The Evolution of Anticancer 3D In Vitro Models: The Potential Role of Machine Learning and AI in the Next Generation of Animal-Free Experiments
Simple Summary
Abstract
1. Introduction
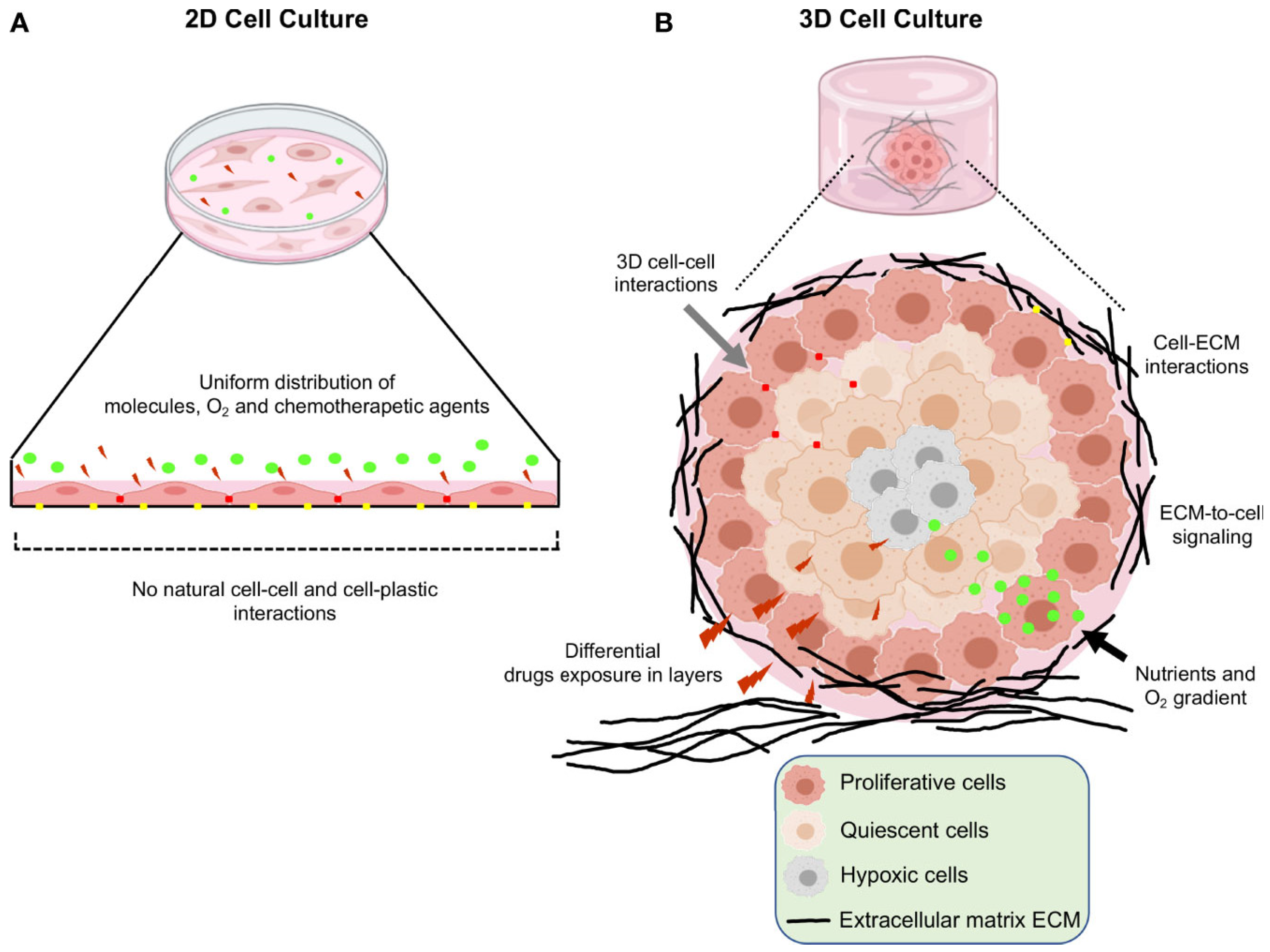
2. Three-Dimensional Models
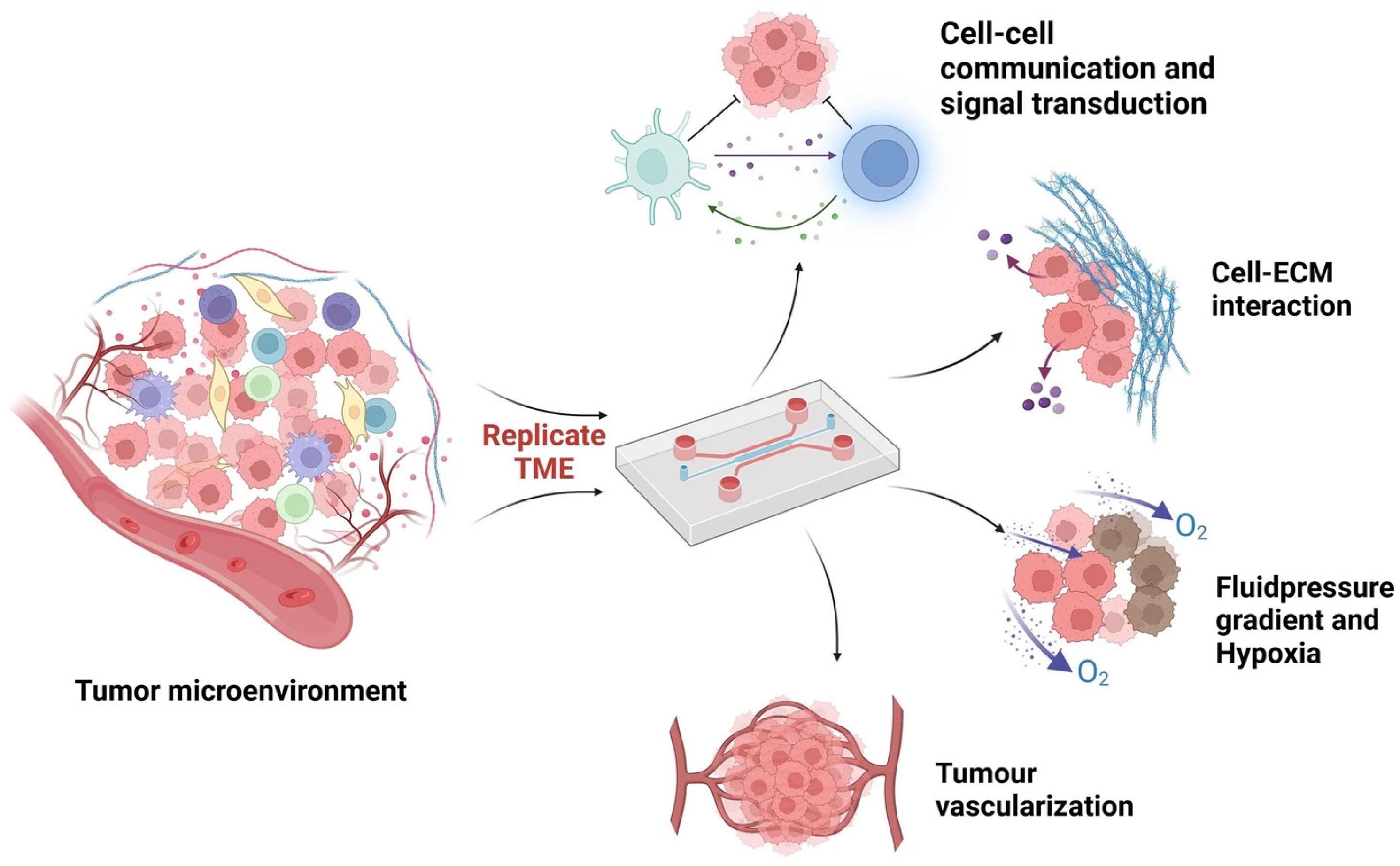
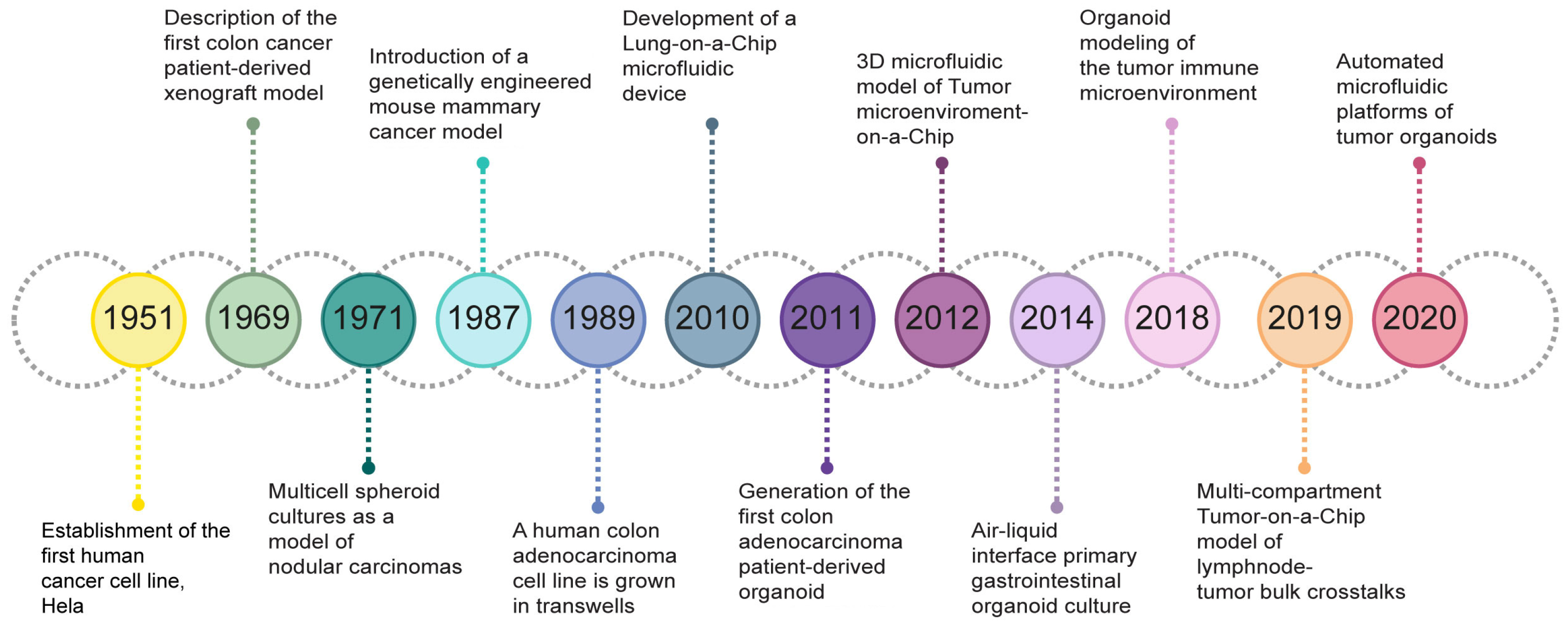
2.1. Development and Relevant TME Features
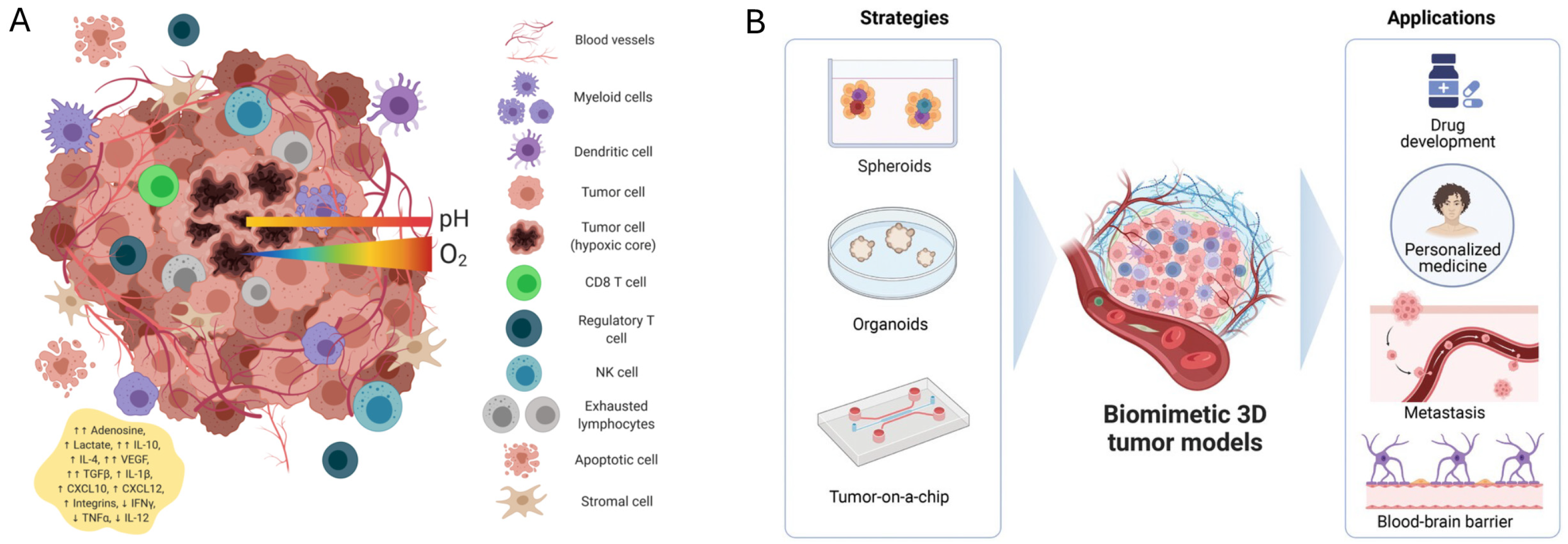
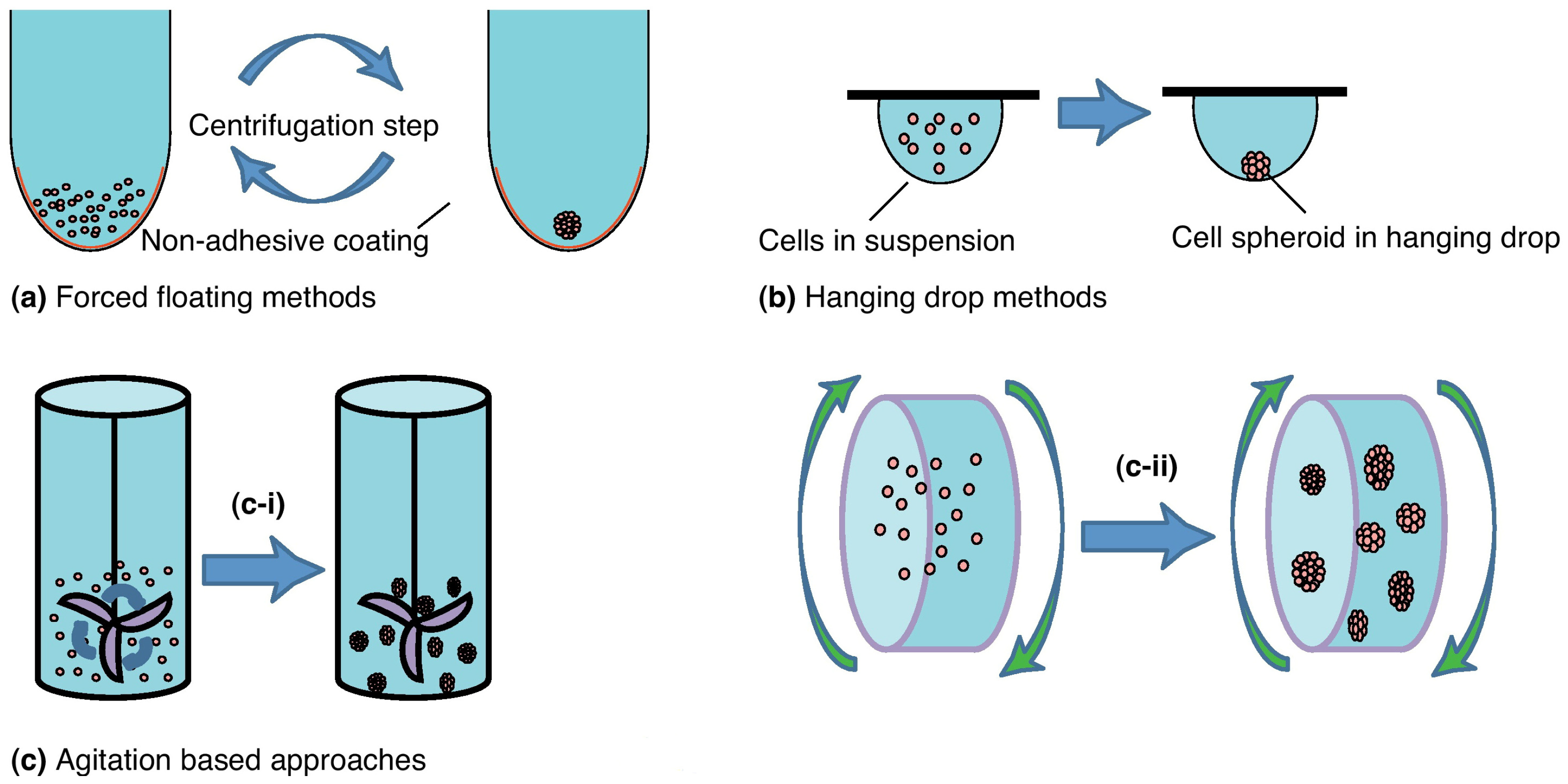
2.2. Spheroids
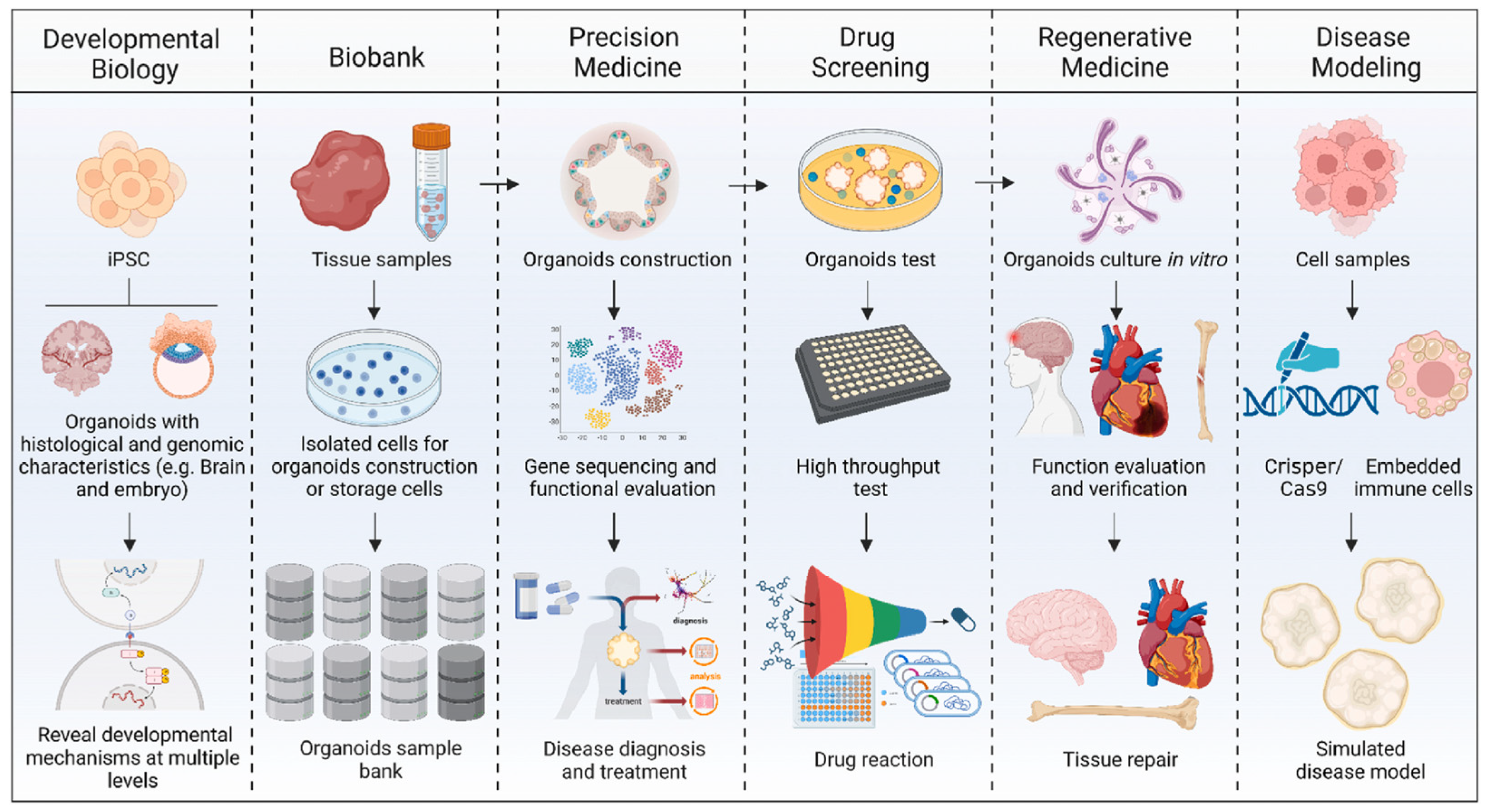
2.3. Organoids
2.4. Tumor-on-a-Chip
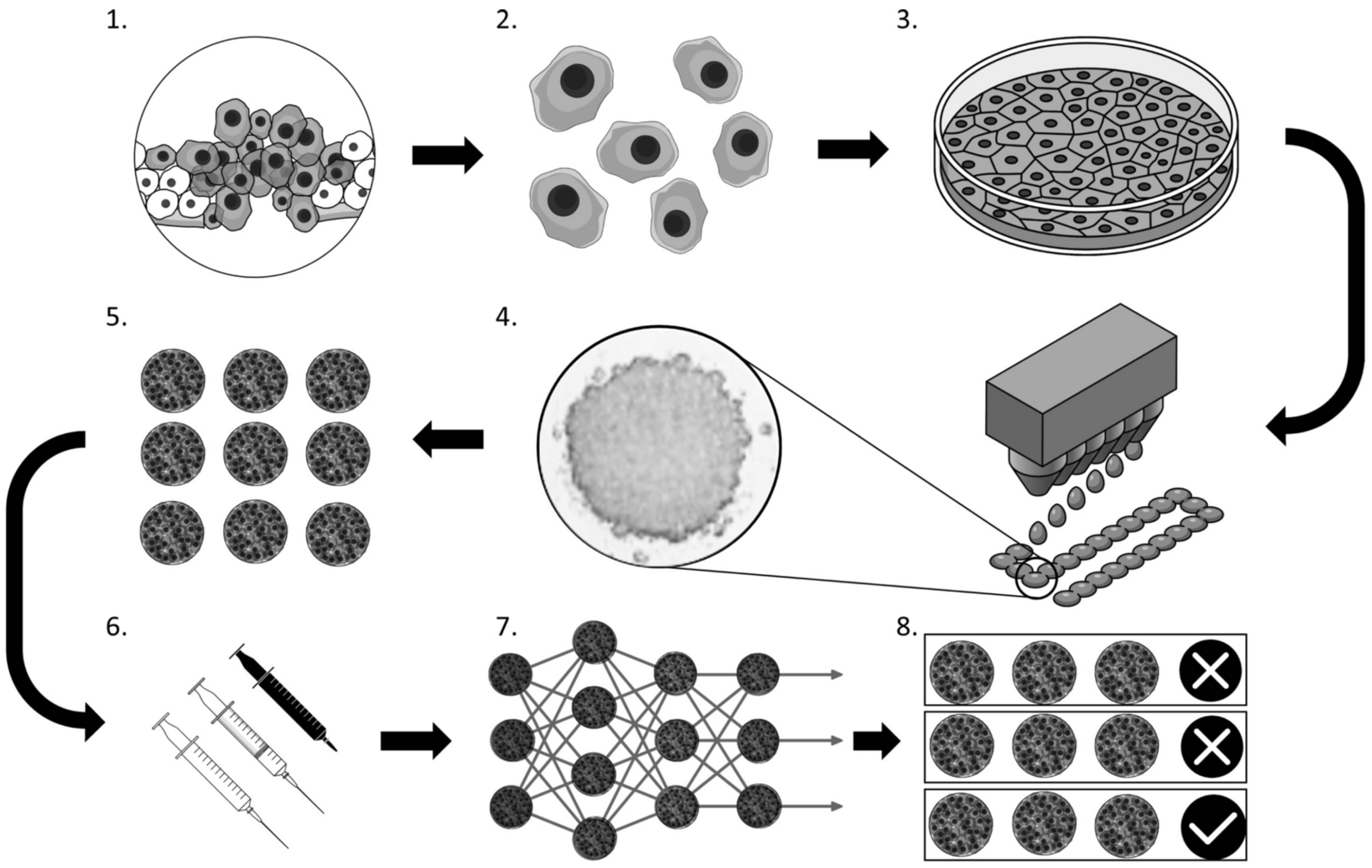
3. Applications of ML and 3D Models
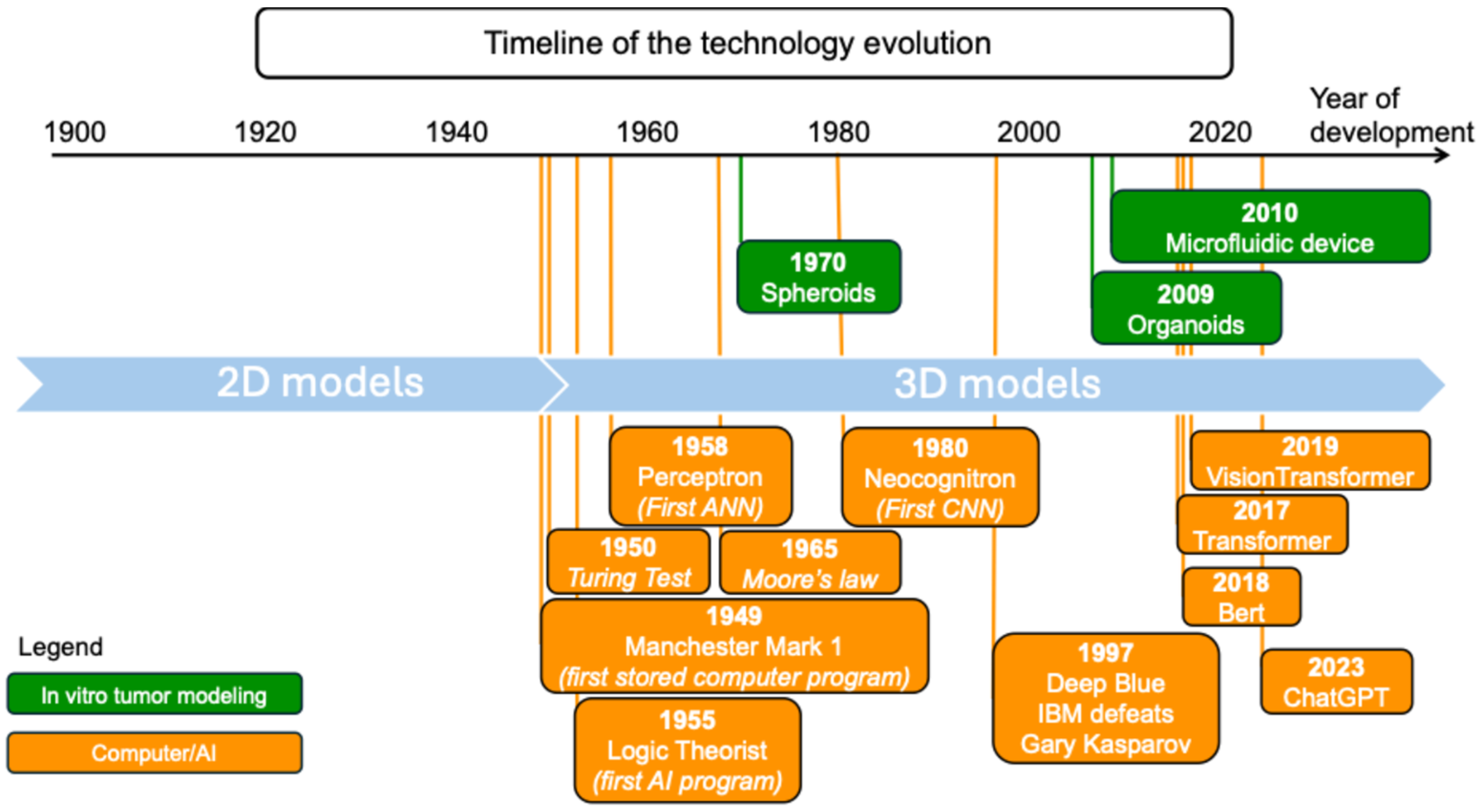
4. Conclusions and Perspectives
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Siegel, R.L.; Giaquinto, A.N.; Jemal, A. Cancer statistics, 2024. CA. Cancer J. Clin. 2024, 74, 12–49. [Google Scholar] [CrossRef]
- Aboulkheyr Es, H.; Montazeri, L.; Aref, A.R.; Vosough, M.; Baharvand, H. Personalized Cancer Medicine: An Organoid Approach. Trends Biotechnol. 2018, 36, 358–371. [Google Scholar] [CrossRef] [PubMed]
- Vitale, I.; Shema, E.; Loi, S.; Galluzzi, L. Intratumoral heterogeneity in cancer progression and response to immunotherapy. Nat. Med. 2021, 27, 212–224. [Google Scholar] [CrossRef]
- Aldea, M.; Andre, F.; Marabelle, A.; Dogan, S.; Barlesi, F.; Soria, J.-C. Overcoming Resistance to Tumor-Targeted and Immune-Targeted Therapies. Cancer Discov. 2021, 11, 874–899. [Google Scholar] [CrossRef] [PubMed]
- Manduca, N.; Maccafeo, E.; De Maria, R.; Sistigu, A.; Musella, M. 3D cancer models: One step closer to in vitro human studies. Front. Immunol. 2023, 14, 1175503. [Google Scholar] [CrossRef] [PubMed]
- Russell, W.M.S.; Burch, R.L.; Hume, C.W. The Principles of Humane Experimental Technique. Med. J. Aust. 1960, 1, 500. [Google Scholar] [CrossRef]
- Vitale, A.; Manciocco, A.; Alleva, E. The 3R principle and the use of non-human primates in the study of neurodegenerative diseases: The case of Parkinson’s disease. Neurosci. Biobehav. Rev. 2009, 33, 33–47. [Google Scholar] [CrossRef] [PubMed]
- Cacciamali, A.; Villa, R.; Dotti, S. 3D Cell Cultures: Evolution of an Ancient Tool for New Applications. Front. Physiol. 2022, 13, 836480. [Google Scholar] [CrossRef] [PubMed]
- Schachtschneider, K.M.; Schwind, R.M.; Newson, J.; Kinachtchouk, N.; Rizko, M.; Mendoza-Elias, N.; Grippo, P.; Principe, D.R.; Park, A.; Overgaard, N.H.; et al. The Oncopig Cancer Model: An Innovative Large Animal Translational Oncology Platform. Front. Oncol. 2017, 7, 190. [Google Scholar] [CrossRef] [PubMed]
- Van Norman, G.A. Limitations of Animal Studies for Predicting Toxicity in Clinical Trials. JACC Basic Transl. Sci. 2019, 4, 845–854. [Google Scholar] [CrossRef]
- Shanks, N.; Greek, R.; Greek, J. Are animal models predictive for humans? Philos. Ethics Humanit. Med. 2009, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Sachdeva, U.M.; Shimonosono, M.; Flashner, S.; Cruz-Acuña, R.; Gabre, J.T.; Nakagawa, H. Understanding the cellular origin and progression of esophageal cancer using esophageal organoids. Cancer Lett. 2021, 509, 39–52. [Google Scholar] [CrossRef] [PubMed]
- Weeber, F.; Van De Wetering, M.; Hoogstraat, M.; Dijkstra, K.K.; Krijgsman, O.; Kuilman, T.; Gadellaa-van Hooijdonk, C.G.M.; Van Der Velden, D.L.; Peeper, D.S.; Cuppen, E.P.J.G.; et al. Preserved genetic diversity in organoids cultured from biopsies of human colorectal cancer metastases. Proc. Natl. Acad. Sci. USA 2015, 112, 13308–13311. [Google Scholar] [CrossRef]
- Schütte, M.; Risch, T.; Abdavi-Azar, N.; Boehnke, K.; Schumacher, D.; Keil, M.; Yildiriman, R.; Jandrasits, C.; Borodina, T.; Amstislavskiy, V.; et al. Molecular dissection of colorectal cancer in pre-clinical models identifies biomarkers predicting sensitivity to EGFR inhibitors. Nat. Commun. 2017, 8, 14262. [Google Scholar] [CrossRef]
- Salinas-Vera, Y.M.; Valdés, J.; Pérez-Navarro, Y.; Mandujano-Lazaro, G.; Marchat, L.A.; Ramos-Payán, R.; Nuñez-Olvera, S.I.; Pérez-Plascencia, C.; López-Camarillo, C. Three-Dimensional 3D Culture Models in Gynecological and Breast Cancer Research. Front. Oncol. 2022, 12, 826113. [Google Scholar] [CrossRef]
- Chen, Z.; Ma, N.; Sun, X.; Li, Q.; Zeng, Y.; Chen, F.; Sun, S.; Xu, J.; Zhang, J.; Ye, H.; et al. Automated evaluation of tumor spheroid behavior in 3D culture using deep learning-based recognition. Biomaterials 2021, 272, 120770. [Google Scholar] [CrossRef] [PubMed]
- Iqbal, M.J.; Javed, Z.; Sadia, H.; Qureshi, I.A.; Irshad, A.; Ahmed, R.; Malik, K.; Raza, S.; Abbas, A.; Pezzani, R.; et al. Clinical applications of artificial intelligence and machine learning in cancer diagnosis: Looking into the future. Cancer Cell Int. 2021, 21, 270. [Google Scholar] [CrossRef] [PubMed]
- Jiménez Pérez, M.; Grande, R.G. Application of artificial intelligence in the diagnosis and treatment of hepatocellular carcinoma: A review. World J. Gastroenterol. 2020, 26, 5617–5628. [Google Scholar] [CrossRef]
- Hwangbo, H.; Chae, S.; Kim, W.; Jo, S.; Kim, G.H. Tumor-on-a-chip models combined with mini-tissues or organoids for engineering tumor tissues. Theranostics 2024, 14, 33–55. [Google Scholar] [CrossRef]
- Huh, D.; Hamilton, G.A.; Ingber, D.E. From 3D cell culture to organs-on-chips. Trends Cell Biol. 2011, 21, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Growth of Multicell Spheroids in Tissue Culture as a Model of Nodular Carcinomas2. JNCI J. Natl. Cancer Inst. 1971. [CrossRef]
- Huh, D.; Matthews, B.D.; Mammoto, A.; Montoya-Zavala, M.; Hsin, H.Y.; Ingber, D.E. Reconstituting organ-level lung functions on a chip. Science 2010, 328, 1662–1668. [Google Scholar] [CrossRef]
- Shi, R.; Tang, Y.; Miao, H. Metabolism in tumor microenvironment: Implications for cancer immunotherapy. MedComm 2020, 1, 47–68. [Google Scholar] [CrossRef] [PubMed]
- Anderson, N.M.; Simon, M.C. The tumor microenvironment. Curr. Biol. 2020, 30, R921–R925. [Google Scholar] [CrossRef] [PubMed]
- Ziółkowska-Suchanek, I. Mimicking Tumor Hypoxia in Non-Small Cell Lung Cancer Employing Three-Dimensional In Vitro Models. Cells 2021, 10, 141. [Google Scholar] [CrossRef]
- Piñeiro Fernández, J.; Luddy, K.A.; Harmon, C.; O’Farrelly, C. Hepatic Tumor Microenvironments and Effects on NK Cell Phenotype and Function. Int. J. Mol. Sci. 2019, 20, 4131. [Google Scholar] [CrossRef]
- Li, W.; Zhou, Z.; Zhou, X.; Khoo, B.L.; Gunawan, R.; Chin, Y.R.; Zhang, L.; Yi, C.; Guan, X.; Yang, M. 3D Biomimetic Models to Reconstitute Tumor Microenvironment In Vitro: Spheroids, Organoids, and Tumor-on-a-Chip. Adv. Healthc. Mater. 2023, 12, 2202609. [Google Scholar] [CrossRef]
- Hirschhaeuser, F.; Menne, H.; Dittfeld, C.; West, J.; Mueller-Klieser, W.; Kunz-Schughart, L.A. Multicellular tumor spheroids: An underestimated tool is catching up again. J. Biotechnol. 2010, 148, 3–15. [Google Scholar] [CrossRef]
- Hanahan, D.; Coussens, L.M. Accessories to the Crime: Functions of Cells Recruited to the Tumor Microenvironment. Cancer Cell 2012, 21, 309–322. [Google Scholar] [CrossRef] [PubMed]
- Costa, E.C.; Moreira, A.F.; de Melo-Diogo, D.; Gaspar, V.M.; Carvalho, M.P.; Correia, I.J. 3D tumor spheroids: An overview on the tools and techniques used for their analysis. Biotechnol. Adv. 2016, 34, 1427–1441. [Google Scholar] [CrossRef] [PubMed]
- Nath, S.; Devi, G.R. Three-dimensional culture systems in cancer research: Focus on tumor spheroid model. Pharmacol. Ther. 2016, 163, 94–108. [Google Scholar] [CrossRef]
- Mayer, B.; Klement, G.; Kaneko, M.; Man, S.; Jothy, S.; Rak, J.; Kerbel, R.S. Multicellular gastric cancer spheroids recapitulate growth pattern and differentiation phenotype of human gastric carcinomas. Gastroenterology 2001, 121, 839–852. [Google Scholar] [CrossRef] [PubMed]
- Breslin, S.; O’Driscoll, L. Three-dimensional cell culture: The missing link in drug discovery. Drug Discov. Today 2013, 18, 240–249. [Google Scholar] [CrossRef] [PubMed]
- Kelm, J.M.; Timmins, N.E.; Brown, C.J.; Fussenegger, M.; Nielsen, L.K. Method for generation of homogeneous multicellular tumor spheroids applicable to a wide variety of cell types. Biotechnol. Bioeng. 2003, 83, 173–180. [Google Scholar] [CrossRef]
- Roy, M.; Alix, C.; Bouakaz, A.; Serrière, S.; Escoffre, J.-M. Tumor Spheroids as Model to Design Acoustically Mediated Drug Therapies: A Review. Pharmaceutics 2023, 15, 806. [Google Scholar] [CrossRef]
- Benien, P.; Swami, A. 3D Tumor Models: History, Advances and Future Perspectives. Future Oncol. 2014, 10, 1311–1327. [Google Scholar] [CrossRef]
- Kim, J.B. Three-dimensional tissue culture models in cancer biology. Semin. Cancer Biol. 2005, 15, 365–377. [Google Scholar] [CrossRef] [PubMed]
- Brancato, V.; Comunanza, V.; Imparato, G.; Corà, D.; Urciuolo, F.; Noghero, A.; Bussolino, F.; Netti, P.A. Bioengineered tumoral microtissues recapitulate desmoplastic reaction of pancreatic cancer. Acta Biomater. 2017, 49, 152–166. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Verwilst, P.; Won, M.; Lee, J.; Sessler, J.L.; Han, J.; Kim, J.S. A Small Molecule Strategy for Targeting Cancer Stem Cells in Hypoxic Microenvironments and Preventing Tumorigenesis. J. Am. Chem. Soc. 2021, 143, 14115–14124. [Google Scholar] [CrossRef] [PubMed]
- Ishiguro, T.; Ohata, H.; Sato, A.; Yamawaki, K.; Enomoto, T.; Okamoto, K. Tumor-derived spheroids: Relevance to cancer stem cells and clinical applications. Cancer Sci. 2017, 108, 283–289. [Google Scholar] [CrossRef]
- Lapidot, T.; Sirard, C.; Vormoor, J.; Murdoch, B.; Hoang, T.; Caceres-Cortes, J.; Minden, M.; Paterson, B.; Caligiuri, M.A.; Dick, J.E. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature 1994, 367, 645–648. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.K.; Clarke, I.D.; Terasaki, M.; Bonn, V.E.; Hawkins, C.; Squire, J.; Dirks, P.B. Identification of a cancer stem cell in human brain tumors. Cancer Res. 2003, 63, 5821–5828. [Google Scholar]
- Dontu, G.; Abdallah, W.M.; Foley, J.M.; Jackson, K.W.; Clarke, M.F.; Kawamura, M.J.; Wicha, M.S. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003, 17, 1253–1270. [Google Scholar] [CrossRef] [PubMed]
- Ricci-Vitiani, L.; Lombardi, D.G.; Pilozzi, E.; Biffoni, M.; Todaro, M.; Peschle, C.; De Maria, R. Identification and expansion of human colon-cancer-initiating cells. Nature 2007, 445, 111–115. [Google Scholar] [CrossRef] [PubMed]
- Masuda, M.; Uno, Y.; Ohbayashi, N.; Ohata, H.; Mimata, A.; Kukimoto-Niino, M.; Moriyama, H.; Kashimoto, S.; Inoue, T.; Goto, N.; et al. TNIK inhibition abrogates colorectal cancer stemness. Nat. Commun. 2016, 7, 12586. [Google Scholar] [CrossRef]
- Tuveson, D.; Clevers, H. Cancer modeling meets human organoid technology. Science 2019, 364, 952–955. [Google Scholar] [CrossRef] [PubMed]
- Corrò, C.; Novellasdemunt, L.; Li, V.S.W. A brief history of organoids. Am. J. Physiol. Cell Physiol. 2020, 319, C151–C165. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Li, X.; Yu, S. Cancer organoid co-culture model system: Novel approach to guide precision medicine. Front. Immunol. 2023, 13, 1061388. [Google Scholar] [CrossRef]
- Bai, L.; Wu, Y.; Li, G.; Zhang, W.; Zhang, H.; Su, J. AI-enabled organoids: Construction, analysis, and application. Bioact. Mater. 2024, 31, 525–548. [Google Scholar] [CrossRef]
- Mun, S.J.; Ryu, J.-S.; Lee, M.-O.; Son, Y.S.; Oh, S.J.; Cho, H.-S.; Son, M.-Y.; Kim, D.-S.; Kim, S.J.; Yoo, H.J.; et al. Generation of expandable human pluripotent stem cell-derived hepatocyte-like liver organoids. J. Hepatol. 2019, 71, 970–985. [Google Scholar] [CrossRef] [PubMed]
- Arjmand, B.; Rabbani, Z.; Soveyzi, F.; Tayanloo-Beik, A.; Rezaei-Tavirani, M.; Biglar, M.; Adibi, H.; Larijani, B. Advancement of Organoid Technology in Regenerative Medicine. Regen. Eng. Transl. Med. 2023, 9, 83–96. [Google Scholar] [CrossRef] [PubMed]
- Sun, N.; Meng, X.; Liu, Y.; Song, D.; Jiang, C.; Cai, J. Applications of brain organoids in neurodevelopment and neurological diseases. J. Biomed. Sci. 2021, 28, 30. [Google Scholar] [CrossRef]
- Sato, T.; Vries, R.G.; Snippert, H.J.; van de Wetering, M.; Barker, N.; Stange, D.E.; van Es, J.H.; Abo, A.; Kujala, P.; Peters, P.J.; et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 2009, 459, 262–265. [Google Scholar] [CrossRef]
- Sachs, N.; de Ligt, J.; Kopper, O.; Gogola, E.; Bounova, G.; Weeber, F.; Balgobind, A.V.; Wind, K.; Gracanin, A.; Begthel, H.; et al. A Living Biobank of Breast Cancer Organoids Captures Disease Heterogeneity. Cell 2018, 172, 373–386.e10. [Google Scholar] [CrossRef] [PubMed]
- van de Wetering, M.; Francies, H.E.; Francis, J.M.; Bounova, G.; Iorio, F.; Pronk, A.; van Houdt, W.; van Gorp, J.; Taylor-Weiner, A.; Kester, L.; et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 2015, 161, 933–945. [Google Scholar] [CrossRef]
- Obreque, J.; Vergara-Gómez, L.; Venegas, N.; Weber, H.; Owen, G.I.; Pérez-Moreno, P.; Leal, P.; Roa, J.C.; Bizama, C. Advances towards the use of gastrointestinal tumor patient-derived organoids as a therapeutic decision-making tool. Biol. Res. 2023, 56, 63. [Google Scholar] [CrossRef] [PubMed]
- Dutta, D.; Clevers, H. Organoid culture systems to study host-pathogen interactions. Curr. Opin. Immunol. 2017, 48, 15–22. [Google Scholar] [CrossRef]
- Alipour, M. Molecular Mechanism of Helicobacter pylori-Induced Gastric Cancer. J. Gastrointest. Cancer 2021, 52, 23–30. [Google Scholar] [CrossRef] [PubMed]
- McCracken, K.W.; Catá, E.M.; Crawford, C.M.; Sinagoga, K.L.; Schumacher, M.; Rockich, B.E.; Tsai, Y.-H.; Mayhew, C.N.; Spence, J.R.; Zavros, Y.; et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature 2014, 516, 400–404. [Google Scholar] [CrossRef]
- Bartfeld, S.; Bayram, T.; van de Wetering, M.; Huch, M.; Begthel, H.; Kujala, P.; Vries, R.; Peters, P.J.; Clevers, H. In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology 2015, 148, 126–136.e6. [Google Scholar] [CrossRef]
- Holokai, L.; Chakrabarti, J.; Broda, T.; Chang, J.; Hawkins, J.A.; Sundaram, N.; Wroblewski, L.E.; Peek, R.M.; Wang, J.; Helmrath, M.; et al. Increased Programmed Death-Ligand 1 is an Early Epithelial Cell Response to Helicobacter pylori Infection. PLoS Pathog. 2019, 15, e1007468. [Google Scholar] [CrossRef]
- Iyer, N.V.; Kotch, L.E.; Agani, F.; Leung, S.W.; Laughner, E.; Wenger, R.H.; Gassmann, M.; Gearhart, J.D.; Lawler, A.M.; Yu, A.Y.; et al. Cellular and developmental control of O 2 homeostasis by hypoxia-inducible factor 1α. Genes Dev. 1998, 12, 149–162. [Google Scholar] [CrossRef] [PubMed]
- Blondel, D.; Lutolf, M.P. Bioinspired Hydrogels for 3D Organoid Culture. Chimia 2019, 73, 81. [Google Scholar] [CrossRef] [PubMed]
- Revete, A.; Aparicio, A.; Cisterna, B.A.; Revete, J.; Luis, L.; Ibarra, E.; Segura González, E.A.; Molino, J.; Reginensi, D. Advancements in the Use of Hydrogels for Regenerative Medicine: Properties and Biomedical Applications. Int. J. Biomater. 2022, 2022, 1–16. [Google Scholar] [CrossRef]
- Peng, W.; Datta, P.; Wu, Y.; Dey, M.; Ayan, B.; Dababneh, A.; Ozbolat, I.T. Challenges in Bio-fabrication of Organoid Cultures. In Cell Biology and Translational Medicine, Volume 3; Turksen, K., Ed.; Advances in Experimental Medicine and Biology; Springer International Publishing: Cham, Switzerland, 2018; Volume 1107, pp. 53–71. ISBN 978-3-030-04184-7. [Google Scholar]
- Yadav, A.; Seth, B.; Chaturvedi, R.K. Brain Organoids: Tiny Mirrors of Human Neurodevelopment and Neurological Disorders. Neuroscientist 2021, 27, 388–426. [Google Scholar] [CrossRef] [PubMed]
- Kozlowski, M.T.; Crook, C.J.; Ku, H.T. Towards organoid culture without Matrigel. Commun. Biol. 2021, 4, 1387. [Google Scholar] [CrossRef] [PubMed]
- Guan, P.-P.; Yu, X.; Guo, J.-J.; Wang, Y.; Wang, T.; Li, J.-Y.; Konstantopoulos, K.; Wang, Z.-Y.; Wang, P. By activating matrix metalloproteinase-7, shear stress promotes chondrosarcoma cell motility, invasion and lung colonization. Oncotarget 2015, 6, 9140–9159. [Google Scholar] [CrossRef]
- Esch, E.W.; Bahinski, A.; Huh, D. Organs-on-chips at the frontiers of drug discovery. Nat. Rev. Drug Discov. 2015, 14, 248–260. [Google Scholar] [CrossRef]
- Tian, C.; Zheng, S.; Liu, X.; Kamei, K. Tumor-on-a-chip model for advancement of anti-cancer nano drug delivery system. J. Nanobiotechnol. 2022, 20, 338. [Google Scholar] [CrossRef]
- Herland, A.; Maoz, B.M.; Das, D.; Somayaji, M.R.; Prantil-Baun, R.; Novak, R.; Cronce, M.; Huffstater, T.; Jeanty, S.S.F.; Ingram, M.; et al. Quantitative prediction of human pharmacokinetic responses to drugs via fluidically coupled vascularized organ chips. Nat. Biomed. Eng. 2020, 4, 421–436. [Google Scholar] [CrossRef]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Fetah, K.L.; DiPardo, B.J.; Kongadzem, E.; Tomlinson, J.S.; Elzagheid, A.; Elmusrati, M.; Khademhosseini, A.; Ashammakhi, N. Cancer Modeling-on-a-Chip with Future Artificial Intelligence Integration. Small 2019, 15, 1901985. [Google Scholar] [CrossRef] [PubMed]
- Bouquerel, C.; Dubrova, A.; Hofer, I.; Phan, D.T.T.; Bernheim, M.; Ladaigue, S.; Cavaniol, C.; Maddalo, D.; Cabel, L.; Mechta-Grigoriou, F.; et al. Bridging the gap between tumor-on-chip and clinics: A systematic review of 15 years of studies. Lab. Chip 2023, 23, 3906–3935. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Wen, J.; Yang, J.; Zhou, S.; Li, Y.; Xu, K.; Li, W.; Li, S. Tumor-microenvironment-on-a-chip: The construction and application. Cell Commun. Signal. 2024, 22, 515. [Google Scholar] [CrossRef]
- Fan, Y.; Nguyen, D.T.; Akay, Y.; Xu, F.; Akay, M. Engineering a Brain Cancer Chip for High-throughput Drug Screening. Sci. Rep. 2016, 6, 25062. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Sun, P.; Yang, L.; Wang, J.; Li, L.; Wang, J. Assay of glioma cell responses to an anticancer drug in a cell-based microfluidic device. Microfluid. Nanofluid. 2010, 9, 717–725. [Google Scholar] [CrossRef]
- Carvalho, M.R.; Barata, D.; Teixeira, L.M.; Giselbrecht, S.; Reis, R.L.; Oliveira, J.M.; Truckenmüller, R.; Habibovic, P. Colorectal tumor-on-a-chip system: A 3D tool for precision onco-nanomedicine. Sci. Adv. 2019, 5, eaaw1317. [Google Scholar] [CrossRef]
- Vidi, P.-A.; Maleki, T.; Ochoa, M.; Wang, L.; Clark, S.M.; Leary, J.F.; Lelièvre, S.A. Disease-on-a-chip: Mimicry of tumor growth in mammary ducts. Lab. Chip 2014, 14, 172–177. [Google Scholar] [CrossRef] [PubMed]
- Albanese, A.; Lam, A.K.; Sykes, E.A.; Rocheleau, J.V.; Chan, W.C.W. Tumour-on-a-chip provides an optical window into nanoparticle tissue transport. Nat. Commun. 2013, 4, 2718. [Google Scholar] [CrossRef] [PubMed]
- Skardal, A.; Devarasetty, M.; Forsythe, S.; Atala, A.; Soker, S. A reductionist metastasis-on-a-chip platform for in vitro tumor progression modeling and drug screening. Biotechnol. Bioeng. 2016, 113, 2020–2032. [Google Scholar] [CrossRef]
- Aleman, J.; Skardal, A. A multi-site metastasis-on-a-chip microphysiological system for assessing metastatic preference of cancer cells. Biotechnol. Bioeng. 2019, 116, 936–944. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.Y.; Wu, Y.; Goh, D.; Tan, V.; Ng, C.W.; Lim, J.C.T.; Lau, M.C.; Yeong, J.P.S. Application of Artificial Intelligence to In Vitro Tumor Modeling and Characterization of the Tumor Microenvironment. Adv. Healthc. Mater. 2023, 12, 2202457. [Google Scholar] [CrossRef] [PubMed]
- Janiesch, C.; Zschech, P.; Heinrich, K. Machine learning and deep learning. Electron. Mark. 2021, 31, 685–695. [Google Scholar] [CrossRef]
- Badillo, S.; Banfai, B.; Birzele, F.; Davydov, I.I.; Hutchinson, L.; Kam-Thong, T.; Siebourg-Polster, J.; Steiert, B.; Zhang, J.D. An Introduction to Machine Learning. Clin. Pharmacol. Ther. 2020, 107, 871–885. [Google Scholar] [CrossRef]
- Vamathevan, J.; Clark, D.; Czodrowski, P.; Dunham, I.; Ferran, E.; Lee, G.; Li, B.; Madabhushi, A.; Shah, P.; Spitzer, M.; et al. Applications of machine learning in drug discovery and development. Nat. Rev. Drug Discov. 2019, 18, 463–477. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Liu, F.; Yang, W.; Peng, S.; Zhou, J. A Survey of Convolutional Neural Networks: Analysis, Applications, and Prospects. IEEE Trans. Neural Netw. Learn. Syst. 2022, 33, 6999–7019. [Google Scholar] [CrossRef] [PubMed]
- Benning, L.; Peintner, A.; Finkenzeller, G.; Peintner, L. Automated spheroid generation, drug application and efficacy screening using a deep learning classification: A feasibility study. Sci. Rep. 2020, 10, 11071. [Google Scholar] [CrossRef] [PubMed]
- Kshatri, S.S.; Singh, D. Convolutional Neural Network in Medical Image Analysis: A Review. Arch. Comput. Methods Eng. 2023, 30, 2793–2810. [Google Scholar] [CrossRef]
- D’Orazio, M.; Corsi, F.; Mencattini, A.; Di Giuseppe, D.; Colomba Comes, M.; Casti, P.; Filippi, J.; Di Natale, C.; Ghibelli, L.; Martinelli, E. Deciphering Cancer Cell Behavior From Motility and Shape Features: Peer Prediction and Dynamic Selection to Support Cancer Diagnosis and Therapy. Front. Oncol. 2020, 10, 580698. [Google Scholar] [CrossRef]
- Mencattini, A.; Di Giuseppe, D.; Comes, M.C.; Casti, P.; Corsi, F.; Bertani, F.R.; Ghibelli, L.; Businaro, L.; Di Natale, C.; Parrini, M.C.; et al. Discovering the hidden messages within cell trajectories using a deep learning approach for in vitro evaluation of cancer drug treatments. Sci. Rep. 2020, 10, 7653. [Google Scholar] [CrossRef]
- Cortesi, M.; Liu, D.; Powell, E.; Barlow, E.; Warton, K.; Ford, C.E. Accurate Identification of Cancer Cells in Complex Pre-Clinical Models Using a Deep-Learning Neural Network: A Transfection-Free Approach. Adv. Biol. 2024, 8, 2400034. [Google Scholar] [CrossRef]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards Real-Time Object Detection with Region Proposal Networks. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 1137–1149. [Google Scholar] [CrossRef]
- Li, Y.; Mahjoubfar, A.; Chen, C.L.; Niazi, K.R.; Pei, L.; Jalali, B. Deep Cytometry: Deep learning with Real-time Inference in Cell Sorting and Flow Cytometry. Sci. Rep. 2019, 9, 11088. [Google Scholar] [CrossRef]
- Oliver, C.R.; Westerhof, T.M.; Castro, M.G.; Merajver, S.D. Quantifying the Brain Metastatic Tumor Micro-Environment using an Organ-On-A Chip 3D Model, Machine Learning, and Confocal Tomography. J. Vis. Exp. 2020, 61654. [Google Scholar] [CrossRef]
- Oliver, C.R.; Altemus, M.A.; Westerhof, T.M.; Cheriyan, H.; Cheng, X.; Dziubinski, M.; Wu, Z.; Yates, J.; Morikawa, A.; Heth, J.; et al. A platform for artificial intelligence based identification of the extravasation potential of cancer cells into the brain metastatic niche. Lab. Chip 2019, 19, 1162–1173. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Hernández, M.A.; Chapresto-Garzón, R.; Cadenas, M.; Navarro-Villarán, E.; Negrete, M.; Gómez-Bravo, M.A.; Victor, V.M.; Padillo, F.J.; Muntané, J. Differential effectiveness of tyrosine kinase inhibitors in 2D/3D culture according to cell differentiation, p53 status and mitochondrial respiration in liver cancer cells. Cell Death Dis. 2020, 11, 339. [Google Scholar] [CrossRef]
- Yoo, J.; Choi, J.W.; Kim, E.; Park, E.-J.; Baek, A.; Kim, J.; Jeong, M.S.; Cho, Y.; Lee, T.G.; Heo, M.B. Evaluating cell growth and hypoxic regions of 3D spheroids via a machine learning approach. Mach. Learn. Sci. Technol. 2024, 5, 035063. [Google Scholar] [CrossRef]
- Kong, J.; Lee, H.; Kim, D.; Han, S.K.; Ha, D.; Shin, K.; Kim, S. Network-based machine learning in colorectal and bladder organoid models predicts anti-cancer drug efficacy in patients. Nat. Commun. 2020, 11, 5485. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.; Jiang, S.; Huang, X.; Ji, C.; Gu, Y.; Qi, Y.; Xiang, Y.; Yao, E.; Zhang, N.; Berman, E.; et al. Integration of 3D bioprinting and multi-algorithm machine learning identified glioma susceptibilities and microenvironment characteristics. Cell Discov. 2024, 10, 39. [Google Scholar] [CrossRef]
- Sammut, S.-J.; Crispin-Ortuzar, M.; Chin, S.-F.; Provenzano, E.; Bardwell, H.A.; Ma, W.; Cope, W.; Dariush, A.; Dawson, S.-J.; Abraham, J.E.; et al. Multi-omic machine learning predictor of breast cancer therapy response. Nature 2022, 601, 623–629. [Google Scholar] [CrossRef] [PubMed]
- Boehm, K.M.; Aherne, E.A.; Ellenson, L.; Nikolovski, I.; Alghamdi, M.; Vázquez-García, I.; Zamarin, D.; Long Roche, K.; Liu, Y.; Patel, D.; et al. Multimodal data integration using machine learning improves risk stratification of high-grade serous ovarian cancer. Nat. Cancer 2022, 3, 723–733. [Google Scholar] [CrossRef] [PubMed]
- Adam, G.; Rampášek, L.; Safikhani, Z.; Smirnov, P.; Haibe-Kains, B.; Goldenberg, A. Machine learning approaches to drug response prediction: Challenges and recent progress. NPJ Precis. Oncol. 2020, 4, 19. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Clayton, E.A.; Matyunina, L.V.; McDonald, L.D.; Benigno, B.B.; Vannberg, F.; McDonald, J.F. Machine learning predicts individual cancer patient responses to therapeutic drugs with high accuracy. Sci. Rep. 2018, 8, 16444. [Google Scholar] [CrossRef] [PubMed]
- Zhavoronkov, A.; Vanhaelen, Q.; Oprea, T.I. Will Artificial Intelligence for Drug Discovery Impact Clinical Pharmacology? Clin. Pharmacol. Ther. 2020, 107, 780–785. [Google Scholar] [CrossRef] [PubMed]
- Stokes, J.M.; Yang, K.; Swanson, K.; Jin, W.; Cubillos-Ruiz, A.; Donghia, N.M.; MacNair, C.R.; French, S.; Carfrae, L.A.; Bloom-Ackermann, Z.; et al. A Deep Learning Approach to Antibiotic Discovery. Cell 2020, 180, 688–702.e13. [Google Scholar] [CrossRef] [PubMed]
- Bonner, S.; Barrett, I.P.; Ye, C.; Swiers, R.; Engkvist, O.; Hoyt, C.T.; Hamilton, W.L. Understanding the performance of knowledge graph embeddings in drug discovery. Artif. Intell. Life Sci. 2022, 2, 100036. [Google Scholar] [CrossRef]
- Artrith, N.; Butler, K.T.; Coudert, F.-X.; Han, S.; Isayev, O.; Jain, A.; Walsh, A. Best practices in machine learning for chemistry. Nat. Chem. 2021, 13, 505–508. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Momoli, C.; Costa, B.; Lenti, L.; Tubertini, M.; Parenti, M.D.; Martella, E.; Varchi, G.; Ferroni, C. The Evolution of Anticancer 3D In Vitro Models: The Potential Role of Machine Learning and AI in the Next Generation of Animal-Free Experiments. Cancers 2025, 17, 700. https://doi.org/10.3390/cancers17040700
Momoli C, Costa B, Lenti L, Tubertini M, Parenti MD, Martella E, Varchi G, Ferroni C. The Evolution of Anticancer 3D In Vitro Models: The Potential Role of Machine Learning and AI in the Next Generation of Animal-Free Experiments. Cancers. 2025; 17(4):700. https://doi.org/10.3390/cancers17040700
Chicago/Turabian StyleMomoli, Carolina, Beatrice Costa, Lorenzo Lenti, Matilde Tubertini, Marco Daniele Parenti, Elisa Martella, Greta Varchi, and Claudia Ferroni. 2025. "The Evolution of Anticancer 3D In Vitro Models: The Potential Role of Machine Learning and AI in the Next Generation of Animal-Free Experiments" Cancers 17, no. 4: 700. https://doi.org/10.3390/cancers17040700
APA StyleMomoli, C., Costa, B., Lenti, L., Tubertini, M., Parenti, M. D., Martella, E., Varchi, G., & Ferroni, C. (2025). The Evolution of Anticancer 3D In Vitro Models: The Potential Role of Machine Learning and AI in the Next Generation of Animal-Free Experiments. Cancers, 17(4), 700. https://doi.org/10.3390/cancers17040700









