,617 (3059)/4732 (144) | 89,790 (2869)/4494 (150) |
| Rwork | 0.167 | 0.166 | 0.163 |
| Rfree | 0.202 | 0.199 | 0.192 |
| RMSD bonds (Å) | 0.010 | 0.004 | 0.007 |
| RMSD angles (°) | 1.595 | 0.655 | 0.866 |
| No. of atoms | 9667 | 9316 | 8962 |
| Protein | 8638 | 8495 | 8456 |
| Ligands | 40 | 12 | 86 |
| Solvent | 989 | 809 | 420 |
| Average B factor (Å2) | 29.7 | 41.1 | 64.0 |
| Protein | 28.7 | 40.7 | 63.7 |
| Ligands | 31.9 | 52.3 | 100.0 |
| Solvent | 38.3 | 44.7 | 62.0 |
| Ramachandran | |||
| Favored (%) | 97.88 | 97.40 | 97.31 |
| Allowed (%) | 2.12 | 2.60 | 2.69 |
| Outliers (%) | 0.0 | 0.0 | 0.0 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Moritzer, A.-C.; Prior, T.; Niemann, H.H. Not Cleaving the His-tag of Thal Results in More Tightly Packed and Better-Diffracting Crystals. Crystals 2020, 10, 1135. https://doi.org/10.3390/cryst10121135
Moritzer A-C, Prior T, Niemann HH. Not Cleaving the His-tag of Thal Results in More Tightly Packed and Better-Diffracting Crystals. Crystals. 2020; 10(12):1135. https://doi.org/10.3390/cryst10121135
Chicago/Turabian StyleMoritzer, Ann-Christin, Tina Prior, and Hartmut H. Niemann. 2020. "Not Cleaving the His-tag of Thal Results in More Tightly Packed and Better-Diffracting Crystals" Crystals 10, no. 12: 1135. https://doi.org/10.3390/cryst10121135
APA StyleMoritzer, A.-C., Prior, T., & Niemann, H. H. (2020). Not Cleaving the His-tag of Thal Results in More Tightly Packed and Better-Diffracting Crystals. Crystals, 10(12), 1135. https://doi.org/10.3390/cryst10121135





