ALK in Non-Small Cell Lung Cancer (NSCLC) Pathobiology, Epidemiology, Detection from Tumor Tissue and Algorithm Diagnosis in a Daily Practice
Abstract
:1. Introduction
2. The ALK Rearrangement in Lung Cancer: Mechanism and Consequences
3. Epidemiology of Lung Cancers with an ALK Rearrangement
4. Methods for Detection of an ALK Rearrangement
4.1. Immunohistochemistry and Immunocytochemistry Methods
4.2. The FISH Method
4.3. RT-PCR Method
4.4. NGS Approach
4.5. Other Methods
4.6. Comparatives Analyses
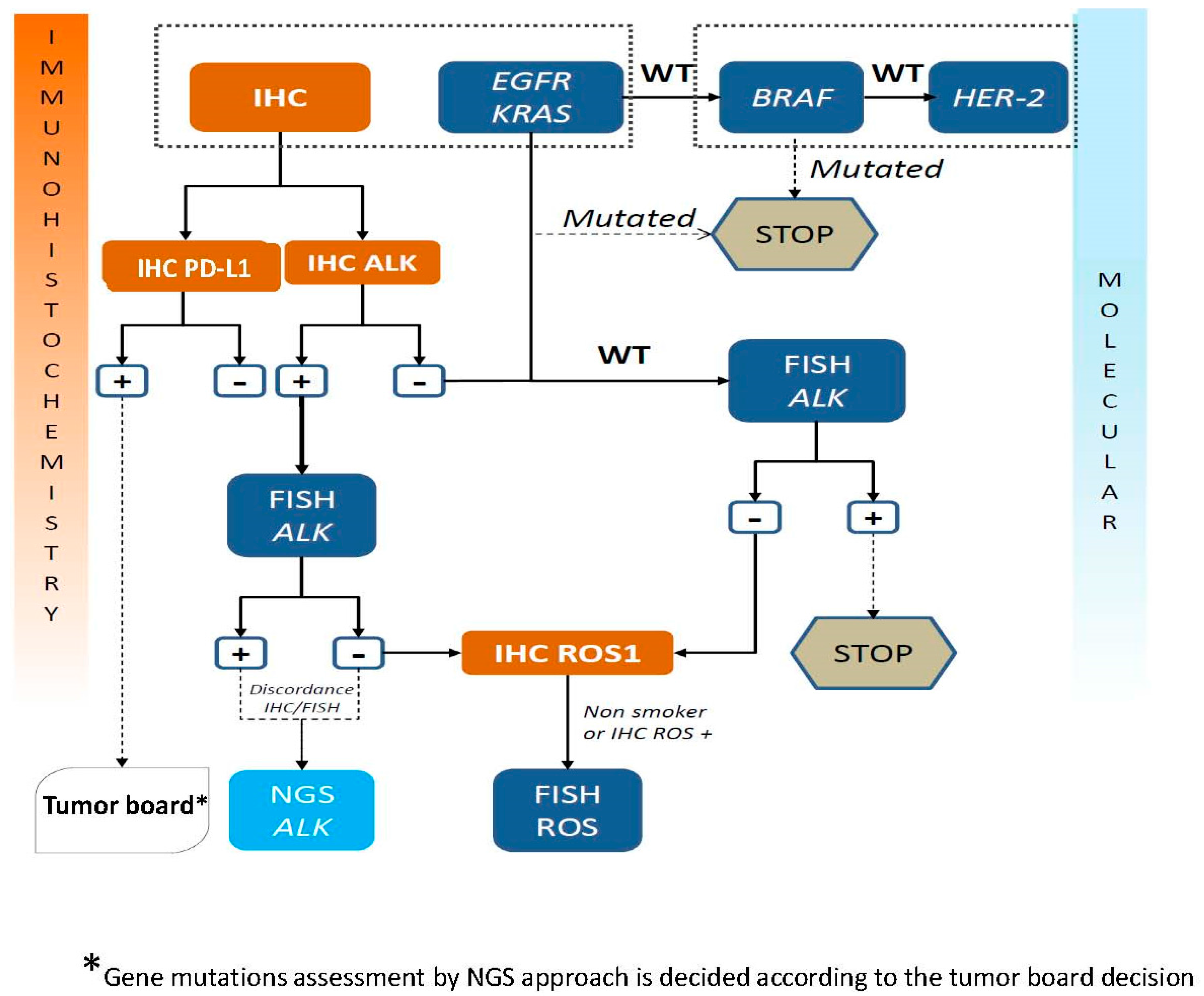
5. Algorithms for Diagnosis of an ALK Rearrangement
6. Conclusions
Acknowledgments
Conflicts of Interest
References
- Shaw, A.T.; Engelman, J.A. ALK in lung cancer: Past, present, and future. J. Clin. Oncol. 2013, 31, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Ilie, M.; Hofman, P. Expanding opportunities for crizotinib resistance in ALK-positive lung cancer patients. Transl. Cancer Res. 2016, 5, 203–205. [Google Scholar] [CrossRef]
- Peters, S.; Camidge, D.R.; Shaw, A.T.; Gadgeel, S.; Ahn, J.S.; Kim, D.W.; Ou, S.I.; Pérol, M.; Dziadziuszko, R.; Rosell, R.; et al. Alectinib versus crizotinib in untreated ALK-positive non-small-cell lung cancer. N. Engl. J. Med. 2017. [Google Scholar] [CrossRef] [PubMed]
- Di Maio, M.; de Marinis, F.; Hirsch, F.R.; Gridelli, C. Diagnostic and therapeutic issues for patients with advanced non-small cell lung cancer harboring anaplastic lymphoma kinase rearrangement: European vs. US perspective (review). Int. J. Oncol. 2014, 45, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Qian, M.; Zhu, B.; Wang, X.; Liebman, M. Drug resistance in ALK-positive non-small cell lung cancer patients. Semin. Cell Dev. Biol. 2017, 64, 150–157. [Google Scholar] [PubMed]
- Khozin, S.; Blumenthal, G.M.; Zhang, L.; Tang, S.; Brower, M.; Fox, E.; Helms, W.; Leong, R.; Song, P.; Pan, Y.; et al. FDA approval: Ceritinib for the treatment of metastatic anaplastic lymphoma kinase-positive non-small cell lung cancer. Clin. Cancer Res. 2015, 21, 2436–2439. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Tiseo, M.; Ahn, M.J.; Reckamp, K.L.; Hansen, K.H.; Kim, S.W.; Huber, R.M.; West, H.L.; Groen, H.J.M.; Hochmair, M.J.; et al. Brigatinib in patients with crizotinib-refractory anaplastic lymphoma kinase-positive non-small-cell lung cancer: A randomized, multicenter phase II trial. J. Clin. Oncol. 2017, 35, 2490–2498. [Google Scholar] [CrossRef] [PubMed]
- Larkins, E.; Blumenthal, G.M.; Chen, H.; He, K.; Agarwal, R.; Gieser, G.; Stephens, O.; Zahalka, E.; Ringgold, K.; Helms, W.; et al. FDA approval: Alectinib for the treatment of metastatic, ALK-positive non-small cell lung cancer following crizotinib. Clin. Cancer Res. 2016, 22, 5171–5176. [Google Scholar] [CrossRef] [PubMed]
- Markham, A. Brigatinib: First global approval. Drugs 2017, 77, 1131–1135. [Google Scholar] [CrossRef] [PubMed]
- Santarpia, M.; Daffinà, M.G.; D’Aveni, A.; Marabello, G.; Liguori, A.; Giovannetti, E.; Karachaliou, N.; Gonzalez Cao, M.; Rosell, R.; Altavilla, G. Spotlight on ceritinib in the treatment of ALK+ NSCLC: Design, development and place in therapy. Drug Des. Devel. Ther. 2017, 11, 2047–2063. [Google Scholar] [CrossRef] [PubMed]
- Barlesi, F.; Mazieres, J.; Merlio, J.P.; Debieuvre, D.; Mosser, J.; Lena, H.; Ouafik, L.; Besse, B.; Rouquette, I.; Westeel, V.; et al. Routine molecular profiling of patients with advanced non-small-cell lung cancer: Results of a 1-year nationwide programme of the French Cooperative Thoracic Intergroup (IFCT). Lancet 2016, 387, 1415–1426. [Google Scholar] [CrossRef]
- Le, T.; Gerber, D.E. ALK alterations and inhibition in lung cancer. Semin. Cancer Biol. 2017, 42, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Schildgen, V.; Lochi, V.; Lüsebrink, J.; Brockmann, M.; Schidgen, O. Epidemiology of EML4-ALK translocations in a small German NSCLC patient cohort. Per. Med. 2012, 9, 801–803. [Google Scholar] [CrossRef]
- Schildgen, V.; Lüsebrink, J.; Schildgen, O.; Stoelben, E.; Brockmann, M. Epidemiology of KRAS, BRAF, and EGFR mutations in lung cancer in a German tertiary hospital in patients with testing indication. Per. Med. 2016, 13, 315–323. [Google Scholar] [CrossRef]
- Schildgen, V.; Schildgen, O. Routine molecular profiling of patients with NSCLC. Lancet 2016, 388, 1054. [Google Scholar]
- Dagogo-Jack, I.; Shaw, A.T. Crizotinib resistance: Implications for therapeutic strategies. Ann. Oncol. 2016, 27, iii42–iii50. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.J.; Riely, G.J.; Shaw, A.T. Targeting ALK: Precision medicine takes on drug resistance. Cancer Discov. 2017, 7, 137–155. [Google Scholar] [CrossRef] [PubMed]
- Gainor, J.F.; Dardaei, L.; Yoda, S.; Friboulet, L.; Leshchiner, I.; Katayama, R.; Dagogo-Jack, I.; Gadgeel, S.; Schultz, K.; Singh, M.; et al. Molecular mechanisms of resistance to first- and second-generation ALK Inhibitors in ALK-rearranged lung cancer. Cancer Discov. 2016, 61118–61133. [Google Scholar]
- Isozaki, H.; Ichihara, E.; Takigawa, N.; Ohashi, K.; Ochi, N.; Yasugi, M.; Ninomiya, T.; Yamane, H.; Hotta, K.; Sakai, K.; et al. Non-small cell lung cancer cells acquire resistance to the ALK inhibitor alectinib by activating alternative receptor tyrosine kinases. Cancer Res. 2016, 76, 1506–1516. [Google Scholar] [CrossRef] [PubMed]
- Ou, S.I.; Lee, T.K.; Young, L.; Fernandez-Rocha, M.Y.; Pavlick, D.; Schrock, A.B.; Zhu, V.W.; Milliken, J.; Ali, S.M.; Gitlitz, B.J. Dual occurrence of ALK G1202R solvent front mutation and small cell lung cancer transformation as resistance mechanisms to second generation ALK inhibitors without prior exposure to crizotinib. Pitfall of solely relying on liquid re-biopsy? Lung Cancer 2017, 106, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Allouche, M. ALK is a novel dependence receptor: Potential implications in development and cancer. Cell Cycle 2007, 6, 1533–1538. [Google Scholar] [CrossRef] [PubMed]
- McDermott, U.; Iafrate, A.J.; Gray, N.S.; Shioda, T.; Classon, M.; Maheswaran, S.; Zhou, W.; Choi, H.G.; Smith, S.L.; Dowell, L.; et al. Genomic alterations of anaplastic lymphoma kinase may sensitize tumors to anaplastic lymphoma kinase inhibitors. Cancer Res. 2008, 68, 3389–3395. [Google Scholar] [CrossRef] [PubMed]
- Soda, M.; Choi, Y.L.; Enomoto, M.; Takada, S.; Yamashita, Y.; Ishikawa, S.; Fujiwara, S.; Watanabe, H.; Kurashina, K.; Hatanaka, H.; et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 2007, 448, 561–566. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, T.; Rodig, S.J.; Chirieac, L.R.; Jänne, P.A. The biology and treatment of EML4-ALK non-small cell lung cancer. Eur. J. Cancer 2010, 46, 1773–1780. [Google Scholar] [CrossRef] [PubMed]
- Woo, C.G.; Seo, S.; Kim, S.W.; Jang, S.J.; Park, K.S.; Song, J.Y.; Lee, B.; Richards, M.W.; Bayliss, R.; Lee, D.H.; et al. Differential protein stability and clinical responses of EML4-ALK fusion variants to various ALK inhibitors in advanced ALK-rearranged non-small cell lung cancer. Ann. Oncol. 2017, 28, 791–797. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, T.; Oya, Y.; Tanaka, K.; Shimizu, J.; Horio, Y.; Kuroda, H.; Sakao, Y.; Hida, T.; Yatabe, Y. Differential crizotinib response duration among ALK fusion variants in ALK-positive non-small-cell lung cancer. J. Clin. Oncol. 2016, 34, 3383–3389. [Google Scholar] [CrossRef] [PubMed]
- Rikova, K.; Guo, A.; Zeng, Q.; Possemato, A.; Yu, J.; Haack, H.; Nardone, J.; Lee, K.; Reeves, C.; Li, Y.; et al. Global survey of phosphotyrosine signaling identifies oncogenic kinases in lung cancer. Cell 2007, 131, 1190–1203. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, K.; Choi, Y.L.; Togashi, Y.; Soda, M.; Hatano, S.; Inamura, K.; Takada, S.; Ueno, T.; Yamashita, Y.; Satoh, Y.; et al. KIF5B-ALK, a novel fusion oncokinase identified by an immunohistochemistry-based diagnostic system for ALK-positive lung cancer. Clin. Cancer Res. 2009, 15, 3143–3149. [Google Scholar] [CrossRef] [PubMed]
- Lovly, C.M.; Pao, W. Escaping ALK inhibition: Mechanisms of and strategies to overcome resistance. Sci. Transl. Med. 2012, 4, 120ps2. [Google Scholar] [CrossRef] [PubMed]
- Van der Wekken, A.J.; Saber, A.; Hiltermann, T.J.; Kok, K.; van den Berg, A.; Groen, H.J. Resistance mechanisms after tyrosine kinase inhibitors afatinib and crizotinib in non-small cell lung cancer, a review of the literature. Crit. Rev. Oncol. Hematol. 2016, 100, 107–116. [Google Scholar] [CrossRef] [PubMed]
- Ha, S.Y.; Choi, S.J.; Cho, J.H.; Choi, H.J.; Lee, J.; Jung, K.; Irwin, D.; Liu, X.; Lira, M.E.; Mao, M.; et al. Lung cancer in never-smoker Asian females is driven by oncogenic mutations, most often involving EGFR. Oncotarget 2015, 6, 5465–5474. [Google Scholar] [PubMed]
- Hong, S.; Fang, W.; Hu, Z.; Zhou, T.; Yan, Y.; Qin, T.; Tang, Y.; Ma, Y.; Zhao, Y.; Xue, C.; et al. A large-scale cross-sectional study of ALK rearrangements and EGFR mutations in non-small-cell lung cancer in Chinese Han population. Sci. Rep. 2014, 4, 7268. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.; Lee, T.; Lee, S.H.; Choi, Y.L.; Han, J. Clinicopathologic characteristics of EGFR, KRAS, and ALK alterations in 6,595 lung cancers. Oncotarget 2016, 7, 23874–23884. [Google Scholar] [CrossRef] [PubMed]
- Vidal, J.; Clavé, S.; de Muga, S.; González, I.; Pijuan, L.; Gimeno, J.; Remón, J.; Reguart, N.; Viñolas, N.; Gironés, R.; et al. Assessment of ALK status by FISH on 1000 Spanish non-small cell lung cancer patients. J. Thorac. Oncol. 2014, 9, 1816–1820. [Google Scholar] [CrossRef] [PubMed]
- Rodig, S.J.; Mino-Kenudson, M.; Dacic, S.; Yeap, B.Y.; Shaw, A.; Barletta, J.A.; Stubbs, H.; Law, K.; Lindeman, N.; Mark, E.; et al. Unique clinicopathologic features characterize ALK-rearranged lung adenocarcinoma in the western population. Clin. Cancer Res. 2009, 15, 5216–5223. [Google Scholar] [CrossRef] [PubMed]
- Sacher, A.G.; Dahlberg, S.E.; Heng, J.; Mach, S.; Jänne, P.A.; Oxnard, G.R. Association between younger age and targetable genomic alterations and prognosis in non-small-cell lung cancer. JAMA Oncol. 2016, 2, 313–320. [Google Scholar] [CrossRef] [PubMed]
- Shaw, A.T.; Yeap, B.Y.; Mino-Kenudson, M.; Digumarthy, S.R.; Costa, D.B.; Heist, R.S.; Solomon, B.; Stubbs, H.; Admane, S.; McDermott, U.; et al. Clinical features and outcome of patients with non-small-cell lung cancer who harbor EML4-ALK. J. Clin. Oncol. 2009, 27, 4247–4253. [Google Scholar] [CrossRef] [PubMed]
- Yang, P.; Kulig, K.; Boland, J.M.; Erickson-Johnson, M.R.; Oliveira, A.M.; Wampfler, J.; Jatoi, A.; Deschamps, C.; Marks, R.; Fortner, C.; et al. Worse disease-free survival in never-smokers with ALK+ lung adenocarcinoma. J. Thorac. Oncol. 2012, 7, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Alrifai, D.; Popat, S.; Ahmed, M.; Gonzalez, D.; Nicholson, A.G.; Parcq, Jd.; Benepal, T. A rare case of squamous cell carcinoma of the lung harbouring ALK and BRAF activating mutations. Lung Cancer 2013, 80, 339–340. [Google Scholar] [CrossRef] [PubMed]
- Bolzacchini, E.; Tuzi, A.; Pinotti, G. ALK-rearranged squamous cell carcinoma of the lung treated with two lines of ALK inhibitors. J. Thorac. Oncol. 2017, 12, e55–e57. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.; Choi, Y.L.; Song, J.Y.; Zhu, Y.; Xu, Q.; Zhang, F.; Jiang, L.; Cheng, J.; Zheng, G.; Mao, M. ALK, ROS1 and RET rearrangements in lung squamous cell carcinoma are very rare. Lung Cancer 2016, 94, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Popat, S.; Gonzalez, D.; Min, T.; Swansbury, J.; Dainton, M.; Croud, J.G.; Rice, A.J.; Nicholson, A.G. ALK translocation is associated with ALK immunoreactivity and extensive signet-ring morphology in primary lung adenocarcinoma. Lung Cancer 2012, 75, 300–305. [Google Scholar] [CrossRef] [PubMed]
- Ochi, N.; Yamane, H.; Yamagishi, T.; Takigawa, N.; Monobe, Y. Can we eliminate squamous cell carcinoma of the lung from testing of EML4-ALK fusion gene? Lung Cancer 2013, 79, 94–95. [Google Scholar] [CrossRef] [PubMed]
- Gainor, J.F.; Varghese, A.M.; Ou, S.H.; Kabraji, S.; Awad, M.M.; Katayama, R.; Pawlak, A.; Mino-Kenudson, M.; Yeap, B.Y.; Riely, G.J.; et al. ALK rearrangements are mutually exclusive with mutations in EGFR or KRAS: An analysis of 1683 patients with non-small cell lung cancer. Clin. Cancer Res. 2013, 19, 4273–4281. [Google Scholar] [CrossRef] [PubMed]
- Martelli, M.P.; Sozzi, G.; Hernandez, L.; Pettirossi, V.; Navarro, A.; Conte, D.; Gasparini, P.; Perrone, F.; Modena, P.; Pastorino, U.; et al. EML4-ALK rearrangement in non-small cell lung cancer and non-tumor lung tissues. Am. J. Pathol. 2009, 174, 661–670. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.M.; Hensing, T.; Schrock, A.B.; Allen, J.; Sanford, E.; Gowen, K.; Kulkarni, A.; He, J.; Suh, J.H.; Lipson, D.; et al. Comprehensive genomic profiling identifies a subset of crizotinib-responsive ALK-rearranged non-small cell lung cancer not detected by fluorescence in situ hybridization. Oncologist 2016, 21, 762–770. [Google Scholar] [CrossRef] [PubMed]
- Abel, H.J.; Al-Kateb, H.; Cottrell, C.E.; Bredemeyer, A.J.; Pritchard, C.C.; Grossmann, A.H.; Wallander, M.L.; Pfeifer, J.D.; Lockwood, C.M.; Duncavage, E.J. Detection of gene rearrangements in targeted clinical next-generation sequencing. J. Mol. Diagn. 2014, 16, 405–417. [Google Scholar] [CrossRef] [PubMed]
- Ilie, M.I.; Bence, C.; Hofman, V.; Long-Mira, E.; Butori, C.; Bouhlel, L.; Lalvée, S.; Mouroux, J.; Poudenx, M.; Otto, J.; et al. Discrepancies between FISH and immunohistochemistry for assessment of the ALK status are associated with ALK ‘borderline’-positive rearrangements or a high copy number: A potential major issue for anti-ALK therapeutic strategies. Ann. Oncol. 2015, 26, 238–244. [Google Scholar] [CrossRef] [PubMed]
- Abe, H.; Kawahara, A.; Azuma, K.; Taira, T.; Takase, Y.; Fukumitsu, C.; Murata, K.; Yamaguchi, T.; Akiba, J.; Ishii, H.; et al. Heterogeneity of anaplastic lymphoma kinase gene rearrangement in non-small-cell lung carcinomas: A comparative study between small biopsy and excision samples. J. Thorac. Oncol. 2015, 10, 800–805. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.W.; Roh, J.; Park, C.S. Immunohistochemistry for Pathologists: Protocols, Pitfalls, and Tips. J. Pathol. Transl. Med. 2016, 50, 411–418. [Google Scholar] [CrossRef] [PubMed]
- Conklin, C.M.; Craddock, K.J.; Have, C.; Laskin, J.; Couture, C.; Ionescu, D.N. Immunohistochemistry is a reliable screening tool for identification of ALK rearrangement in non-small-cell lung carcinoma and is antibody dependent. J. Thorac. Oncol. 2013, 8, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Hofman, P.; Ilie, M.; Hofman, V.; Roux, S.; Valent, A.; Bernheim, A.; Alifano, M.; Leroy-Ladurie, F.; Vaylet, F.; Rouquette, I.; et al. Immunohistochemistry to identify EGFR mutations or ALK rearrangements in patients with lung adenocarcinoma. Ann. Oncol. 2012, 23, 1738–1743. [Google Scholar] [CrossRef] [PubMed]
- Hutarew, G.; Hauser-Kronberger, C.; Strasser, F.; Llenos, I.C.; Dietze, O. Immunohistochemistry as a screening tool for ALK rearrangement in NSCLC: Evaluation of five different ALK antibody clones and ALK FISH. Histopathology 2014, 65, 398–407. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, M.; Parry, S.; Wilkinson, D.; Bilbe, N.; Allen, D.; Forrest, S.; Maxwell, P.; O’Grady, A.; Starczynski, J.; Tanier, P.; et al. ALK immunohistochemistry in NSCLC: Discordant staining can impact patient treatment regimen. J. Thorac. Oncol. 2016, 11, 2241–2247. [Google Scholar] [CrossRef] [PubMed]
- Gruber, K.; Kohlhäufl, M.; Friedel, G.; Ott, G.; Kalla, C. A novel, highly sensitive ALK antibody 1A4 facilitates effective screening for ALK rearrangements in lung adenocarcinomas by standard immunohistochemistry. J. Thorac. Oncol. 2015, 10, 713–716. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhao, L.; Yang, X.; Wei, S.; Zeng, Y.; Mao, C.; Lin, L.; Fu, P.; Lyu, L.; Li, Z.; et al. Antibody 1A4 with routine immunohistochemistry demonstrates high sensitivity for ALK rearrangement screening of Chinese lung adenocarcinoma patients: A single-center large-scale study. Lung Cancer 2016, 95, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Betz, B.L.; Dixon, C.A.; Weigelin, H.C.; Knoepp, S.M.; Roh, M.H. The use of stained cytologic direct smears for ALK gene rearrangement analysis of lung adenocarcinoma. Cancer Cytopathol. 2013, 121, 489–499. [Google Scholar] [CrossRef] [PubMed]
- Proietti, A.; Alì, G.; Pelliccioni, S.; Lupi, C.; Sensi, E.; Boldrini, L.; Servadio, A.; Chella, A.; Ribechini, A.; Cappuzzo, F.; et al. Anaplastic lymphoma kinase gene rearrangements in cytological samples of non-small cell lung cancer: Comparison with histological assessment. Cancer Cytopathol. 2014, 122, 445–453. [Google Scholar] [CrossRef] [PubMed]
- Zhong, J.; Li, X.; Bai, H.; Zhao, J.; Wang, Z.; Duan, J.; An, T.; Wu, M.; Wang, Y.; Wang, S.; et al. Malignant pleural effusion cell blocks are substitutes for tissue in EML4-ALK rearrangement detection in patients with advanced non-small-cell lung cancer. Cytopathology 2016, 27, 433–443. [Google Scholar] [CrossRef] [PubMed]
- Zito Marino, F.; Rossi, G.; Brunelli, M.; Malzone, M.G.; Liguori, G.; Bogina, G.; Morabito, A.; Rocco, G.; Franco, R.; Botti, G. Diagnosis of anaplastic lymphoma kinase rearrangement in cytological samples through a fluorescence in situ hybridization-based assay: Cytological smears versus cell blocks. Cancer 2017, 125, 303–312. [Google Scholar] [CrossRef] [PubMed]
- Long-Mira, E.; Washetine, K.; Hofman, P. Sense and nonsense in the process of accreditation of a pathology laboratory. Virchows Arch. 2016, 468, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Minca, E.C.; Portier, B.P.; Wang, Z.; Lanigan, C.; Farver, C.F.; Feng, Y.; Ma, P.C.; Arrossi, V.A.; Pennell, N.A.; Tubbs, R.R. ALK status testing in non-small cell lung carcinoma: Correlation between ultrasensitive IHC and FISH. J. Mol. Diagn. 2013, 15, 341–346. [Google Scholar] [CrossRef] [PubMed]
- Minca, E.C.; Lanigan, C.P.; Reynolds, J.P.; Wang, Z.; Ma, P.C.; Cicenia, J.; Almeida, F.A.; Pennell, N.A.; Tubbs, R.R. ALK status testing in non-small-cell lung carcinoma by FISH on thin prep slides with cytology material. J. Thorac. Oncol. 2014, 9, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Martin, V.; Bernasconi, B.; Merlo, E.; Balzarini, P.; Vermi, W.; Riva, A.; Chiaravalli, A.M.; Frattini, M.; Sahnane, N.; Facchetti, F.; et al. ALK testing in lung adenocarcinoma: Technical aspects to improve FISH evaluation in daily practice. J. Thorac. Oncol. 2015, 10, 595–602. [Google Scholar] [CrossRef] [PubMed]
- Von Laffert, M.; Stenzinger, A.; Hummel, M.; Weichert, W.; Lenze, D.; Warth, A.; Penzel, R.; Herbst, H.; Kellner, U.; Jurmeister, P.; et al. ALK-FISH borderline cases in non-small cell lung cancer: Implications for diagnostics and clinical decision making. Lung Cancer 2015, 90, 465–471. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Sholl, L.M.; Nishino, M.; Heng, J.C.; Jänne, P.A.; Oxnard, G.R. Clinical implications of variant ALK FISH rearrangement patterns. J. Thorac. Oncol. 2015, 10, 1648–1652. [Google Scholar] [CrossRef] [PubMed]
- Selinger, C.; Cooper, W.; Lum, T.; McNeil, C.; Morey, A.; Waring, P.; Amanuel, B.; Millward, M.; Peverall, J.; van Vliet, C.; et al. Equivocal ALK fluorescence in situ hybridization (FISH) cases may benefit from ancillary ALK FISH probe testing. Histopathology 2015, 67, 654–663. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Maus, M.K.; Desai, S.J.; Beckett, L.A.; Stephens, C.; Huang, E.; Hsiang, J.; Zeger, G.; Danenberg, K.D.; Astrow, S.H.; et al. Large-scale screening and molecular characterization of EML4-ALK fusion variants in archival non-small-cell lung cancer tumor specimens using quantitative reverse transcription polymerase chain reaction assays. J. Thorac. Oncol. 2014, 9, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhou, J.G.; Wu, H.L.; Ma, H.; Jiang, Z.X. Diagnostic accuracy of PCR for detecting ALK gene rearrangement in NSCLC patients: A systematic review and meta-analysis. Oncotarget 2017. [Google Scholar] [CrossRef] [PubMed]
- Karachaliou, N.; Rosell, R. Optimal detection of ALK rearranged lung adenocarcinomas. J. Thorac. Oncol. 2013, 8, 255–256. [Google Scholar] [CrossRef] [PubMed]
- Inamura, K.; Takeuchi, K.; Togashi, Y.; Nomura, K.; Ninomiya, H.; Okui, M.; Satoh, Y.; Okumura, S.; Nakagawa, K.; Soda, M.; et al. EML4-ALK fusion is linked to histological characteristics in a subset of lung cancers. J. Thorac. Oncol. 2008, 3, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Hiley, C.T.; Le Quesne, J.; Santis, G.; Sharpe, R.; de Castro, D.G.; Middleton, G.; Swanton, C. Challenges in molecular testing in non-small-cell lung cancer patients with advanced disease. Lancet 2016, 388, 1002–1011. [Google Scholar] [CrossRef]
- Rozenblum, A.B.; Ilouze, M.; Dudnik, E.; Dvir, A.; Soussan-Gutman, L.; Geva, S.; Peled, N. Clinical impact of hybrid capture-based next-generation sequencing on changes in treatment decisions in lung cancer. J. Thorac. Oncol. 2017, 12, 258–268. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Luthra, R.; Goswami, R.S.; Singh, R.R.; Roy-Chowdhuri, S. Analysis of pre-analytic factors affecting the success of clinical next-generation sequencing of solid organ malignancies. Cancers 2015, 7, 1699–1715. [Google Scholar] [CrossRef] [PubMed]
- Devarakonda, S.; Masood, A.; Govindan, R. Next-generation sequencing of lung cancers: Lessons learned and future directions. Hematol. Oncol. Clin. North. Am. 2017, 31, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Hagemann, I.S.; Devarakonda, S.; Lockwood, C.M.; Spencer, D.H.; Guebert, K.; Bredemeyer, A.J.; Al-Kateb, H.; Nguyen, T.T.; Duncavage, E.J.; Cottrell, C.E.; et al. Clinical next-generation sequencing in patients with non-small cell lung cancer. Cancer 2015, 121, 631–639. [Google Scholar] [CrossRef] [PubMed]
- Luthra, R.; Chen, H.; Roy-Chowdhuri, S.; Singh, R.R. Next-generation sequencing in clinical molecular diagnostics of cancer: Advantages and challenges. Cancers 2015, 7, 2023–2036. [Google Scholar] [CrossRef] [PubMed]
- Dagogo-Jack, I.; Shaw, A.T. Screening for ALK rearrangements in lung cancer: Time for a new generation of diagnostics? Oncologist 2016, 21, 662–663. [Google Scholar] [CrossRef] [PubMed]
- Goswami, R.S.; Luthra, R.; Singh, R.R.; Patel, K.P.; Routbort, M.J.; Aldape, K.D.; Yao, H.; Dang, H.D.; Barkoh, B.A.; Manekia, J.; et al. Identification of factors affecting the success of next-generation sequencing testing in solid tumors. Am. J.Clin. Pathol. 2016, 145, 222–237. [Google Scholar] [CrossRef] [PubMed]
- Dacic, S.; Villaruz, L.C.; Abberbock, S.; Mahaffey, A.; Incharoen, P.; Nikiforova, M.N. ALK FISH patterns and the detection of ALK fusions by next generation sequencing in lung adenocarcinoma. Oncotarget 2016, 7, 82943–82952. [Google Scholar] [CrossRef] [PubMed]
- Koitzsch, U.; Heydt, C.; Attig, H.; Immerschitt, I.; Merkelbach-Bruse, S.; Fammartino, A.; Büttner, R.H.; Kong, Y.; Odenthal, M. Use of the GeneReader NGS System in a clinical pathology laboratory: A comparative study. J. Clin. Pathol. 2017, 70, 725–728. [Google Scholar] [CrossRef] [PubMed]
- Lindeman, N.I.; Cagle, P.T.; Beasley, M.B.; Chitale, D.A.; Dacic, S.; Giaccone, G.; Jenkins, R.B.; Kwiatkowski, D.J.; Saldivar, J.S.; Squire, J.; et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: Guideline from the college of american pathologists, international association for the study of lung cancer, and association for molecular pathology. J. Thorac. Oncol. 2013, 8, 823–859. [Google Scholar] [PubMed]
- Evangelista, A.F.; Zanon, M.F.; Carloni, A.C.; de Paula, F.E.; Morini, M.A.; Ferreira-Neto, M.; Soares, I.C.; Miziara, J.E.; de Marchi, P.; Scapulatempo-Neto, C.; et al. Detection of ALK fusion transcripts in FFPE lung cancer samples by NanoString Technology. BMC Pulm. Med. 2017, 17, 86. [Google Scholar] [CrossRef] [PubMed]
- Geiss, G.K.; Bumgarner, R.E.; Birditt, B.; Dahl, T.; Dowidar, N.; Dunaway, D.L.; Fell, H.P.; Ferree, S.; George, R.D.; Grogan, T.; et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat. Biotechnol. 2008, 26, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Lira, M.E.; Choi, Y.L.; Lim, S.M.; Deng, S.; Huang, D.; Ozeck, M.; Han, J.; Jeong, J.Y.; Shim, H.S.; Cho, B.C.; et al. A single-tube multiplexed assay for detecting ALK, ROS1, and RET fusions in lung cancer. J. Mol. Diagn. 2014, 16, 229–243. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, N.; Yoshizawa, A.; Kondo, K.; Rokutan-Kurata, M.; Hirata, M.; Furuhata, A.; Sumiyoshi, S.; Sonobe, M.; Menju, T.; Momose, M.; et al. Evaluating the effectiveness of RNA in situ hybridization for detecting lung adenocarcinoma with anaplastic lymphoma kinase rearrangement. Histopathology 2017, 71, 143–149. [Google Scholar] [CrossRef] [PubMed]
- Schildhaus, H.U.; Deml, K.F.; Schmitz, K.; Meiboom, M.; Binot, E.; Hauke, S.; Merkelbach-Bruse, S.; Büttner, R. Chromogenic in situ hybridization is a reliable assay for detection of ALK rearrangements in adenocarcinomas of the lung. Mod. Pathol. 2013, 26, 1468–1477. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Flanagan, J.; Su, N.; Wang, L.C.; Bui, S.; Nielson, A.; Wu, X.; Vo, H.T.; Ma, X.J.; Luo, Y. RNAscope: A novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J. Mol. Diagn. 2012, 14, 22–29. [Google Scholar] [CrossRef] [PubMed]
- Roussel, H.; de Guillebon, E.; Biard, L.; Mandavit, M.; Gibault, L.; Fabre, E.; Antoine, M.; Hofman, P.; Beau-Faller, M.; Blons, H.; et al. Composite biomarkers defined by multiparametric immunofluorescence analysis identify ALK-positive adenocarcinoma as a potential target for immunotherapy. Oncoimmunology 2017, 6, e1286437. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Yoo, S.B.; Choe, J.Y.; Paik, J.H.; Xu, X.; Nitta, H.; Zhang, W.; Grogan, T.M.; Lee, C.T.; Jheon, S.; et al. Detection of ALK gene rearrangement in non-small cell lung cancer: A comparison of fluorescence in situ hybridization and chromogenic in situ hybridization with correlation of ALK protein expression. J. Thorac. Oncol. 2011, 6, 1359–1366. [Google Scholar] [CrossRef] [PubMed]
- Martinez, P.; Hernández-Losa, J.; Montero, M.Á.; Cedrés, S.; Castellví, J.; Martinez-Marti, A.; Tallada, N.; Murtra-Garrell, N.; Navarro-Mendivill, A.; Rodriguez-Freixinos, V.; et al. Fluorescence in situ hybridization and immunohistochemistry as diagnostic methods for ALK positive non-small cell lung cancer patients. PLoS ONE 2013, 8, e52261. [Google Scholar] [CrossRef] [PubMed]
- McLeer-Florin, A.; Moro-Sibilot, D.; Melis, A.; Salameire, D.; Lefebvre, C.; Ceccaldi, F.; de Fraipont, F.; Brambilla, E.; Lantuejoul, S. Dual IHC and FISH testing for ALK gene rearrangement in lung adenocarcinomas in a routine practice: A French study. J. Thorac. Oncol. 2012, 7, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Mino-Kenudson, M.; Chirieac, L.R.; Law, K.; Hornick, J.L.; Lindeman, N.; Mark, E.J.; Cohen, D.W.; Johnson, B.E.; Jänne, P.A.; Iafrate, A.J.; et al. A novel, highly sensitive antibody allows for the routine detection of ALK-rearranged lung adenocarcinomas by standard immunohistochemistry. Clin. Cancer Res. 2010, 16, 1561–1571. [Google Scholar] [CrossRef] [PubMed]
- Ying, J.; Guo, L.; Qiu, T.; Shan, L.; Ling, Y.; Liu, X.; Lu, N. Diagnostic value of a novel fully automated immunochemistry assay for detection of ALK rearrangement in primary lung adenocarcinoma. Ann. Oncol. 2013, 24, 2589–2593. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Yang, H.; He, P.; Liang, W.; Zhang, J.; Li, J.; Liu, Y.; He, J. Improving selection criteria for ALK inhibitor therapy in non-small cell lung cancer: A pooled-data analysis on diagnostic operating characteristics of immunohistochemistry. Am. J. Surg. Pathol. 2016, 40, 697–703. [Google Scholar] [CrossRef] [PubMed]
- Alì, G.; Proietti, A.; Pelliccioni, S.; Niccoli, C.; Lupi, C.; Sensi, E.; Giannini, R.; Borrelli, N.; Menghi, M.; Chella, A.; et al. ALK rearrangement in a large series of consecutive non-small cell lung cancers: Comparison between a new immunohistochemical approach and fluorescence in situ hybridization for the screening of patients eligible for crizotinib treatment. Arch. Pathol. Lab. Med. 2014, 138, 1449–1458. [Google Scholar] [CrossRef] [PubMed]
- Cabillic, F.; Gros, A.; Dugay, F.; Begueret, H.; Mesturoux, L.; Chiforeanu, D.C.; Dufrenot, L.; Jauffret, V.; Dachary, D.; Corre, R.; et al. Parallel FISH and immunohistochemical studies of ALK status in 3244 non-small-cell lung cancers reveal major discordances. J. Thorac. Oncol. 2014, 9, 295–306. [Google Scholar] [CrossRef] [PubMed]
- Heuckmann, J.M.; Pauwels, P.; Thunnissen, E. Comprehensive hybrid capture-based next-generation sequencing identifies a double ALK gene fusion in a patient previously identified to be false-negative by FISH. J. Thorac. Oncol. 2017, 12, e22–e24. [Google Scholar] [CrossRef] [PubMed]
- Lantuejoul, S.; Rouquette, I.; Blons, H.; Le Stang, N.; Ilie, M.; Begueret, H.; Grégoire, V.; Hofman, P.; Gros, A.; Garcia, S.; et al. French multicentric validation of ALK rearrangement diagnostic in 547 lung adenocarcinomas. Eur. Respir. J. 2015, 46, 207–218. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhang, J.; Guo, L.; Chuai, S.; Shan, L.; Ying, J. Combinational analysis of FISH and immunohistochemistry reveals rare genomic events in ALK fusion patterns in NSCLC that responds to crizotinib treatment. J. Thorac. Oncol. 2017, 12, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Xu, X.; Yoo, S.B.; Sun, P.L.; Jin, Y.; Paik, J.H.; Choe, G.; Jheon, S.; Lee, C.T.; Chung, J.H. Discordance between anaplastic lymphoma kinase status in primary non-small-cell lung cancers and their corresponding metastases. Histopathology 2013, 62, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Trejo Bittar, H.E.; Luvison, A.; Miller, C.; Dacic, S. A comparison of ALK gene rearrangement and ALK protein expression in primary lung carcinoma and matched metastasis. Histopathology 2017. [Google Scholar] [CrossRef] [PubMed]
- Wiesner, T.; Lee, W.; Obenauf, A.C.; Ran, L.; Murali, R.; Zhang, Q.F.; Wong, E.W.; Hu, W.; Scott, S.N.; Shah, R.H.; et al. Alternative transcription initiation leads to expression of a novel ALK isoform in cancer. Nature 2015, 526, 453–457. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, A.; Di Lorito, A.; Pace, M.V.; Iezzi, M.; Felicioni, L.; D’Antuono, T.; Filice, G.; Guetti, L.; Mucilli, F.; Buttitta, F. ALK protein analysis by IHC staining after recent regulatory changes: A comparison of two widely used approaches, revision of the literature, and a new testing algorithm. J. Thorac. Oncol. 2016, 11, 487–495. [Google Scholar] [CrossRef] [PubMed]
- Rosoux, A.; Pauwels, P.; Duplaquet, F.; D’Haene, N.; Weynand, B.; Delos, M.; Menon, R.; Heukamp, L.C.; Thunnissen, E.; Ocak, S. Effectiveness of crizotinib in a patient with ALK IHC-positive/FISH-negative metastatic lung adenocarcinoma. Lung Cancer 2016, 98, 118–121. [Google Scholar] [CrossRef] [PubMed]
- Ilie, M.; Hofman, P. Reply to the letter to the editor ’ALK FISH rearranged and amplified tumor with negative immunohistochemistry: A rare and challenging case concerning ALK status screening in lung cancer’ by Uguen et al. Ann. Oncol. 2015, 26, 1802. [Google Scholar] [CrossRef] [PubMed]
- Uguen, A.; Talagas, M.; Andrieu-Key, S.; Costa, S.; Quintin-Roué, I.; De Braekeleer, M.; Marcorelles, P. ALK FISH rearranged and amplified tumor with negative immunohistochemistry: A rare and challenging case concerning ALK status screening in lung cancer. Ann. Oncol. 2015, 26, 1514–1515. [Google Scholar] [CrossRef] [PubMed]
- Roth, A.; Streubel, A.; Grah, C.; Stephan-Falkenau, S.; Mairinger, T.; Wagner, F. A rare case of an EML4-ALK-rearranged lung adenocarcinoma missed by in situ-hybridization but detected by RT-PCR. J. Clin. Pathol. 2014, 67, 839–840. [Google Scholar] [CrossRef] [PubMed]
- Pekar-Zlotin, M.; Hirsch, F.R.; Soussan-Gutman, L.; Ilouze, M.; Dvir, A.; Boyle, T.; Wynes, M.; Miller, V.A.; Lipson, D.; Palmer, G.A.; et al. Fluorescence in situ hybridization, immunohistochemistry, and next-generation sequencing for detection of EML4-ALK rearrangement in lung cancer. Oncologist 2015, 20, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Peled, N.; Palmer, G.; Hirsch, F.R.; Wynes, M.W.; Ilouze, M.; Varella-Garcia, M.; Soussan-Gutman, L.; Otto, G.A.; Stephens, P.J.; Ross, J.S.; et al. Next-generation sequencing identifies and immunohistochemistry confirms a novel crizotinib-sensitive ALK rearrangement in a patient with metastatic non-small-cell lung cancer. J. Thorac. Oncol. 2012, 7, e14–e16. [Google Scholar] [CrossRef] [PubMed]
- Teixidó, C.; Karachaliou, N.; Peg, V.; Gimenez-Capitan, A.; Rosell, R. Concordance of IHC, FISH and RT-PCR for EML4-ALK rearrangements. Transl. Lung Cancer Res. 2014, 3, 70–74. [Google Scholar] [PubMed]
- Reguart, N.; Teixidó, C.; Giménez-Capitán, A.; Paré, L.; Galván, P.; Viteri, S.; Rodríguez, S.; Peg, V.; Aldeguer, E.; Viñolas, N.; et al. Identification of ALK, ROS1, and RET fusions by a multiplexed mRNA-based assay in formalin-fixed, paraffin-embedded samples from advanced non-small-cell lung cancer patients. Clin. Chem. 2017, 63, 751–760. [Google Scholar] [CrossRef] [PubMed]
- Rogers, T.M.; Arnau, G.M.; Ryland, G.L.; Huang, S.; Lira, M.E.; Emmanuel, Y.; Perez, O.D.; Irwin, D.; Fellowes, A.P.; Wong, S.Q.; et al. Multiplexed transcriptome analysis to detect ALK, ROS1 and RET rearrangements in lung cancer. Sci. Rep. 2017, 7, 42259. [Google Scholar] [CrossRef] [PubMed]
- Sunami, K.; Furuta, K.; Tsuta, K.; Sasada, S.; Izumo, T.; Nakaoku, T.; Shimada, Y.; Saito, M.; Nokihara, H.; Watanabe, S.; et al. Multiplex diagnosis of oncogenic fusion and MET exon skipping by molecular counting using formalin-fixed paraffin embedded lung adenocarcinoma tissues. J. Thorac. Oncol. 2016, 11, 203–212. [Google Scholar] [CrossRef] [PubMed]
- Wynes, M.W.; Sholl, L.M.; Dietel, M.; Schuuring, E.; Tsao, M.S.; Yatabe, Y.; Tubbs, R.R.; Hirsch, F.R. An international interpretation study using the ALK IHC antibody D5F3 and a sensitive detection KIT demonstrates high concordance between ALK IHC and ALK FISH and between evaluators. J. Thorac. Oncol. 2014, 9, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Shan, L.; Lian, F.; Guo, L.; Yang, X.; Ying, J.; Lin, D. Combination of conventional immunohistochemistry and qRT-PCR to detect ALK rearrangement. Diagn. Pathol. 2014, 14, 3. [Google Scholar] [CrossRef] [PubMed]
- Kerr, K.M.; López-Ríos, F. Precision medicine in NSCLC and pathology: How does ALK fit in the pathway? Ann. Oncol. 2016, 27, III16–III24. [Google Scholar] [CrossRef] [PubMed]
- Aisner, D.L.; Rumery, M.D.; Merrick, D.T.; Kondo, K.L.; Nijmeh, H.; Linderman, D.J.; Doebele, R.C.; Thomas, N.; Chesnut, P.C.; Varella-Garcia, M.; et al. Do more with less: Tips and techniques for maximizing small biopsy and cytology specimens for molecular and ancillary testing: The university of colorado experience. Arch. Pathol. Lab. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Yatabe, Y. ALK FISH and IHC: You cannot have one without the other. J. Thorac. Oncol. 2015, 10, 548–550. [Google Scholar] [CrossRef] [PubMed]
- Houang, M.; Toon, C.W.; Clarkson, A.; Sioson, L.; Watson, N.; Farzin, M.; Selinger, C.I.; Chou, A.; Morey, A.L.; Cooper, W.A.; et al. Reflex ALK immunohistochemistry is feasible and highly specific for ALK gene rearrangements in lung cancer. Pathology 2014, 46, 383–388. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, A.; Pace, M.V.; Di Lorito, A.; Canarecci, S.; Felicioni, L.; D’Antuono, T.; Liberatore, M.; Filice, G.; Guetti, L.; Mucilli, F.; et al. Validation of a new algorithm for a quick and easy RT-PCR-based ALK test in a large series of lung adenocarcinomas: Comparison with FISH, immunohistochemistry and next generation sequencing assays. Lung Cancer 2016, 99, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Thorne-Nuzzo, T.; Williams, C.; Catallini, A.; Clements, J.; Singh, S.; Amberson, J.; Dickinson, K.; Gatalica, Z.; Ho, S.N.; Loftin, I.; et al. A sensitive ALK immunohistochemistry companion diagnostic test identifies patients eligible for treatment with crizotinib. J. Thorac. Oncol. 2017, 12, 804–813. [Google Scholar] [CrossRef] [PubMed]
- Van der Wekken, A.J.; Pelgrim, R.; ’t Hart, N.; Werner, N.; Mastik, M.F.; Hendriks, L.; Van der Heijden, E.H.F.M.; Looijen-Salamon, M.; De Langen, A.J.; Staal-van den Brekel, J.; et al. Dichotomous ALK-IHC is a better predictor for ALK inhibition outcome than traditional ALK-FISH in advanced non-small cell lung cancer. Clin. Cancer Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Von Laffert, M.; Schirmacher, P.; Warth, A.; Weichert, W.; Büttner, R.; Huber, R.M.; Wolf, J.; Griesinger, F.; Dietel, M.; Grohé, C. ALK-Testing in non-small cell lung cancer (NSCLC): Immunohistochemistry (IHC) and/or fluorescence in situ Hybridisation (FISH)?: Statement of the Germany Society for Pathology (DGP) and the Working Group Thoracic Oncology (AIO) of the German Cancer Society e.V. (Stellungnahme der Deutschen Gesellschaft für Pathologie und der AG Thorakale Onkologie der Arbeitsgemeinschaft Onkologie/Deutsche Krebsgesellschaft e.V.). Lung Cancer 2017, 103, 1–5. [Google Scholar] [PubMed]
- Vinciguerra, G.L.R.; Scarpino, S.; Pini, B.; Cippitelli, C.; Fochetti, F.; Ruco, L. Optimized immunohistochemistry using the D5F3 antibody provides a reliable test for identification of ALK-positive lung adenocarcinomas. Virchows Arch. 2017, 471, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, J.T.; Hersom, M. Companion diagnostics-a tool to improve pharmacotherapy. Ann. Transl. Med. 2016, 4, 482. [Google Scholar] [CrossRef] [PubMed]
- Jürgens, J.; Engel-Riedel, W.; Stoelben, E.; Schildgen, V.; Schildgen, O.; Brockmann, M. The (con-) fusion in ALK diagnostics: When food and drug administration-approved algorithms fail. J. Clin. Oncol. 2016, 34, 1961–1962. [Google Scholar] [CrossRef] [PubMed]
- Thunnissen, E.; Allen, T.C.; Adam, J.; Aisner, D.L.; Beasley, M.B.; Borczuk, A.C.; Cagle, P.T.; Capelozzi, V.L.; Cooper, W.; Hariri, L.P.; et al. Immunohistochemistry of pulmonary biomarkers: A perspective from members of the Pulmonary Pathology Society. Arch. Pathol. Lab. Med. 2017, 140, 341–344. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhang, R.; Peng, R.; Ding, J.; Han, Y.; Wang, G.; Zhang, K.; Lin, G.; Li, J. Reliability assurance of detection of EML4-ALK rearrangement in non-small cell lung cancer: The results of proficiency testing in China. J. Thorac. Oncol. 2016, 11, 924–929. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, A.; Barberis, M.; Papotti, M.; Rossi, G.; Franco, R.; Malatesta, S.; Buttitta, F.; Ardizzoni, A.; Crinò, L.; Gridelli, C.; et al. ALK rearrangement testing by FISH analysis in non-small-cell lung cancer patients: Results of the first italian external quality assurance scheme. J. Thorac. Oncol. 2014, 9, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, S. External quality assessment for immunohistochemistry: Experiences from NordiQC. Biotech. Histochem. 2015, 90, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Tembuyser, L.; Tack, V.; Zwaenepoel, K.; Pauwels, P.; Miller, K.; Bubendorf, L.; Kerr, K.; Schuuring, E.; Thunnissen, E.; Dequeker, E.M. The relevance of external quality assessment for molecular testing for ALK positive non-small cell lung cancer: Results from two pilot rounds show room for optimization. PLoS ONE 2014, 9, e112159. [Google Scholar] [CrossRef] [PubMed]
| Parameters | FISH | IHC | qRT-PCR | NGS | NanoString |
|---|---|---|---|---|---|
| Sample criteria | |||||
| RNA input required | NA | NA | + | +++ | ++ |
| RNA quality | ++ | +++ | + | + | ++ |
| % of tumor cells | + | +++ | ++ | + | + |
| Sensitivity | +++ | +++ | ++ | ++ | ++ |
| Specificity | +++ | +++ | ++ | +++ | ++ |
| Costs | |||||
| Reagents | ++ | +++ | ++ | + | +++ |
| Device/hardware/software | ++ | +++ | ++ | + | + |
| Resolution | |||||
| Quantitative precision | + | + | ++ | +++ | +++ |
| Accuracy at low concentrations | NA | NA | ++ | + | +++ |
| Variants detection | NA | NA | ++ | +++ | ++ |
| TAT | |||||
| Hands on time | + | +++ | ++ | + | +++ |
| Results analysis | + | +++ | ++ | + | ++ |
| Interpretation | + | +++ | ++ | ++ | ++ |
| Throughput | + | ++ | ++ | +++ | +++ |
© 2017 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hofman, P. ALK in Non-Small Cell Lung Cancer (NSCLC) Pathobiology, Epidemiology, Detection from Tumor Tissue and Algorithm Diagnosis in a Daily Practice. Cancers 2017, 9, 107. https://doi.org/10.3390/cancers9080107
Hofman P. ALK in Non-Small Cell Lung Cancer (NSCLC) Pathobiology, Epidemiology, Detection from Tumor Tissue and Algorithm Diagnosis in a Daily Practice. Cancers. 2017; 9(8):107. https://doi.org/10.3390/cancers9080107
Chicago/Turabian StyleHofman, Paul. 2017. "ALK in Non-Small Cell Lung Cancer (NSCLC) Pathobiology, Epidemiology, Detection from Tumor Tissue and Algorithm Diagnosis in a Daily Practice" Cancers 9, no. 8: 107. https://doi.org/10.3390/cancers9080107
APA StyleHofman, P. (2017). ALK in Non-Small Cell Lung Cancer (NSCLC) Pathobiology, Epidemiology, Detection from Tumor Tissue and Algorithm Diagnosis in a Daily Practice. Cancers, 9(8), 107. https://doi.org/10.3390/cancers9080107






