Characterization of a Metagenome-Derived β-Glucosidase and Its Application in Conversion of Polydatin to Resveratrol
Abstract
:1. Introduction
2. Results and Discussion
2.1. Construction of the Metagenomic Library and Screening for Clones with β-Glucosidase Activity
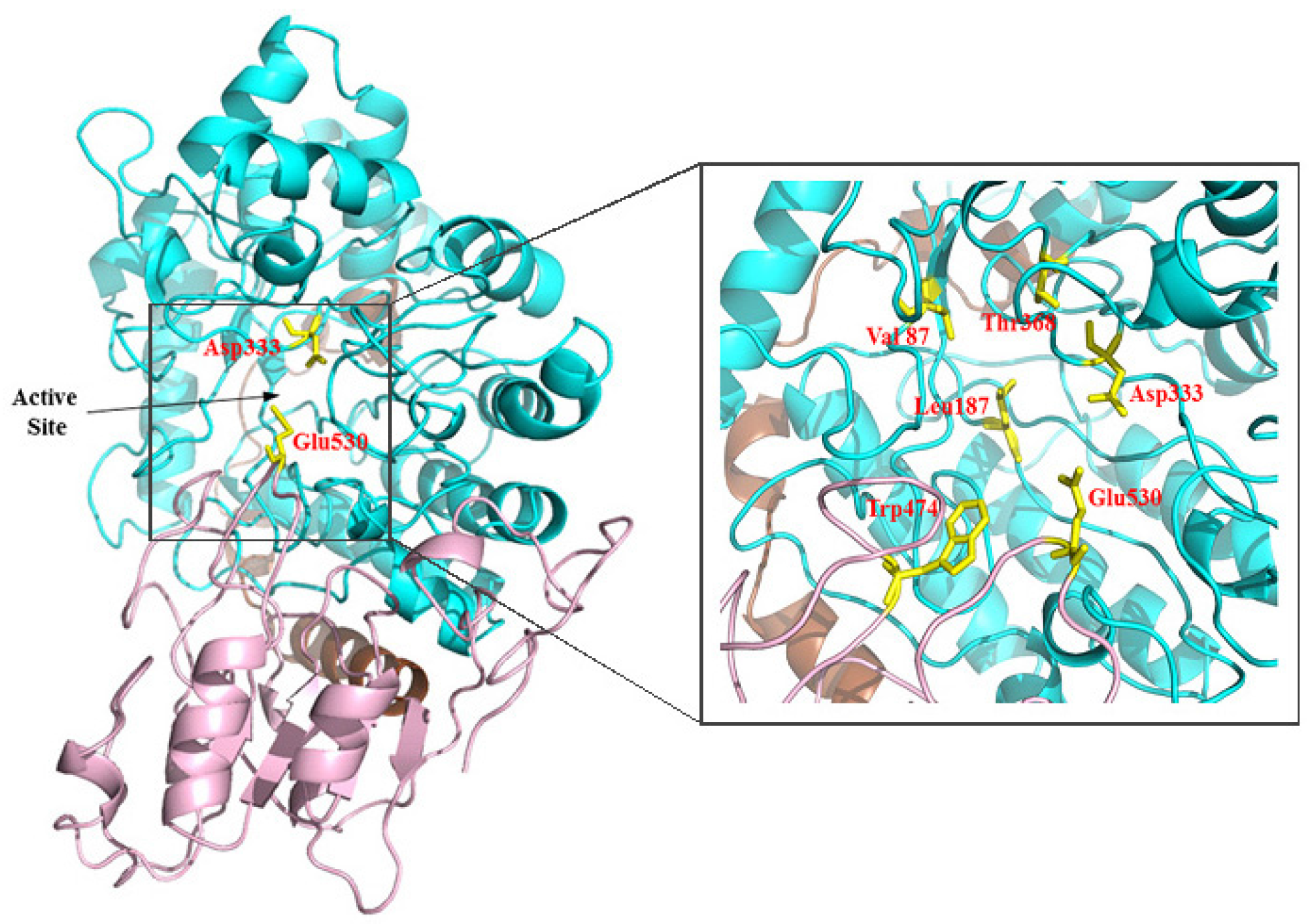
2.2. Sequence Analysis
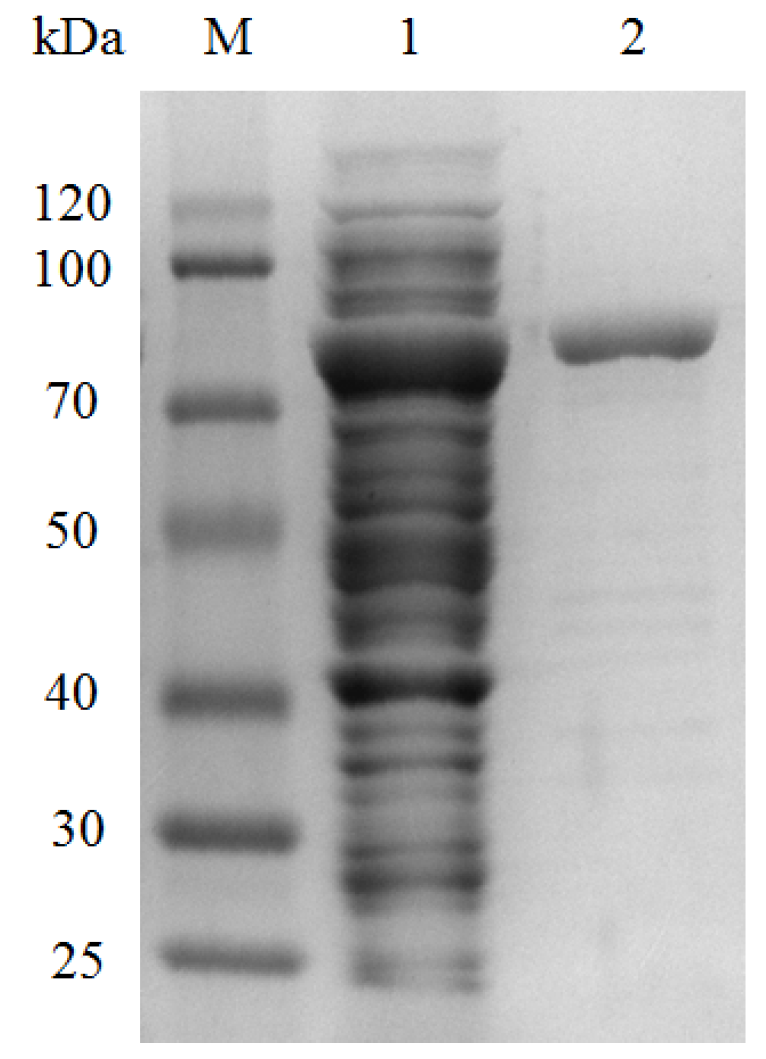
2.3. Enzyme Overexpression and Purification
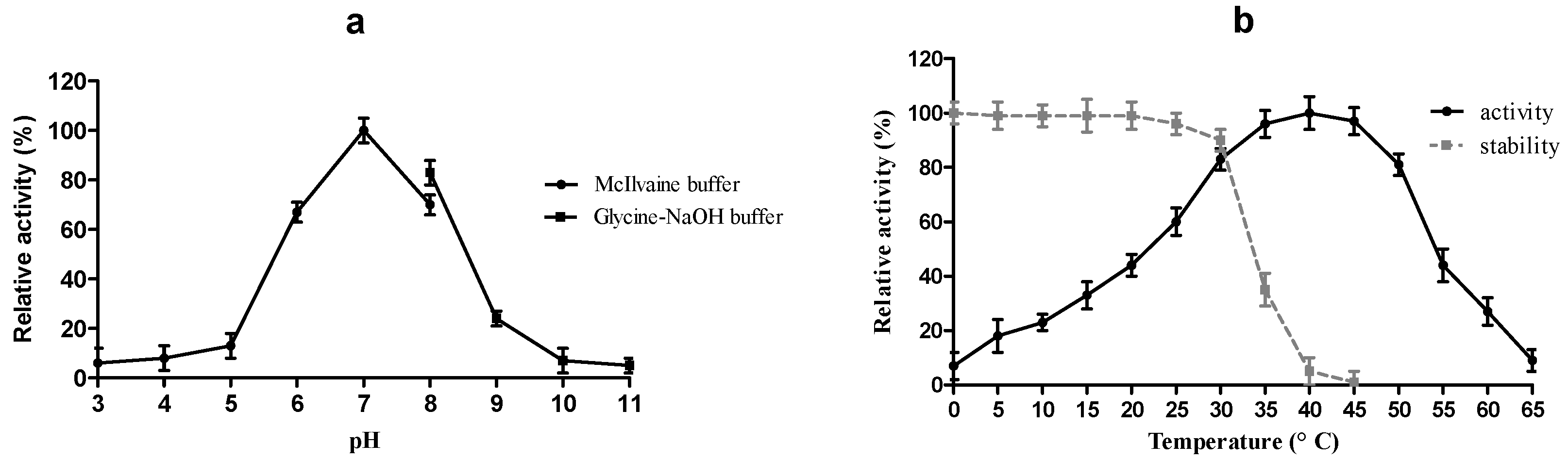
2.4. Physico-Chemical Characterization of r-MlBgl
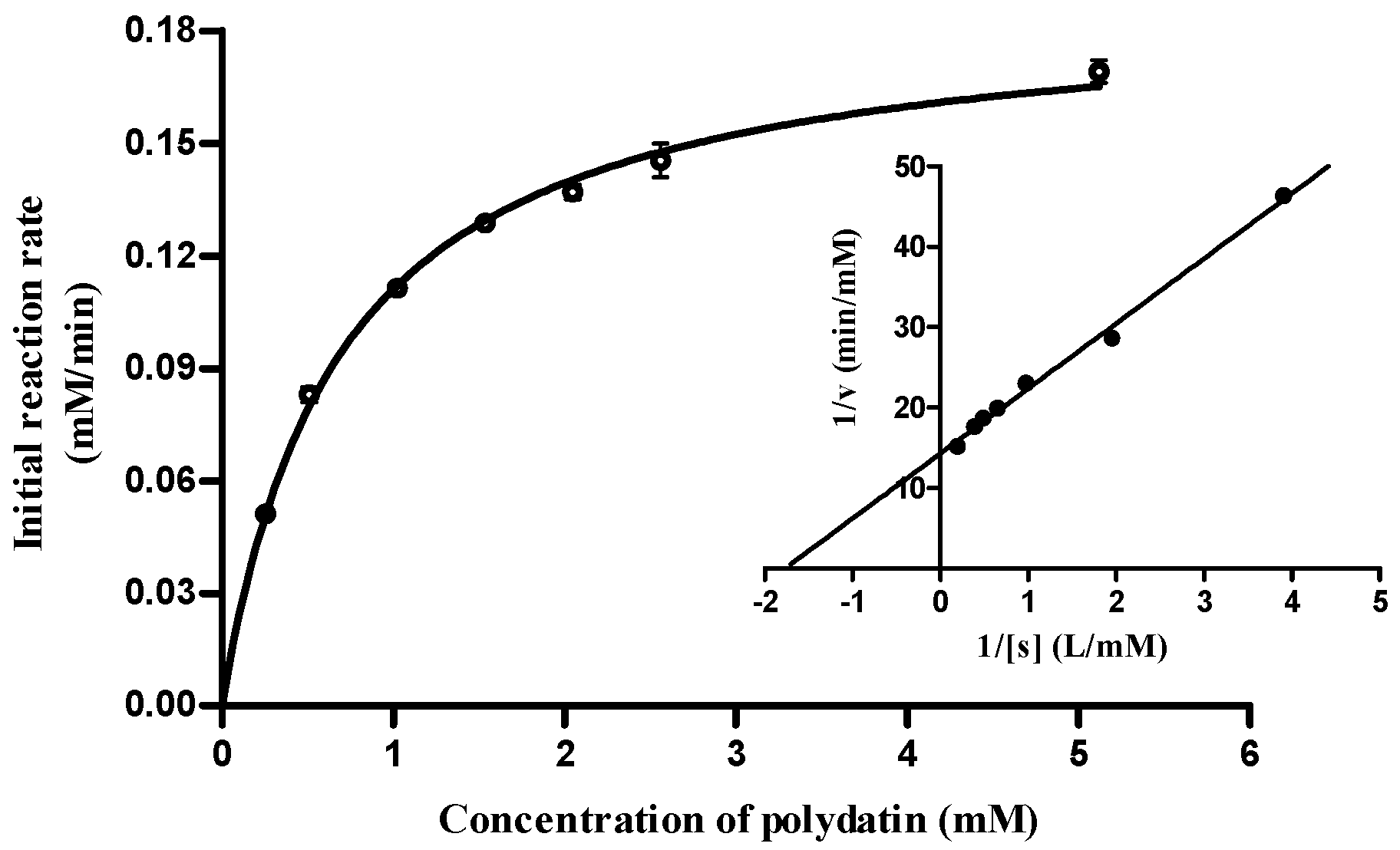
2.5. Substrate Specificity and Enzymatic Kinetics
2.6. Enzymatic Conversion of Polydatin to Resveratrol
3. Experimental Section
3.1. Strains, Vectors and Reagents
3.2. Metagenomic Library Construction
3.3. Screening of β-Glucosidase Gene MlBgl
3.4. Sequence Analysis
3.5. Gene Expression and Purification of Recombinant β-Glucosidase r-MlBgl
3.6. Enzyme Activity Assay
3.7. Physico-Chemical Characterization of r-MlBgl
3.8. Substrate Specificity
3.9. Enzymatic Conversion of Polydatin to Resveratrol
3.10. Nucleotide Sequence Accession Number
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bertelli, A.; Ferrara, F.; Diana, G.; Fulgenzi, A.; Corsi, M.; Ponti, W.; Ferrero, M.; Bertelli, A. Resveratrol, a natural stilbene in grapes and wine, enhances intraphagocytosis in human promonocytes: A co-factor in antiinflammatory and anticancer chemopreventive activity. Int. J. Tissue React. 1998, 21, 93–104. [Google Scholar]
- Fremont, L. Biological effects of resveratrol. Life Sci. 2000, 66, 663–673. [Google Scholar] [CrossRef]
- Jiang, L.; Gu, Y.; Ye, J.; Liu, F.; Zhao, Y.; Wang, C.; Xu, Y.; Cao, X.; Zhang, L.; Dong, W. Resveratrol prevents hepatic steatosis induced by hepatitis C virus core protein. Biotechnol. Lett. 2012, 34, 2205–2212. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Patán, F.; Monagas, M.; Moreno-Arribas, M.V.; Bartolomé, B. Determination of microbial phenolic acids in human faeces by UPLC-ESI-TQ MS. J. Agric. Food Chem. 2011, 59, 2241–2247. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Liu, L.; Guo, Y.X.; Dong, Y.S.; Zhang, D.J.; Xiu, Z.L. Biotransformation of piceid in Polygonum cuspidatum to resveratrol by Aspergillus oryzae. Appl. Microbiol. Biotechnol. 2007, 75, 763–768. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.; Yan, A.; Yang, X.; Cai, Y.; Chen, J. An optimum fermentation model established by genetic algorithm for biotransformation from crude polydatin to resveratrol. Appl. Biochem. Biotechnol. 2012, 166, 446–457. [Google Scholar] [CrossRef] [PubMed]
- Voorhorst, W.; Eggen, R.; Luesink, E.J.; De Vos, W. Characterization of the celB gene coding for beta-glucosidase from the hyperthermophilic archaeon Pyrococcus furiosus and its expression and site-directed mutation in Escherichia coli. J. Bacteriol. 1995, 177, 7105–7111. [Google Scholar] [PubMed]
- Shewale, J. β-Glucosidase: Its role in cellulase synthesis and hydrolysis of cellulose. Int. J. Biochem. 1982, 14, 435–443. [Google Scholar] [CrossRef]
- Hu, Y.; Luan, H.; Zhou, K.; Ge, G.; Yang, S.; Yang, L. Purification and characterization of a novel glycosidase from the china white jade snail (Achatina fulica) showing transglycosylation activity. Enzym. Microb. Technol. 2008, 43, 35–42. [Google Scholar] [CrossRef]
- González-Pombo, P.; Fariña, L.; Carrau, F.; Batista-Viera, F.; Brena, B.M. A novel extracellular β-glucosidase from Issatchenkia terricola: Isolation, immobilization and application for aroma enhancement of white Muscat wine. Process Biochem. 2011, 46, 385–389. [Google Scholar] [CrossRef]
- Du, L.; Wang, Z.; Zhao, Y.; Huang, J.; Pang, H.; Wei, Y.; Lin, L.; Huang, R. A β-glucosidase from Novosphingobium sp. GX9 with high catalytic efficiency toward isoflavonoid glycoside hydrolysis and (+)-catechin transglycosylation. Appl. Microbiol. Biotechnol. 2014, 98, 7069–7079. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Li, D.; Gao, Z.; Zhang, C. Enzymatic transformation of polydatin to resveratrol by piceid-β-d-glucosidase from Aspergillus oryzae. Bioprocess Biosyst. Eng. 2014, 37, 1411–1416. [Google Scholar] [CrossRef] [PubMed]
- Loughlin, W.A. Biotransformations in organic synthesis. Bioresour. Technol. 2000, 74, 49–62. [Google Scholar] [CrossRef]
- Tian, T.; Sun, Q.; Shen, J.; Zhang, T.; Gao, P.; Sun, Q. Microbial transformation of polydatin and emodin-8-β-d-glucoside of Polygonum cuspidatum Sieb. et Zucc into resveratrol and emodin respectively by Rhizopus microsporus. World J. Microbiol. Biotechnol. 2008, 24, 861–866. [Google Scholar] [CrossRef]
- Jin, S.; Luo, M.; Wang, W.; Zhao, C.J.; Gu, C.B.; Li, C.Y.; Zu, Y.G.; Fu, Y.J.; Guan, Y. Biotransformation of polydatin to resveratrol in Polygonum cuspidatum roots by highly immobilized edible Aspergillus niger and Yeast. Bioresour. Technol. 2013, 136, 766–770. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Li, D.; Yu, H.; Zhang, B.; Jin, F. Purification and characterization of piceid-β-d-glucosidase from Aspergillus oryzae. Process Biochem. 2007, 42, 83–88. [Google Scholar] [CrossRef]
- Ko, J.A.; Park, J.; Kwon, H.; Ryu, Y.; Jeong, H.; Park, S.; Kim, C.; Oh, H.; Park, C.; Lim, Y. Purification and functional characterization of the first stilbene glucoside-specific β-glucosidase isolated from Lactobacillus kimchi. Enzyme Microbiol. Technol. 2014, 67, 59–66. [Google Scholar] [CrossRef]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
- Jiang, C.; Li, S.X.; Luo, F.F.; Jin, K.; Wang, Q.; Hao, Z.Y.; Wu, L.L.; Zhao, G.C.; Ma, G.F.; Shen, P.H. Biochemical characterization of two novel β-glucosidase genes by metagenome expression cloning. Bioresour. Technol. 2011, 102, 3272–3278. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Jiang, Y.; Fan, X.J.; Liu, Y.H. Molecular cloning and characterization of a novel β-glucosidase with high hydrolyzing ability for soybean isoflavone glycosides and glucose-tolerance from soil metagenomic library. Bioresour. Technol. 2012, 123, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Duan, C.J.; Liu, L.; Tang, J.L.; Feng, J.X. Properties of a metagenome-derived β-glucosidase from the contents of rabbit cecum. Biosci. Biotechnol. Biochem. 2009, 73, 1470–1473. [Google Scholar] [CrossRef] [PubMed]
- Nacke, H.; Engelhaupt, M.; Brady, S.; Fischer, C.; Tautzt, J.; Daniel, R. Identification and characterization of novel cellulolytic and hemicellulolytic genes and enzymes derived from German grassland soil metagenomes. Biotechnol. Lett. 2012, 34, 663–675. [Google Scholar] [CrossRef] [PubMed]
- Mai, Z.; Su, H.; Li, L.; Zhang, S. Construction of a mangrove soil metagenome library and identification of two novel β-glucosidase genes. Biotechnol. Bull. 2014, 6, 168–172. [Google Scholar]
- Kim, S.J.; Lee, C.M.; Han, B.R.; Kim, M.Y.; Yeo, Y.S.; Yoon, S.H.; Koo, B.S.; Jun, H.K. Characterization of a gene encoding cellulase from uncultured soil bacteria. FEMS Microbiol. Lett. 2008, 282, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Hao, Z.Y.; Jin, K.; Li, S.X.; Che, Z.Q.; Ma, G.F.; Wu, B. Identification of a metagenome-derived β-glucosidase from bioreactor contents. J. Mol. Catal. B 2010, 63, 11–16. [Google Scholar] [CrossRef]
- Mao, X.; Hong, Y.; Shao, Z.; Zhao, Y.; Liu, Z. A novel cold-active and alkali-stable β-glucosidase gene isolated from the marine bacterium Martelella mediterrânea. Appl. Biochem. Biotechnol. 2010, 162, 2136–2148. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.K.; Chir, J.; Tanaka, S.; Chen, F.Y. Identification of the general acid/base catalyst of a family 3β-glucosidase from Flavobacterium meningosepticum. Biochemistry 2002, 41, 2751–2759. [Google Scholar] [CrossRef] [PubMed]
- Vocadlo, D.J.; Mayer, C.; He, S.; Withers, S.G. Mechanism of action and identification of Asp242 as the catalytic nucleophile of Vibrio furnisii N-acetyl-β-d-glucosaminidase using 2-acetamido-2-deoxy-5-fluoro-α-l-idopyranosyl fluoride. Biochemistry 2000, 39, 117–126. [Google Scholar] [CrossRef] [PubMed]
- Zechel, D.L.; Withers, S.G. Glycosidase mechanisms: anatomy of a finely tuned catalyst. Acc. Chem. Res. 2000, 33, 11–18. [Google Scholar] [PubMed]
- Varghese, J.N.; Hrmova, M.; Fincher, G.B. Three-dimensional structure of a barley β-d-glucan exohydrolase, a family 3 glycosyl hydrolase. Structure 1999, 7, 179–190. [Google Scholar] [CrossRef]
- Todaro, A.; Palmeri, R.; Barbagallo, R.N.; Pifferi, P.G.; Spagna, G. Increase of trans-resveratrol in typical Sicilian wine using β-glucosidase from various sources. Food Chem. 2008, 107, 1570–1575. [Google Scholar] [CrossRef]
- Shipkowski, S.; Brenchley, J.E. Characterization of an unusual cold-active β-glucosidase belonging to family 3 of the glycoside hydrolases from the psychrophilic isolate Paenibacillus sp. strain C7. Appl. Environ. Microbiol. 2005, 71, 4225–4232. [Google Scholar] [CrossRef] [PubMed]
- Siddiqui, K.S.; Cavicchioli, R. Cold-adapted enzymes. Annu. Rev. Biochem. 2006, 75, 403–433. [Google Scholar] [CrossRef] [PubMed]
- Crespim, E.; Zanphorlin, L.M.; de Souza, F.H.; Diogo, J.A.; Gazolla, A.C.; Machado, C.B.; Figueiredo, F.; Sousa, A.S.; Nóbrega, F.; Pellizari, V.H. A novel cold-adapted and glucose-tolerant GH1 β-glucosidase from Exiguobacterium antarcticum B7. Int. J. Biol. Macromol. 2016, 82, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.X.; Miao, L.L.; Liu, Y.; Liu, H.C.; Liu, Z.P. Gene cloning and characterization of a cold-adapted β-glucosidase belonging to glycosyl hydrolase family 1 from a psychrotolerant bacterium Micrococcus antarcticus. Enzym. Microbiol. Technol. 2011, 49, 94–99. [Google Scholar] [CrossRef] [PubMed]
- Gu, N.Y.; Kim, J.L.; Kim, H.J.; You, D.J.; Kim, H.W.; Jeon, S.J. Gene cloning and enzymatic properties of hyperthermostable β-glycosidase from Thermus thermophilus HJ6. J. Biosci. Bioeng. 2009, 107, 21–26. [Google Scholar] [CrossRef] [PubMed]
- Parry, N.; Beever, D.; Owen, E.; Vandenberghe, I.; Van Beeumen, J.; Bhat, M. Biochemical characterization and mechanism of action of a thermostable β-glucosidase purified from Thermoascus aurantiacus. Biochem. J. 2001, 353, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Suen, W.C.; Windsor, W.; Xiao, L.; Madison, V.; Zaks, A. Improving tolerance of Candida antarctica lipase B towards irreversible thermal inactivation through directed evolution. Protein Eng. 2003, 16, 599–605. [Google Scholar] [CrossRef] [PubMed]
- Kudo, K.; Watanabe, A.; Ujiie, S.; Shintani, T.; Gomi, K. Purification and enzymatic characterization of secretory glycoside hydrolase family 3 (GH3) aryl β-glucosidases screened from Aspergillus oryzae genome. J. Biosci. Bioeng. 2015, 120, 614–623. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, Y.; Mishra, S.; Bisaria, V. Microbial β-glucosidases: Cloning, properties, and applications. Crit. Rev. Biotechnol. 2002, 22, 375–407. [Google Scholar] [CrossRef] [PubMed]
- Yeom, S.J.; Kim, B.N.; Kim, Y.S.; Oh, D.K. Hydrolysis of isoflavone glycosides by a thermostable β-glucosidase from Pyrococcus furiosus. J. Agric. Food Chem. 2012, 60, 1535–1541. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Mai, Z.; Yang, J.; Tian, X.; Li, J.; Zhang, S. Gene cloning and characterization of a novel salt-tolerant and glucose-enhanced β-glucosidase from a marine Streptomycete. Appl. Biochem. Biotechnol. 2013, 169, 1512–1522. [Google Scholar] [CrossRef] [PubMed]
- Somogyi, M. Notes on sugar determination. J. Biol. Chem. 1952, 195, 19–23. [Google Scholar]

| Reagents | Relative Activity (%) a | |
|---|---|---|
| Concentration (1 mM) | Concentration (10 mM) | |
| None | 100.0 ± 2.6 b | 100.0 ± 4.2 |
| Na+ | 98.9 ± 5.4 | 81.53 ± 3.4 |
| K+ | 104.5 ± 2.5 | 85.6 ± 3.2 |
| Li+ | 91 ± 3.6 | 80 ± 4.1 |
| NH4+ | 102.4 ± 4.2 | 93.3 ± 4.8 |
| Mg2+ | 104.1 ± 4.3 | 88.1 ± 3.5 |
| Fe3+ | 96.5 ± 3.2 | 58.8 ± 3.6 |
| Zn2+ | 12.3 ± 2.4 | 5.7 ± 1.1 |
| Ca2+ | 58.4 ± 4.3 | 45.6 ± 2.9 |
| Cu2+ | 8.2 ± 2.3 | 5.9 ± 1.4 |
| Ni2+ | 82.4 ± 4.2 | 59.3 ± 3.8 |
| Ba2+ | 44.5 ± 3.4 | 23.4 ± 3.1 |
| Co2+ | 66.1 ± 3.5 | 44.7 ± 4.6 |
| EDTA | 56.4 ± 4.7 | 25.9 ± 3.8 |
| SDS | 76.4 ± 4.8 | 55.2 ± 3.1 |
| Substrates | Configuration of Glycoside Linkage | Specific Activity (U·mg−1) a |
|---|---|---|
| Aryl β-glycosides | - | - |
| p-Nitrophenyl-β-d-glucopyranoside | β-Glucose | 248.26 |
| p-Nitrophenyl-β-d-galactopyranoside | β-Galactose | 28.79 |
| p-Nitrophenyl-β-d-cellobioside | β-Cellobiose | ND |
| p-Nitrophenyl-β-d-xylopyranoside | β-Xylose | 18.87 |
| Polydatin | β-Glucose | 422.21 |
| Esculin | β-Glucose | 126.69 |
| Oligosaccharides and polysaccharides | - | - |
| Cellobiose | β-(1,4)-Glucose | ND |
| Cellotriose | β-(1,4)-Glucose | ND |
| Carboxymethyl cellulose | β-(1,4)-Glucose | ND |
| Avicel | β-(1,4)-Glucose | ND |
| Starch | α-(1,4) and α-(1,6)-Glucose | ND |
| Enzyme Origin | Amounts | Time (h) | Productivity (mM/h) | Km (mM) | kcat (s−1) | kcat/Km (mM−1·s−1) | References |
|---|---|---|---|---|---|---|---|
| Lactobacillus kimchi | 3.28 μg/mL | 40 | 0.64 | 0.20 | 1.29 | 6.45 | [17] |
| Aspergillus oryzae | NR a | 2 | NR | 0.74 | NR | NR | [16] |
| Aspergillus oryzae | 5 U/mL | 4 | 5.12 | NR | NR | NR | [12] |
| Metagenomic library | 0.13 U/mL | 1 | 1.28 b | 0.67 | 989 | 1476 | This study |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mai, Z.; Su, H.; Zhang, S. Characterization of a Metagenome-Derived β-Glucosidase and Its Application in Conversion of Polydatin to Resveratrol. Catalysts 2016, 6, 35. https://doi.org/10.3390/catal6030035
Mai Z, Su H, Zhang S. Characterization of a Metagenome-Derived β-Glucosidase and Its Application in Conversion of Polydatin to Resveratrol. Catalysts. 2016; 6(3):35. https://doi.org/10.3390/catal6030035
Chicago/Turabian StyleMai, Zhimao, Hongfei Su, and Si Zhang. 2016. "Characterization of a Metagenome-Derived β-Glucosidase and Its Application in Conversion of Polydatin to Resveratrol" Catalysts 6, no. 3: 35. https://doi.org/10.3390/catal6030035
APA StyleMai, Z., Su, H., & Zhang, S. (2016). Characterization of a Metagenome-Derived β-Glucosidase and Its Application in Conversion of Polydatin to Resveratrol. Catalysts, 6(3), 35. https://doi.org/10.3390/catal6030035





